Efficient Extraction of Deep Image Features Using a Convolutional Neural Network (CNN) for Detecting Ventricular Fibrillation and Tachycardia
Abstract
:1. Introduction
1.1. Related Work
1.2. Proposed Work
2. Deep Learning Algorithms
2.1. Fundamental Concepts of Convolutional Neural Networks
- The convolutional layer (CONV), which processes the received input data;
- The pooling layer (POOL), which allows compressing the information by reducing the size of the intermediate image (often by subsampling);
- The Fully Connected Layer (FCL) layer, which is a perceptron-type layer;
- The classification layer (Softmax), which predicts the class of the input image.
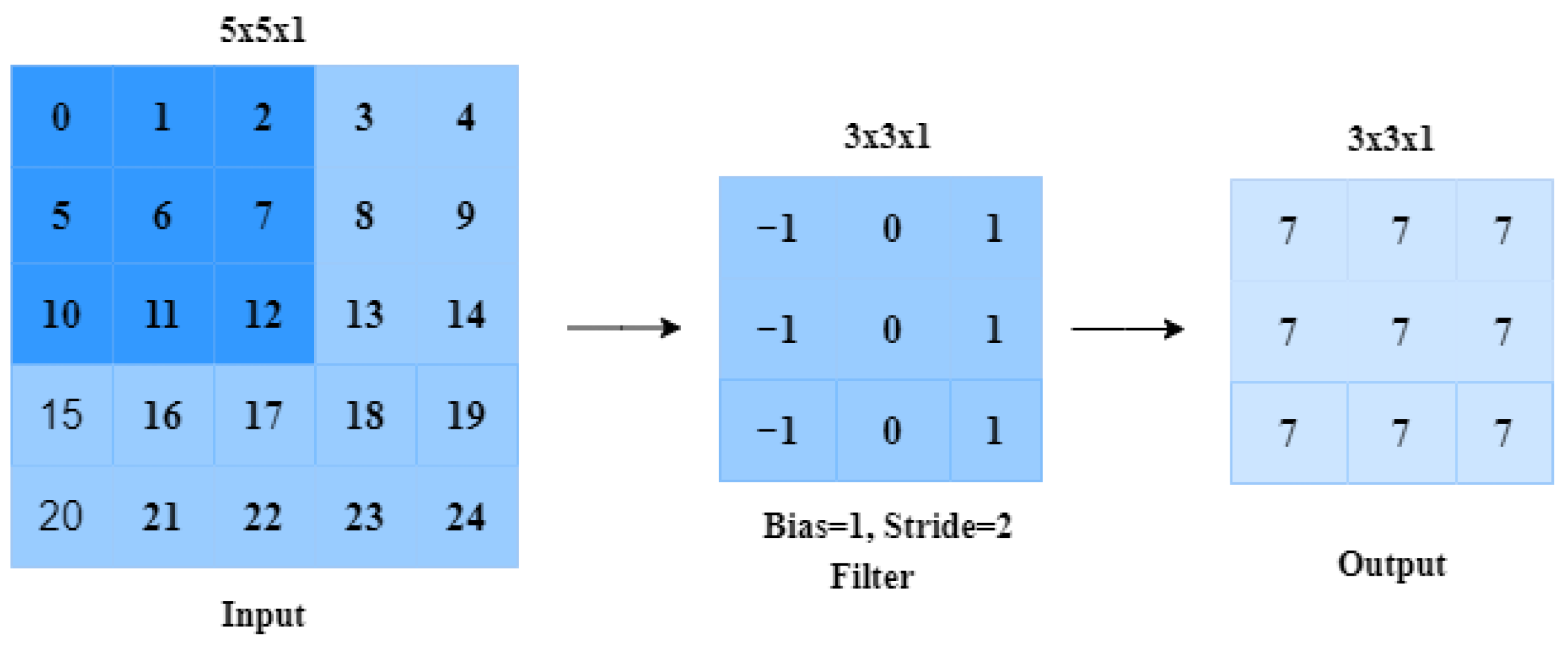
2.1.1. Convolutional Layer
2.1.2. Nonlinear Activation Function
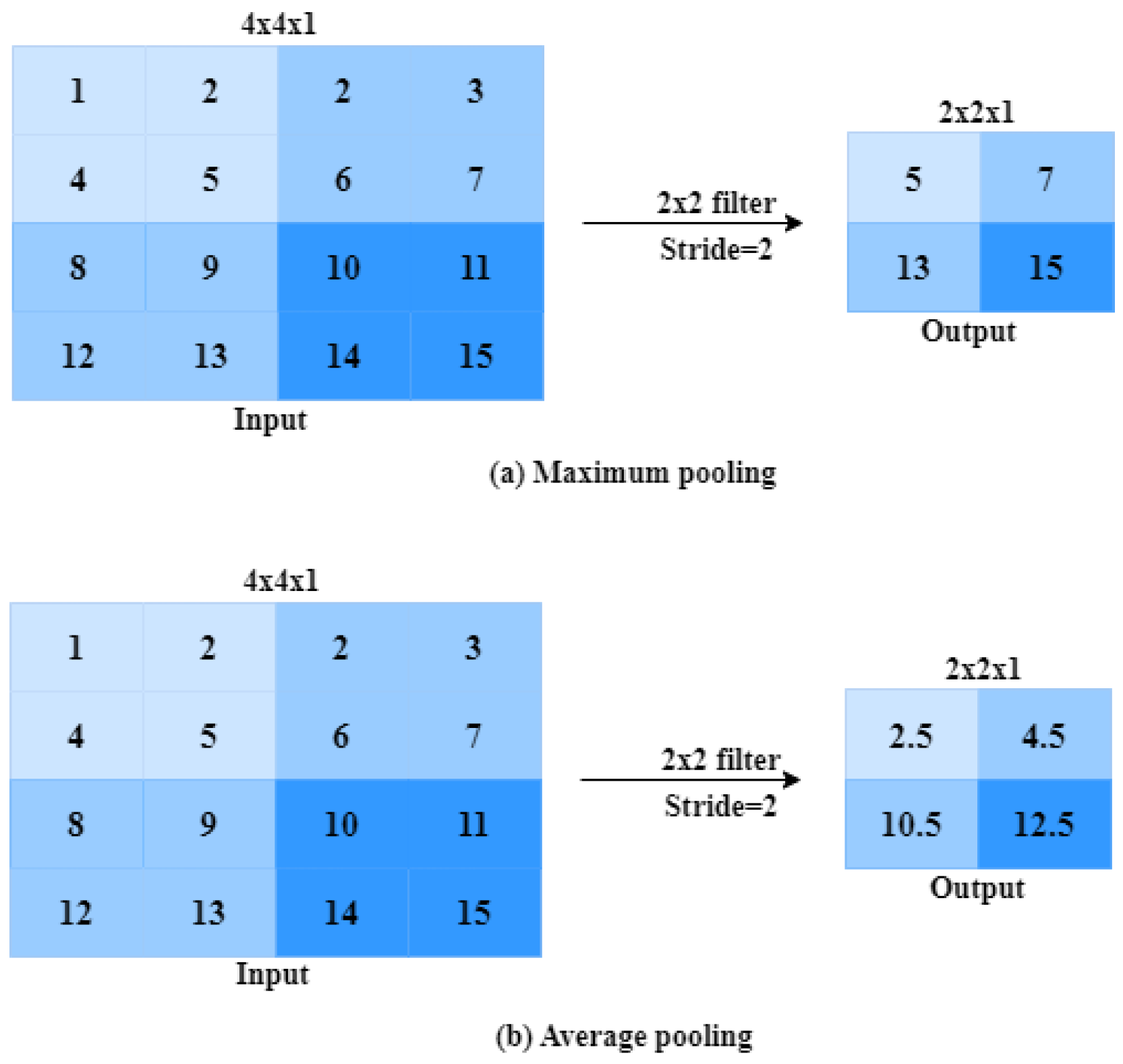
2.1.3. Pooling Layer
2.1.4. Fully Connected Layer
2.1.5. Loss Function
2.2. Optimization of Hyperparameters
- Number of layers [42]: A conventional CNN typically consists of multiple layers, including convolutional layers, activation layers (e.g., ReLU), pooling layers, and fully connected layers.
