Low-Cost Probabilistic 3D Denoising with Applications for Ultra-Low-Radiation Computed Tomography
Abstract
1. Introduction
2. Materials and Methods
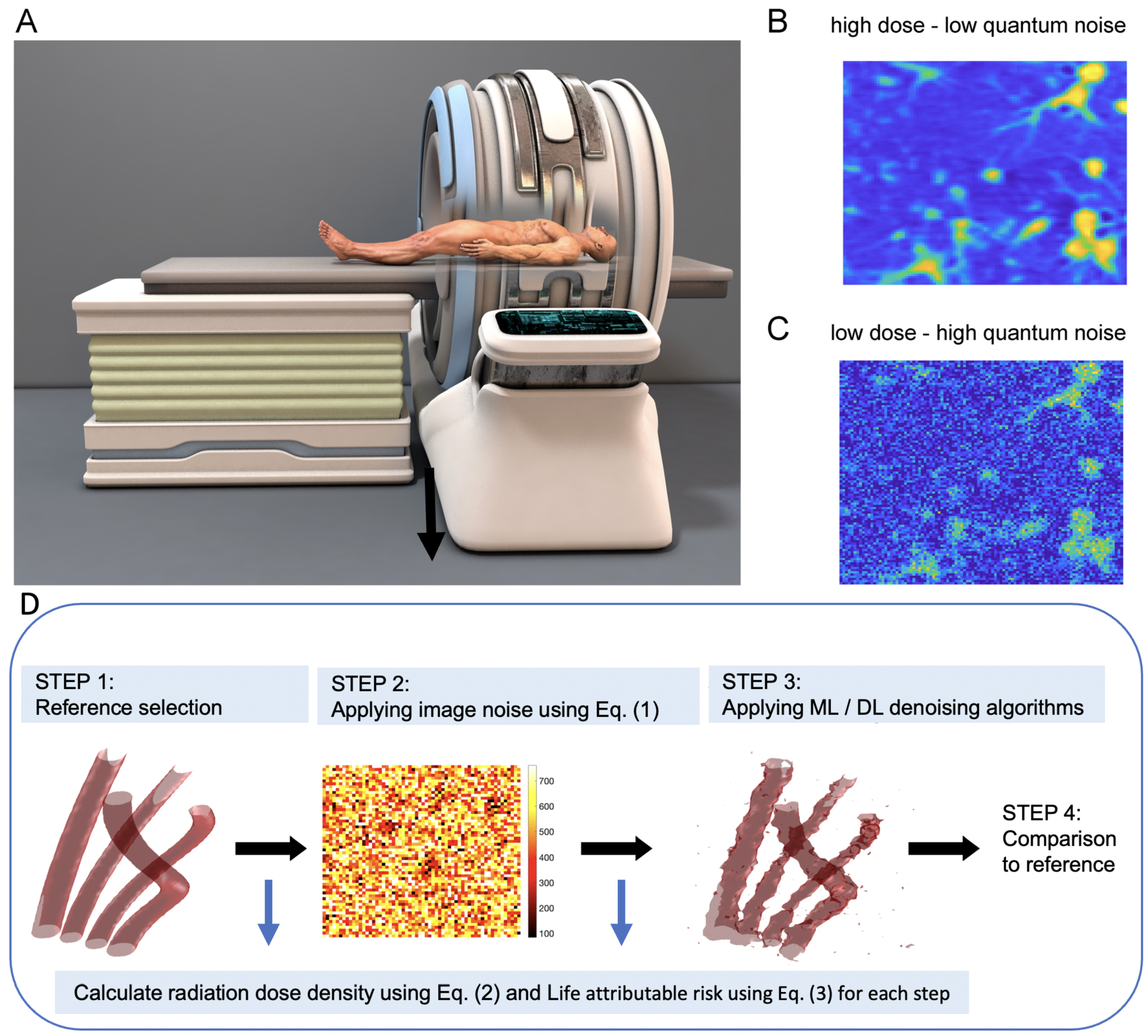
2.1. Patient-Specific Generation of Synthetic CT Images, Radiation Exposure Estimation, and LAR Computations
