Detection of HER2 from Haematoxylin-Eosin Slides Through a Cascade of Deep Learning Classifiers via Multi-Instance Learning
Abstract
1. Introduction
2. Materials and Methods
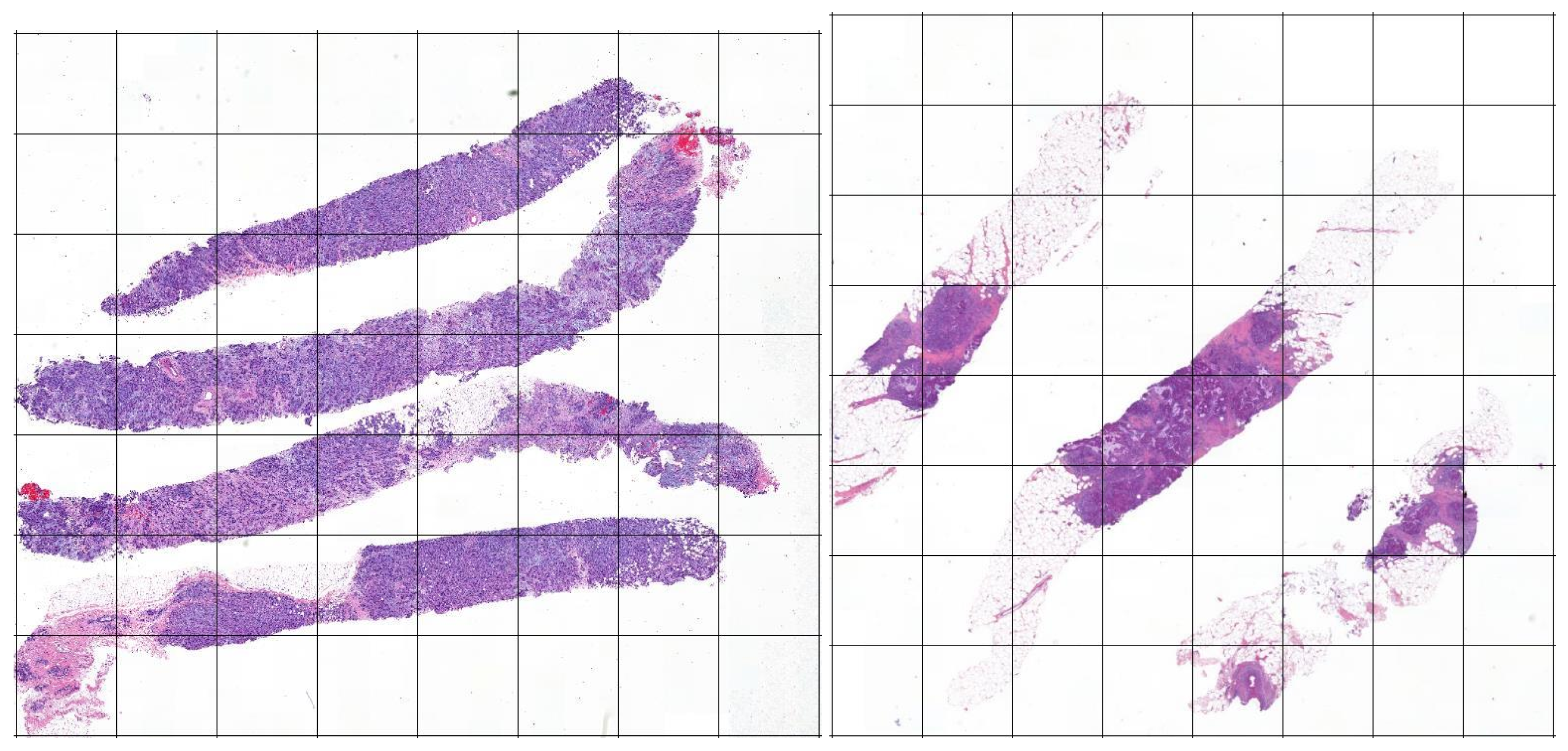
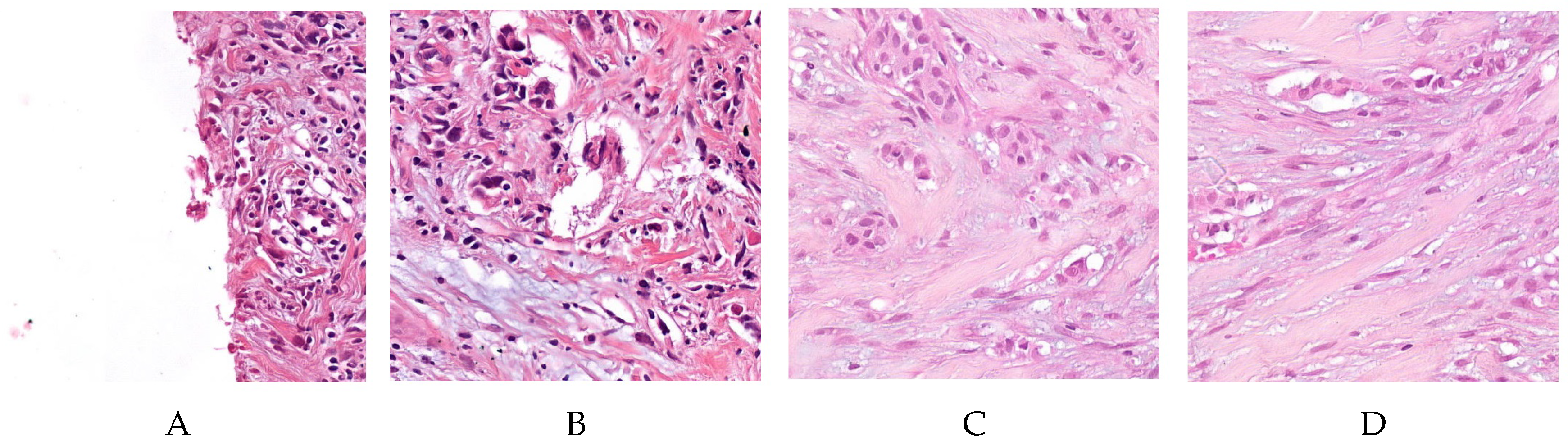
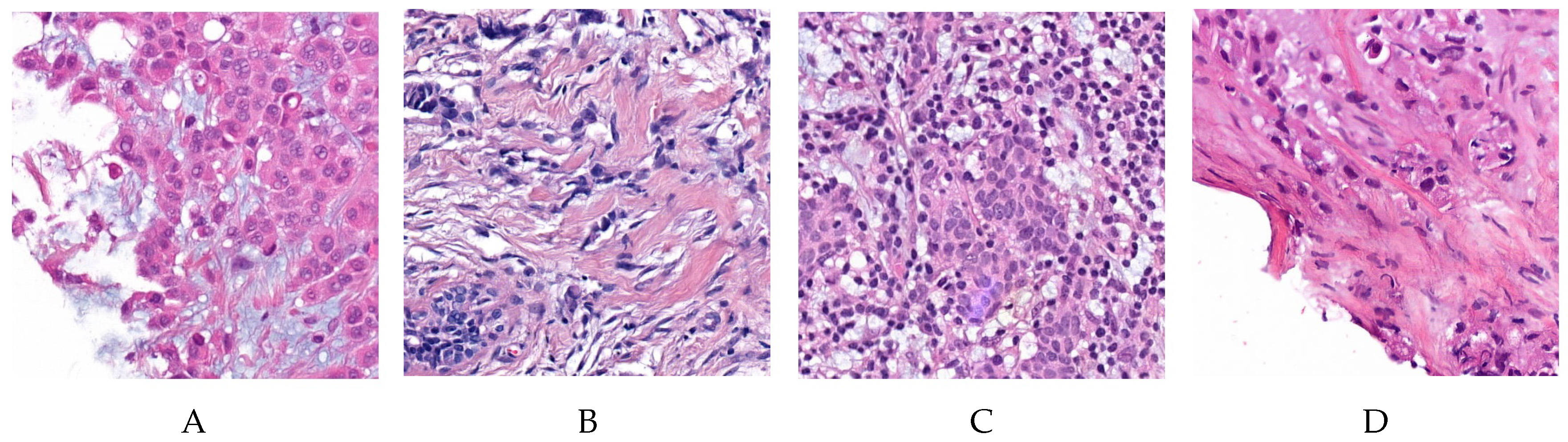
2.1. Dataset and Materials
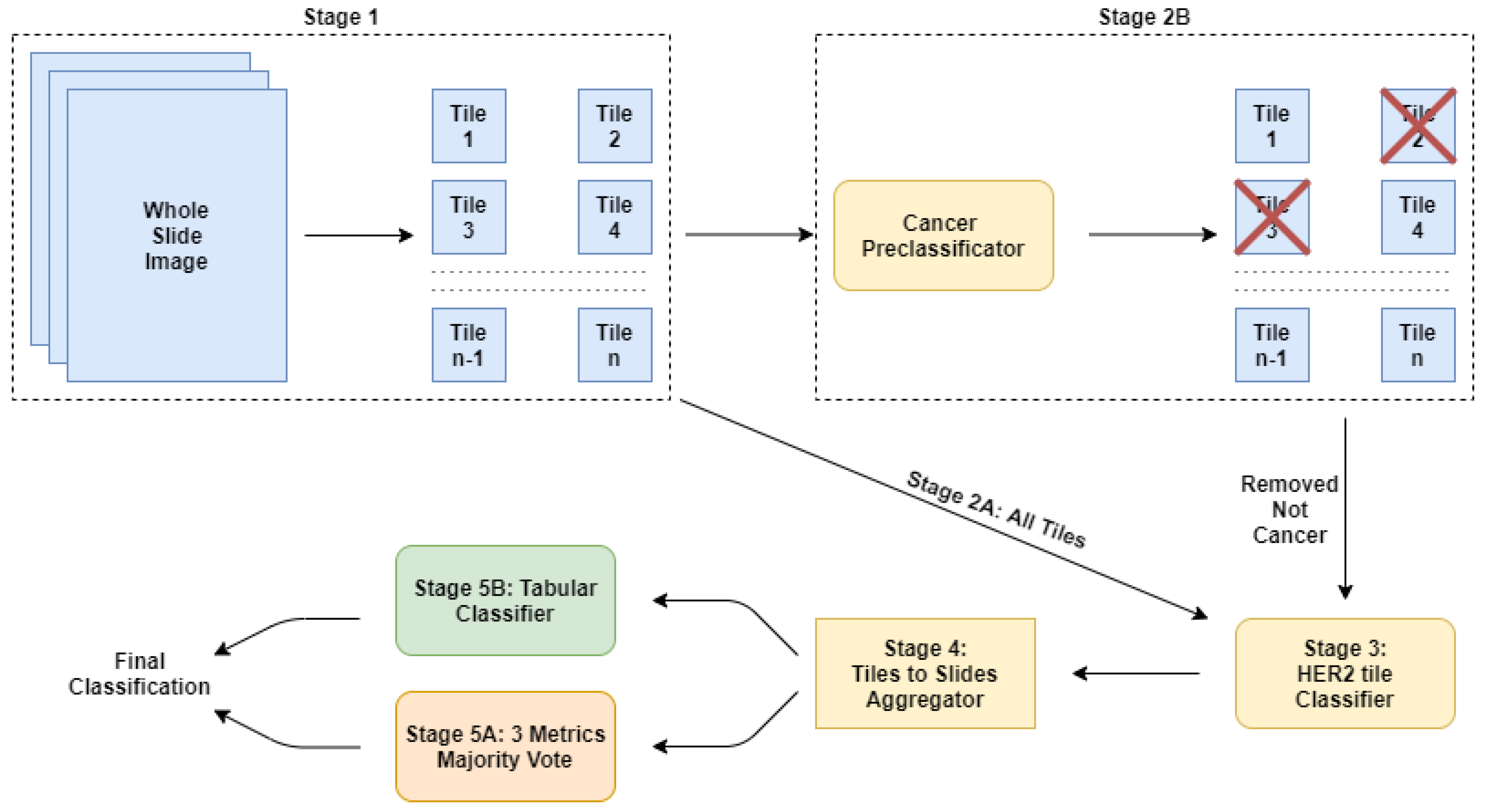
2.2. Our Approach: A Cascade of Deep Learning Classifiers
2.2.1. Stage 1: The Tiles Extractor
2.2.2. Stage 2A: Using All the Tiles
2.2.3. Stage 2B: Using a Subset of Tiles
2.2.4. Stage 3: The Main Classifier
2.2.5. Stage 4: Computing the Features
2.2.6. Stage 5A: Post-Trainer Aggregation Using Majority Vote
2.2.7. Stage 5B: Post-Trainer Aggregation Using a Tabular Learner
3. Results
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CNN | Convolutional Neural Network |
| DL | Deep Learning |
| HER2 | Human Epidermal growth factor Receptor 2 |
| MIL | Multiple Instance Learning |
| ML | Machine Learning |
| WSI | Whole Slide Image |
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA A Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Wolff, A.C.; Hammond, M.E.H.; Allison, K.H.; Harvey, B.E.; Mangu, P.B.; Bartlett, J.M.S.; Bilous, M.; Ellis, I.O.; Fitzgibbons, P.; Hanna, W.; et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. Arch. Pathol. Lab. Med. 2018, 142, 1364–1382. [Google Scholar] [CrossRef] [PubMed]
- Dean-Colomb, W.; Esteva, F.J. Her2-positive breast cancer: Herceptin and beyond. Eur. J. Cancer 2008, 44, 2806–2812. [Google Scholar] [CrossRef] [PubMed]
- Nahta, R.; Esteva, F.J. HER-2-Targeted Therapy. Clin. Cancer Res. 2003, 9, 5078–5084. [Google Scholar]
- Gutierrez, C.; Schiff, R. HER2: Biology, detection, and clinical implications. Arch. Pathol. Lab. Med. 2011, 135, 55–62. [Google Scholar]
