Faster R-CNN-Based Glomerular Detection in Multistained Human Whole Slide Images
Abstract
1. Introduction
1.1. Detection of Glomeruli in Whole Slide Images
1.2. Digital Pathology Analysis with Deeply Multilayered Neural Network (DNN)
1.3. Previous Work
1.3.1. Hand-Crafted Feature-Based Methods
1.3.2. Convolutional Neural Network (CNN) Based Methods
1.4. Objective
- In glomerular detection from human WSIs, recent publications have reported that a CNN-based approach showed the F-measures 0.937 and a handcraft feature-based approaches (mrcLBP with SVM) showed 0.832 in PAS stain.
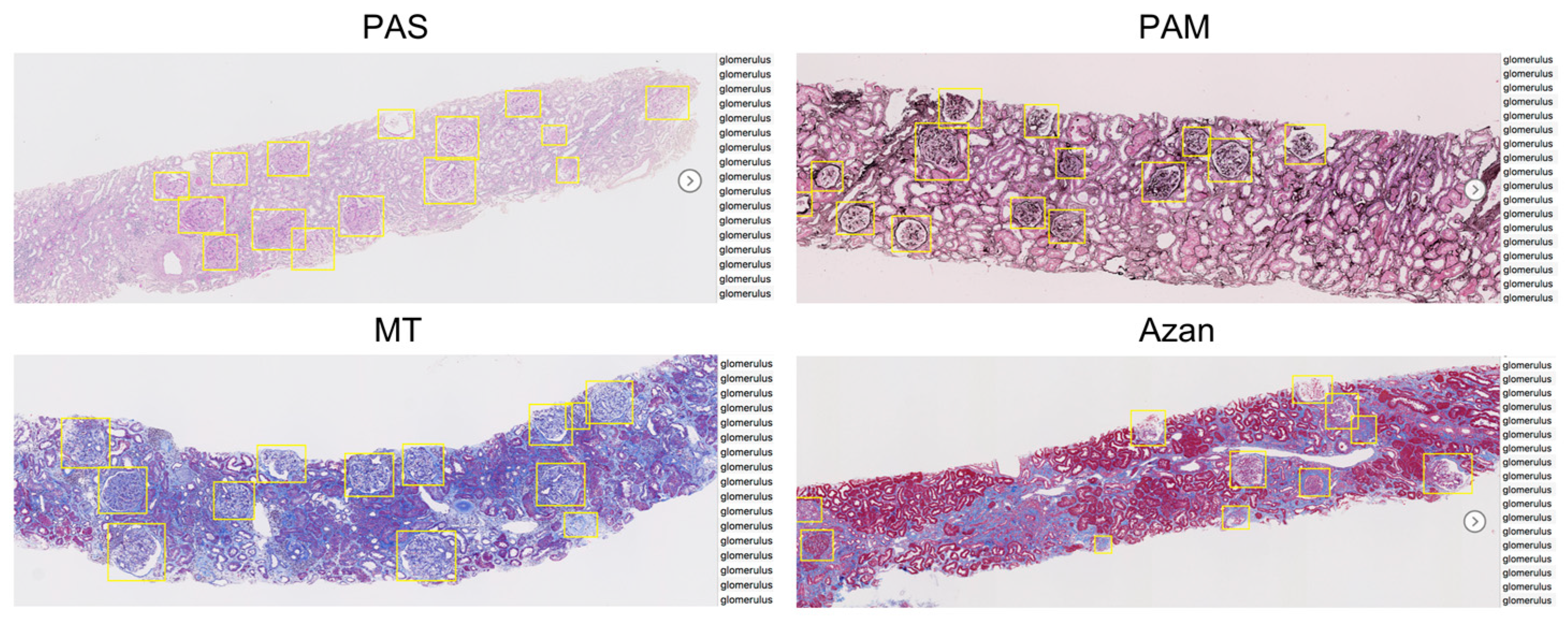