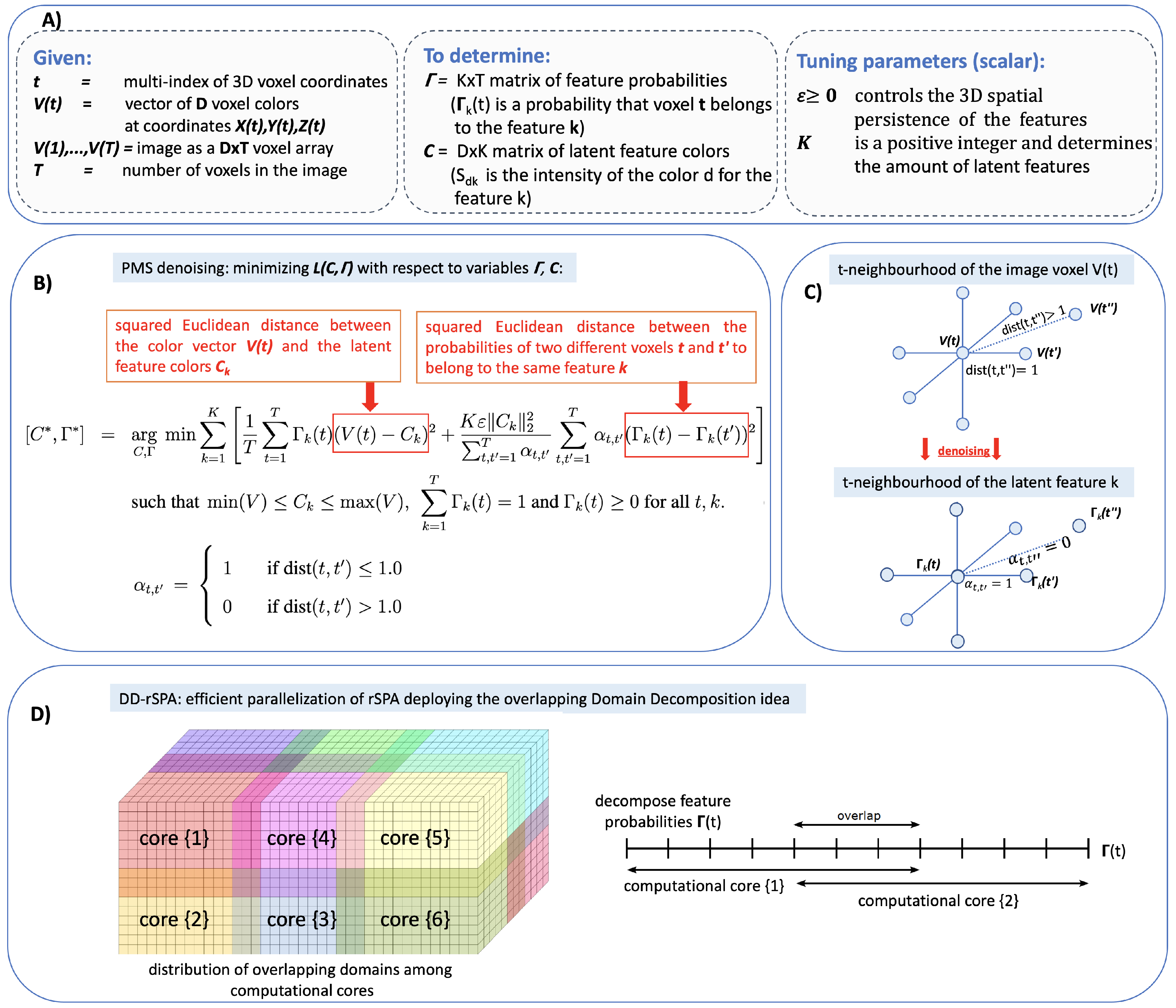
2.2. Probabilistic Mumford–Shah Model Formulation
2.3. Relation of Probabilistic Mumford–Shah and rSPA Algorithm to Regularized Mumford–Shah Framework (MS) and Rudin–Osher–Fatemi (ROF) Total Variation Model
2.4. Practical Implementation
2.4.1. Synthetic CT Image Generation Model