- Filter size (Kernel Size) [43]: The size of the filters used in the convolutional layers is an important parameter. Common filter sizes are 3 × 3, 5 × 5, and 7 × 7.
- Number of filters [44]: The number of filters in each convolutional layer determines the depth of the feature maps generated. More filters lead to more expressive power but also increase computation requirements.
- Stride [45]: The stride determines the step size at which the filter is moved across the input image. Common values are 1 and 2, with larger strides reducing the size of the output feature maps.
- Padding [45]: Padding can be used to preserve the spatial dimensions of the input when convolving with filters. Common padding values are ’same’ and ’valid’.
- Activation function [46]: Common activation functions include ReLU (rectified linear unit), leaky ReLU, and Sigmoid. ReLU is widely used due to its simplicity and effectiveness.
- Pooling [47]: Pooling layers downsample the feature maps reduces the spatial dimensions. Common pooling types are Max pooling and average pooling, typically with a pool size of .
- Fully connected layers [48]: The number of neurons in the fully connected layers can vary based on the complexity of the task. The output layer size depends on the number of classes in the classification task.
- Dropout [49]: Dropout is a regularization technique that randomly sets a fraction of neurons to zero during training, preventing overfitting. Common dropout rates are between 0.2 and 0.5.
- Batch size [50]: The number of samples used in each iteration during training. Smaller batch sizes are computationally more expensive but can lead to better convergence.
- Number of epochs [51]: This is the number of times the entire training dataset is passed through the network during training.
- Learning rate [52]: The learning rate controls the step size during optimization. A small learning rate leads to slow convergence, while a large learning rate can cause instability.
- Optimizer: Common optimizers used in CNNs include Stochastic Gradient Descent (SGD) [53], Adam, and RMSprop.
2.3. CNN Architectures
2.3.1. AlexNet
2.3.2. VGGNet
2.3.3. Inception V3
2.3.4. MobileNet
3. Time–Frequency Representation
4. Material and Methods
- First phase: The dataset used is described.
- Second phase: The ECG data undergoes filtering to reduce baseline interference. Once filtered, the Window Reference Mark (WRM) of the ECG signal is obtained. Each WRM indicates the start of a time window (tw) within the ECG signal.
- Third phase: Information extraction is performed by applying the Hilbert transform (Ht) to each window tw obtained in the first phase. Subsequently, the TFR matrix is computed using the Pseudo Wigner–Ville method, resulting in the Time–Frequency Representation Image (TFRI).
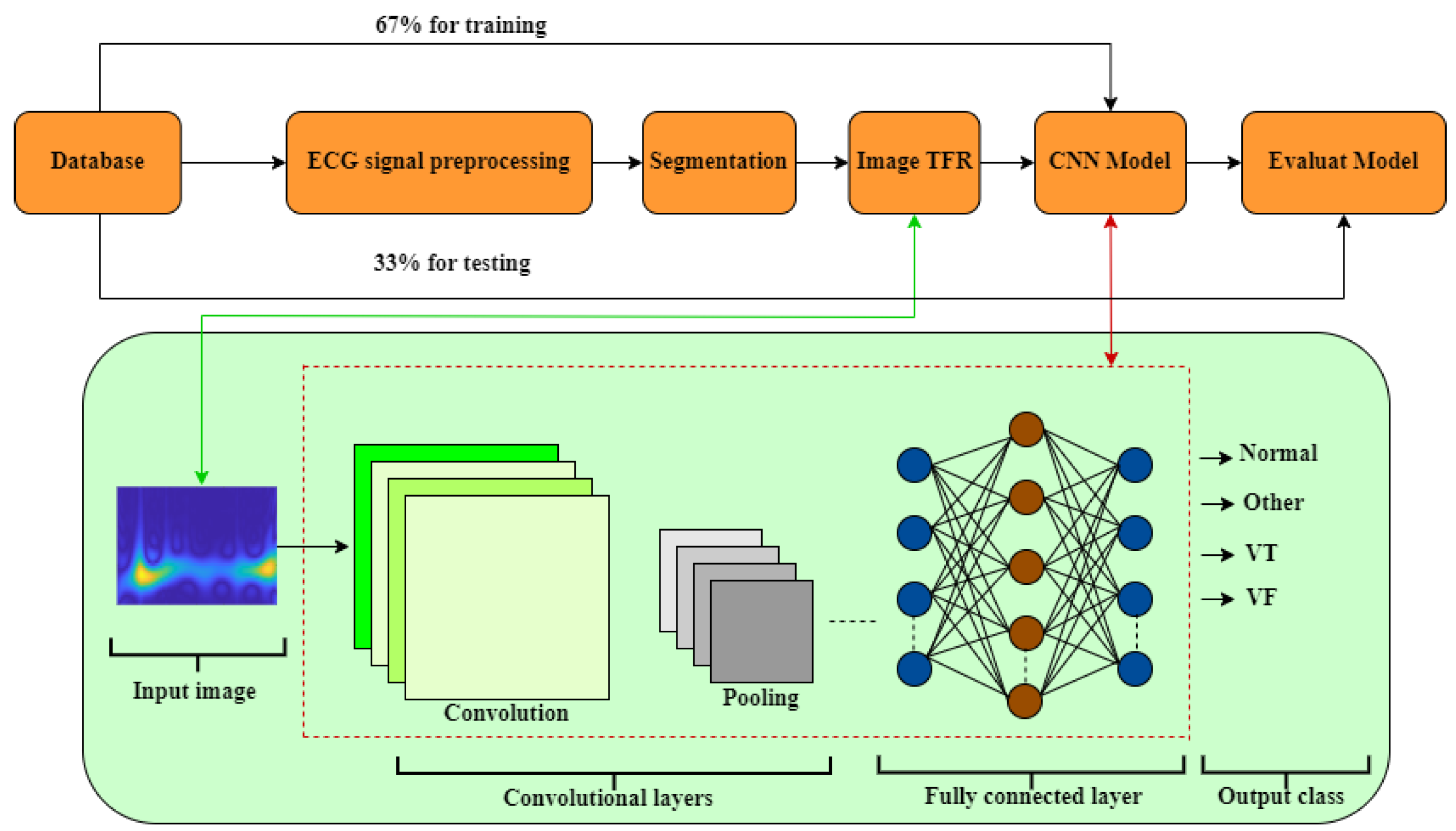
- Fourth phase: The TFRI matrices obtained in the previous step are used as input for a deep learning CNN (CNN1, CNN2, InceptionV3, MobilNet, VGGNet, and AlexNet), as detailed in Section 2.3 and Section 4.4.1. The success of ventricular fibrillation (VF) detection relies on signal processing techniques and the structure of the classifiers employed. To achieve optimal performance, it is necessary to adjust the CNN parameters to better adapt to the data.