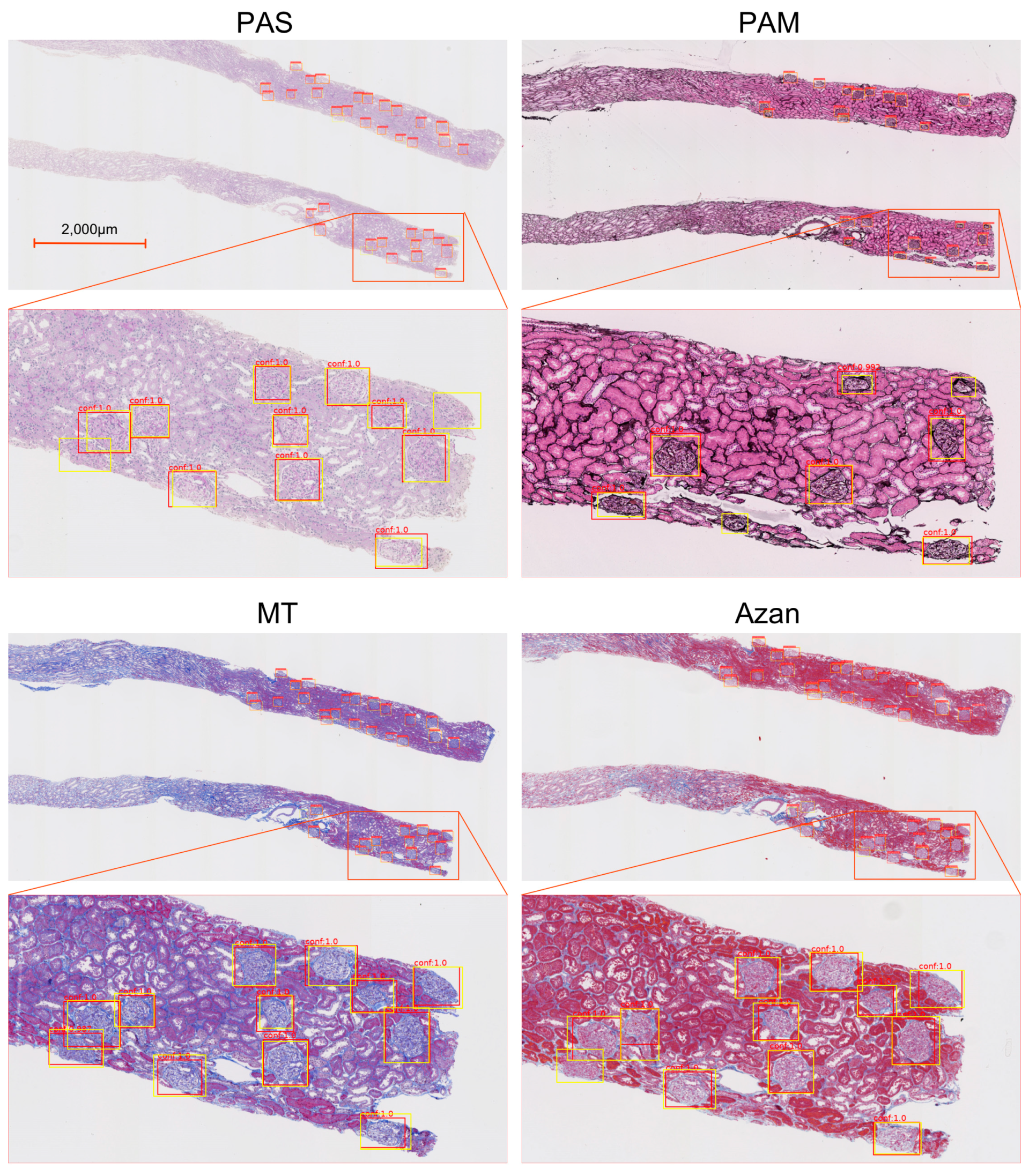
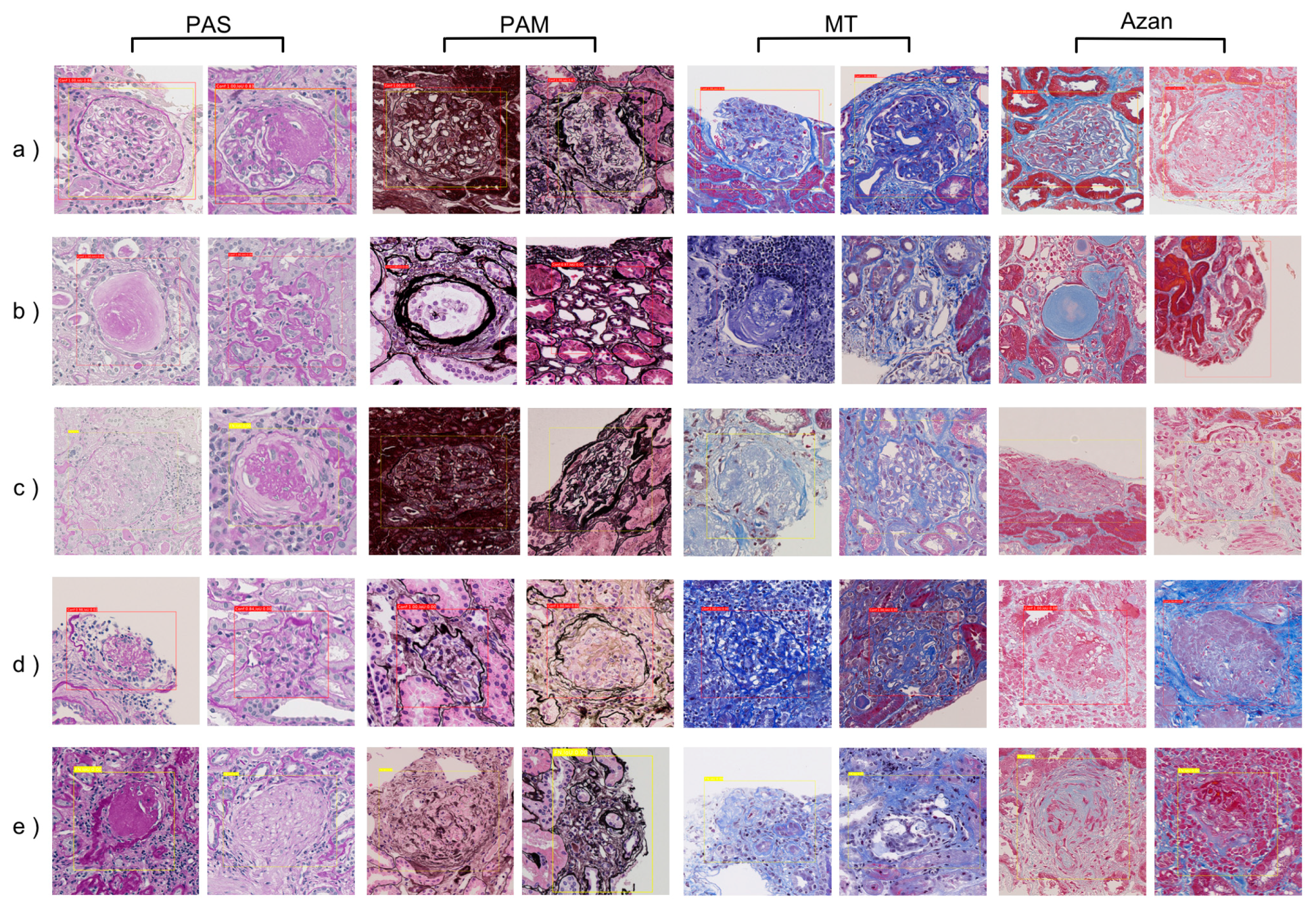
- Our approach based on a Faster-RCNN showed the F-measures 0.925 in PAS stain. It also showed equally high performance can be obtained not only for PAS stain but also for PAM (0.928), MT (0.898), and Azan (0.877) stains.