2.4.2. Common CT Image Denoising and Image Quality Assessment Methods
3. Results and Discussion
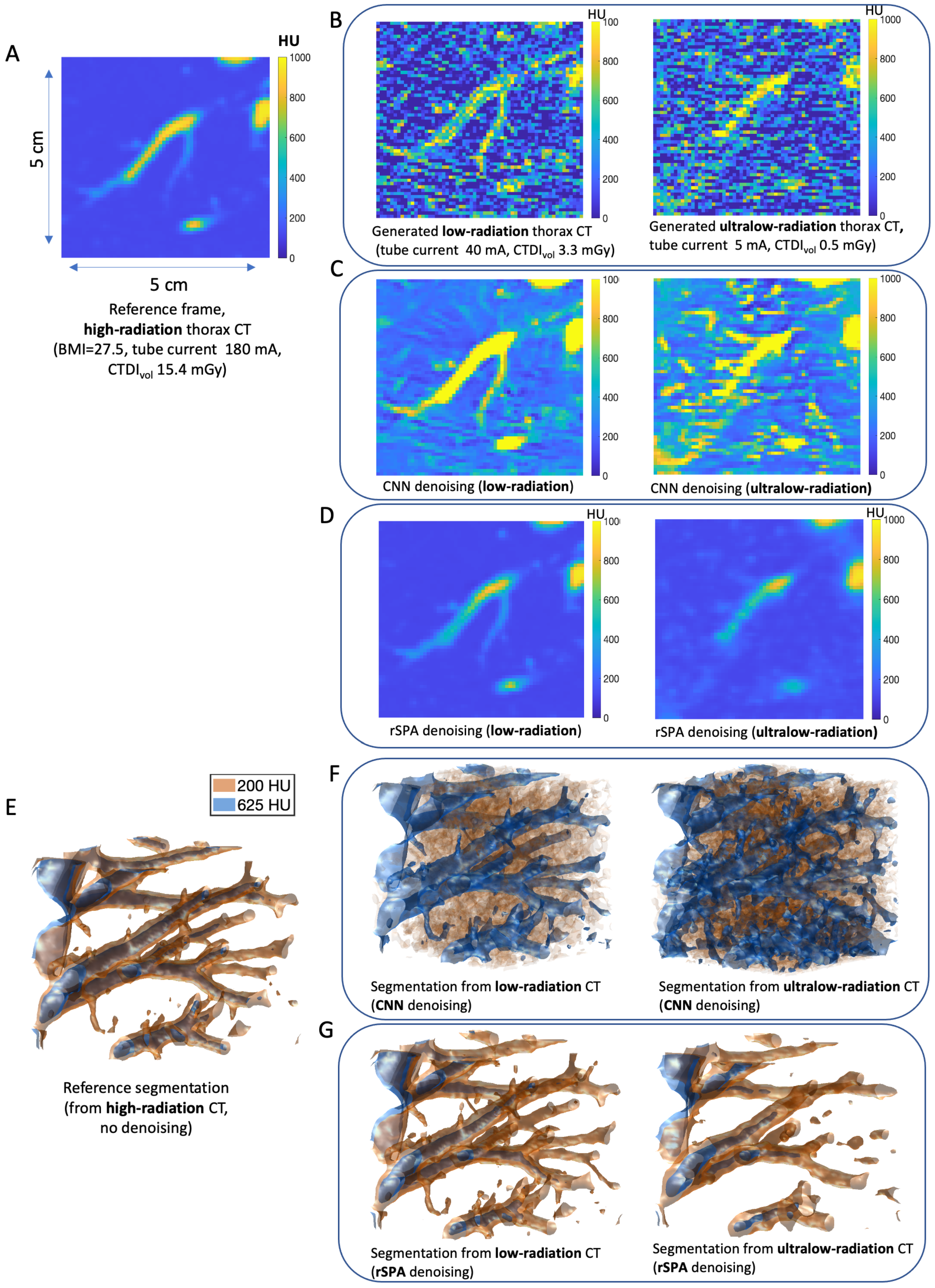
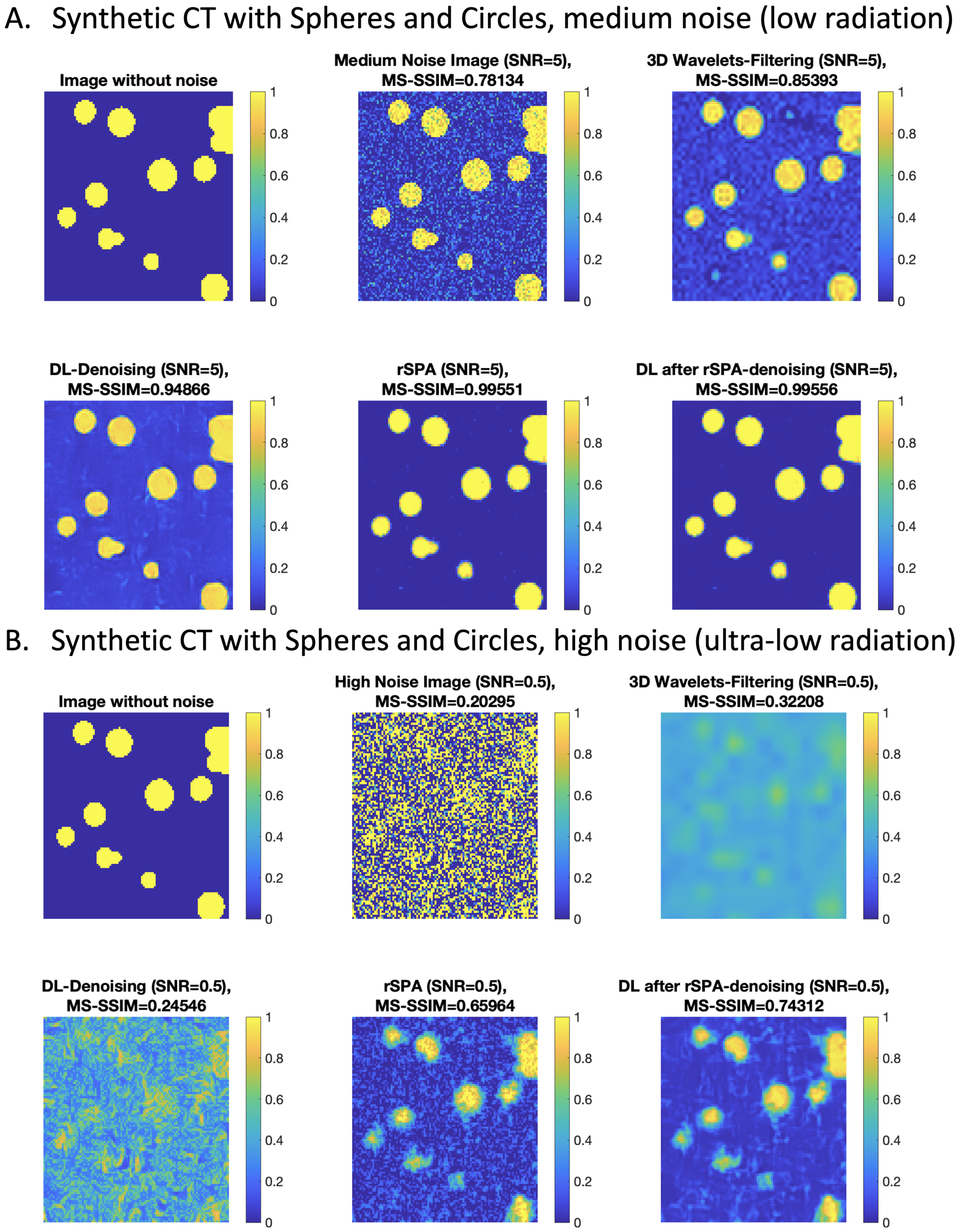
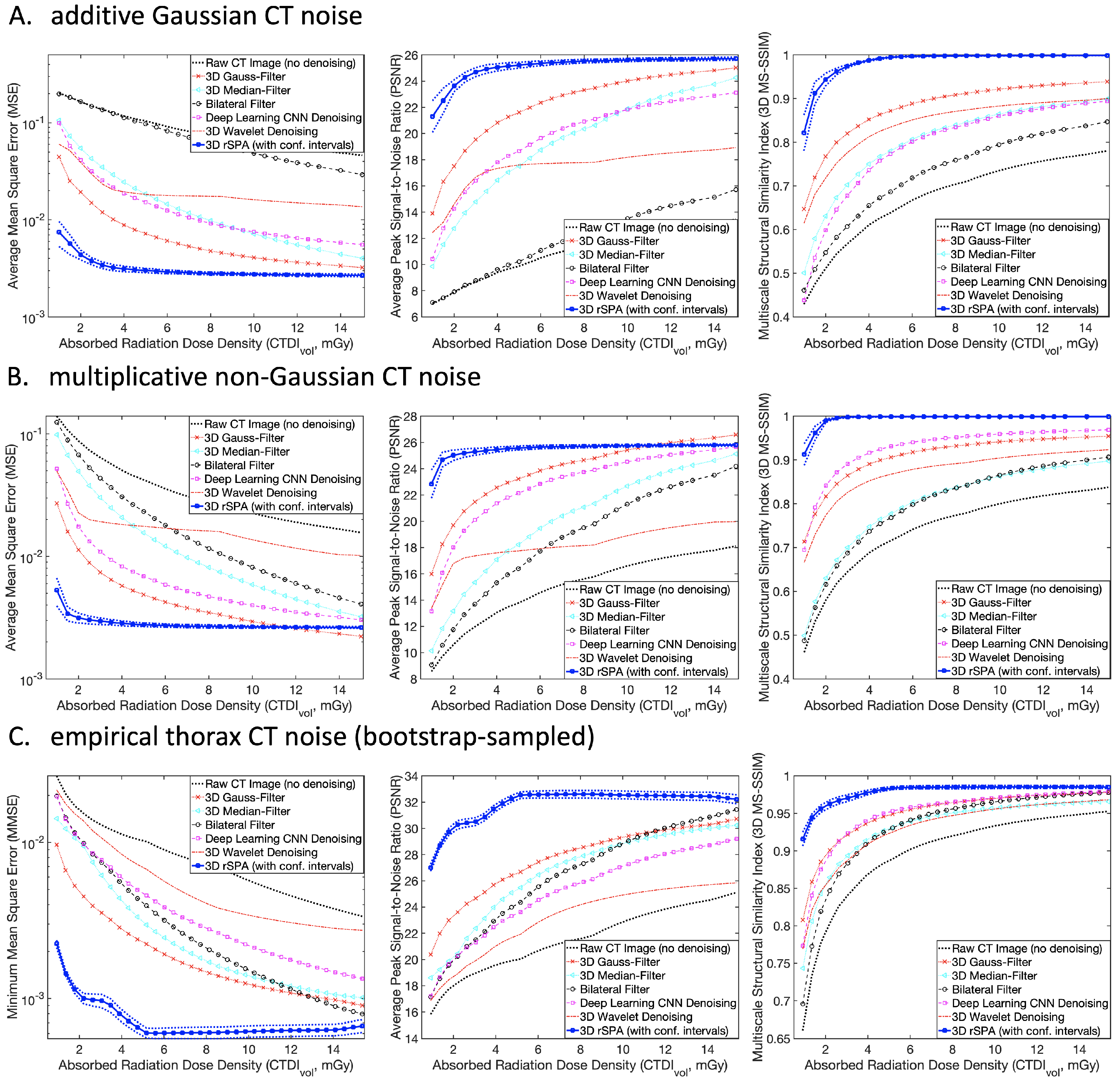
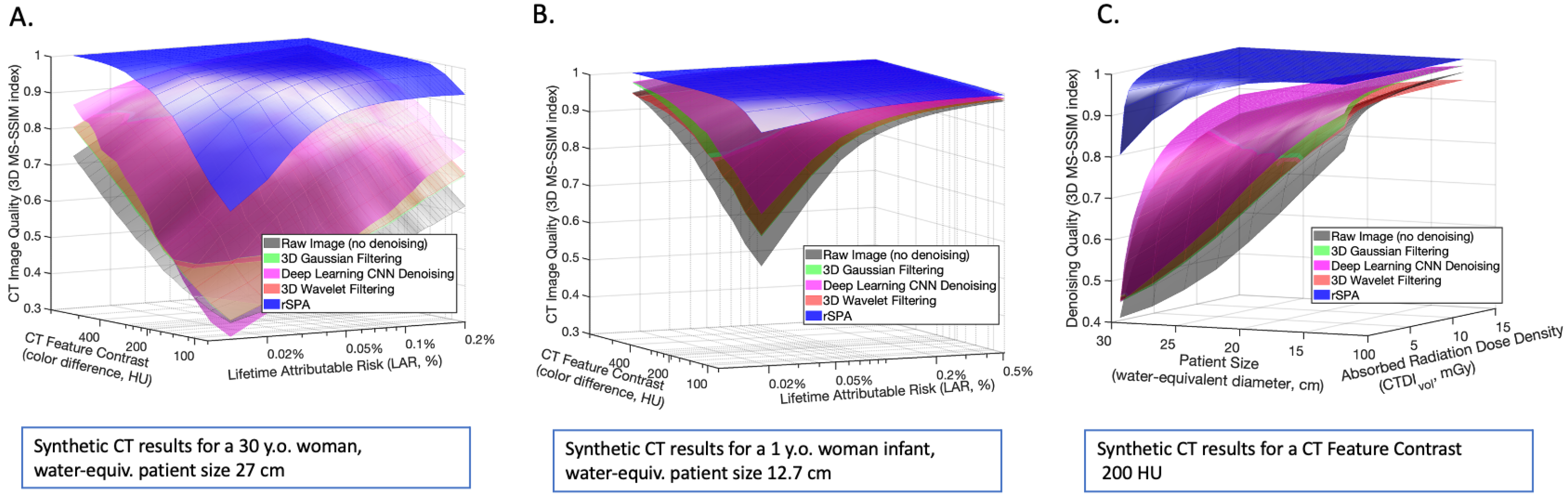
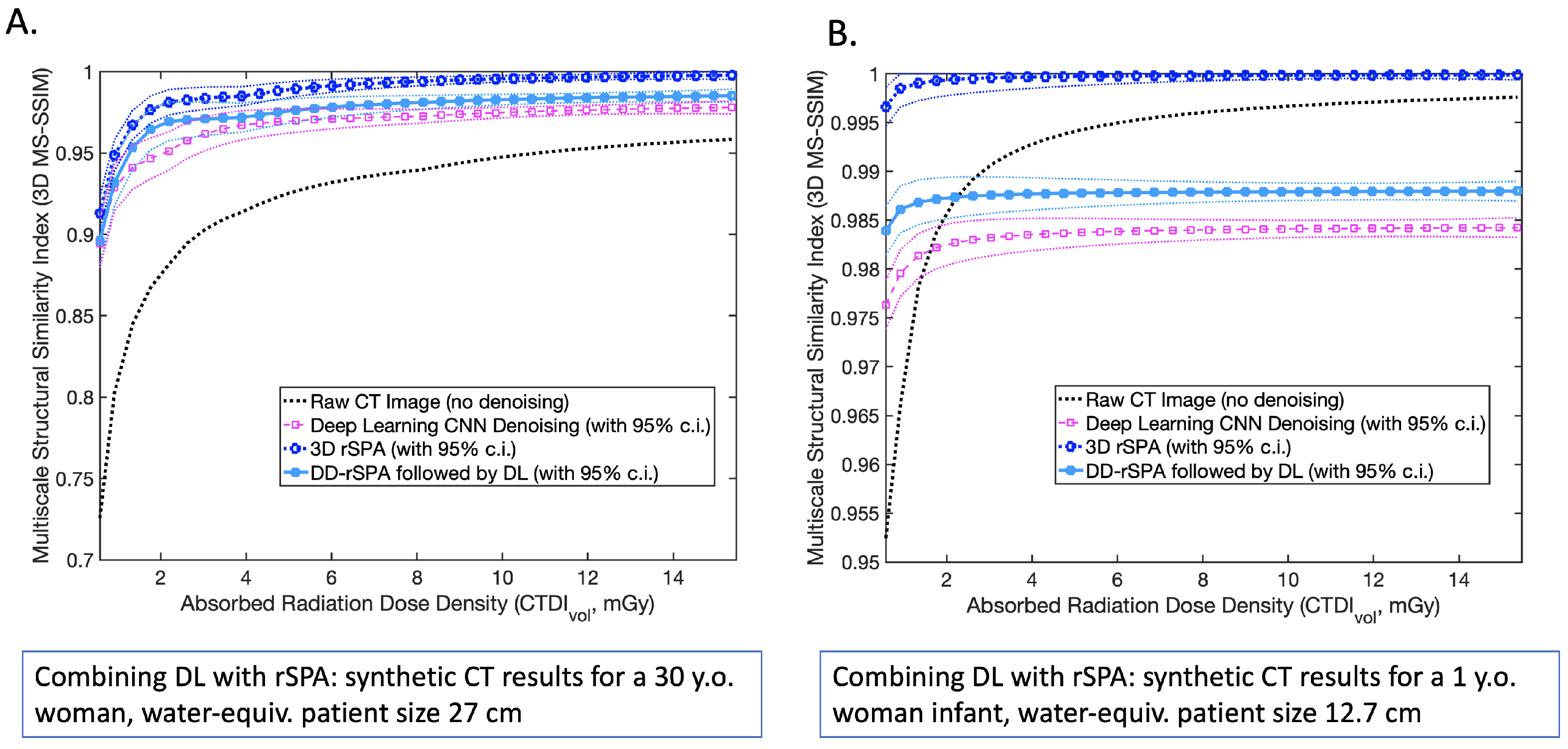
3.1. Application and Comparison of the PMS Model with Standard Methods
3.2. Implementation Details
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Communicating Radiation Risks in Paediatric Imaging: Information to Support Health Care Discussions about Benefit and Risk; World Health Organization: Geneva, Switzerland, 2016; 88p.
