1. Introduction
Geometric morphometrics aims to provide a mathematical description of biological shapes [
1,
2,
3,
4,
5]. Three-dimensional (3D) surface scanning [
6] is a technique that allows one to capture the 3D shape, e.g., of the human face, as shown in
Figure 1 for the author’s face. It is a useful tool in understanding dental and maxillofacial diagnostics, treatment planning, and the effects of treatment [
7]. Such biological shapes may be described by a set of landmark points, illustrated also in
Figure 1. Methods such as Procrustes transformation [
1] are often used to standardise centring, orientation, and scale in a dataset of such 3D shapes.
Multivariate data contains more than one “outcome” variable, such as the
x-,
y- and
z-components of the Cartesian landmark points (again, as shown in
Figure 1). These variables tend to be highly correlated and so multivariate statistical methods such as principal components analysis (PCA) [
1] are needed to in order to analyse such data. Between-group PCA (bgPCA) [
8,
9] is an extension of standard PCA that carries out separate PCAs on (between-group) covariance matrices based on “group means” and (within group) covariance matrices based on individual shapes around these means. It has much in common with (though it is not the same as) canonical variates analysis/linear discriminant analysis [
10]. Multilevel PCA (mPCA) has been used by us [
11,
12,
13,
14,
15,
16,
17] to analyse 3D facial shapes obtained from 3D facial scans; note that two-level multilevel PCA (mPCA) is equivalent to bgPCA. mPCA has been used by us to investigate changes by ethnicity and sex [
11,
12], the act of smiling [
13,
14], facial shape changes in adolescents due to age [
15,
16], and the effects of maternal smoking and alcohol consumption on the facial shape of English adolescents [
17].
Recent articles [
8,
9,
18,
19,
20] have pointed out a number of “pathologies” in techniques such as bgPCA (and therefore also mPCA). Perhaps the most notable pathology [
8,
