Matrix Nanopatterning Regulates Mesenchymal Differentiation through Focal Adhesion Size and Distribution According to Cell Fate
Abstract
1. Introduction
2. Materials and Methods
2.1. Production of Nanopatterned Substrates
2.2. Cell Culture
2.3. Immunostaining and Image Acquisition
2.4. Analysis of Focal Adhesions and Nuclei Morphology
2.5. Analysis of Differentiation Markers
2.6. Statistical Analysis
3. Results
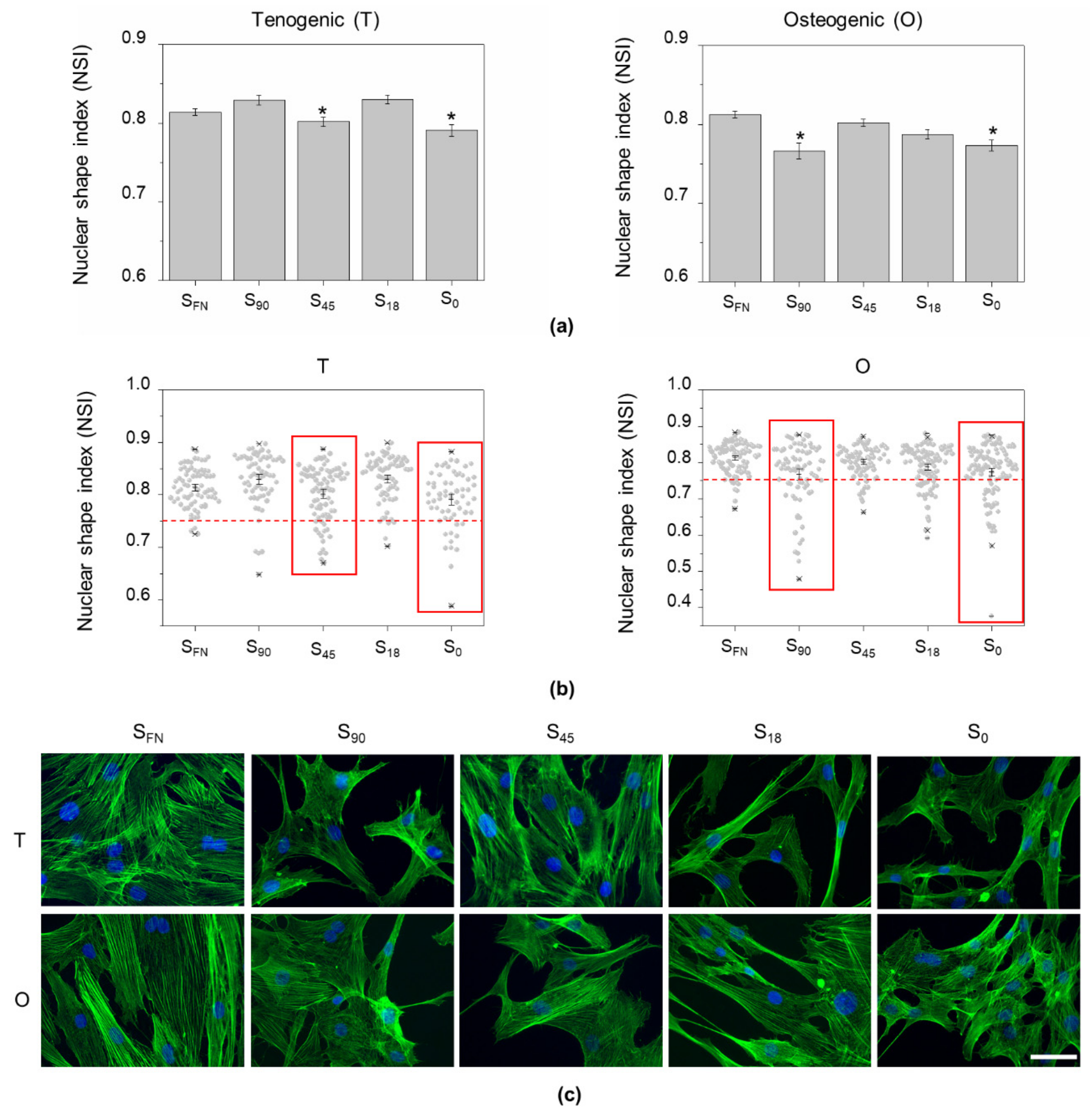
3.1. Nuclear Remodelling on the Nanopatterns
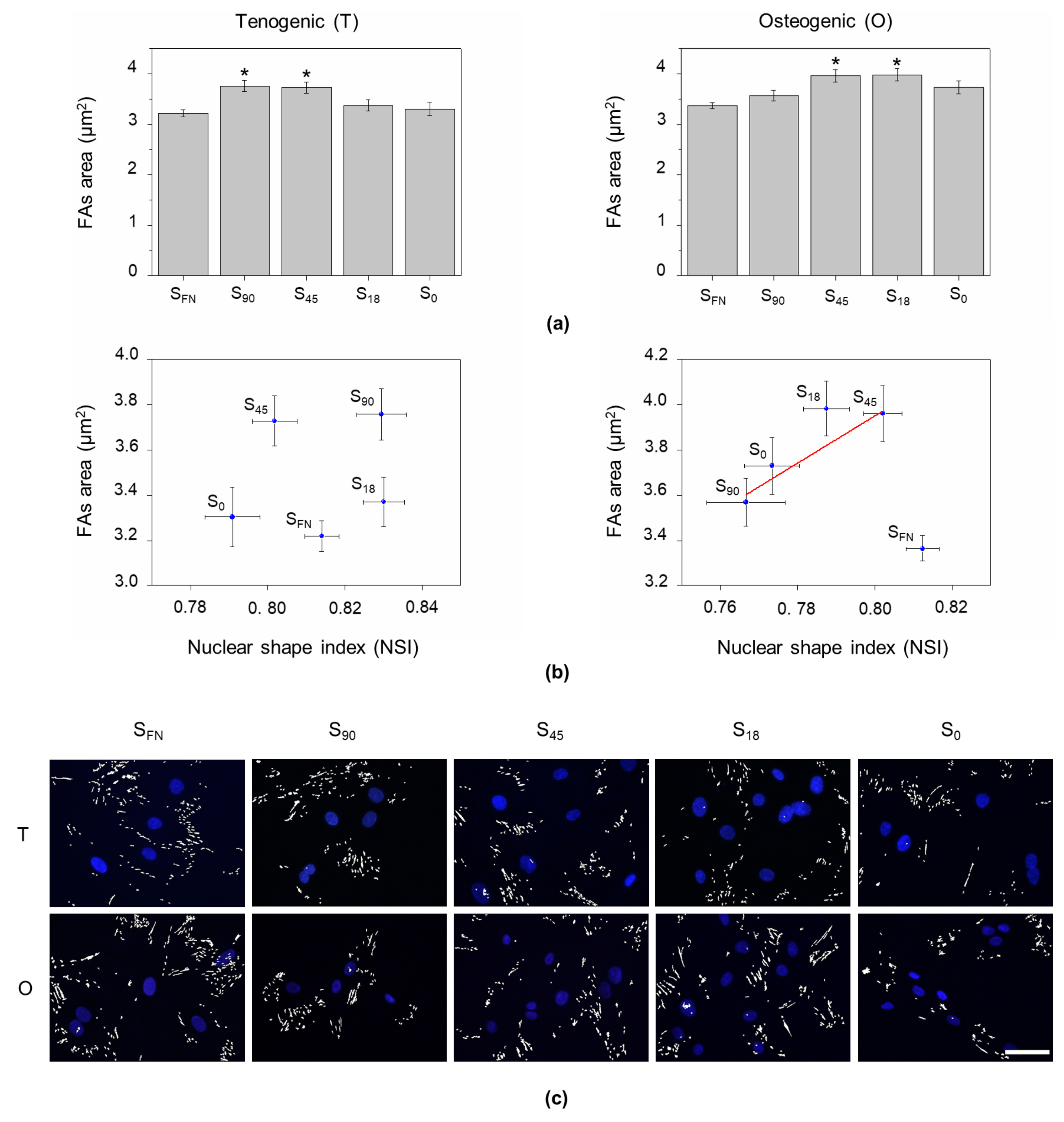
3.2. Cytoskeletal Tension Influence on Nuclear Remodelling
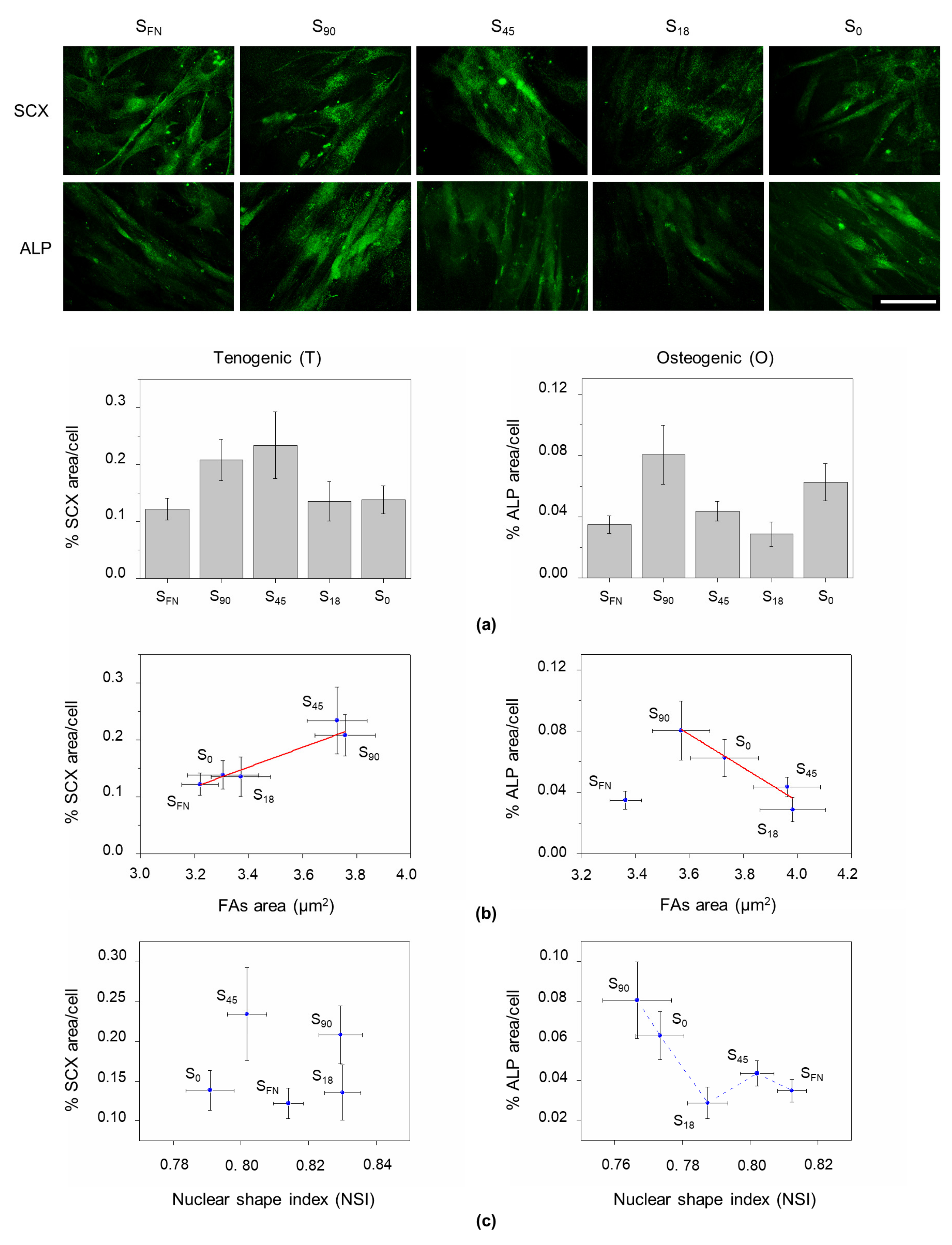
3.3. Nuclear Shape and Lineage Commitment
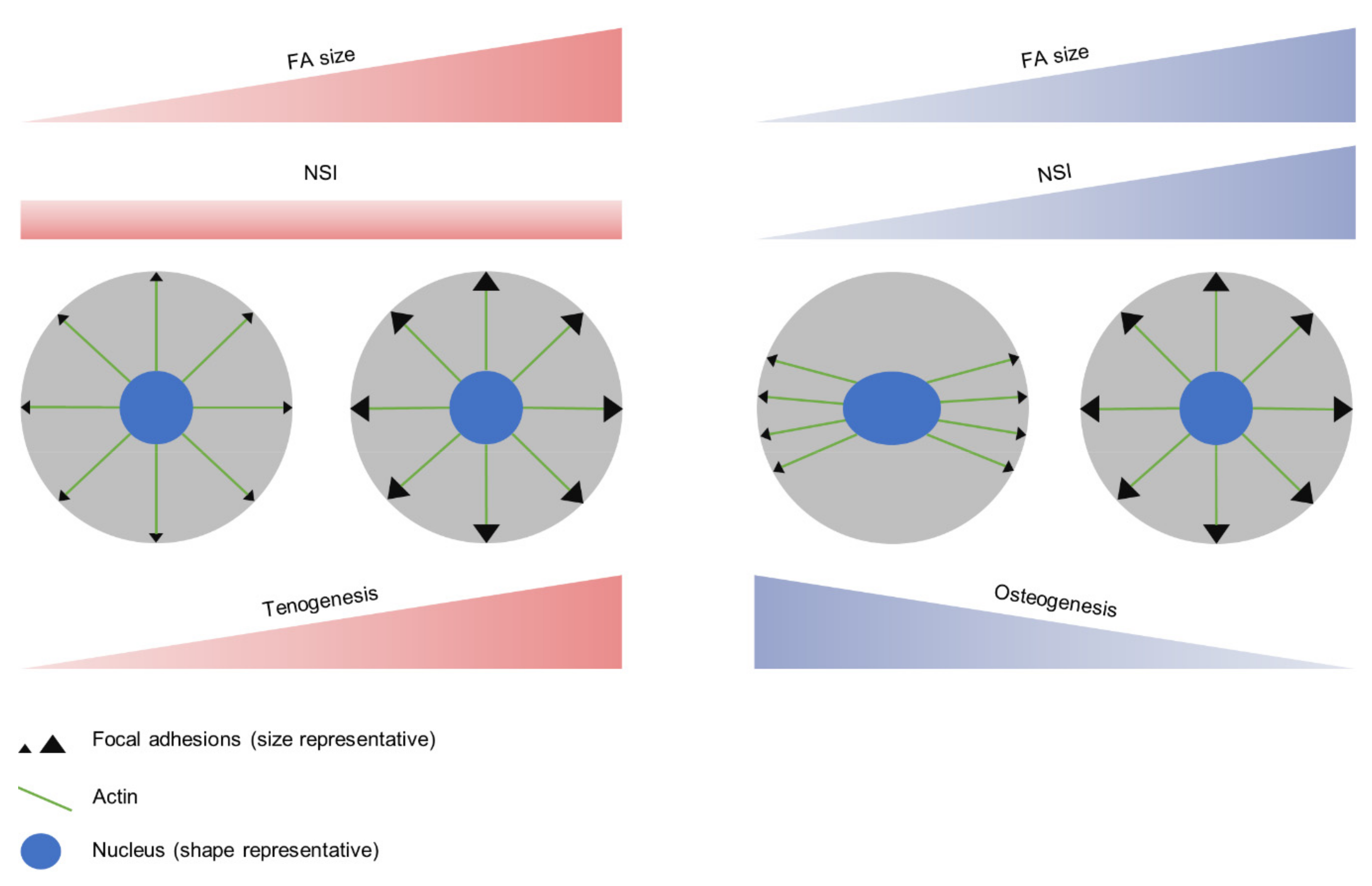
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bonnans, C.; Chou, J.; Werb, Z. Remodelling the extracellular matrix in development and disease. Nat. Rev. Mol. Cell Biol. 2014, 15, 786–801. [Google Scholar] [CrossRef] [PubMed]
- Miwa, H.; Era, T. Tracing the destiny of mesenchymal stem cells from embryo to adult bone marrow and white adipose tissue via Pdgfrα expression. Development 2018, 145, dev155879. [Google Scholar] [CrossRef] [PubMed]
- Assis-Ribasa, T.; Fornib, M.F.; Winnischoferc, S.M.B.; Sogayara, M.C.; Trombetta-Lima, M. Extracellular matrix dynamics during mesenchymal stem cells differentiation. Dev. Biol. 2018, 437, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Ingber, D.E.; Wang, N.; Stamenovic, D. Tensegrity, cellular biophysics, and the mechanics of living systems. Rep. Prog. Phys. 2014, 77, 046603–046624. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Guo, S.S.; Fässler, R. Integrin-mediated mechanotransduction. J. Cell Biol. 2016, 215, 445–456. [Google Scholar] [CrossRef] [PubMed]