- Ai, T.; Yang, Z.; Hou, H.; Zhan, C.; Chen, C.; Lv, W.; Tao, Q.; Sun, Z.; Xia, L. Correlation of Chest CT and RT-PCR Testing in Coronavirus Disease 2019 (COVID-19) in China: A Report of 1014 Cases. Radiology 2020, 296, E32–E40. [Google Scholar] [CrossRef] [PubMed]
- Bernheim, A.; Mei, X.; Huang, M.; Yang, Y.; Fayad, Z.A.; Zhang, N.; Diao, K.; Lin, B.; Zhu, X.; Li, K.; et al. Chest CT Findings in Coronavirus Disease-19 (COVID-19): Relationship to Duration of Infection. Radiology 2020, 295, 200463. [Google Scholar] [CrossRef] [PubMed]
- Brenner, D.J.; Doll, R.; Goodhead, D.T.; Hall, E.J.; Land, C.E.; Little, J.B.; Lubin, J.H.; Preston, D.L.; Preston, R.J.; Puskin, J.S.; et al. Cancer risks attributable to low doses of ionizing radiation: Assessing what we really know. Proc. Natl. Acad. Sci. USA 2003, 100, 13761–13766. [Google Scholar] [CrossRef] [PubMed]
- Radiation, committee and research, board and studies, division and council, national. In Health Risks from Exposure to Low Levels of Ionizing Radiation: BEIR VII Phase 2; The National Academies Press: Washington, DC, USA, 2006; pp. 1–406. [CrossRef]
- Brenner, D.J.; Hall, E.J. Computed Tomography—An Increasing Source of Radiation Exposure. N. Engl. J. Med. 2007, 357, 2277–2284. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, D.T. A diffusional bimolecular propensity function. J. Chem. Phys. 2009, 131, 164109. [Google Scholar] [CrossRef] [PubMed]
- Siegel, J.A.; Greenspan, B.S.; Maurer, A.H.; Taylor, A.T.; Phillips, W.T.; Van Nostrand, D.; Sacks, B.; Silberstein, E.B. The BEIR VII Estimates of Low-Dose Radiation Health Risks Are Based on Faulty Assumptions and Data Analyses: A Call for Reassessment. J. Nucl. Med. 2018, 59, 1017–1019. [Google Scholar] [CrossRef]
- de González, A.B.; Mahesh, M.; Kim, K.P.; Bhargavan, M.; Lewis, R.; Mettler, F.; Land, C. Projected Cancer Risks From Computed Tomographic Scans Performed in the United States in 2007. Arch. Intern. Med. 2009, 169, 2071–2077. [Google Scholar] [CrossRef]
- Miglioretti, D.L.; Johnson, E.; Williams, A.; Greenlee, R.T.; Weinmann, S.; Solberg, L.I.; Feigelson, H.S.; Roblin, D.; Flynn, M.J.; Vanneman, N.; et al. The Use of Computed Tomography in Pediatrics and the Associated Radiation Exposure and Estimated Cancer Risk. JAMA Pediatr. 2013, 167, 700–707. [Google Scholar] [CrossRef]
- Duncan, J.R.; Lieber, M.R.; Adachi, N.; Wahl, R.L. Radiation Dose Does Matter: Mechanistic Insights into DNA Damage and Repair Support the Linear No-Threshold Model of Low-Dose Radiation Health Risks. J. Nucl. Med. 2018, 59, 1014–1016. [Google Scholar] [CrossRef]
- Huang, R.; Liu, X.; He, L.; Zhou, P.K. Radiation Exposure Associated With Computed Tomography in Childhood and the Subsequent Risk of Cancer: A Meta-Analysis of Cohort Studies. Dose-Response Publ. Int. Hormesis Soc. 2020, 18, 1559325820923828. [Google Scholar] [CrossRef]
- Choy, G.; Khalilzadeh, O.; Michalski, M.; Do, S.; Samir, A.E.; Pianykh, O.S.; Geis, J.R.; Pandharipande, P.V.; Brink, J.A.; Dreyer, K.J. Current Applications and Future Impact of Machine Learning in Radiology. Radiology 2018, 288, 318–328. [Google Scholar] [CrossRef] [PubMed]
- Koziol, P.; Raczkowska, M.K.; Skibinska, J.; Urbaniak-Wasik, S.; Paluszkiewicz, C.; Kwiatek, W.; Wrobel, T.P. Comparison of spectral and spatial denoising techniques in the context of High Definition FT-IR imaging hyperspectral data. Sci. Rep. 2018, 8, 14351. [Google Scholar] [CrossRef] [PubMed]
- Roels, J.; Vernaillen, F.; Kremer, A.; Gonçalves, A.; Aelterman, J.; Luong, H.Q.; Goossens, B.; Philips, W.; Lippens, S.; Saeys, Y. An interactive ImageJ plugin for semi-automated image denoising in electron microscopy. Nat. Commun. 2020, 11, 771. [Google Scholar] [CrossRef] [PubMed]
- Wirjadi, O.; Breuel, T. Approximate separable 3D anisotropic Gauss filter. In Proceedings of the IEEE International Conference on Image Processing 2005, Genova, Italy, 14 September 2005; Volume 2, pp. 11–149. [Google Scholar] [CrossRef]
- Tomasi, C.; Manduchi, R. Bilateral Filtering for Gray and Color Images. In Proceedings of the Sixth International Conference on Computer Vision, Bombay, India, 7 January 1998; IEEE Computer Society: Washington, DC, USA, 1998; p. 839. [Google Scholar]
- Harms, J.; Wang, T.; Petrongolo, M.; Niu, T.; Zhu, L. Noise suppression for dual-energy CT via penalized weighted least-square optimization with similarity-based regularization. Med. Phys. 2016, 43, 2676–2686. [Google Scholar] [CrossRef] [PubMed]
- Donoho, D.L. De-noising by soft-thresholding. IEEE Trans. Inf. Theory 1995, 41, 613–627. [Google Scholar] [CrossRef]
- Arias-Castro, E.; Donoho, D.L. Does median filtering truly preserve edges better than linear filtering? Ann. Stat. 2009, 37, 1172–1206. [Google Scholar] [CrossRef]
- Wang, Y.; Che, X.; Ma, S. Nonlinear filtering based on 3D wavelet transform for MRI denoising. EURASIP J. Adv. Signal Process. 2012, 2012, 40. [Google Scholar] [CrossRef]
- Tang, S.; Tang, X. Statistical CT noise reduction with multiscale decomposition and penalized weighted least squares in the projection domain. Med. Phys. 2012, 39, 5498–5512. [Google Scholar] [CrossRef][Green Version]
- Yang, Q.; Yan, P.; Zhang, Y.; Yu, H.; Shi, Y.; Mou, X.; Kalra, M.K.; Zhang, Y.; Sun, L.; Wang, G. Low-Dose CT Image Denoising Using a Generative Adversarial Network With Wasserstein Distance and Perceptual Loss. IEEE Trans. Med. Imaging 2018, 37, 1348–1357. [Google Scholar] [CrossRef]
- Konefal, A.; Tang, C.; Li, J.; Wang, L.; Li, Z.; Jiang, L.; Cai, A.; Zhang, W.; Liang, N.; Li, L.; et al. Unpaired Low-Dose CT Denoising Network Based on Cycle-Consistent Generative Adversarial Network with Prior Image Information. Comput. Math. Methods Med. 2019, 2019, 8639825. [Google Scholar] [CrossRef]