9] is that spurious conclusions about differences between groups can occur when the number of parameters in the model is much larger than sample sizes used in the model. Another limitation occurs when sample sizes are not balanced between groups [
8]. Here, we wish to explore these pathologies by carrying out Monte Carlo simulated experiments firstly using uncorrelated normally distributed variables and secondly for correlated multivariate normally distributed data based on “real” data using the 21 landmark points in
Figure 1.
3. Results
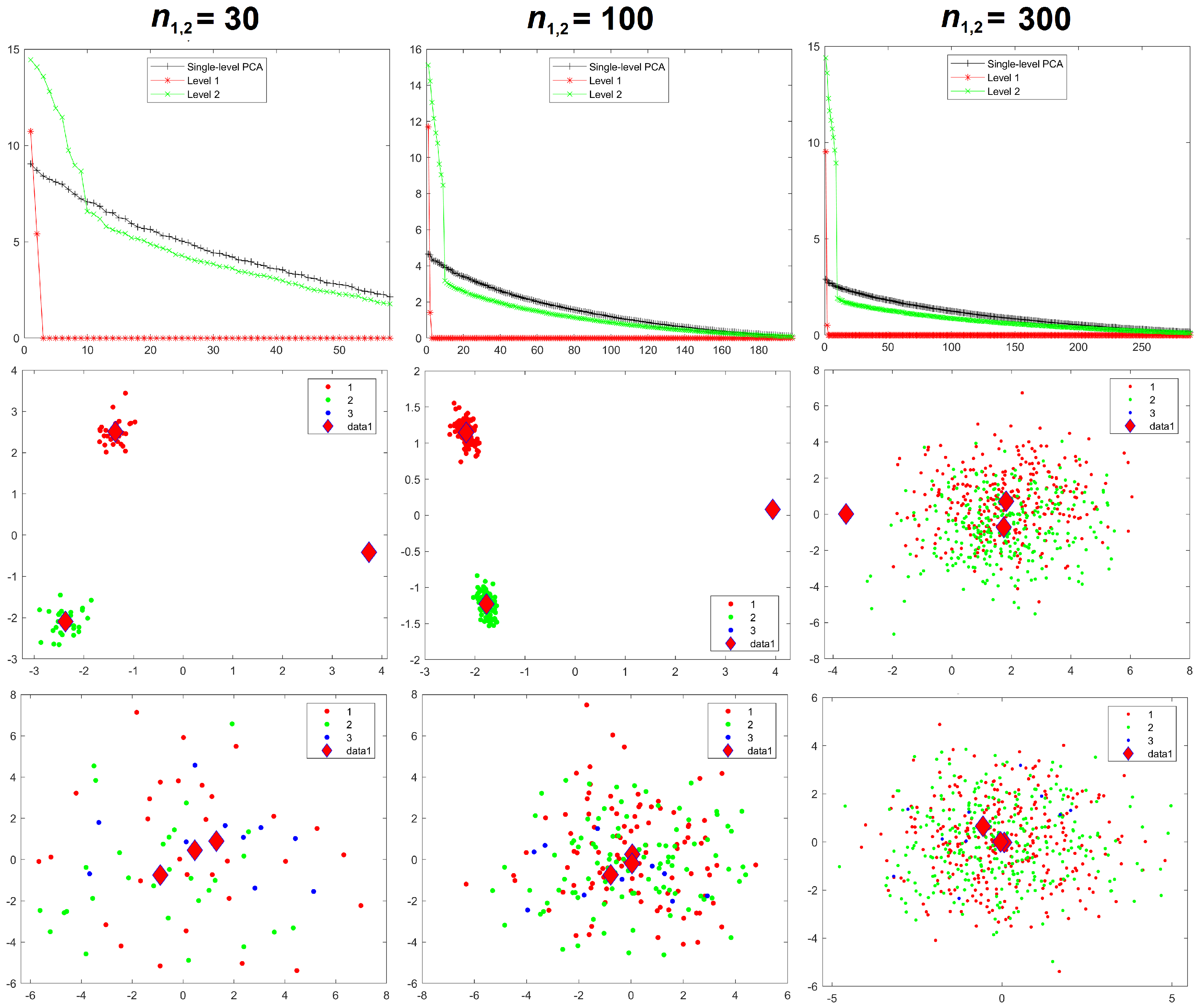
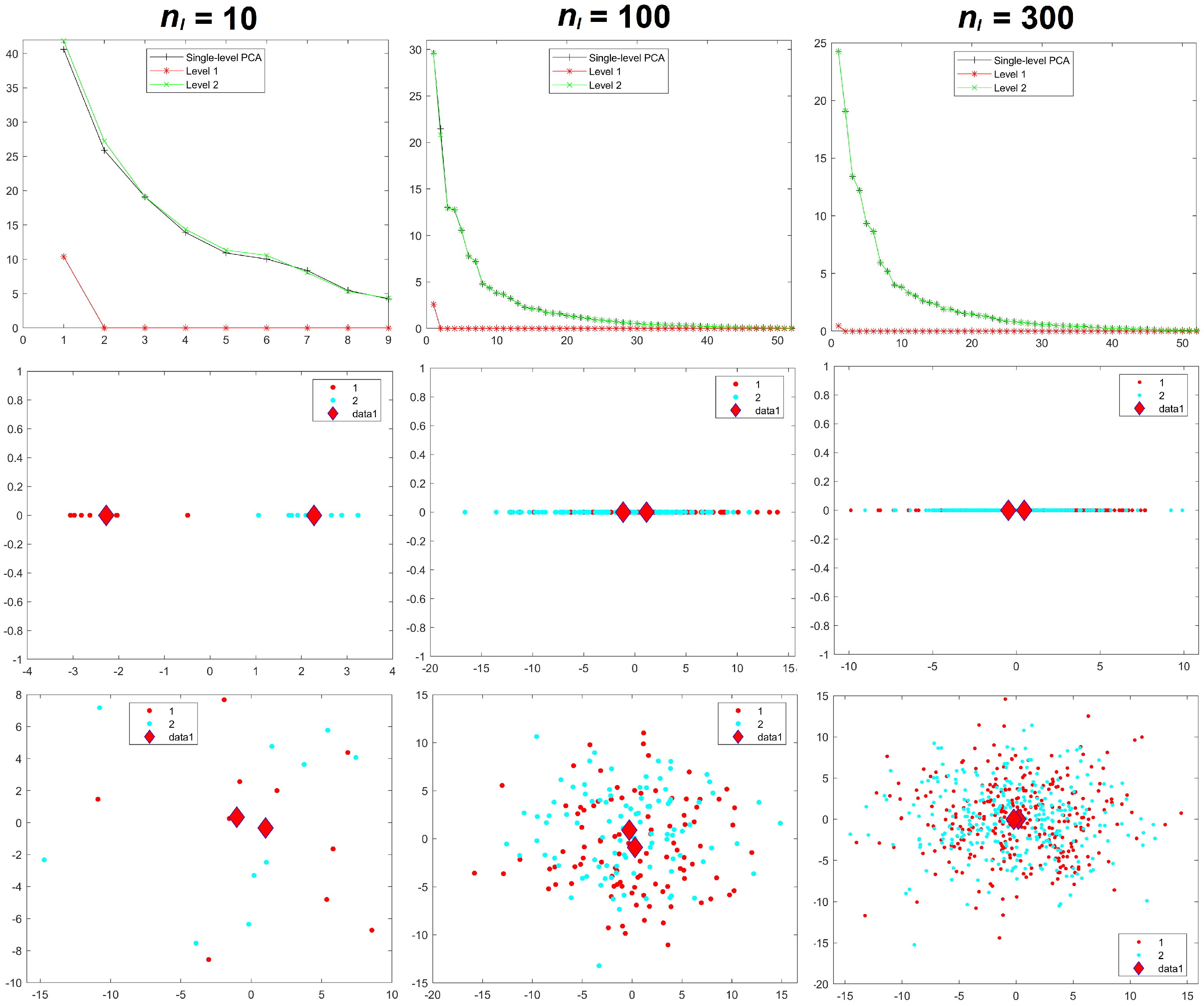
Results for the eigenvalues from mPCA and single-level PCA for Experiment 1 are shown in
Figure 3. The magnitude of these eigenvalues at level 1 via mPCA (with respect to the total variation) reduces strongly with increasing sample size per group
, as shown in
Figure 3. The percentage variation at level 1 via mPCA also reduces strongly with increasing values of
, as shown in