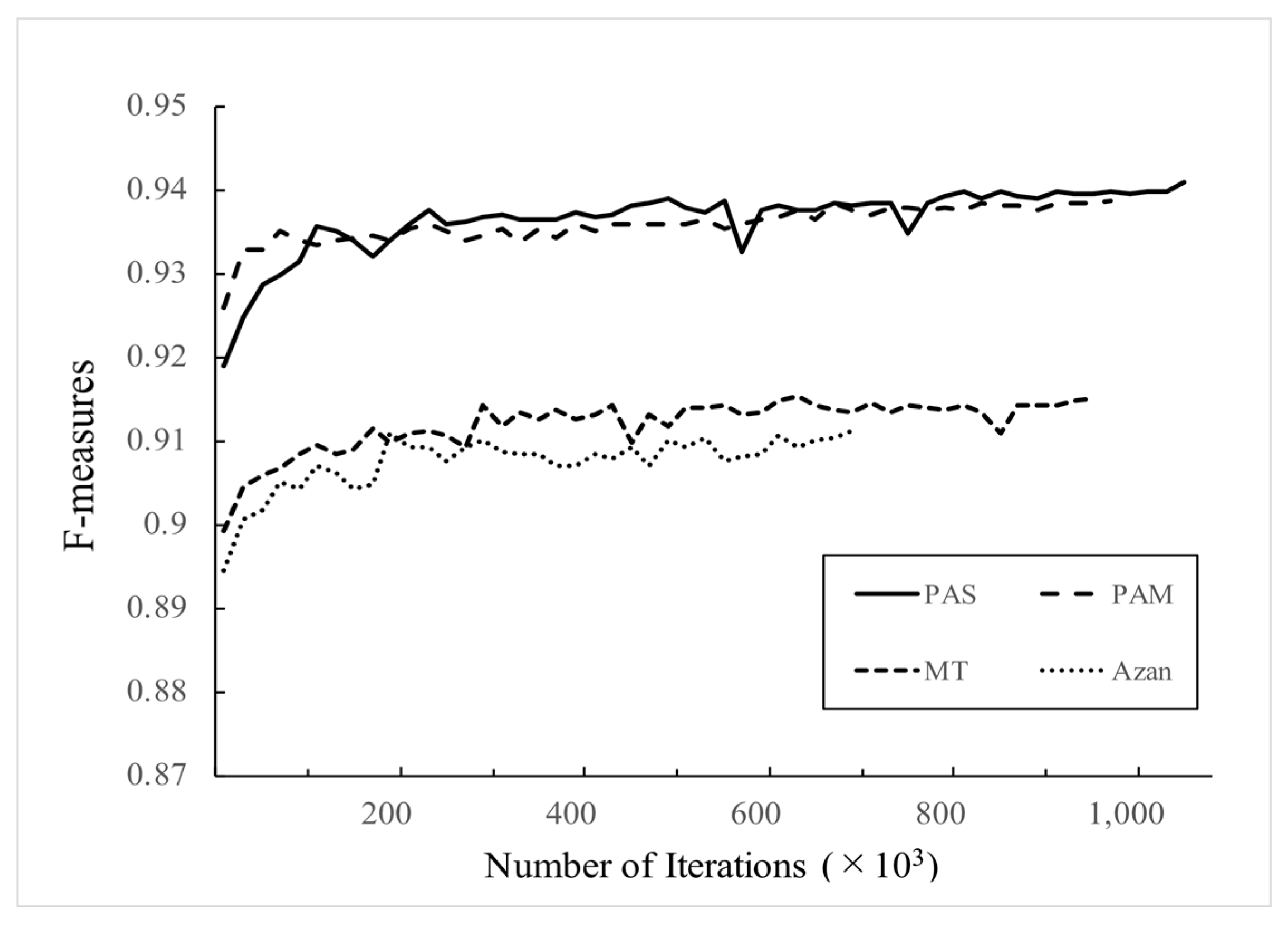
- As for the required number of WSIs used for the network training, the F-measures were saturated with 60 WSIs in PAM, MT, and Azan stains. However, it was not saturated with 120 WSIs in the PAS stain.