- Kanchanawong, P.; Shtengel, G.; Pasapera, A.M.; Ramko, E.B.; Davidson, M.W.; Hess, H.F.; Waterman, C.M. Nanoscale architecture of integrin-based cell adhesions. Nature 2010, 468, 580–584. [Google Scholar] [CrossRef] [PubMed]
- Rahil, Z.; Pedron, S.; Wang, X.; Ha, T.; Harley, B.; Leckband, D. Nanoscale mechanics guides cellular decision making. Integr. Biol. 2016, 8, 929–935. [Google Scholar] [CrossRef]
- Arnold, M.; Cavalcanti-Adam, E.A.; Glass, R.; Blümmel, J.; Eck, W.; Kantlehner, M.; Kessler, H.; Spatz, J.P. Activation of integrin function by nanopatterned adhesive interfaces. Chem. Phys. Chem. 2004, 5, 383–388. [Google Scholar] [CrossRef] [PubMed]
- Cavalcanti-Adam, E.A.; Micoulet, A.; Blümmel, J.; Auernheimer, J.; Kessler, H.; Spatz, J.P. Lateral spacing of integrin ligands influences cell spreading and focal adhesion assembly. Eur. J. Cell. Biol. 2006, 85, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Cavalcanti-Adam, E.A.; Volberg, T.; Micoulet, A.; Kessler, H.; Geiger, B.; Spatz, J.P. Cell spreading and focal adhesion dynamics are regulated by spacing of integrin ligands. Biophys. J. 2007, 92, 2964–2974. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yan, C.; Ye, K.; He, Y.; Li, Z.; Ding, J. Effect of RGD nanospacing on differentiation of stem cells. Biomaterials 2013, 34, 2865–2874. [Google Scholar] [CrossRef] [PubMed]