4.1. Materials
4.2. Electrocardiographic Signal Preprocessing
4.2.1. Denoising
4.2.2. Segmentation
4.3. Extraction of Image from TFR
4.4. Model Training and Evaluation
4.4.1. Model Architecture
- In the CNN1 method, 2 fully connected layers utilize the output from the TFR and predict the class of the image based on the vector calculated in previous stages.
- In the CNN2 method, the network consists of 6 layers, including 2 convolution layers, 2 max-pooling layers, and 2 fully connected layers. Each convolution layer (layers 1 and 2) applies convolution with its respective kernel size (layers 3 and 4). Following each convolution layer, a max-pooling operation is performed on the generated feature maps. The purpose of max-pooling is to reduce the dimensionality of the feature maps, aiding in the extraction of essential features.
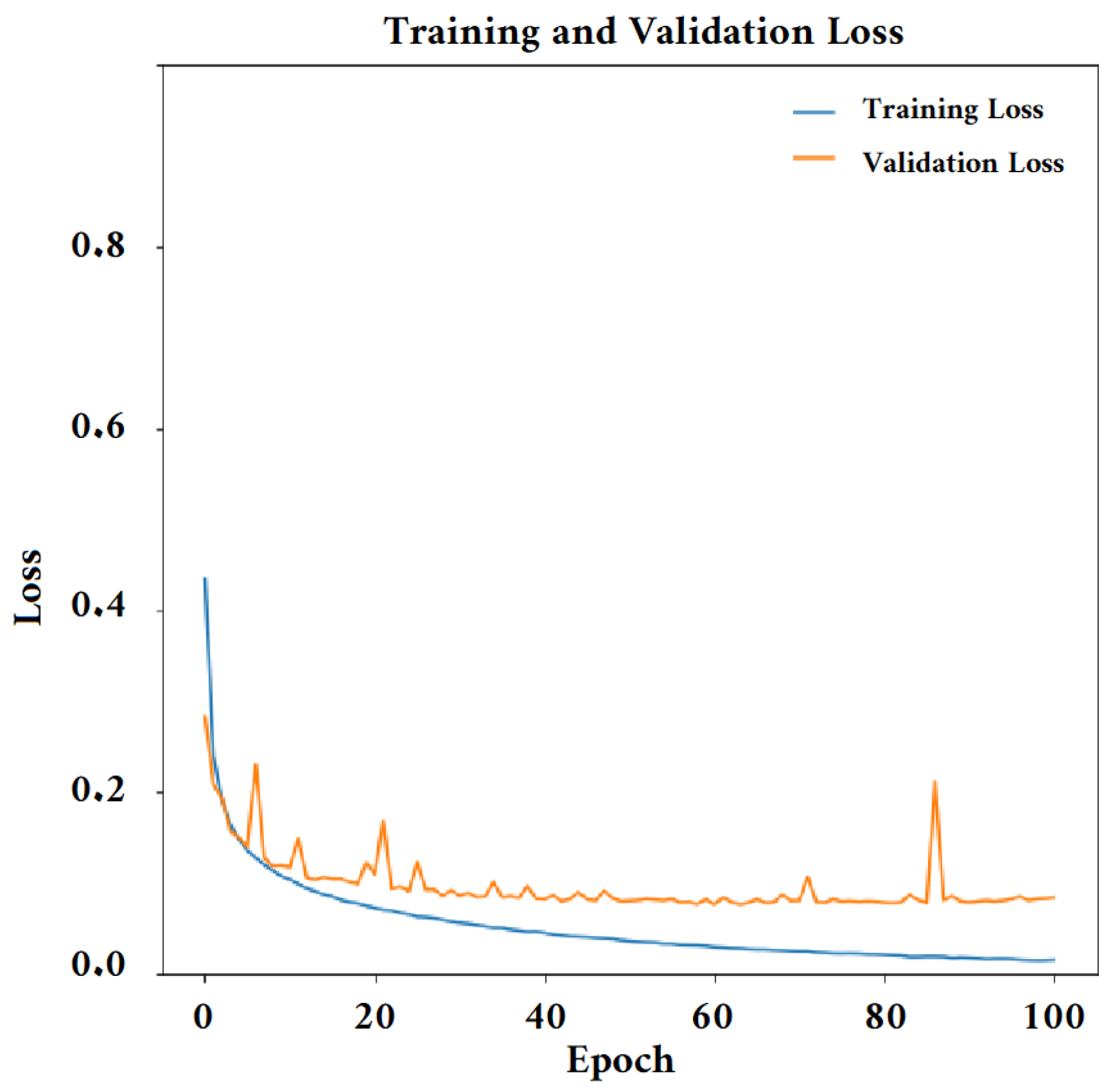
4.4.2. Training the Convolutional Neural Network Model
4.5. Performance Metrics for Classification
5. Results
- In the TFR_CNN1 approach, we initially transformed each tw into a time–frequency Representation Image (TFRI) utilizing the Pseudo Wigner–Ville transform, without using the Hilbert transform (Ht). The resulting image was then converted into a feature vector, which served as input for the Fully Connected Layer (FCL) of the classifier.
- In the Ht_TFR_CNN1 method, information extraction involved applying the Hilbert transform to each window’s tw obtained in the first phase, followed by the assessment of the Time–Frequency Representation (TFR) matrix using the Pseudo Wigner–Ville transform. The resulting TFR matrix was used to generate the TFRI, which was then used as input for the FCL.
- In the Ht_TFR_CNN2 method, the parameters were extracted using CNN2 by combining the Hilbert transform (Ht) and the TFRI. The extracted vectors were then used as input for the FCL.