2. Materials and Methods
2.1. Datasets
2.2. Faster R-CNN
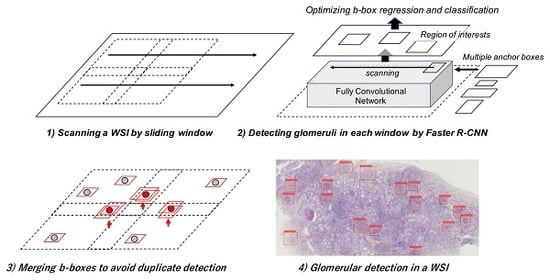
2.3. Glomerular Detection Process from Whole Slide Images (WSIs)
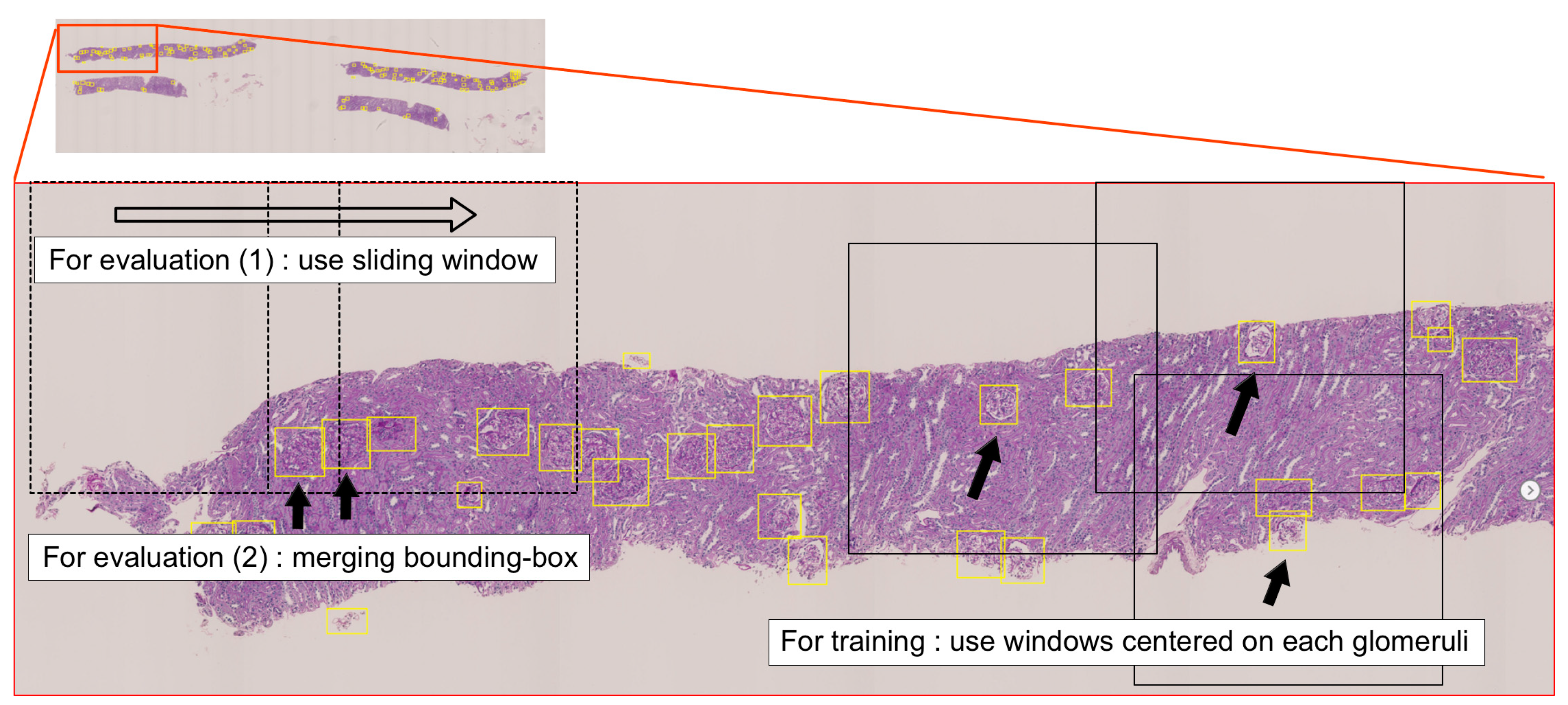
2.3.1. Sliding Window Method for WSIs
2.3.2. Evaluation Metrics
2.3.3. Faster R-CNN Training
2.4. Experimental Settings
3. Results
3.1. Detection Performance with Different Stains
3.2. Detection Performance Corresponding to the Number of WSIs to be Used for Training
3.3. Post-Evaluation
4. Discussion
4.1. Glomerular Detection Performance
4.2. Processing Speed
4.3. Quality Assessment of the Annotation
4.4. Mutual Utilization among Hospitals
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A
- In our experiment, we used the Tensorflow Object Detection API which is an open source object detection framework that is licensed under Apache License 2.0. An overview and usage of the Tensorflow Object Detection API is described in the following URL: (https://github.com/tensorflow/models/blob/master/research/object_detection/README.md)
- We also used a pre-trained model of Faster R-CNN with Inception-ResNet which had been trained on the COCO dataset. This pre-trained model can be downloaded from the following URL: (http://download.tensorflow.org/models/object_detection/faster_rcnn_inception_resnet_v2_atrous_coco_11_06_2017.tar.gz)
- To facilitate further research to build upon our results, the source code, network configurations, and the trained network-derived results are available at the following URLs. By using these materials, it is possible to perform glomerular detection on WSIs. We also provided a few WSIs and annotations to validate them: (https://github.com/jinseikenai/glomeruli_detection/blob/master/README.md; https://github.com/jinseikenai/glomeruli_detection/blob/master/config/glomerulus_model.config)
Appendix B
Appendix C
| PAS | Number of WSIs to Be Used for Training | ||
|---|---|---|---|
| 60 | 90 | 120 | |
| Training iterations | 780,000 | 640,000 | 1,060,000 |
| Confidence thresholds | 0.300 | 0.700 | 0.950 |
| F-measure | 0.907 | 0.905 | 0.925 |
| Precision | 0.921 | 0.916 | 0.931 |
| Recall | 0.894 | 0.896 | 0.919 |
| PAM | Number of WSIs to Be Used for Training | ||
|---|---|---|---|
| 60 | 90 | 120 | |
| Training iterations | 640,000 | 740,000 | 980,000 |
| Confidence thresholds | 0.950 | 0.900 | 0.975 |
| F-measure | 0.927 | 0.926 | 0.928 |
| Precision | 0.951 | 0.950 | 0.939 |
| Recall | 0.904 | 0.904 | 0.918 |
| MT | Number of WSIs to Be Used for Training | ||
|---|---|---|---|
| 60 | 90 | 120 | |