- Al-Khafaji, Q.; Harris, M.; Tombelli, S.; Laschi, S.; Turner, A.; Mascini, M.; Marrazza, G. An electrochemical immunoassay for HER2 detection. Electroanalysis 2012, 24, 735–742. [Google Scholar] [CrossRef]
- Riethdorf, S.; Müller, V.; Zhang, L.; Rau, T.; Loibl, S.; Komor, M.; Roller, M.; Huober, J.; Fehm, T.; Schrader, I. Detection and HER2 expression of circulating tumor cells: Prospective monitoring in breast cancer patients treated in the neoadjuvant GeparQuattro trial. Clin. Cancer Res. 2010, 16, 2634–2645. [Google Scholar] [CrossRef]
- Gevensleben, H.; Garcia-Murillas, I.; Graeser, M.K.; Schiavon, G.; Osin, P.; Parton, M.; Smith, I.E.; Ashworth, A.; Turner, N.C. Noninvasive detection of HER2 amplification with plasma DNA digital PCR. Clin. Cancer Res. 2013, 19, 3276–3284. [Google Scholar] [CrossRef]
- Gohring, J.T.; Dale, P.S.; Fan, X. Detection of HER2 breast cancer biomarker using the opto-fluidic ring resonator biosensor. Sens. Actuators B Chem. 2010, 146, 226–230. [Google Scholar] [CrossRef]
- Vandenberghe, M.E.; Scott, M.L.; Scorer, P.W.; Söderberg, M.; Balcerzak, D.; Barker, C. Relevance of deep learning to facilitate the diagnosis of HER2 status in breast cancer. Sci. Rep. 2017, 7, 45938. [Google Scholar] [CrossRef]
- Yaziji, H.; Goldstein, L.C.; Barry, T.S.; Werling, R.; Hwang, H.; Ellis, G.K.; Gralow, J.R.; Livingston, R.B.; Gown, A.M. HER-2 testing in breast cancer using parallel tissue-based methods. JAMA 2004, 291, 1972–1977. [Google Scholar] [CrossRef] [PubMed]
- Basavanhally, A.N.; Ganesan, S.; Agner, S.; Monaco, J.P.; Feldman, M.D.; Tomaszewski, J.E.; Bhanot, G.; Madabhushi, A. Computerized image-based detection and grading of lymphocytic infiltration in HER2+ breast cancer histopathology. IEEE Trans. Biomed. Eng. 2009, 57, 642–653. