body2vec: 3D Point Cloud Reconstruction for Precise Anthropometry with Handheld Devices
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Segmentation Model
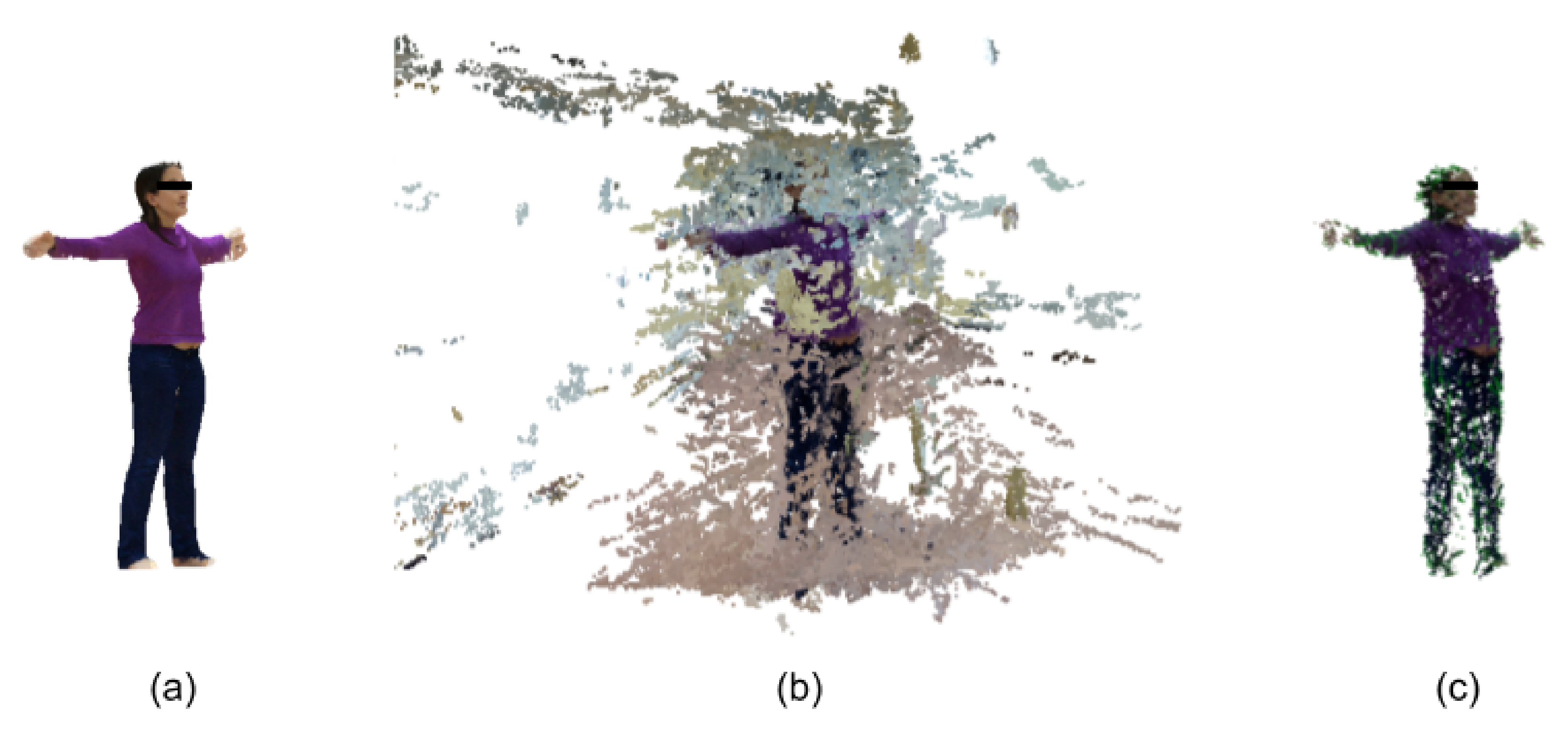
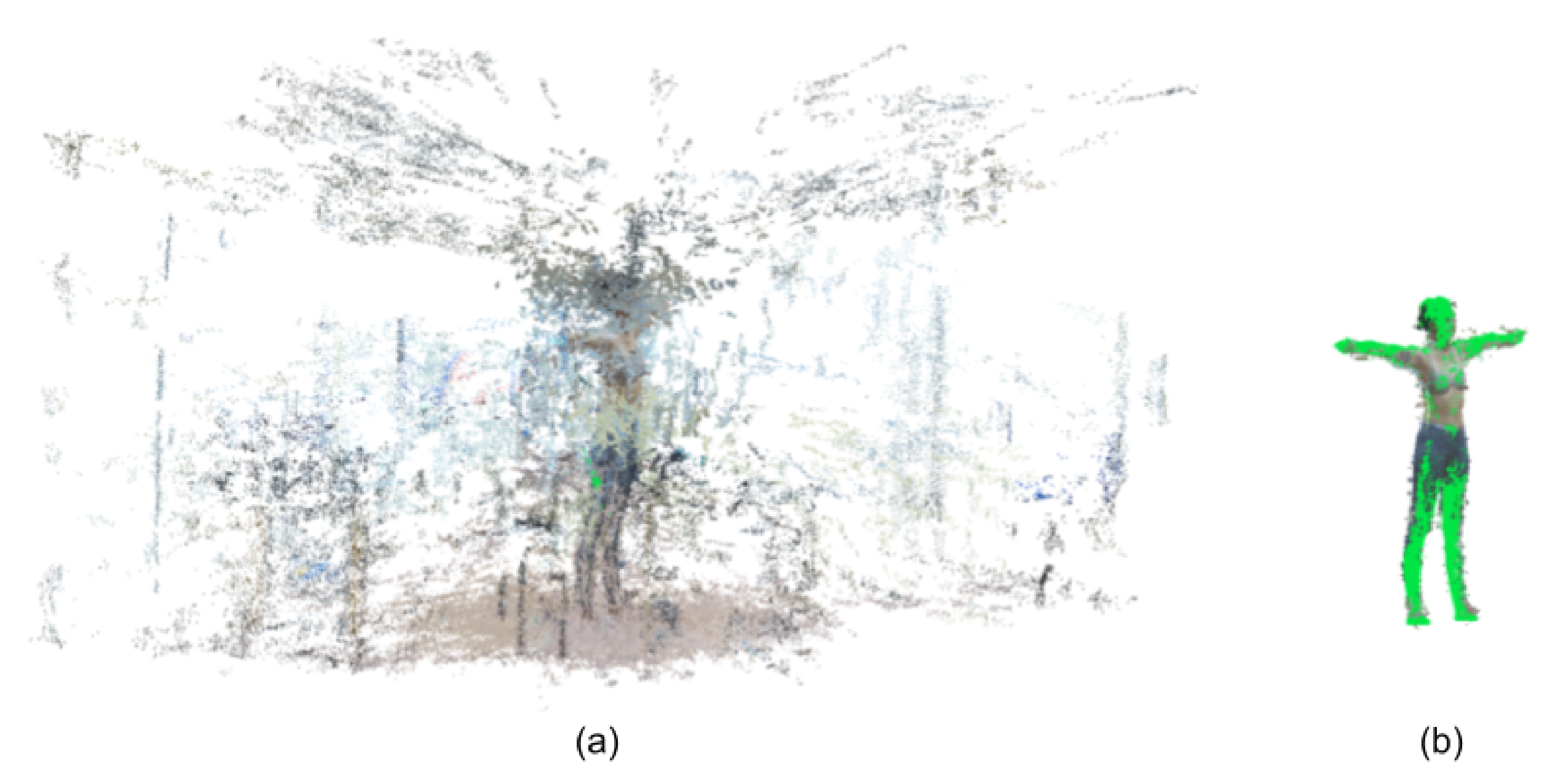
2.3. 3D Reconstruction and Measurement
3. Results
3.1. Mask Segmentation
3.2. Segmented Point Cloud Evaluation
3.3. Abdominal Perimeter Measurements
4. Discussion and Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| LiDAR | Light Detection and Ranging o Laser Imaging Detection and Ranging |
| SfM | Structure from motion |
References
- Santos, D.A.; Dawson, J.A.; Matias, C.N.; Rocha, P.M.; Minderico, C.S.; Allison, D.B.; Sardinha, L.B.; Silva, A.M. Reference values for body composition and anthropometric measurements in athletes. PLoS ONE 2014, 9, e97846. [Google Scholar] [CrossRef] [PubMed]
- Maessen, M.F.; Eijsvogels, T.M.; Verheggen, R.J.; Hopman, M.T.; Verbeek, A.L.; de Vegt, F. Entering a new era of body indices: The feasibility of a body shape index and body roundness index to identify cardiovascular health status. PLoS ONE 2014, 9, e107212. [Google Scholar] [CrossRef] [PubMed]
- Zakaria, N.; Gupta, D. Anthropometry, Apparel Sizing and Design; Woodhead Publishing: Sawston, UK, 2019. [Google Scholar]
- Schmitz, A.; Gäbel, H.; Weiss, H.; Schmitt, O. Anthropometric 3D-body scanning in idiopathic scoliosis. Zeitschrift fur Orthopadie und ihre Grenzgebiete 2002, 140, 632. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Obesity: Preventing and Managing the Global Epidemic; World Health Organization: Geneva, Switzerland, 2000. [Google Scholar]
- Ruderman, A.; Pérez, L.O.; Adhikari, K.; Navarro, P.; Ramallo, V.; Gallo, C.; Poletti, G.; Bedoya, G.; Bortolini, M.C.; Acuña-Alonzo, V.; et al. Obesity, genomic ancestry, and socioeconomic variables in Latin American mestizos. Am. J. Hum. Biol. 2019, 31, e23278. [Google Scholar] [CrossRef]