- Zhang, K.; Zuo, W.; Chen, Y.; Meng, D.; Zhang, L. Beyond a Gaussian Denoiser: Residual Learning of Deep CNN for Image Denoising. Trans. Img. Proc. 2017, 26, 3142–3155. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zhang, Y.; Zhang, W.; Liao, P.; Li, K.; Zhou, J.; Wang, G. Low-dose CT via convolutional neural network. Biomed. Opt. Express 2017, 8, 679–694. [Google Scholar] [CrossRef] [PubMed]
- Topal, E.; Löffler, M.; Zschech, E. Deep Learning-based Inaccuracy Compensation in Reconstruction of High Resolution XCT Data. Sci. Rep. 2020, 10, 7682. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Fei, L.; Zheng, W.; Xu, Y.; Zuo, W.; Lin, C.W. Deep learning on image denoising: An overview. Neural Netw. 2020, 131, 251–275. [Google Scholar] [CrossRef]
- Bhadra, S.; Kelkar, V.A.; Brooks, F.J.; Anastasio, M.A. On Hallucinations in Tomographic Image Reconstruction. IEEE Trans. Med. Imaging 2021, 40, 3249–3260. [Google Scholar] [CrossRef]
- Kaur, P.; Singh, G.; Kaur, P. A review of denoising medical images using machine learning approaches. Curr. Med. Imaging 2018, 14, 675–685. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Lundervold, A.S.; Lundervold, A. An overview of deep learning in medical imaging focusing on MRI. Z. Med. Phys. 2019, 29, 102–127. [Google Scholar] [CrossRef]
- Razzak, M.I.; Naz, S.; Zaib, A. Deep learning for medical image processing: Overview, challenges and the future. Classif. Bioapps 2018, 26, 323–350. [Google Scholar]
- Liu, S.; Deng, W. Very deep convolutional neural network based image classification using small training sample size. In Proceedings of the 2015 3rd IAPR Asian Conference on Pattern Recognition (ACPR), Kuala Lumpur, Malaysia, 3–6 November 2015; IEEE: Piscataway, NJ, USA, 2015; pp. 730–734. [Google Scholar]
- Gaonkar, B.; Hovda, D.; Martin, N.; Macyszyn, L. Deep learning in the small sample size setting: Cascaded feed forward neural networks for medical image segmentation. In Proceedings of the Medical Imaging 2016: Computer-Aided Diagnosis, San Diego, CA, USA, 27 February–3 March 2016; International Society for Optics and Photonics: Bellingham, WA, USA, 2016; Volume 9785, p. 97852I. [Google Scholar]
- Zhao, W. Research on the deep learning of the small sample data based on transfer learning. AIP Conf. Proc. 2017, 1864, 020018. [Google Scholar]
- Keshari, R.; Ghosh, S.; Chhabra, S.; Vatsa, M.; Singh, R. Unravelling small sample size problems in the deep learning world. In Proceedings of the 2020 IEEE Sixth International Conference on Multimedia Big Data (BigMM), New Delhi, India, 24–26 September 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 134–143. [Google Scholar]
- D’souza, R.N.; Huang, P.Y.; Yeh, F.C. Structural analysis and optimization of convolutional neural networks with a small sample size. Sci. Rep. 2020, 10, 834. [Google Scholar] [CrossRef] [PubMed]
- Dietterich, T. Overfitting and undercomputing in machine learning. ACM Comput. Surv. 1995, 27, 326–327. [Google Scholar] [CrossRef]
- Zhang, C.; Vinyals, O.; Munos, R.; Bengio, S. A study on overfitting in deep reinforcement learning. arXiv 2018, arXiv:1804.06893. [Google Scholar]
- Rice, L.; Wong, E.; Kolter, Z. Overfitting in adversarially robust deep learning. In Proceedings of the International Conference on Machine Learning, Virtual, 13–18 July 2020; pp. 8093–8104. [Google Scholar]
- Hosseini, M.; Powell, M.; Collins, J.; Callahan-Flintoft, C.; Jones, W.; Bowman, H.; Wyble, B. I tried a bunch of things: The dangers of unexpected overfitting in classification of brain data. Neurosci. Biobehav. Rev. 2020, 119, 456–467. [Google Scholar] [CrossRef] [PubMed]
- Gerber, S.; Pospisil, L.; Sys, S.; Hewel, C.; Torkamani, A.; Horenko, I. Co-inference of data mislabeling reveals improved models in genomics and breast cancer diagnostics. Front. Artif. Intell. 2022, 4, 739432. [Google Scholar] [CrossRef]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Ying, X. An overview of overfitting and its solutions. J. Phys. Conf. Ser. 2019, 1168, 022022. [Google Scholar] [CrossRef]
- Pan, S.J.; Yang, Q. A survey on transfer learning. IEEE Trans. Knowl. Data Eng. 2009, 22, 1345–1359. [Google Scholar] [CrossRef]
- Weiss, K.; Khoshgoftaar, T.M.; Wang, D. A survey of transfer learning. J. Big Data 2016, 3, 9. [Google Scholar] [CrossRef]
- Jang, Y.; Lee, H.; Hwang, S.J.; Shin, J. Learning what and where to transfer. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 9–15 June 2019; pp. 3030–3039. [Google Scholar]
- Raghu, M.; Zhang, C.; Kleinberg, J.; Bengio, S. Transfusion: Understanding transfer learning for medical imaging. Adv. Neural Inf. Process. Syst. 2019, 32, 3347–3357. [Google Scholar]
- Alzubaidi, L.; Fadhel, M.A.; Al-Shamma, O.; Zhang, J.; Santamaría, J.; Duan, Y.; Oleiwi, S.R. Towards a better understanding of transfer learning for medical imaging: A case study. Appl. Sci. 2020, 10, 4523. [Google Scholar] [CrossRef]
- Alzubaidi, L.; Al-Amidie, M.; Al-Asadi, A.; Humaidi, A.J.; Al-Shamma, O.; Fadhel, M.A.; Zhang, J.; Santamaría, J.; Duan, Y. Novel Transfer Learning Approach for Medical Imaging with Limited Labeled Data. Cancers 2021, 13, 1590. [Google Scholar] [CrossRef] [PubMed]
- Tartaglione, E.; Barbano, C.A.; Berzovini, C.; Calandri, M.; Grangetto, M. Unveiling covid-19 from chest X-ray with deep learning: A hurdles race with small data. Int. J. Environ. Res. Public Health 2020, 17, 6933. [Google Scholar] [CrossRef] [PubMed]