- Oria, R.; Wiegand, T.; Escribano, J.; Elosegui-Artola, A.; Uriarte, J.J.; Moreno-Pulido, C.; Platzman, I.; Delcanale, P.; Albertazzi, L.; Navajas, D.; et al. Force loading explains spatial sensing of ligands by cells. Nature 2017, 552, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Chambliss, A.B.; Wirtz, D. The multi-faceted role of the actin cap in cellular mechanosensation and mechanotransduction. Soft Matter 2013, 9, 5516–5523. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.H.; Collier, J.H.; Sfeir, C.S.; Healy, K.E. Engineering gene expression and protein synthesis by modulation of nuclear shape. Proc. Natl. Acad. Sci. USA 2002, 99, 1972–1977. [Google Scholar] [CrossRef] [PubMed]
- Lagunas, A.; Castaño, A.G.; Artés, J.M.; Vida, Y.; Collado, D.; Pérez-Inestrosa, E.; Gorostiza, P.; Claros, S.; Andrades, J.A.; Samitier, J. Large-scale dendrimer-based uneven nanopatterns for the study of local arginine–glycine–aspartic acid (RGD) density effects on cell adhesion. Nano Res. 2014, 7, 399–409. [Google Scholar] [CrossRef]
- Casanellas, I.; Lagunas, A.; Tsintzou, I.; Vida, Y.; Collado, D.; Pérez-Inestrosa, E.; Rodríguez-Pereira, C.; Magalhaes, J.; Gorostiza, P.; Andrades, J.A.; et al. Dendrimer-based uneven nanopatterns to locally control surface adhesiveness: a method to direct chondrogenic differentiation. J. Vis. Exp. 2018, 20. [Google Scholar] [CrossRef] [PubMed]
- Lagunas, A.; Tsintzou, I.; Vida, Y.; Collado, D.; Pérez-Inestrosa, E.; Rodríguez Pereira, C.; Magalhaes, J.; Andrades, J.A.; Samitier, J. Tailoring RGD local surface density at the nanoscale toward adult stem cell chondrogenic commitment. Nano Res. 2017, 10, 1959–1971. [Google Scholar] [CrossRef]