Analysis Based on Different CNN Algorithms
6. Discussion
7. Application in a Real Clinical Setting
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Doolan, A.; Semsarian, C.; Langlois, N. Causes of sudden cardiac death in young Australians. Med. J. Aust. 2004, 180, 110–112. [Google Scholar] [CrossRef] [PubMed]
- Beck, C.S.; Pritchard, W.H.; Feil, H.S. Ventricular Fibrillation of Long Duration Abolished by Electric Shock. J. Am. Med. Assoc. 1947, 135, 985–986. [Google Scholar] [CrossRef] [PubMed]
- Kerber, R.E.; Becker, L.B.; Bourland, J.D.; Cummins, R.O.; Hallstrom, A.P.; Michos, M.B.; Nichol, G.; Ornato, J.P.; Thies, W.H.; White, R.D.; et al. Automatic external defibrillators for public access defibrillation: Recommendations for specifying and reporting arrhythmia analysis algorithm performance, incorporating new waveforms, and enhancing safety: A statement for health professionals from the American Heart Association Task Force on Automatic External Defibrillation, Subcommittee on AED Safety and Efficacy. Circulation 1997, 95, 1677–1682. [Google Scholar] [PubMed]
- Jin, D.; Dai, C.; Gong, Y.; Lu, Y.; Zhang, L.; Quan, W.; Li, Y. Does the choice of definition for defibrillation and CPR success impact the predictability of ventricular fibrillation waveform analysis? Resuscitation 2017, 111, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Amann, A.; Tratnig, R.; Unterkofler, K. Reliability of old and new ventricular fibrillation detection algorithms for automated external defibrillators. Biomed. Eng. Online 2005, 4, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Pourmand, A.; Galvis, J.; Yamane, D. The controversial role of dual sequential defibrillation in shockable cardiac arrest. Am. J. Emerg. Med. 2018, 36, 1674–1679. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Chung, K.Y.; Lim, J.S. Detection of ventricular fibrillation using Hilbert transforms, phase-space reconstruction, and time-domain analysis. Pers. Ubiquitous Comput. 2014, 18, 1315–1324. [Google Scholar] [CrossRef]
- Othman, M.A.; Safri, N.M.; Ghani, I.A.; Harun, F.K.C. Characterization of ventricular tachycardia and fibrillation using semantic mining. Comput. Inf. Sci. 2012, 5, 35. [Google Scholar] [CrossRef]
- Shyu, L.Y.; Wu, Y.H.; Hu, W. Using wavelet transform and fuzzy neural network for VPC detection from the Holter ECG. IEEE Trans. Biomed. Eng. 2004, 51, 1269–1273. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.S. Finding features for real-time premature ventricular contraction detection using a fuzzy neural network system. IEEE Trans. Neural Netw. 2009, 20, 522–527. [Google Scholar] [CrossRef] [PubMed]
- Rosado-Munoz, A.; Martínez-Martínez, J.M.; Escandell-Montero, P.; Soria-Olivas, E. Visual data mining with self-organising maps for ventricular fibrillation analysis. Comput. Methods Programs Biomed. 2013, 111, 269–279. [Google Scholar] [CrossRef]
- Orozco-Duque, A.; Rúa, S.; Zuluaga, S.; Redondo, A.; Restrepo, J.V.; Bustamante, J. Support Vector Machine and Artificial Neural Network Implementation in Embedded Systems for Real Time Arrhythmias Detection. In Proceedings of the International Conference on Bio-Inspired Systems and Signal Processing, Barcelona, Spain, 11–14 February 2013; pp. 310–313. [Google Scholar]
- Pooyan, M.; Akhoondi, F. Providing an efficient algorithm for finding R peaks in ECG signals and detecting ventricular abnormalities with morphological features. J. Med. Signals Sens. 2016, 6, 218. [Google Scholar] [CrossRef]
- Tripathy, R.; Sharma, L.; Dandapat, S. Detection of shockable ventricular arrhythmia using variational mode decomposition. J. Med. Syst. 2016, 40, 1–13. [Google Scholar] [CrossRef]
- Jekova, I.; Krasteva, V. Real time detection of ventricular fibrillation and tachycardia. Physiol. Meas. 2004, 25, 1167. [Google Scholar] [CrossRef] [PubMed]
- Mohanty, M.; Sahoo, S.; Biswal, P.; Sabut, S. Efficient classification of ventricular arrhythmias using feature selection and C4.5 classifier. Biomed. Signal Process. Control 2018, 44, 200–208. [Google Scholar] [CrossRef]
- Jothiramalingam, R.; Jude, A.; Patan, R.; Ramachandran, M.; Duraisamy, J.H.; Gandomi, A.H. Machine learning-based left ventricular hypertrophy detection using multi-lead ECG signal. Neural Comput. Appl. 2021, 33, 4445–4455. [Google Scholar] [CrossRef]
- Tang, J.; Li, J.; Liang, B.; Huang, X.; Li, Y.; Wang, K. Using Bayesian decision for ontology mapping. J. Web Semant. 2006, 4, 243–262. [Google Scholar] [CrossRef]
- Kuzilek, J.; Kremen, V.; Soucek, F.; Lhotska, L. Independent component analysis and decision trees for ECG holter recording de-noising. PLoS ONE 2014, 9, e98450. [Google Scholar] [CrossRef] [PubMed]
- Ayachi, R.; Said, Y.E.; Atri, M. To perform road signs recognition for autonomous vehicles using cascaded deep learning pipeline. Artif. Intell. Adv. 2019, 1, 1–10. [Google Scholar] [CrossRef]
- Afif, M.; Ayachi, R.; Said, Y.; Pissaloux, E.; Atri, M. Indoor Image Recognition and Classification via Deep Convolutional Neural Network; Springer: Berlin/Heidelberg, Germany, 2020; pp. 364–371. [Google Scholar]
- Afif, M.; Ayachi, R.; Said, Y.; Pissaloux, E.; Atri, M. An evaluation of retinanet on indoor object detection for blind and visually impaired persons assistance navigation. Neural Process. Lett. 2020, 51, 2265–2279. [Google Scholar] [CrossRef]
- Virmani, D.; Girdhar, P.; Jain, P.; Bamdev, P. FDREnet: Face detection and recognition pipeline. Eng. Technol. Appl. Sci. Res. 2019, 9, 3933–3938. [Google Scholar] [CrossRef]
- Khan, U.; Khan, K.; Hassan, F.; Siddiqui, A.; Afaq, M. Towards achieving machine comprehension using deep learning on non-GPU machines. Eng. Technol. Appl. Sci. Res. 2019, 9, 4423–4427. [Google Scholar] [CrossRef]
- Moon, H.M.; Seo, C.H.; Pan, S.B. A face recognition system based on convolution neural network using multiple distance face. Soft Comput. 2017, 21, 4995–5002. [Google Scholar] [CrossRef]
- Khalajzadeh, H.; Mansouri, M.; Teshnehlab, M. Face recognition using convolutional neural network and simple logistic classifier. In Proceedings of the Soft Computing in Industrial Applications: Proceedings of the 17th Online World Conference on Soft Computing in Industrial Applications; Springer: Berlin/Heidelberg, Germany, 2014; pp. 197–207. [Google Scholar]
- Yale Face Database. Available online: http://vision.ucsd.edu/content/yale-face-database (accessed on 14 June 2023).
- Yan, K.; Huang, S.; Song, Y.; Liu, W.; Fan, N. Face recognition based on convolution neural network. In Proceedings of the IEEE 2017 36th Chinese Control Conference (CCC), Dalian, China, 26–28 July 2017; pp. 4077–4081. [Google Scholar]
- AT&T Database of Faces: ORL Face Database. Available online: http://cam-orl.co.uk/facedatabase.html (accessed on 14 June 2023).