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Roa, A.; Gilmore, H.; Basavanhally, A.; Feldman, M.; Ganesan, S.; Shih, N.N.; Tomaszewski, J.; González, F.A.; Madabhushi, A. Accurate and reproducible invasive breast cancer detection in whole-slide images: A Deep Learning approach for quantifying tumor extent. Sci. Rep. 2017, 7, 46450. [Google Scholar] [CrossRef]
- Ravì, D.; Wong, C.; Deligianni, F.; Berthelot, M.; Andreu-Perez, J.; Lo, B.; Yang, G.Z. Deep learning for health informatics. IEEE J. Biomed. Health Inform. 2016, 21, 4–21. [Google Scholar] [CrossRef] [PubMed]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Shen, D.; Wu, G.; Suk, H.I. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 2017, 19, 221–248. [Google Scholar] [CrossRef]
- Zha, Z.J.; Hua, X.S.; Mei, T.; Wang, J.; Qi, G.J.; Wang, Z. Joint multi-label multi-instance learning for image classification. In Proceedings of the 2008 IEEE Conference on Computer Vision and Pattern Recognition, Anchorage, AK, USA, 23–28 June 2008; pp. 1–8. [Google Scholar]
- Zhou, Z.H.; Xu, J.M. On the relation between multi-instance learning and semi-supervised learning. In Proceedings of the 24th International Conference on Machine Learning, Corvallis, OR, USA, 20–24 June 2007; pp. 1167–1174. [Google Scholar]
- Zhou, Z.H.; Zhang, M.L. Neural networks for multi-instance learning. In Proceedings of the International Conference on Intelligent Information Technology, Beijing, China, 22–25 September 2002; pp. 455–459. [Google Scholar]
- Zhou, Z.H. Multi-instance learning: A survey. Dep. Comput. Sci. Technol. Nanjing Univ. Tech. Rep. 2004, 2. [Google Scholar]
- Zhou, Z.H.; Zhang, M.L. Ensembles of multi-instance learners. In Proceedings of the 14th European Conference on Machine Learning, Cavtat-Dubrovnik, Croatia, 22–26 September 2003; Springer: Berlin/Heidelberg, Germany, 2003; pp. 492–502. [Google Scholar]