Table 2. Indeed, both measures are clearly tending towards the correct asymptotic value of zero in the limit of “infinite” sample size per group. The average sum of eigenvalues for single-level PCA over all MC simulations is (to within expected sampling error) equal to 300 for all values of sample size per group
. It is stated in [
9] that mPCA underestimates the variation due to within-group effects (i.e., at level 2 of the mPCA model). However, we find that the sum of eigenvalues at level 2 for the mPCA model averaged over all MC simulations is also (again to within expected sampling error) equal to 300 in all simulations, which seems apparently to contradict this statement.
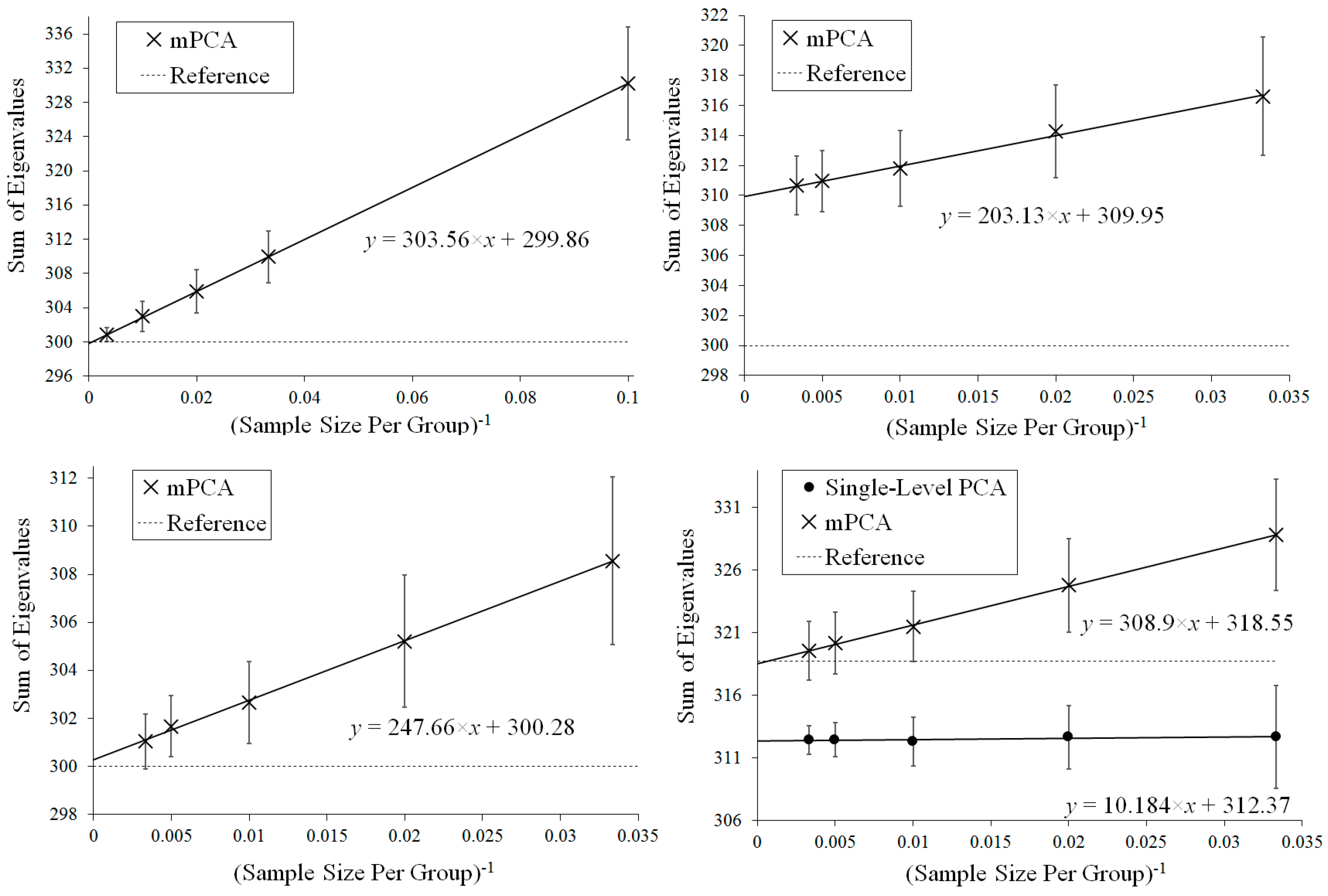
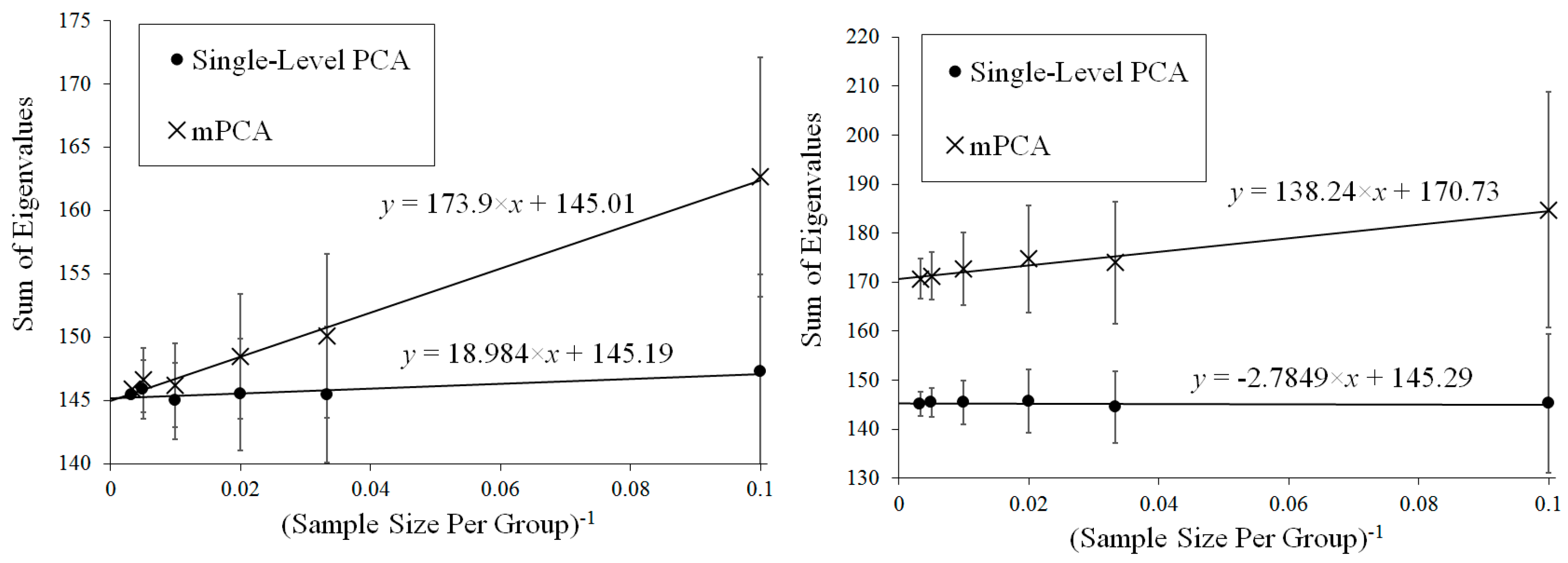
Figure 4 shows that results for the sum of eigenvalues over both levels via mPCA extrapolate to the correct value of 300 in the limit
. We see also from
Figure 3 that the curves for the eigenvalues via single-level PCA and level 2 mPCA become flatter as we increase the sample size per group
, which agrees with the Marchenko–Pastur theorem [
21].