- Navarro, P.; Ramallo, V.; Cintas, C.; Ruderman, A.; de Azevedo, S.; Paschetta, C.; Pérez, O.; Pazos, B.; Delrieux, C.; González-José, R. Body shape: Implications in the study of obesity and related traits. Am. J. Human Biol. 2020, 32, e23323. [Google Scholar] [CrossRef]
- Anguelov, D.; Srinivasan, P.; Koller, D.; Thrun, S.; Rodgers, J.; Davis, J. SCAPE: Shape Completion and Animation of People. ACM Trans. Graph. 2005, 24, 408–416. [Google Scholar] [CrossRef]
- Ulijaszek, S.J.; Kerr, D.A. Anthropometric measurement error and the assessment of nutritional status. Br. J. Nutr. 1999, 82, 165–177. [Google Scholar] [CrossRef]
- Gordon, C.C.; Bradtmiller, B. Interobserver error in a large scale anthropometric survey. Am. J. Hum. Biol. 1992, 4, 253–263. [Google Scholar] [CrossRef]
- Perini, T.A.; de Oliveira, G.L.; Ornellas, J.d.S.; de Oliveira, F.P. Technical error of measurement in anthropometry. Rev. Bras. Med. Esporte 2005, 11, 81–85. [Google Scholar] [CrossRef]
- Grellety, E.; Golden, M.H. The effect of random error on diagnostic accuracy illustrated with the anthropometric diagnosis of malnutrition. PLoS ONE 2016, 11, e0168585. [Google Scholar] [CrossRef]
- Krishan, K.; Kanchan, T. Measurement error in anthropometric studies and its significance in forensic casework. Ann. Med. Health Sci. Res. 2016, 6, 62. [Google Scholar] [CrossRef]
- Daniell, N.; Olds, T.; Tomkinson, G. Volumetric differences in body shape among adults with differing body mass index values: An analysis using three-dimensional body scans. Am. J. Hum. Biol. 2014, 26, 156–163. [Google Scholar] [CrossRef] [PubMed]
- Jaeschke, L.; Steinbrecher, A.; Pischon, T. Measurement of waist and hip circumference with a body surface scanner: Feasibility, validity, reliability, and correlations with markers of the metabolic syndrome. PLoS ONE 2015, 10, e0119430. [Google Scholar] [CrossRef] [PubMed]