| Training iterations | 760,000 | 720,000 | 960,000 |
| Confidence thresholds | 0.925 | 0.975 | 0.950 |
| F-measure | 0.898 | 0.892 | 0.896 |
| Precision | 0.927 | 0.905 | 0.915 |
| Recall | 0.871 | 0.879 | 0.878 |
| Azan | Number of WSIs to Be Used for Training | ||
|---|---|---|---|
| 60 | 90 | 120 | |
| Training iterations | 560,000 | 420,000 | 680,000 |
| Confidence thresholds | 0.700 | 0.800 | 0.950 |
| F-measure | 0.877 | 0.876 | 0.876 |
| Precision | 0.892 | 0.892 | 0.904 |
| Recall | 0.863 | 0.860 | 0.849 |
References
- Gurcan, M.N.; Boucheron, L.E.; Can, A.; Madabhushi, A.; Rajpoot, N.M.; Yener, B. Histopathological Image Analysis: A Review. IEEE Rev. Biomed. Eng. 2009, 2, 147–171. [Google Scholar] [CrossRef] [PubMed]
- Pantanowitz, L.; Evans, A.; Pfeifer, J.; Collins, L.; Valenstein, P.; Kaplan, K.; Wilbur, D.; Colgan, T. Review of the current state of whole slide imaging in pathology. J. Pathol. Inform. 2011, 2, 36. [Google Scholar] [CrossRef] [PubMed]
- Madabhushi, A.; Lee, G. Image analysis and machine learning in digital pathology: Challenges and opportunities. Med. Image Anal. 2016, 33, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Adv. Neural Inf. Process. Syst. 2012, 1, 1097–1105. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv, 2014; arXiv:1409.1556. [Google Scholar]
- Hannun, A.; Case, C.; Casper, J.; Catanzaro, B.; Diamos, G.; Elsen, E.; Prenger, R.; Satheesh, S.; Sengupta, S.; Coates, A.; et al. DeepSpeech: Scaling up end-to-end speech recognition. arXiv, 2014; arXiv:1412.5567. [Google Scholar]
- Amodei, D.; Anubhai, R.; Battenberg, E.; Case, C.; Casper, J.; Catanzaro, B.; Chen, J.; Chrzanowski, M.; Coates, A.; Diamos, G.; et al. Deep Speech 2: End-to-End Speech Recognition in English and Mandarin. arXiv, 2015; arXiv:1512.02595. [Google Scholar]
- Devlin, J.; Zbib, R.; Huang, Z.; Lamar, T.; Schwartz, R.; Makhoul, J. Fast and Robust Neural Network Joint Models for Statistical Machine Translation. In Proceedings of the 52nd Annual Meeting of the Association for Computational Linguistics, Baltimore, MD, USA, 22–27 June 2014; Volume 1. [Google Scholar]
- Wu, Y.; Schuster, M.; Chen, Z.; Le, Q.V.; Norouzi, M.; Macherey, W.; Krikun, M.; Cao, Y.; Gao, Q.; Macherey, K.; et al. Google’s Neural Machine Translation System: Bridging the Gap between Human and Machine Translation. arXiv, 2016; arXiv:1609.08144. [Google Scholar]