- Martinez, A.; Benavente, R. The AR Face Database; Technical Report Series; Report #24; CVC Tech: Fontana, CA, USA, 1998. [Google Scholar]
- Li, L.; Jun, Z.; Fei, J.; Li, S. An incremental face recognition system based on deep learning. In Proceedings of the IEEE 2017 15th IAPR International Conference on Machine Vision Applications (MVA), Nagoya, Japan, 8–12 May 2017; pp. 238–241. [Google Scholar]
- Schroff, F.; Kalenichenko, D.; Philbin, J. Facenet: A unified embedding for face recognition and clustering. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 815–823. [Google Scholar]
- Nakada, M.; Wang, H.; Terzopoulos, D. AcFR: Active face recognition using convolutional neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition Workshops, Honolulu, HI, USA, 21–26 July 2017; pp. 35–40. [Google Scholar]
- Gross, R.; Matthews, I.; Cohn, J.; Kanade, T.; Baker, S. Multi-pie. Image Vis. Comput. 2010, 28, 807–813. [Google Scholar] [CrossRef]
- Li, J.; Qiu, T.; Wen, C.; Xie, K.; Wen, F.Q. Robust face recognition using the deep C2D-CNN model based on decision-level fusion. Sensors 2018, 18, 2080. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Tan, L.; Jiang, H. Review on convolutional neural network (CNN) applied to plant leaf disease classification. Agriculture 2021, 11, 707. [Google Scholar] [CrossRef]
- Guan, S.; Kamona, N.; Loew, M. Segmentation of thermal breast images using convolutional and deconvolutional neural networks. In Proceedings of the 2018 IEEE Applied Imagery Pattern Recognition Workshop (AIPR), Washington, DC, USA, 9–11 October 2018; pp. 1–7. [Google Scholar]
- Rahman, T.; Chowdhury, M.E.; Khandakar, A.; Islam, K.R.; Islam, K.F.; Mahbub, Z.B.; Kadir, M.A.; Kashem, S. Transfer learning with deep convolutional neural network (CNN) for pneumonia detection using chest X-ray. Appl. Sci. 2020, 10, 3233. [Google Scholar] [CrossRef]
- Desai, M.; Shah, M. An anatomization on breast cancer detection and diagnosis employing multi-layer perceptron neural network (MLP) and Convolutional neural network (CNN). Clin. eHealth 2021, 4, 1–11. [Google Scholar] [CrossRef]
- Budak, Ü.; Cömert, Z.; Çıbuk, M.; Şengür, A. DCCMED-Net: Densely connected and concatenated multi Encoder-Decoder CNNs for retinal vessel extraction from fundus images. Med. Hypotheses 2020, 134, 109426. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.P.; Choi, S.; Park, S.J.; Park, S.H.; Yoon, J. Inspecting method for defective casting products with convolutional neural network (CNN). Int. J. Precis. Eng. Manuf.-Green Technol. 2021, 8, 583–594. [Google Scholar] [CrossRef]
- Victoria, A.H.; Maragatham, G. Automatic tuning of hyperparameters using Bayesian optimization. Evol. Syst. 2021, 12, 217–223. [Google Scholar] [CrossRef]
- Young, S.R.; Rose, D.C.; Karnowski, T.P.; Lim, S.H.; Patton, R.M. Optimizing deep learning hyper-parameters through an evolutionary algorithm. In Proceedings of the Workshop on Machine Learning in High-Performance Computing Environments, Austin, TX, USA, 15 November 2015; pp. 1–5. [Google Scholar]
- Cui, H.; Bai, J. A new hyperparameters optimization method for convolutional neural networks. Pattern Recognit. Lett. 2019, 125, 828–834. [Google Scholar] [CrossRef]
- Lee, W.Y.; Park, S.M.; Sim, K.B. Optimal hyperparameter tuning of convolutional neural networks based on the parameter-setting-free harmony search algorithm. Optik 2018, 172, 359–367. [Google Scholar] [CrossRef]
- Kiliçarslan, S.; Celik, M. RSigELU: A nonlinear activation function for deep neural networks. Expert Syst. Appl. 2021, 174, 114805. [Google Scholar] [CrossRef]
- Zou, X.; Wang, Z.; Li, Q.; Sheng, W. Integration of residual network and convolutional neural network along with various activation functions and global pooling for time series classification. Neurocomputing 2019, 367, 39–45. [Google Scholar] [CrossRef]
- Basha, S.S.; Vinakota, S.K.; Dubey, S.R.; Pulabaigari, V.; Mukherjee, S. Autofcl: Automatically tuning fully connected layers for handling small dataset. Neural Comput. Appl. 2021, 33, 8055–8065. [Google Scholar] [CrossRef]
- Dahl, G.E.; Sainath, T.N.; Hinton, G.E. Improving deep neural networks for LVCSR using rectified linear units and dropout. In Proceedings of the 2013 IEEE International Conference on Acoustics, Speech and Signal Processing, Vancouver, BC, Canada, 26–31 May 2013; pp. 8609–8613. [Google Scholar]
- Radiuk, P.M. Impact of Training Set Batch Size on the Performance of Convolutional Neural Networks for Diverse Datasets; Information Technology and Management Science: Riga, Latvia, 2017. [Google Scholar]
- Utama, A.B.P.; Wibawa, A.P.; Muladi, M.; Nafalski, A. PSO based Hyperparameter tuning of CNN Multivariate Time-Series Analysis. J. Online Inform. 2022, 7, 193–202. [Google Scholar] [CrossRef]
- Zeiler, M.D. Adadelta: An adaptive learning rate method. arXiv 2012, arXiv:1212.5701. [Google Scholar]
- Gulcehre, C.; Moczulski, M.; Bengio, Y. Adasecant: Robust adaptive secant method for stochastic gradient. arXiv 2014, arXiv:1412.7419. [Google Scholar]
- Dhar, P.; Dutta, S.; Mukherjee, V. Cross-wavelet assisted convolution neural network (AlexNet) approach for phonocardiogram signals classification. Biomed. Signal Process. Control 2021, 63, 102142. [Google Scholar] [CrossRef]
- Anand, R.; Sowmya, V.; Gopalakrishnan, E.; Soman, K. Modified Vgg deep learning architecture for Covid-19 classification using bio-medical images. Iop Conf. Ser. Mater. Sci. Eng. 2021, 1084, 012001. [Google Scholar] [CrossRef]
- Vijayan, T.; Sangeetha, M.; Karthik, B. Efficient analysis of diabetic retinopathy on retinal fundus images using deep learning techniques with inception v3 architecture. J. Green Eng. 2020, 10, 9615–9625. [Google Scholar]
- Gómez, J.C.V.; Incalla, A.P.Z.; Perca, J.C.C.; Padilla, D.I.M. Diferentes Configuraciones Para MobileNet en la Detección de Tumores Cerebrales: Different Configurations for MobileNet in the Detection of Brain Tumors. In Proceedings of the 2021 IEEE 1st International Conference on Advanced Learning Technologies on Education & Research, Lima, Peru, 16–18 December 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 1–4. [Google Scholar]
- Mjahad, A.; Rosado-Muñoz, A.; Bataller-Mompeán, M.; Francés-Víllora, J.; Guerrero-Martínez, J. Ventricular Fibrillation and Tachycardia detection from surface ECG using time–frequency representation images as input dataset for machine learning. Comput. Methods Programs Biomed. 2017, 141, 119–127. [Google Scholar] [CrossRef] [PubMed]
- PhysioNet. Available online: http://physionet.org (accessed on 14 June 2023).