- Medina-Inojosa, J.; Somers, V.K.; Ngwa, T.; Hinshaw, L.; Lopez-Jimenez, F. Reliability of a 3D body scanner for anthropometric measurements of central obesity. Obes. Open Access 2016, 2. [Google Scholar] [CrossRef]
- Mikić, I.; Trivedi, M.; Hunter, E.; Cosman, P. Human body model acquisition and tracking using voxel data. Int. J. Comput. Vis. 2003, 53, 199–223. [Google Scholar] [CrossRef]
- Balan, A.O.; Sigal, L.; Black, M.J.; Davis, J.E.; Haussecker, H.W. Detailed human shape and pose from images. In Proceedings of the 2007 IEEE Conference on Computer Vision and Pattern Recognition, Minneapolis, MN, USA, 17–22 June 2007; pp. 1–8. [Google Scholar]
- Ng, B.; Hinton, B.; Fan, B.; Kanaya, A.; Shepherd, J. Clinical anthropometrics and body composition from 3D whole-body surface scans. Eur. J. Clin. Nutr. 2016, 70, 1265–1270. [Google Scholar] [CrossRef]
- Chiu, C.Y.; Pease, D.L.; Fawkner, S.; Sanders, R.H. Automated body volume acquisitions from 3D structured-light scanning. Comput. Biol. Med. 2018, 101, 112–119. [Google Scholar] [CrossRef]
- Saba, M.; Sorrentino, F.; Muntoni, A.; Casti, S.; Cherchi, G.; Carcangiu, A.; Corda, F.; Murru, A.; Spano, L.D.; Scateni, R.; et al. A Seamless Pipeline for the Acquisition of the Body Shape: The Virtuoso Case Study. In Proceedings of the Eurographics Italian Chapter Conference, Catania, Italy, 11–12 September 2017; pp. 71–80. [Google Scholar]
- Westoby, M.J.; Brasington, J.; Glasser, N.F.; Hambrey, M.J.; Reynolds, J.M. ‘Structure-from-Motion’ photogrammetry: A low-cost, effective tool for geoscience applications. Geomorphology 2012, 179, 300–314. [Google Scholar] [CrossRef]
- Genchi, S.A.; Vitale, A.J.; Perillo, G.M.; Delrieux, C.A. Structure-from-motion approach for characterization of bioerosion patterns using UAV imagery. Sensors 2015, 15, 3593–3609. [Google Scholar] [CrossRef]