- Sze, V.; Chen, Y.H.; Emer, J.; Suleiman, A.; Zhang, Z. Hardware for machine learning: Challenges and opportunities. In Proceedings of the 2017 IEEE Custom Integrated Circuits Conference (CICC), Austin, TX, USA, 30 April–3 May 2017; pp. 1–8. [Google Scholar]
- Carbonneau, M.A.; Cheplygina, V.; Granger, E.; Gagnon, G. Multiple Instance Learning: A Survey of Problem Characteristics and Applications. Pattern Recognit. 2016. [Google Scholar] [CrossRef]
- Herrera, F.; Ventura, S.; Bello, R.; Cornelis, C.; Zafra, A.; Sanchez Tarrago, D.; Vluymans, S. Multiple Instance Learning. Foundations and Algorithms; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar] [CrossRef]
- Foulds, J.; Frank, E. A Review of Multi-Instance Learning Assumptions. Knowl. Eng. Rev. 2010, 25. [Google Scholar] [CrossRef]
- Campanella, G.; Hanna, M.; Geneslaw, L.; Miraflor, A.; Silva, V.; Busam, K.; Brogi, E.; Reuter, V.; Klimstra, D.; Fuchs, T. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 2019, 25, 1. [Google Scholar] [CrossRef]
- Sudharshan, P.J.; Petitjean, C.; Spanhol, F.; Oliveira, L.; Heutte, L.; Honeine, P. Multiple Instance Learning for Histopathological Breast Cancer Image Classification. Expert Syst. Appl. 2018. [Google Scholar] [CrossRef]
- Couture, H.D.; Williams, L.A.; Geradts, J.; Nyante, S.J.; Butler, E.N.; Marron, J.S.; Perou, C.M.; Troester, M.A.; Niethammer, M. Image analysis with deep learning to predict breast cancer grade, ER status, histologic subtype, and intrinsic subtype. NPJ Breast Cancer 2018. [Google Scholar] [CrossRef]