- Park, A.; Hogan, M.V.; Kesturu, G.S.; James, R.; Balian, G.; Chhabra, A.B. Adipose-derived mesenchymal stem cells treated with growth differentiation factor-5 express tendon-specific markers. Tissue Eng. Part A. 2010, 9, 2941–2951. [Google Scholar] [CrossRef]
- Xiong, J.P.; Stehle, T.; Zhang, R.G.; Joachimiak, A.; Frech, M.; Goodman, S.L.; Arnaout, M.A. Crystal structure of the extracellular segment of integrin αVβ3 in complex with an Arg-Gly-Asp ligand. Science 2002, 296, 151–155. [Google Scholar] [CrossRef]
- George, E.L.; Georges-Labouesse, E.N.; Patel-King, R.S.; Rayburn, H.; Hynes, R.O. Defects in mesoderm, neural tube and vascular development in mouse embryos lacking fibronectin. Development 1993, 119, 1079–1091. [Google Scholar]
- Versaevel, M.; Grevessse, T.; Gabriele, S. Spatial coordination between cell and nuclear shape within micropatterned endothelial cells. Nature Commun. 2012, 3, 671–681. [Google Scholar] [CrossRef] [PubMed]
- Sims, J.R.; Karp, S.; Ingber, D.E. Altering the cellular mechanical force balance results in integrated changes in cell, cytoskeletal and nuclear shape. J. Cell Sci. 1992, 103, 1215–1222. [Google Scholar] [PubMed]
- Jean, R.P.; Gray, D.S.; Spector, A.A.; Chen, C.S. Characterization of the nuclear deformation caused by changes in endothelial cell shapes. J. Biomech. Eng. 2004, 126, 552–558. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.-H.; Cho, S.; Wirtz, D. Tight coupling between nucleus and cell migration through the perinuclear actin cap. J. Cell Sci. 2014, 127, 2528–2541. [Google Scholar] [CrossRef] [PubMed]