- American Heart Association ECG Database. Available online: http://ecri.org (accessed on 14 June 2023).
- Kaur, M.; Singh, B. Comparison of different approaches for removal of baseline wander from ECG signal. In Proceedings of the International Conference & Workshop on Emerging Trends in Technology, Mumbai, India, 25–26 February 2011; pp. 1290–1294. [Google Scholar]
- Narwaria, R.P.; Verma, S.; Singhal, P. Removal of baseline wander and power line interference from ECG signal-a survey approach. Int. J. Electron. Eng. 2011, 3, 107–111. [Google Scholar]
- Ramezan, C.A.; Warner, T.A.; Maxwell, A.E. Evaluation of Sampling and Cross-Validation Tuning Strategies for Regional-Scale Machine Learning Classification. Remote. Sens. 2019, 11, 185. [Google Scholar] [CrossRef]
- Labatut, V.; Cherifi, H. Accuracy measures for the comparison of classifiers. arXiv 2012, arXiv:1207.3790. [Google Scholar]
- Rebouças Filho, P.P.; Peixoto, S.A.; da Nóbrega, R.V.M.; Hemanth, D.J.; Medeiros, A.G.; Sangaiah, A.K.; de Albuquerque, V.H.C. Automatic histologically-closer classification of skin lesions. Comput. Med Imaging Graph. 2018, 68, 40–54. [Google Scholar] [CrossRef]
- Arafat, M.A.; Chowdhury, A.W.; Hasan, M.K. A simple time domain algorithm for the detection of ventricular fibrillation in electrocardiogram. Signal Image Video Process. 2011, 5, 1–10. [Google Scholar] [CrossRef]
- Roopaei, M.; Boostani, R.; Sarvestani, R.R.; Taghavi, M.A.; Azimifar, Z. Chaotic based reconstructed phase space features for detecting ventricular fibrillation. Biomed. Signal Process. Control 2010, 5, 318–327. [Google Scholar] [CrossRef]
- Alonso-Atienza, F.; Morgado, E.; Fernandez-Martinez, L.; Garcia-Alberola, A.; Rojo-Alvarez, J.L. Detection of life-threatening arrhythmias using feature selection and support vector machines. IEEE Trans. Biomed. Eng. 2013, 61, 832–840. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Rajagopalan, C.; Clifford, G.D. Ventricular fibrillation and tachycardia classification using a machine learning approach. IEEE Trans. Biomed. Eng. 2013, 61, 1607–1613. [Google Scholar] [PubMed]
- Ibtehaz, N.; Rahman, M.S.; Rahman, M.S. VFPred: A fusion of signal processing and machine learning techniques in detecting ventricular fibrillation from ECG signals. Biomed. Signal Process. Control 2019, 49, 349–359. [Google Scholar] [CrossRef]
- Acharya, U.R.; Fujita, H.; Lih, O.S.; Hagiwara, Y.; Tan, J.H.; Adam, M. Automated detection of arrhythmias using different intervals of tachycardia ECG segments with convolutional neural network. Inf. Sci. 2017, 405, 81–90. [Google Scholar] [CrossRef]
- Xia, D.; Meng, Q.; Chen, Y.; Zhang, Z. Classification of ventricular tachycardia and fibrillation based on the lempel-ziv complexity and EMD. In Proceedings of the Intelligent Computing in Bioinformatics: 10th International Conference, ICIC 2014, Taiyuan, China, 3–6 August 2014; Springer: Berlin/Heidelberg, Germany, 2014. Proceedings 10. pp. 322–329. [Google Scholar]
- Mjahad, A.; Frances-Villora, J.V.; Bataller-Mompean, M.; Rosado-Muñoz, A. Ventricular Fibrillation and Tachycardia Detection Using Features Derived from Topological Data Analysis. Appl. Sci. 2022, 12, 7248. [Google Scholar] [CrossRef]
- Kaur, L.; Singh, V. Ventricular fibrillation detection using emprical mode decomposition and approximate entropy. Int. J. Emerg. Technol. Adv. Eng. 2013, 3, 260–268. [Google Scholar]
- Xie, H.B.; Zhong-Mei, G.; Liu, H. Classification of ventricular tachycardia and fibrillation using fuzzy similarity-based approximate entropy. Expert Syst. Appl. 2011, 38, 3973–3981. [Google Scholar] [CrossRef]
- Acharya, U.R.; Fujita, H.; Oh, S.L.; Raghavendra, U.; Tan, J.H.; Adam, M.; Gertych, A.; Hagiwara, Y. Automated identification of shockable and non-shockable life-threatening ventricular arrhythmias using convolutional neural network. Future Gener. Comput. Syst. 2018, 79, 952–959. [Google Scholar] [CrossRef]
- Buscema, P.M.; Grossi, E.; Massini, G.; Breda, M.; Della Torre, F. Computer aided diagnosis for atrial fibrillation based on new artificial adaptive systems. Comput. Methods Programs Biomed. 2020, 191, 105401. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Pachori, R.B.; Acharya, U.R. Automated diagnosis of atrial fibrillation ECG signals using entropy features extracted from flexible analytic wavelet transform. Biocybern. Biomed. Eng. 2018, 38, 564–573. [Google Scholar] [CrossRef]
- Cheng, P.; Dong, X. Life-threatening ventricular arrhythmia detection with personalized features. IEEE Access 2017, 5, 14195–14203. [Google Scholar] [CrossRef]
- Xu, Y.; Wang, D.; Zhang, W.; Ping, P.; Feng, L. Detection of ventricular tachycardia and fibrillation using adaptive variational mode decomposition and boosted-CART classifier. Biomed. Signal Process. Control 2018, 39, 219–229. [Google Scholar] [CrossRef]
- Brown, G.; Conway, S.; Ahmad, M.; Adegbie, D.; Patel, N.; Myneni, V.; Alradhawi, M.; Kumar, N.; Obaid, D.R.; Pimenta, D.; et al. Role of artificial intelligence in defibrillators: A narrative review. Open Heart 2022, 9, e001976. [Google Scholar] [CrossRef]