- Janowczyk, A.; Madabhushi, A. Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases. J. Pathol. Inform. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.W.M.; van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, X.; Müller, H.; Zhang, S. Large-scale retrieval for medical image analytics: A comprehensive review. Med. Image Anal. 2018, 43, 66–84. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhang, M.; Zhou, Z.; Chu, J.; Cao, F. Automatic detection and classification of leukocytes using convolutional neural networks. Med. Biol. Eng. Comput. 2017, 55, 1287–1301. [Google Scholar] [CrossRef] [PubMed]
- Sirinukunwattana, K.; Raza, S.E.A.; Tsang, Y.W.; Snead, D.R.J.; Cree, I.A.; Rajpoot, N.M. Locality Sensitive Deep Learning for Detection and Classification of Nuclei in Routine Colon Cancer Histology Images. IEEE Trans. Med. Imaging 2016, 35, 1196–1206. [Google Scholar] [CrossRef] [PubMed]
- Roux, L.; Racoceanu, D.; Loménie, N.; Kulikova, M.; Irshad, H.; Klossa, J.; Capron, F.; Genestie, C.; Naour, G.; Gurcan, M. Mitosis detection in breast cancer histological images An ICPR 2012 contest. J. Pathol. Inform. 2013, 4, 8. [Google Scholar] [CrossRef] [PubMed]
- Ciresan, D.C.; Giusti, A.; Gambardella, L.M.; Schmidhuber, J. Mitosis Detection in Breast Cancer Histology Images using Deep Neural Networks. Med. Image Comput. Comput. Interv. 2013, 16, 411–418. [Google Scholar]
- Veta, M.; van Diest, P.J.; Willems, S.M.; Wang, H.; Madabhushi, A.; Cruz-Roa, A.; Gonzalez, F.; Larsen, A.B.L.; Vestergaard, J.S.; Dahl, A.B.; et al. Assessment of algorithms for mitosis detection in breast cancer histopathology images. Med. Image Anal. 2015, 20, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Kainz, P.; Pfeiffer, M.; Urschler, M. Semantic Segmentation of Colon Glands with Deep Convolutional Neural Networks and Total Variation Segmentation. arXiv, 2017; arXiv:1511.06919v2. [Google Scholar]
- Dalal, N.; Triggs, B. Histograms of oriented gradients for human detection. In Proceedings of the 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition, San Diego, CA, USA, 20–25 June 2005; Volume 1. [Google Scholar]
- Kakimoto, T.; Okada, K.; Hirohashi, Y.; Relator, R.; Kawai, M.; Iguchi, T.; Fujitaka, K.; Nishio, M.; Kato, T.; Fukunari, A.; et al. Automated image analysis of a glomerular injury marker desmin in spontaneously diabetic Torii rats treated with losartan. J. Endocrinol. 2014, 222, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Relator, R.; Ngouv, H.; Hirohashi, Y.; Takaki, O.; Kakimoto, T.; Okada, K. Segmental HOG: New descriptor for glomerulus detection in kidney microscopy image. BMC Bioinform. 2015, 16, 316. [Google Scholar] [CrossRef] [PubMed]