- Paschetta, C.; Ramallo, V.; Teodoroff, T.; Navarro, P.; Pazos, B.; Trujillo Jiménez, M.A.; Morales, L.; Pérez, O.; De Azevedo, S.; González-José, R. RAICES: Una Experiencia de Muestreo Patagónico, 1st ed.; Libro de Resúmenes de las Decimocuartas Jornadas Nacionales de Antropología Biológica: Buenos Aires, Argentina, 2019; Volume 1. (In Spanish) [Google Scholar]
- Wei, X.S.; Xie, C.W.; Wu, J.; Shen, C. Mask-CNN: Localizing parts and selecting descriptors for fine-grained bird species categorization. Patter Recognit. 2018, 76, 704–714. [Google Scholar] [CrossRef]
- De Brabandere, B.; Neven, D.; Van Gool, L. Semantic instance segmentation with a discriminative loss function. arXiv 2017, arXiv:1708.02551. [Google Scholar]
- Lin, T.Y.; Maire, M.; Belongie, S.; Hays, J.; Perona, P.; Ramanan, D.; Dollár, P.; Zitnick, C.L. Microsoft coco: Common objects in context. In Proceedings of the European Conference on Computer Vision, Zürich, Switzerland, 6–12 September 2014; pp. 740–755. [Google Scholar]
- Dutta, A.; Gupta, A.; Zissermann, A. VGG Image Annotator (VIA). 2016. Available online: http://www.robots.ox.ac.uk/vgg/software/via (accessed on 1 September 2020).
- Wu, C. VisualSFM: A Visual Structure from Motion System. 2011. Available online: http://www.cs.washington.edu/homes/ccwu/vsfm (accessed on 1 September 2020).
- Cignoni, P.; Callieri, M.; Corsini, M.; Dellepiane, M.; Ganovelli, F.; Ranzuglia, G. Meshlab: An open-source mesh processing tool. In Proceedings of the Eurographics Italian Chapter Conference, Salerno, Italy, 2–4 July 2008; Volume 2008, pp. 129–136. [Google Scholar]
- Girardeau-Montaut, D. CloudCompare. 2016. Available online: http://pcp2019.ifp.uni-stuttgart.de/presentations/04-CloudCompare_PCP_2019_public.pdf (accessed on 1 September 2020).