- Galbraith, C.G.; Sheetz, M.P. A micromachined device provides a new bend on fibroblast traction forces. Proc. Natl. Acad. Sci. USA 1997, 94, 9114–9118. [Google Scholar] [CrossRef]
- Balaban, N.Q.; Schwarz, U.S.; Riveline, D.; Goichberg, P.; Tzur, G.; Sabanay, I.; Mahalu, D.; Safran, S.; Bershadsky, A.; Addadi, L.; et al. Force and focal adhesion assembly: a close relationship studied using elastic micropatterned substrates. Nat. Cell Biol. 2001, 3, 466–473. [Google Scholar] [CrossRef]
- Kim, D.H.; Wirtz, D. Focal adhesion size uniquely predicts cell migration. FASEB J. 2013, 27, 1351–1361. [Google Scholar] [CrossRef]
- Vergani, L.; Grattarola, M.; Nicolini, C. Modifications of chromatin structure and gene expression following induced alterations of cellular shape. Int. J. Biochem. Cell Biol. 2004, 36, 1447–1461. [Google Scholar] [CrossRef]
- Schweitzer, R.; Chyung, J.H.; Murtaugh, L.C.; Brent, A.E.; Rosen, V.; Olson, E.N.; Lassar, A.; Tabin, C.J. Analysis of the tendon cell fate using Scleraxis, a specific marker for tendons and ligaments. Development 2001, 128, 3855–3866. [Google Scholar]
- Tan, S.-L.; Ahmad, T.S.; Ng, W.-M.; Azlina, A.A.; Azhar, M.M.; Selvaratnam, L.; Kamarul, T. Identification of pathways mediating growth differentiation Factor5-induced tenogenic differentiation in human bone marrow stromal cells. PLoS ONE 2015, 10, e0140869. [Google Scholar] [CrossRef]
- Zhang, W.; Yang, N.; Shi, X.-M. Regulation of mesenchymal stem cell osteogenic differentiation by glucocorticoid-induced leucine Zipper (GILZ). J. Biol. Chem. 2008, 283, 4723–4729. [Google Scholar] [CrossRef] [PubMed]