- Tsymbal, A. The problem of concept drift: Definitions and related work. Comput. Sci. Dep. Trinity Coll. Dublin 2004, 106, 58. [Google Scholar]
- Žliobaitė, I. Learning under concept drift: An overview. arXiv 2010, arXiv:1010.4784. [Google Scholar]
- Gama, J.; Žliobaitė, I.; Bifet, A.; Pechenizkiy, M.; Bouchachia, A. A survey on concept drift adaptation. ACM Comput. Surv. 2014, 46, 1–37. [Google Scholar] [CrossRef]
- Alippi, C. Learning in Non-stationary Environments. In Proceedings of the ECTA 2014—Proceedings of the International Conference on Evolutionary Computation Theory and Applications, part of IJCCI 2014, Rome, Italy, 22–24 October 2014; Rosa, A.C., Guervós, J.J.M., Filipe, J., Eds.; SciTePress: Setúbal, Portugal, 2014; p. IS-11. [Google Scholar]
- Souza, V.M.A.; dos Reis, D.M.; Maletzke, A.; Batista, G.E.A.P.A. Challenges in benchmarking stream learning algorithms with real-world data. Data Min. Knowl. Discov. 2020, 34, 1805–1858. [Google Scholar] [CrossRef]
- Horenko, I. On a Scalable Entropic Breaching of the Overfitting Barrier for Small Data Problems in Machine Learning. Neural Comput. 2020, 32, 1563–1579. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long Short-term Memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef]
- Horenko, I. Finite Element Approach to Clustering of Multidimensional Time Series. SIAM J. Sci. Comput. 2010, 32, 62–83. [Google Scholar] [CrossRef]
- Metzner, P.; Putzig, L.; Horenko, I. Analysis of persistent nonstationary time series and applications. Commun. Appl. Math. Comput. Sci. 2012, 7, 175–229. [Google Scholar] [CrossRef]
- Gerber, S.; Horenko, I. Improving clustering by imposing network information. Sci. Adv. 2015, 1, e1500163. [Google Scholar] [CrossRef] [PubMed]
- Pospisil, L.; Gagliardini, P.; Sawyer, W.; Horenko, I. On a scalable nonparametric denoising of time series signals. Commun. Appl. Math. Comput. Sci. 2018, 13, 107–138. [Google Scholar] [CrossRef]
- Rodrigues, D.R.; Everschor-Sitte, K.; Gerber, S.; Horenko, I. A deeper look into natural sciences with physics-based and data-driven measures. Iscience 2021, 24, 102171. [Google Scholar] [CrossRef]
- Gerber, S.; Horenko, I. Towards a direct and scalable identification of reduced models for categorical processes. Proc. Natl. Acad. Sci. USA 2017, 114, 4863–4868. [Google Scholar] [CrossRef]
- Wackerly, D.; Mendenhall, W., III; Scheaffer, R.L. Mathematical Statistics with Applications, 6th ed.; Duxbury Advanced Series; Cengage Learning: Boston, MA, USA, 2002. [Google Scholar]
- Huynh-Thu, Q.; Ghanbari, M. The accuracy of PSNR in predicting video quality for different video scenes and frame rates. Telecommun. Syst. 2012, 49, 35–48. [Google Scholar] [CrossRef]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image Quality Assessment: From Error Visibility to Structural Similarity. Trans. Img. Proc. 2004, 13, 600–612. [Google Scholar] [CrossRef]
- Hallgrímsson, B.; Hall, B. Variation: A Central Concept in Biology; Elsevier Science: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Meyer, M.; Ronald, J.; Vernuccio, F.; Nelson, R.C.; Ramirez-Giraldo, J.C.; Solomon, J.; Patel, B.N.; Samei, E.; Marin, D. Reproducibility of CT Radiomic Features within the Same Patient: Influence of Radiation Dose and CT Reconstruction Settings. Radiology 2019, 293, 583–591. [Google Scholar] [CrossRef]
- De Man, B.; Basu, S.; Chandra, N.; Dunham, B.; Edic, P.; Iatrou, M.; McOlash, S.; Sainath, P.; Shaughnessy, C.; Tower, B.; et al. CatSim: A new computer assisted tomography simulation environment. In Proceedings of the Medical Imaging 2007: Physics of Medical Imaging, San Diego, CA, USA, 17–22 February 2007; International Society for Optics and Photonics: Bellingham, WA, USA, 2007; Volume 6510, p. 65102. [Google Scholar]
- Yu, L.; Shiung, M.; Jondal, D.; McCollough, C.H. Development and validation of a practical lower-dose-simulation tool for optimizing computed tomography scan protocols. J. Comput. Assist. Tomogr. 2012, 36, 477–487. [Google Scholar] [CrossRef]
- McCollough, C.H.; Bartley, A.C.; Carter, R.E.; Chen, B.; Drees, T.A.; Edwards, P.; Holmes, D.R., III; Huang, A.E.; Khan, F.; Leng, S.; et al. Low-dose CT for the detection and classification of metastatic liver lesions: Results of the 2016 low dose CT grand challenge. Med. Phys. 2017, 44, e339–e352. [Google Scholar] [CrossRef]
- Moen, T.R.; Chen, B.; Holmes, D.R., III; Duan, X.; Yu, Z.; Yu, L.; Leng, S.; Fletcher, J.G.; McCollough, C.H. Low-dose CT image and projection dataset. Med. Phys. 2021, 48, 902–911. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Samei, E. Comparison of patient size-based methods for estimating quantum noise in CT images of the lung. Med. Phys. 2009, 36, 541–546. [Google Scholar] [CrossRef] [PubMed]
- Solomon, J.; Lyu, P.; Marin, D.; Samei, E. Noise and spatial resolution properties of a commercially available deep? Learning based CT reconstruction algorithm. Med. Phys. 2020, 47, 3961–3971. [Google Scholar] [CrossRef] [PubMed]
- Samei, E.; Kinahan, P.; Nishikawa, R.M.M.; Maidment, A. Virtual Clinical Trials: Why and What (Special Section Guest Editorial). J. Med. Imaging 2020, 7, 042801. [Google Scholar] [CrossRef] [PubMed]
- Anam, C.; Haryanto, F.; Widita, R.; Arif, I.; Dougherty, G.; McLean, D. Volume computed tomography dose index (CTDIvol) and size-specific dose estimate (SSDE) for tube current modulation (TCM) in CT scanning. Int. J. Radiat. Res. 2018, 16, 289–297. [Google Scholar]
- Karimi, D.; Deman, P.; Ward, R.; Ford, N. A sinogram denoising algorithm for low-dose computed tomography. BMC Med. Imaging 2016, 16, 11. [Google Scholar] [CrossRef]