- Qaiser, T.; Mukherjee, A.; Reddy Pb, C.; Munugoti, S.D.; Tallam, V.; Pitkäaho, T.; Lehtimäki, T.; Naughton, T.; Berseth, M.; Pedraza, A.; et al. HER2 challenge contest: A detailed assessment of automated HER2 scoring algorithms in whole slide images of breast cancer tissues. Histopathology 2017, 72, 227–238. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, M.R.; Teerapakpinyo, C.; Shuangshoti, S.; Keelawat, S. Comparison between digital image analysis and visual assessment of immunohistochemical HER2 expression in breast cancer. Pathol.-Res. Pract. 2018, 214, 2087–2092. [Google Scholar] [CrossRef] [PubMed]
- Mukundan, R. Analysis of Image Feature Characteristics for Automated Scoring of HER2 in Histology Slides. J. Imaging 2019, 5, 35. [Google Scholar] [CrossRef]
- Qaiser, T.; Rajpoot, N.M. Learning Where to See: A Novel Attention Model for Automated Immunohistochemical Scoring. IEEE Trans. Med. Imaging 2019, 38, 2620–2631. [Google Scholar] [CrossRef]
- Yim, K.; Park, H.S.; Kim, D.M.; Lee, Y.S.; Lee, A. Image Analysis of HER2 Immunohistochemical Staining of Surgical Breast Cancer Specimens. Yonsei Med. J. 2019, 60, 158. [Google Scholar] [CrossRef]
- Li, A.C.; Zhao, J.; Zhao, C.; Ma, Z.; Hartage, R.; Zhang, Y.; Li, X.; Parwani, A.V. Quantitative digital imaging analysis of HER2 immunohistochemistry predicts the response to anti-HER2 neoadjuvant chemotherapy in HER2-positive breast carcinoma. Breast Cancer Res. Treat. 2020, 180, 321–329. [Google Scholar] [CrossRef]
- Güneş, A.; Kalkan, H.; Durmuş, E. Optimizing the color-to-grayscale conversion for image classification. Signal Image Video Process. 2016, 10, 853–860. [Google Scholar] [CrossRef]
- Della Mea, V.; Pilutti, D. Classification of Histologic Images Using a Single Staining: Experiments With Deep Learning on Deconvolved Images. Stud. Health Technol. Inform. 2020, 270, 1223–1224. [Google Scholar]
- Iandola, F.; Moskewicz, M.; Karayev, S.; Girshick, R.; Darrell, T.; Keutzer, K. Densenet: Implementing efficient convnet descriptor pyramids. arXiv 2014, arXiv:1404.1869. [Google Scholar]