- Lagunas, A.; Comelles, J.; Oberhansl, S.; Hortigüela, V.; Martínez, E.; Samitier, J. Continuous bone morphogenetic protein-2 gradients for concentration effect studies on C2C12 osteogenic fate. Nanomedicine 2013, 9, 694–701. [Google Scholar] [CrossRef] [PubMed]
- Tai, G.; Christodoulou, J.; Bishop, A.E.; Polak, J.M. Use of green fluorescent fusion protein to track activation of the transcription factor osterix during early osteoblast differentiation. Biochem. Biophys. Res. Commun. 2005, 333, 1116–1122. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, S.; Yan, C.; Liu, P.; Ding, J. Fabrication of RGD micro/nanopattern and corresponding study of stem cell differentiation. Nano Lett. 2015, 15, 1457–1467. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.S.; Mrksich, M.; Huang, S.; Whitesides, G.M.; Ingber, D.E. Geometric control of cell life and death. Science 1997, 276, 1425–1428. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.B.; Gao, C.Y.; Liu, X.Y.; He, T.; Shen, J.C. Immobilization of biomacromolecules onto aminolyzed poly(L-lactic acid) toward acceleration of endothelium regeneration. Tissue Eng. 2004, 10, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Elosegui-Artola, A.; Trepat, X.; Roca-Cusachs, P. Control of mechanotransduction by molecular clutch dynamics. Trends Cell Biol. 2018, 28, 356–367. [Google Scholar] [CrossRef]