Results for component scores are also given in
Figure 3. As in [
8], we find that strong apparent differences seem to occur between groups occurs at level 1 of the mPCA model. We see from
Figure 3 that this occurs also via single-level PCA, albeit to a lesser extent. Again, these differences are due to random sampling effects and so are spurious. (Note that group centroids of component scores at level 2 via mPCA were indeed congruent with the origin for all MC simulated datasets and in all experiments carried out here). Differences between groups in
Figure 3 become less pronounced for both mPCA and single-level PCA as the sample size per group is increased. Indeed, very strong overlap occurs in components scores and spurious differences between groups are quite small for a sample size per group of
= 300 at level 1 via mPCA, as shown by the group centroids in this figure. Experiment 1 shows that random differences between groups that are spread over all 300 variables (and therefore probably also over possible principal components via traditional single-level PCA) are now being concentrated in just two components at the level 1 of the mPCA model. Experiment 1 indicates (as a “rule of thumb” only) that the sample sizes per group should at least be of similar magnitude to the number of variables, i.e., 300, in order to obtain reasonable results.
Results for the eigenvalues from mPCA and single-level PCA for Experiment 2a are shown in
Figure 5. In this case, the sample sizes per group are varied for groups 1 and 2 only, whereas group 3 has
= 10 in all simulations. The magnitude of these eigenvalues at level 1 mPCA shown in
Figure 5 and percentage variation shown in
Table 2 reduce with increasing sample size per group,
n, although they are clearly “saturating” by
n = 100 for groups 1 and 2. It is clear that the covariance matrices at both level 1 and 2 via mPCA are being very strongly affected by the small sample size in group 3. For example, eigenvalues at level 2 via mPCA exhibit a strange “spike” for low values of eigenvalue number. Eigenvalues at level 1 via mPCA are higher than those in
Figure 3, where sample sizes are equal across all groups. However, we again find that the sum of eigenvalues for both single-level PCA and mPCA at level 2 is equal to 300 (again to within expected sampling error).
Table 2 shows that the percentage variance does not tend to correct value of zero percent; a result is driven by the small sample size in group 3.
Figure 4 shows that results for the sum of eigenvalues at both levels via mPCA do not extrapolate to the correct value of 300 in the limit
in Experiment 2a.
Component scores in
Figure 5 show that group 3 is also a clear outlier for both mPCA, level 1 and single-level PCA. (Again, group centroids for mPCA at level 2 are congruent with the origin). It is noticeable that group 3 produces an outlying result in
Figure 5 even for single-level PCA for
= 300. Spurious differences are exaggerated for mPCA compared to single-level PCA, especially between groups 1 and 2 compared to group 3 for mPCA. Reasonable results for component scores via mPCA are never achieved in Experiment 2a due to the low sample size in group 3.
Results for the eigenvalues from mPCA and single-level PCA for Experiment 2b are shown in
Figure 6. Again, the sample sizes per group are varied for groups 1 and 2 only, whereas group 3 has
= 10 in all simulations. We see that eigenvalues for level 2 mPCA now are of very similar magnitude to results of single-level PCA. Indeed, we see that problems with leading eigenvalues for both levels 1 and 2 mPCA due to imbalances in sample sizes appear to have been removed in
Figure 6 by the weighted form of the covariance matrices, which is an encouraging result. Eigenvalues for level 2, mPCA are very slightly lower in magnitude in
Figure 6 than single-level PCA because Equations (7) and (8) are essentially population rather than sample covariance matrices, although this effect reduces quickly with increasing sample size per group
.