- Podbregar, M.; Kovačič, M.; Podbregar-Marš, A.; Brezocnik, M. Predicting defibrillation success by ‘genetic’ programming in patients with out-of-hospital cardiac arrest. Resuscitation 2003, 57, 153–159. [Google Scholar] [CrossRef] [PubMed]
















| Model | CNN1 | ||
|---|---|---|---|
| Layer | Kernel Size | Filter Number | #Parameters |
| FC1 | 512 | - | 16589312 |
| FC2 | 256 | - | 131328 |
| Softmax | 4 | - | 1285 |
| Model | CNN2 | ||
| Layer | Kernel Size | Filter Number | #Parameters |
| Conv1 | 3 × 3 | 32 | 320 |
| Max Pooling1 | 4 × 4 | - | 0 |
| Conv2 | 3 × 3 | 64 | 18496 |
| Max Pooling2 | 4 × 4 | - | 0 |
| FC1 | 128 | - | 991360 |
| FC2 | 256 | - | 33024 |
| Softmax | 4 | - | 1028 |
| Class | Normal | ||||||
|---|---|---|---|---|---|---|---|
| Algorithms | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| Normal | Global | VF | VT | Other | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 50) | 89.70 | 98.57 | 99.53 | 99.48 | 97.73 | 98.76 | 93.92 |
| Ht_TFR_CNN1 (Epochs = 100) | 99.29 | 98.62 | 98.88 | 99.33 | 98.03 | 98.91 | 98.95 |
| Ht_TFR_CNN2 (Epochs = 100) | 99.34 | 98.35 | 99.59 | 99.83 | 99.59 | 98.89 | 98.84 |
| TFR_CNN1 (Epochs = 50) | 98.70 | 98.59 | 99.46 | 98.73 | 97.73 | 98.65 | 98.64 |
| Class | Other | ||||||
|---|---|---|---|---|---|---|---|
| Algorithms | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| Other | Global | VT | Normal | VF | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 50) | 97.24 | 99.41 | 99.82 | 99.29 | 99.65 | 98.95 | 98.31 |
| Ht_TFR_CNN1 (Epochs = 100) | 97.74 | 99.62 | 99.83 | 99.60 | 99.58 | 99.22 | 98.67 |
| Ht_TFR_CNN2 (Epochs = 100) | 96.98 | 99.68 | 99.96 | 99.61 | 99.79 | 99.11 | 98.31 |
| TFR_CNN1 (Epochs = 50) | 97.24 | 99.47 | 100 | 99.33 | 99.73 | 98.98 | 98.34 |
| Class | VT | ||||||
|---|---|---|---|---|---|---|---|
| Algorithms | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| VT | Global | VF | Other | Normal | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 50) | 89.70 | 99.70 | 96.71 | 99.84 | 99.94 | 99.00 | 94.43 |
| Ht_TFR_CNN1 (Epochs = 100) | 92.70 | 99.53 | 97.78 | 99.94 | 99.92 | 99.06 | 95.99 |
| Ht_TFR_CNN2 (Epochs = 100) | 90.45 | 99.73 | 96.92 | 99.94 | 99.98 | 99.09 | 94.86 |
| TFR_CNN1 (Epochs = 50) | 95.84 | 97.19 | 98.55 | 99.84 | 99.84 | 97.90 | 96.51 |
| Class | VF | ||||||
|---|---|---|---|---|---|---|---|
| Algorithms | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| VF | Global | VT | Other | Normal | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 50) | 98.04 | 98.94 | 90.96 | 99.64 | 99.68 | 98.77 | 98.48 |
| Ht_TFR_CNN1 (Epochs = 100) | 96.44 | 99.28 | 94.01 | 99.74 | 99.76 | 98.75 | 97.83 |
| Ht_TFR_CNN2 (Epochs = 100) | 98.16 | 99.07 | 91.56 | 99.74 | 99.83 | 98.91 | 98.61 |
| TFR_CNN1 (Epochs = 50) | 85.88 | 99.30 | 96.58 | 99.64 | 99.52 | 96.82 | 92.10 |
| Class | Normal | ||||||
|---|---|---|---|---|---|---|---|
| Techniques | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| Normal | Global | VF | VT | Other | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 100) | 99.29 | 98.62 | 98.88 | 99.33 | 98.03 | 98.91 | 98.95 |
| Ht_TFR_CNN2 (Epochs = 100) | 99.34 | 98.35 | 99.59 | 99.83 | 99.59 | 98.89 | 98.84 |
| InceptionV3 (Epochs = 6) | 77.99 | 99.65 | 99.92 | 39.30 | 99.32 | 87.17 | 87.49 |
| MobilNet (Epochs = 6) | 79.42 | 99.44 | 99.08 | 99.36 | 99.64 | 88.39 | 88.30 |
| VGGnet (Epochs = 6) | 96.61 | 98.32 | 97.97 | 100 | 98.59 | 97.39 | 97.45 |
| AlexNet (Epochs = 6) | 99.43 | 97.29 | 98.69 | 100 | 95.83 | 98.45 | 98.34 |
| Class | Other | ||||||
|---|---|---|---|---|---|---|---|
| Techniques | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| Other | Global | VT | Normal | VF | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 100) | 97.74 | 99.62 | 99.83 | 99.60 | 99.58 | 99.22 | 98.67 |
| Ht_TFR_CNN2 (Epochs = 100) | 96.98 | 99.68 | 99.96 | 99.61 | 99.79 | 99.11 | 98.31 |
| InceptionV3 (Epochs = 6) | 88.42 | 99.81 | 100 | 99.72 | 100 | 96.96 | 93.77 |
| MobilNet (Epochs = 6) | 99.64 | 85.08 | 98.41 | 79.60 | 97.68 | 88.21 | 91.78 |
| VGGnet (Epochs = 6) | 98.54 | 97.57 | 100 | 96.74 | 99.26 | 97.39 | 98.05 |
| AlexNet (Epochs = 6) | 95.78 | 99.57 | 100 | 99.58 | 99.40 | 98.77 | 97.63 |
| Class | VT | ||||||
|---|---|---|---|---|---|---|---|
| Techniques | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| VT | Global | VF | Other | Normal | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 100) | 92.70 | 99.53 | 97.78 | 99.94 | 99.92 | 99.06 | 95.99 |
| HT_TFR_CNN2 (Epochs = 100) | 90.45 | 99.73 | 96.92 | 99.94 | 99.98 | 99.09 | 94.86 |
| InceptionV3 (Epochs = 6) | 98.15 | 83.55 | 99.11 | 99.04 | 80.18 | 84.59 | 90.26 |
| MobilNet (Epochs = 6) | 95.53 | 97.66 | 98.89 | 100 | 99.90 | 97.49 | 96.58 |
| VGGnet (Epochs = 6) | 90.15 | 99.15 | 97.07 | 99.94 | 100 | 98.77 | 94.43 |
| AlexNet (Epochs = 6) | 91.84 | 99.47 | 97.54 | 99.94 | 100 | 98.94 | 95.50 |
| Class | VF | ||||||
|---|---|---|---|---|---|---|---|
| Techniques | Sensitivity (%) | Specificity (%) | Accuracy (%) | F Score (%) | |||
| VF | Global | VT | Other | Normal | Total | Total | |
| Ht_TFR_CNN1 (Epochs = 100) | 96.44 | 99.28 | 94.01 | 99.74 | 99.76 | 98.75 | 97.83 |