- Huang, J.; Gräter, S.V.; Corbellini, F.; Rinck, S.; Bock, E.; Kemkemer, R.; Kessler, H.; Ding, J.; Spatz, J.P. Impact of order and disorder in RGD nanopatterns on cell adhesion. Nano Lett. 2009, 9, 1111–1116. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, W.; Liu, Z.; Zhu, T.; Shen, W.; Ran, J.; Tang, O.; Gong, X.; Backman, L.J.; Chen, X.; et al. Characterization and comparison of post-natal rat Achilles tendonderived stem cells at different development stages. Sci. Rep. 2016, 6, 22946. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Casanellas, I.; Lagunas, A.; Vida, Y.; Pérez-Inestrosa, E.; Andrades, J.A.; Becerra, J.; Samitier, J. Matrix Nanopatterning Regulates Mesenchymal Differentiation through Focal Adhesion Size and Distribution According to Cell Fate. Biomimetics 2019, 4, 43. https://doi.org/10.3390/biomimetics4020043
Casanellas I, Lagunas A, Vida Y, Pérez-Inestrosa E, Andrades JA, Becerra J, Samitier J. Matrix Nanopatterning Regulates Mesenchymal Differentiation through Focal Adhesion Size and Distribution According to Cell Fate. Biomimetics. 2019; 4(2):43. https://doi.org/10.3390/biomimetics4020043
Chicago/Turabian StyleCasanellas, Ignasi, Anna Lagunas, Yolanda Vida, Ezequiel Pérez-Inestrosa, José A. Andrades, José Becerra, and Josep Samitier. 2019. "Matrix Nanopatterning Regulates Mesenchymal Differentiation through Focal Adhesion Size and Distribution According to Cell Fate" Biomimetics 4, no. 2: 43. https://doi.org/10.3390/biomimetics4020043
APA StyleCasanellas, I., Lagunas, A., Vida, Y., Pérez-Inestrosa, E., Andrades, J. A., Becerra, J., & Samitier, J. (2019). Matrix Nanopatterning Regulates Mesenchymal Differentiation through Focal Adhesion Size and Distribution According to Cell Fate. Biomimetics, 4(2), 43. https://doi.org/10.3390/biomimetics4020043