- Yu, X.; Zeng, N.; Liu, S.; Zhang, Y.D. Utilization of DenseNet201 for diagnosis of breast abnormality. Mach. Vis. Appl. 2019, 30, 1135–1144. [Google Scholar] [CrossRef]
- Yilmaz, F.; Kose, O.; Demir, A. Comparison of two different deep learning architectures on breast cancer. In Proceedings of the 2019 Medical Technologies Congress (TIPTEKNO), Izmir, Turkey, 3–5 October 2019; pp. 1–4. [Google Scholar]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar]
- Aresta, G.; Araújo, T.; Kwok, S.; Chennamsetty, S.S.; Safwan, M.; Alex, V.; Marami, B.; Prastawa, M.; Chan, M.; Donovan, M. Bach: Grand challenge on breast cancer histology images. Med. Image Anal. 2019, 56, 122–139. [Google Scholar] [CrossRef] [PubMed]
- Della Mea, V.; Baroni, G.L.; Pilutti, D.; Loreto, C.D. SlideJ: An ImageJ plugin for automated processing of whole slide images. PLoS ONE 2017, 12, e0180540. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the 29th IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A.A. Inception-v4, inception-resnet and the impact of residual connections on learning. In Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017. [Google Scholar]
- Targ, S.; Almeida, D.; Lyman, K. Resnet in resnet: Generalizing residual architectures. arXiv 2016, arXiv:1603.08029. [Google Scholar]
- Wu, Z.; Shen, C.; Van Den Hengel, A. Wider or deeper: Revisiting the resnet model for visual recognition. Pattern Recognit. 2019, 90, 119–133. [Google Scholar] [CrossRef]
- Smith, L.N. A disciplined approach to neural network hyper-parameters: Part 1—Learning rate, batch size, momentum, and weight decay. arXiv 2018, arXiv:1803.09820. [Google Scholar]
- Nguyen, L.D.; Lin, D.; Lin, Z.; Cao, J. Deep CNNs for microscopic image classification by exploiting transfer learning and feature concatenation. In Proceedings of the 2018 IEEE International Symposium on Circuits and Systems (ISCAS), Florence, Italy, 27–30 May 2018; pp. 1–5. [Google Scholar]