Figure 4 shows that results for the sum of eigenvalues via mPCA extrapolate to the correct value of 300 in the limit
for Experiment 2b.
We see from
Figure 6 that problems of spurious differences between groups are not removed by the weighted forms of Equations (7) and (8). Differences between groups that are contained in all variables (and again are probably spread over all components via single-level PCA) are again being concentrated in just two components at level 1 via mPCA. These spurious differences reduce strongly between groups 1 and 2 with increasing sample sizes in these groups, although they persist between groups 1 and 2 compared to group 3 even up to
= 300, as shown in
Figure 6. Reasonable results for component scores via mPCA are never achieved in Experiment 2b due to the low sample size in group 3, even when the weighted forms of the covariance matrices are used.
Figure 7 shows results for Experiment 3 in which between-group variation is added to the data, where the means of each group now follow a normal distribution with means equal to zero and a standard deviation of 0.25.
Table 2 shows that mPCA is clearly tending towards the theoretical value of 5.9% with increasing sample size per group. Note that the sum of eigenvalues at level 2 via mPCA is again equal to 300 (within expected sampling error).
Figure 4 demonstrates that the sum of eigenvalues over both levels via mPCA scale approximately linearly with
to a value of 318.55 in the limit
. This is good agreement with the aysmptotic value of 318.75, although even better correspondence would presumably also be obtained by including higher values of
in the regression data in
Figure 4.
Figure 4 shows also that the values for the sum of all eigenvalues via single-level PCA is approximately flat with respect to
. Indeed, the total variation captured by single-level PCA is clearly well below the asymptotic overall total value of 318.75.
Figure 7 shows that both mPCA and single-level PCA overestimate differences between groups in component scores for small sample sizes per group. However, the broad pattern in the component scores for mPCA (and single-level PCA) has largely converged for sample size per group
= 200 (not shown here) and it has certainly converged by the time that
= 300 is reached (shown in