- Halır, R.; Flusser, J. Numerically stable direct least squares fitting of ellipses. In Proceedings of the 6th International Conference in Central Europe on Computer Graphics and Visualization (WSCG), Citeseer, Bory, Czech Republic, 9–13 February 1998; Volume 98, pp. 125–132. [Google Scholar]
- Utkualp, N.; Ercan, I. Anthropometric measurements usage in medical sciences. BioMed Res. Int. 2015, 2015, 404261. [Google Scholar] [CrossRef]






| Measure | Mean | Standard Deviation | Min | Max | ||||
|---|---|---|---|---|---|---|---|---|
| BRemNet | Mask R-CNN | BRemNet | Mask R-CNN | BRemNet | Mask R-CNN | BRemNet | Mask R-CNN | |
| Hamming loss | 0.04149 | 0.04734 | 0.00559 | 0.00693 | 0.03428 | 0.03889 | 0.05578 | 0.06111 |
| Jaccard | 0.86457 | 0.84577 | 0.01913 | 0.02830 | 0.83373 | 0.80468 | 0.90798 | 0.89994 |
| F-measure | 0.92726 | 0.91620 | 0.01096 | 0.01655 | 0.90933 | 0.89177 | 0.95177 | 0.94733 |
| Accuracy | 0.95851 | 0.95266 | 0.00559 | 0.00693 | 0.94422 | 0.93889 | 0.96572 | 0.96111 |
| FPR | 0.02929 | 0.03226 | 0.04308 | 0.04979 | 0.03844 | 0.03938 | 0.03918 | 0.04252 |
| FNR | 0.07050 | 0.08385 | 0.09068 | 0.11291 | 0.05415 | 0.06017 | 0.06929 | 0.08992 |
| Video 1 | Video 2 | Video 3 | Video 4 | Video 5 | Video 6 | Video 7 | Video 8 | Video 9 | Video 10 | |
|---|---|---|---|---|---|---|---|---|---|---|
| min Jaccard | 0.6000 | 0.6606 | 0.7035 | 0.5057 | 0.5942 | 0.5967 | 0.3723 | 0.6195 | 0.4757 | 0.4541 |
| Jaccard < 0.8 | 25.57% | 11.11% | 3.23% | 29.73% | 12.24% | 30.43% | 21.43% | 21.16% | 34.56% | 38.57% |
| max FN | 110,746 | 67,498 | 77,635 | 96,063 | 254,296 | 59,932 | 152,988 | 59,032 | 87,076 | 84,309 |
| Measure | Mean | Standard Deviation | Min | Max | ||||
|---|---|---|---|---|---|---|---|---|
| BRemNet | Mask R-CNN | BRemNet | Mask R-CNN | BRemNet | Mask R-CNN | BRemNet | Mask R-CNN | |
| Jaccard | 0.73543 | 0.28999 | 0.11578 | 0.13299 | 0.51644 | 0.06965 | 0.84800 | 0.45189 |
| F-measure | 0.84264 | 0.43349 | 0.08188 | 0.17417 | 0.68112 | 0.13023 | 0.91775 | 0.62249 |
| FPR | 0.15062 | 0.50837 | 0.17849 | 0.50000 | 0.10318 | 0.56022 | 0.13559 | 0.51065 |
| FNR | 0.15077 | 0.63428 | 0.17222 | 0.74144 | 0.09727 | 0.74409 | 0.13376 | 0.73075 |
| Mean Error (cm) | Standard Deviation (cm) | |||
|---|---|---|---|---|
| Hip | Waist | Hip | Waist | |
| LiDAR-based meshes | 7.935 | 0.910 | 6.864 | 1.808 |
| Unsegmented point clouds. | 271.708 | 302.718 | 87.375 | 123.548 |
| BRemNet-segmented point clouds | 6.701 | 4.128 | 4.419 | 3.148 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trujillo-Jiménez, M.A.; Navarro, P.; Pazos, B.; Morales, L.; Ramallo, V.; Paschetta, C.; De Azevedo, S.; Ruderman, A.; Pérez, O.; Delrieux, C.; et al. body2vec: 3D Point Cloud Reconstruction for Precise Anthropometry with Handheld Devices. J. Imaging 2020, 6, 94. https://doi.org/10.3390/jimaging6090094
Trujillo-Jiménez MA, Navarro P, Pazos B, Morales L, Ramallo V, Paschetta C, De Azevedo S, Ruderman A, Pérez O, Delrieux C, et al. body2vec: 3D Point Cloud Reconstruction for Precise Anthropometry with Handheld Devices. Journal of Imaging. 2020; 6(9):94. https://doi.org/10.3390/jimaging6090094
Chicago/Turabian StyleTrujillo-Jiménez, Magda Alexandra, Pablo Navarro, Bruno Pazos, Leonardo Morales, Virginia Ramallo, Carolina Paschetta, Soledad De Azevedo, Anahí Ruderman, Orlando Pérez, Claudio Delrieux, and et al. 2020. "body2vec: 3D Point Cloud Reconstruction for Precise Anthropometry with Handheld Devices" Journal of Imaging 6, no. 9: 94. https://doi.org/10.3390/jimaging6090094
APA StyleTrujillo-Jiménez, M. A., Navarro, P., Pazos, B., Morales, L., Ramallo, V., Paschetta, C., De Azevedo, S., Ruderman, A., Pérez, O., Delrieux, C., & Gonzalez-José, R. (2020). body2vec: 3D Point Cloud Reconstruction for Precise Anthropometry with Handheld Devices. Journal of Imaging, 6(9), 94. https://doi.org/10.3390/jimaging6090094