- Koyuncu, H.; Ceylan, R. Elimination of white Gaussian noise in arterial phase CT images to bring adrenal tumours into the forefront. Comput. Med. Imaging Graph. 2018, 65, 46–57. [Google Scholar] [CrossRef]
- Sheppard, J.P.; Nguyen, T.; Alkhalid, Y.; Beckett, J.S.; Salamon, N.; Yang, I. Risk of Brain Tumor Induction from Pediatric Head CT Procedures: A Systematic Literature Review. Brain Tumor Res. Treat. 2018, 6, 1–7. [Google Scholar] [CrossRef]
- Bezdek, J.C.; Ehrlich, R.; Full, W. FCM: The Fuzzy c-MEANS Clustering Algorithm. Comput. Geosci. 1984, 10, 191–203. [Google Scholar] [CrossRef]
- Höppner, F.; Klawonn, F.; Kruse, R.; Runkler, T. Fuzzy Cluster Analysis: Methods for Classification, Data Analysis and Image Recognition; John Wiley & Sons: Hoboken, NJ, USA, 1999. [Google Scholar]
- Jain, A.K. Data clustering: 50 years beyond K-means. Pattern Recognit. Lett. 2010, 31, 651–666. [Google Scholar] [CrossRef]
- Mumford, D.; Shah, J. Optimal approximations by piecewise smooth functions and associated variational problems. Comm. Pure Appl. Math. 1989, 42, 577–685. [Google Scholar] [CrossRef]
- Gerber, S.; Pospisil, L.; Navandar, M.; Horenko, I. Low-cost scalable discretization, prediction, and feature selection for complex systems. Sci. Adv. 2020, 6, eaaw0961. [Google Scholar] [CrossRef] [PubMed]
- de Wiljes, J.; Majda, A.; Horenko, I. An Adaptive Markov Chain Monte Carlo Approach to Time Series Clustering of Processes with Regime Transition Behavior. SIAM Multiscale Model. Simul. 2013, 11, 415–441. [Google Scholar] [CrossRef]
- Rudin, L.I.; Osher, S.; Fatemi, E. Nonlinear total variation based noise removal algorithms. Phys. D Nonlinear Phenom. 1992, 60, 259–268. [Google Scholar] [CrossRef]
- Chambolle, A. Finite-differences discretizations of the mumford-shah functional. ESAIM Math. Model. Numer. Anal. 1999, 33, 261–288. [Google Scholar] [CrossRef]
- Lysaker, O.M.; Lundervold, A.; Tai, X. Noise removal using fourth-order partial differential equation with applications to medical magnetic resonance images in space and time. IEEE Trans. Image Process. 2003, 12, 1579–1590. [Google Scholar] [CrossRef]
- Chan, T.F.; Shen, J. Image Processing and Analysis: Variational, PDE, Wavelet, and Stochastic Methods; SIAM: Philadelphia, PA, USA, 2005. [Google Scholar]
- Pock, T.; Cremers, D.; Bischof, H.; Chambolle, A. An algorithm for minimizing the Mumford-Shah functional. In Proceedings of the 2009 IEEE 12th International Conference on Computer Vision, Kyoto, Japan, 29 September–2 October 2009; pp. 1133–1140. [Google Scholar] [CrossRef]
- Hohm, K.; Storath, M.; Weinmann, A. An algorithmic framework for Mumford–Shah regularization of inverse problems in imaging. Inverse Probl. 2015, 31, 115011. [Google Scholar] [CrossRef]
- Paragios, N.; Duncan, J.; Ayache, N. Handbook of Biomedical Imaging: Methodologies and Clinical Research; Springer: Berlin/Heidelberg, Germany, 2015; p. 590. [Google Scholar] [CrossRef]
- Barzilai, J.; Borwein, J.M. Two point step size gradient methods. IMA J. Numer. Anal. 1988, 8, 141–148. [Google Scholar]
- Birgin, E.G.; Martínez, J.M.; Raydan, M.M. Nonmonotone spectral projected gradient methods on convex sets. SIAM J. Optim. 2000, 10, 1196–1211. [Google Scholar]
- Boyd, L.; Vandenberghe, L. Convex Optimization, 1st ed.; Cambridge University Press: New York, NY, USA, 2004. [Google Scholar]
- Chen, Y.; Ye, X. Projection onto a simplex. arXiv 2011, Unpublished manuscript. arXiv:101.6081. [Google Scholar]
- Grippo, L.; Lampariello, F.; Lucidi, S. nonmonotone line search technique for Newton’s method. SIAM J. Numer. Anal. 1986, 23, 707–716. [Google Scholar]

| Image Quality Loss (3D MS-SSIM, in %) | |||||
|---|---|---|---|---|---|
| Reduction of LAR (in %) | Raw Image | 3D Gaussian Filtering | DL CNN Denoising | 3D Wavelet Filtering | rSPA |
| 6 | 1.16 | 0.77 | 0.54 | 0.79 | 0.01 |
| 16 | 1.26 | 0.84 | 0.59 | 0.86 | 0.01 |
| 23 | 1.34 | 0.90 | 0.63 | 0.92 | 0.01 |
| 29 | 1.44 | 0.96 | 0.67 | 0.99 | 0.01 |
| 36 | 1.56 | 1.04 | 0.73 | 1.07 | 0.01 |
| 42 | 1.70 | 1.13 | 0.79 | 1.16 | 0.01 |
| 49 | 1.88 | 1.25 | 0.88 | 1.29 | 0.01 |
| 56 | 2.12 | 1.41 | 0.99 | 1.45 | 0.02 |
| 63 | 2.45 | 1.63 | 1.14 | 1.67 | 0.02 |
| 70 | 2.92 | 1.95 | 1.36 | 2.00 | 0.02 |
| 77 | 3.69 | 2.46 | 1.72 | 2.52 | 0.03 |
| 83 | 5.11 | 3.40 | 2.38 | 3.49 | 0.04 |
| 90 | 8.72 | 5.81 | 4.07 | 5.97 | 0.06 |
| 97 | 25.96 | 18.20 | 13.37 | 18.16 | 0.17 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Horenko, I.; Pospíšil, L.; Vecchi, E.; Albrecht, S.; Gerber, A.; Rehbock, B.; Stroh, A.; Gerber, S. Low-Cost Probabilistic 3D Denoising with Applications for Ultra-Low-Radiation Computed Tomography. J. Imaging 2022, 8, 156. https://doi.org/10.3390/jimaging8060156
Horenko I, Pospíšil L, Vecchi E, Albrecht S, Gerber A, Rehbock B, Stroh A, Gerber S. Low-Cost Probabilistic 3D Denoising with Applications for Ultra-Low-Radiation Computed Tomography. Journal of Imaging. 2022; 8(6):156. https://doi.org/10.3390/jimaging8060156
Chicago/Turabian StyleHorenko, Illia, Lukáš Pospíšil, Edoardo Vecchi, Steffen Albrecht, Alexander Gerber, Beate Rehbock, Albrecht Stroh, and Susanne Gerber. 2022. "Low-Cost Probabilistic 3D Denoising with Applications for Ultra-Low-Radiation Computed Tomography" Journal of Imaging 8, no. 6: 156. https://doi.org/10.3390/jimaging8060156
APA StyleHorenko, I., Pospíšil, L., Vecchi, E., Albrecht, S., Gerber, A., Rehbock, B., Stroh, A., & Gerber, S. (2022). Low-Cost Probabilistic 3D Denoising with Applications for Ultra-Low-Radiation Computed Tomography. Journal of Imaging, 8(6), 156. https://doi.org/10.3390/jimaging8060156