| Ht_TFR_CNN2 (Epochs = 100) | 98.16 | 99.07 | 91.56 | 99.74 | 99.83 | 98.91 | 98.61 |
| InceptionV3 (Epochs = 6) | 77.28 | 94.90 | 98.15 | 89.72 | 96.86 | 91.28 | 85.18 |
| MobilNet (Epochs = 6) | 86.97 | 99.62 | 97.33 | 100 | 99.80 | 97.01 | 92.86 |
| VGGnet (Epochs = 6) | 93.34 | 99.25 | 92.28 | 100 | 99.85 | 98.14 | 96.20 |
| AlexNet (Epochs = 6) | 95.58 | 99.34 | 93.42 | 100 | 99.84 | 98.64 | 97.42 |
| Class | VF | VT | Other | Normal | Data Base | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Techniques | Sens (%) | Spe (%) | Acc (%) | Sens (%) | Spe (%) | Acc (%) | Sens (%) | Spe (%) | Acc (%) | Sens (%) | Spe (%) | Acc (%) | |
| This work, Ht_TFR_CNN1 (Epochs = 50) | 98.04 | 98.94 | 98.77 | 89.7 | 99.70 | 99 | 97.24 | 99.41 | 98.95 | 89.7 | 98.57 | 98.76 | MITBIH, AHA |
| This work, Ht_TFR_CNN1 (Epochs = 100) | 96.44 | 99.28 | 98.75 | 92.70 | 99.53 | 99.06 | 97.74 | 99.62 | 99.22 | 99.29 | 98.62 | 98.91 | MITBIH, AHA |
| This work, Ht_TFR_CNN2 (Epochs = 100) | 98.16 | 99.07 | 98.91 | 90.45 | 99.73 | 99.09 | 96.98 | 99.68 | 99.11 | 99.34 | 98.35 | 98.89 | MITBIH, AHA |
| This work, InceptionV3 (Epochs = 6) | 77.28 | 94.9 | 91.28 | 98.15 | 83.55 | 84.59 | 88.42 | 99.81 | 96.96 | 77.99 | 99.65 | 87.17 | MITBIH, AHA |
| This work, MobilNet (Epochs = 6) | 86.97 | 99.62 | 97.01 | 95.53 | 97.66 | 97.49 | 99.64 | 85.08 | 88.21 | 79.42 | 99.44 | 88.39 | MITBIH, AHA |
| This work, VGGnet (Epochs = 6) | 93.34 | 99.25 | 98.14 | 90.15 | 99.15 | 98.77 | 98.54 | 97.57 | 97.39 | 96.61 | 98.32 | 97.39 | MITBIH, AHA |
| This work, AlexNet (Epochs = 6) | 95.58 | 99.34 | 98.64 | 91.84 | 99.47 | 98.94 | 95.78 | 99.57 | 98.77 | 99.43 | 97.29 | 98.45 | MITBIH, AHA |
| [58] SSVR, TFR | 91 | 97 | 92.8 | 98.7 | 92.3 | 99.2 | 96.6 | 96.3 | MITBIH, AHA | ||||
| [58] BAGG, TFR | 95.2 | 96.4 | 88.8 | 99.7 | 88.6 | 99.8 | 96.6 | 94.1 | MITBIH, AHA | ||||
| [58] I2-RLR and TFR | 89.6 | 96.7 | 91 | 98.1 | 92.5 | 98.1 | 94.9 | 96.4 | MITBIH, AHA | ||||
| [58] ANNC and TFR | 92.8 | 97 | 91.8 | 98.7 | 92.9 | 99 | 96.2 | 96.7 | MITBIH, AHA | ||||
| [66] TCSC algorithm | 80.97 | 98.51 | 98.14 | MITBIH, CUDB | |||||||||
| [67] Chaotic based | 88.6 | MITBIH, CCU | |||||||||||
| [68] SVM and FS | 91.9 | 97.1 | 96.8 | MITBIH, CUDB | |||||||||
| [69] SVM and Genetic algorithm | 98.4 | 98 | 96.3 | CUDB, AHA | |||||||||
| [70] SVM and EMD | 99.99 | 98.4 | 99.19 | MITBIH, CUDB | |||||||||
| [71] CNN neural network | 56.44 | 98.19 | 97.88 | MITBIH, CUDB | |||||||||
| [72] EMD and Lempel-Ziv | 98.15 | 96.01 | 96.01 | 98.15 | MITBIH, CUDB | ||||||||
| [73] TDA | 97.07 | 99.25 | 98.68 | 92.72 | 99.53 | 99.05 | 97.43 | 99.54 | 99.09 | 99.05 | 98.45 | 98.76 | MITBIH, AHA |
| [73] PDI | 84.34 | 96.77 | 94.26 | 82.25 | 98.53 | 97.38 | 92.86 | 97.15 | 96.19 | 93.09 | 92.14 | 92.65 | MITBIH, AHA |
| [74] App Entropy and EMD | 90.47 | 91.66 | 91.17 | 90.62 | 91.11 | 90.8 | MITBIH | ||||||
| [75] Approximated entropy | 97.98 | 97.03 | 97.03 | 97.98 | MITBIH, CUDB | ||||||||
| Class | Shockable (VT+VF) | Data Base | ||
|---|---|---|---|---|
| Technique | Sensitivity (%) | Specificity (%) | Accuracy (%) | |
| This work, Ht_TFR_CNN1 | 98.53 | 99.69 | 99.39 | MITBIH, AHA |
| This work, Ht_TFR_CNN2 | 99.23 | 99.74 | 99.61 | MITBIH, AHA |
| [73] TDA | 99.03 | 99.67 | 99.51 | MITBIH, AHA |
| [73] PDI | 89.63 | 96.96 | 95.12 | MITBIH, AHA |
| [76] CNN | 95.32 | 91.04 | 93.2 | MITDB, CUDB, VFDB |
| [14] VMD with Random Forest | 96.54 | 97.97 | 97.23 | MITBIH, CUDB |
| [77] RNN | 99.72 | MITBIH | ||
| [78] CNN and IENN | 98.6 | 98.9 | 98.8 | MITBIH, AFDB |
| [68] FS and SVM | 95 | 99 | 98.6 | MITBIH, CUDB |
| [79] Personalized features SVM | 95.6 | 95.5 | MITBIH, CUDB, VFDB | |
| [16] C4.5 classifier | 90.97 | 97.86 | 97.02 | MITBIH, CUDB |
| [69] SVM and bootstrap | 98.4 | 98 | 98.1 | MITBIH, AHA, CUDB |
| [80] Adaptive variational and boosted CART | 97.32 | 98.95 | 98.29 | MITBIH, CUDB |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mjahad, A.; Saban, M.; Azarmdel, H.; Rosado-Muñoz, A. Efficient Extraction of Deep Image Features Using a Convolutional Neural Network (CNN) for Detecting Ventricular Fibrillation and Tachycardia. J. Imaging 2023, 9, 190. https://doi.org/10.3390/jimaging9090190
Mjahad A, Saban M, Azarmdel H, Rosado-Muñoz A. Efficient Extraction of Deep Image Features Using a Convolutional Neural Network (CNN) for Detecting Ventricular Fibrillation and Tachycardia. Journal of Imaging. 2023; 9(9):190. https://doi.org/10.3390/jimaging9090190
Chicago/Turabian StyleMjahad, Azeddine, Mohamed Saban, Hossein Azarmdel, and Alfredo Rosado-Muñoz. 2023. "Efficient Extraction of Deep Image Features Using a Convolutional Neural Network (CNN) for Detecting Ventricular Fibrillation and Tachycardia" Journal of Imaging 9, no. 9: 190. https://doi.org/10.3390/jimaging9090190
APA StyleMjahad, A., Saban, M., Azarmdel, H., & Rosado-Muñoz, A. (2023). Efficient Extraction of Deep Image Features Using a Convolutional Neural Network (CNN) for Detecting Ventricular Fibrillation and Tachycardia. Journal of Imaging, 9(9), 190. https://doi.org/10.3390/jimaging9090190