Figure 7). Again, Experiment 3 indicates again (as a rough-and-ready “rule of thumb”) that reasonable results are obtained when the sample sizes in all groups are of similar magnitude to the number of variables, i.e., 300 here.
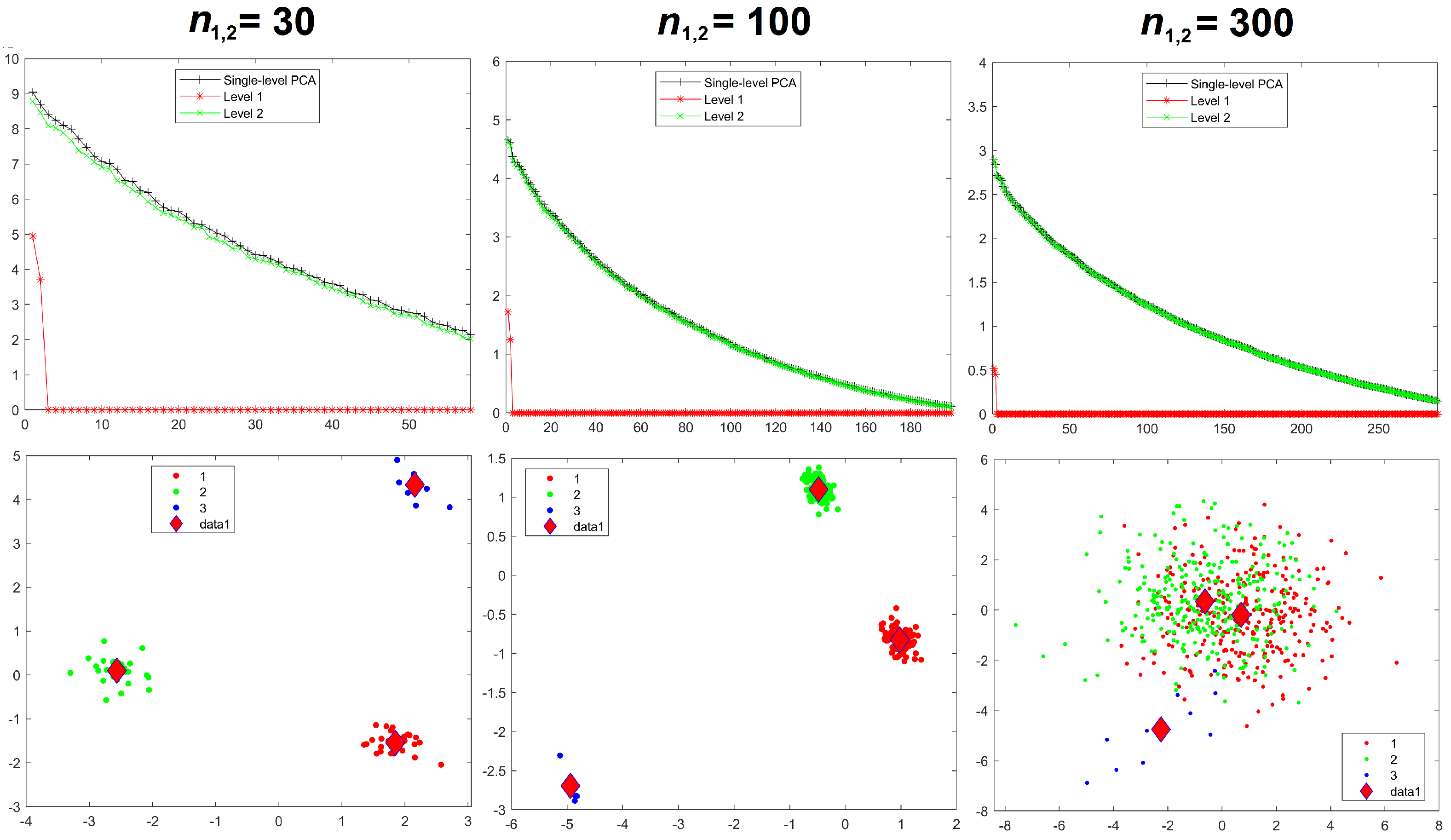
Results for the eigenvalues from mPCA and single-level PCA for Experiments 4 and 5 are shown in
Figure 8 and
Figure 9. These results for the correlated data in Experiments 4 and 5 are very similar to those earlier results from Experiments 1 to 3, which involved uncorrelated data. We see from
Figure 8 and from
Table 2 that variances at level 1 mPCA for Experiment 4 reduces with increasing sample size per group.
Figure 10 shows that the sum of eigenvalues at both levels via mPCA scales approximately linearly with
for Experiment 4 and that this line extrapolates to a value that is very close that of single-level PCA in the limit
. By contrast,
Figure 9 shows that eigenvalues at level 1 do not tend to zero as the sample size per group increases for Experiment 5 and we note again that between-group variation has been explicitly added to the MC data in this case.
Figure 10 shows that the sum of eigenvalues at both levels via mPCA scales approximately linearly with
for Experiment 5 and that it extrapolates in the limit
to a value that is much larger than that from single-level PCA.
Table 2 shows that the percentage of variance explained by level 1 via mPCA for Experiment 5 converges to a non-zero value probably near to about 10%.
Component scores in
Figure 8 and
Figure 9 for Experiments 4 and 5 again also show a very similar pattern to those results in Experiments 1 to 3. Strong initial differences between groups in component scores via mPCA (and single-level PCA to some extent also) reduce strongly as sample sizes per group increased in Experiment 4. Indeed, it is noticeable in
Figure 8 that differences between groups via mPCA are fairly small for
= 100. By contrast, differences in component scores between groups via mPCA are observed for all sample sizes per group in Experiment 5, where between-group variation has been added explicitly to the MC data, for both single-level PCA and mPCA. Indeed,
Figure 9 shows that differences between groups via mPCA are fairly similar for
= 100 compared to
= 300. Experiments 4 and 5 indicate again (and very broadly) that reasonable results are obtained when the sample sizes in all groups are of similar magnitude to the number of variables, i.e., 63 components for Experiments 4 and 5.