| P | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ||
| ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ||
| All Tiles | Subset of Tiles | Best | ||||||
|---|---|---|---|---|---|---|---|---|
| Majority Vote | Tabular Classifier | Majority Vote | Tabular Classifier | HEROHE | ||||
| Accuracy | 0.687 | 0.673 | 0.667 | 0.707 | - | |||
| Precision | 0.570 | 0.560 | 0.580 | 0.603 | 0.5682 | |||
| Recall | 0.883 | 0.864 | 0.783 | 0.797 | 0.8333 | |||
| F1-Score | 0.693 | 0.680 | 0.667 | 0.687 | 0.6757 | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
La Barbera, D.; Polónia, A.; Roitero, K.; Conde-Sousa, E.; Della Mea, V. Detection of HER2 from Haematoxylin-Eosin Slides Through a Cascade of Deep Learning Classifiers via Multi-Instance Learning. J. Imaging 2020, 6, 82. https://doi.org/10.3390/jimaging6090082
La Barbera D, Polónia A, Roitero K, Conde-Sousa E, Della Mea V. Detection of HER2 from Haematoxylin-Eosin Slides Through a Cascade of Deep Learning Classifiers via Multi-Instance Learning. Journal of Imaging. 2020; 6(9):82. https://doi.org/10.3390/jimaging6090082
Chicago/Turabian StyleLa Barbera, David, António Polónia, Kevin Roitero, Eduardo Conde-Sousa, and Vincenzo Della Mea. 2020. "Detection of HER2 from Haematoxylin-Eosin Slides Through a Cascade of Deep Learning Classifiers via Multi-Instance Learning" Journal of Imaging 6, no. 9: 82. https://doi.org/10.3390/jimaging6090082
APA StyleLa Barbera, D., Polónia, A., Roitero, K., Conde-Sousa, E., & Della Mea, V. (2020). Detection of HER2 from Haematoxylin-Eosin Slides Through a Cascade of Deep Learning Classifiers via Multi-Instance Learning. Journal of Imaging, 6(9), 82. https://doi.org/10.3390/jimaging6090082