4. Discussion
An exploration of “pathologies” of mPCA was carried out here by considering a two-level mPCA model that is equivalent to bgPCA. It was clear that spurious differences between groups due to random sampling effects contained in all variables in Experiments 1, 2, and 4 were concentrated in the (relatively few) components at level 1 for mPCA. This effect meant that mPCA therefore falsely gave an impression of strong differences in component scores where in truth there were none. As stated in [
8,
9,
17,
18,
19,
20], pathologies of bgPCA (and therefore also mPCA) do exist, mostly strikingly in terms of interpretation of these component scores when sample sizes are low in any of the groups. However, these spurious differences in component scores via mPCA decreased significantly as the sample sizes per group were increased.
Note that 3D facial scanning of human subjects can be costly because it can be time consuming and/or labour intensive. Sample sizes might often measured in tens of subjects only, where such “pathologies” are likely to manifest. However, sample sizes per group can be a problem even for large-scale epidemiological studies in humans if one is interested for example in subsets of subjects with rare syndromes that can affect facial appearance and shape. Similar problems might occur in archaeology or palaeontology, where the number of samples to be scanned might naturally be constrained. Another limitation of mPCA is that the rank of covariance matrices at higher levels of the model are limited to the number of groups minus one. In practice, this places a limit on the number of non-zero eigenvalues at these levels.
Imbalances in sample sizes in different groups can be addressed by using a form weighted covariance matrices inspired by the maximum likelihood solution, previously also suggested in [
9]. Our results suggested that this “weighting” had a beneficial effect on covariance matrices and eigenvalues. However, weighting did not solve all of the problems of spurious differences in component scores between groups, which were due here to a very small sample size in one group. Experiment 2 demonstrates that misleading results for component scores persist via mPCA if the sample sizes are low in any of the groups. Notably, the usefulness of such “weighting” was questioned also in [
8]. However, such weighting schemes might be useful when such imbalances occur and when sample sizes per group are sufficiently large enough in all groups. This topic requires more investigation in future.
Our calculations also indicated that single-level PCA underestimated the total amount of variance when between-group variation was introduced explicitly to the data generation model in Experiment 3. For example,
Figure 4 showed that mPCA results extrapolated to a value that was very close to the theoretical asymptotic value for the total variation in Experiment 3, whereas single-level PCA did not. These results were also supported by evidence in
Table 2. However, this is exactly what one would expect as the models used in MC data generation and via mPCA were essentially identical. Very similar results were seen in Experiment 5 where between-group variation was introduced to correlated 3D MC data representing 3D facial shape. Traditional PCA is essentially just a single-level method and so one would not expect it to capture the effects of such multilevel structures and/or of “clustering.” Interestingly, the sums of eigenvalues via mPCA scaled approximately linearly with the inverse of the sample size per group in all Experiments 1 to 5, which is another potentially important result of this research. We speculate that this might be another manifestation of the Marchenko–Pastur theorem [
21].
The simulations presented here for the uncorrelated normally distributed variables in Experiments 1 to 3 are the most severe (and artificial) test of both mPCA and single-level PCA as any apparent structure to the data is due purely to random sampling effects. It was noted (e.g., in [
9]) that these problems of spurious differences between group for bgPCA are reduced when the multivariate data is correlated, which generally is the case in reality, e.g., for shape data. The evidence from Experiments 4 and 5, in which correlated multivariate normally distributed data was generated, were inconclusive in relation to this claim specifically, although they do not contradict it. However, the total number of variables was much lower in Experiments 4 and 5 compared to Experiments 1 to 3, which makes it harder to compare results on an equal footing. Experiments 4 and 5 do underline that the results presented here are clearly relevant to modelling shape, such as those illustrated by
Figure 1 for 3D facial shape and as described in [
11,
12,
13,
14,
15,
16,
17].
The results of this work show broadly that reasonable results ought to be obtained when sufficiently large sample sizes per group are used in all groups. As a “rule of thumb” only, sample sizes per group in all groups should be at least equal to the number of variables. However, modes of variation from mPCA should also always be examined critically and they should be compared to known results in the literature where they are known to exist, e.g., known changes in facial shapes in humans due to sex [
12]. The author of [
8] presents a detailed list of recommendations about the use (or refraining from use) of bgPCA in relation to biological morphometrics. The authors of [
19] propose cross-validation as a method of overcoming these problems, whereas the authors of [
20] propose a mixture of permutation tests and cross-validation. The interested reader is referred to [
8,
19,
20] for more details.