- Simon, O.; Yacoub, R.; Jain, S.; Tomaszewski, J.E.; Sarder, P. Multi-radial LBP Features as a Tool for Rapid Glomerular Detection and Assessment in Whole Slide Histopathology Images. Sci. Rep. 2018, 8, 2032. [Google Scholar] [CrossRef] [PubMed]
- Temerinac-Ott, M.; Forestier, G.; Schmitz, J.; Hermsen, M.; Braseni, J.H.; Feuerhake, F.; Wemmert, C. Detection of glomeruli in renal pathology by mutual comparison of multiple staining modalities. In Proceedings of the 10th International Symposium on Image and Signal Processing and Analysis, Ljubljana, Slovenia, 18–20 September 2017. [Google Scholar]
- Gallego, J.; Pedraza, A.; Lopez, S.; Steiner, G.; Gonzalez, L.; Laurinavicius, A.; Bueno, G. Glomerulus Classification and Detection Based on Convolutional Neural Networks. J. Imaging 2018, 4, 20. [Google Scholar] [CrossRef]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards Real-Time Object Detection with Region Proposal Networks. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 1137–1149. [Google Scholar] [CrossRef] [PubMed]
- Satyanarayanan, M.; Goode, A.; Gilbert, B.; Harkes, J.; Jukic, D. OpenSlide: A vendor-neutral software foundation for digital pathology. J. Pathol. Inform. 2013, 4, 27. [Google Scholar] [CrossRef] [PubMed]
- Girshick, R.; Donahue, J.; Darrell, T.; Malik, J. Rich feature hierarchies for accurate object detection and semantic segmentation. arXiv, 2014; arXiv:1311.2524. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Spatial Pyramid Pooling in Deep Convolutional Networks for Visual Recognition. arXiv, 2014; arXiv:1406.4729. [Google Scholar]
- Wang, X.; Shrivastava, A.; Gupta, A. A-Fast-RCNN: Hard Positive Generation via Adversary for Object Detection. arXiv, 2017; arXiv:1704.03414. [Google Scholar]
- Joseph, R.; Santosh, D.; Ross, G.; Ali, F. You Only Look Once: Unified, Real-Time Object Detection. arXiv, 2015; arXiv:1506.02640. [Google Scholar]
- Liu, W.; Anguelov, D.; Erhan, D.; Szegedy, C.; Reed, S.; Fu, C.Y.; Berg, A.C. SSD: Single shot multibox detector. arXiv, 2016; arXiv:1512.02325. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A. Inception-v4, Inception-ResNet and the Impact of Residual Connections on Learning. arXiv, 2016; arXiv:1602.07261. [Google Scholar]
- Everingham, M.; Eslami, S.M.A.; Van Gool, L.; Williams, C.K.I.; Winn, J.; Zisserman, A. The Pascal Visual Object Classes Challenge: A Retrospective. Int. J. Comput. Vis. 2014, 111, 98–136. [Google Scholar] [CrossRef]
- Lin, T.Y.; Zitnick, C.L.; Doll, P. Microsoft COCO: Common Objects in Context. arXiv, 2015; arXiv:1405.0312. [Google Scholar]
- Tajbakhsh, N.; Shin, J.Y.; Gurudu, S.R.; Hurst, R.T.; Kendall, C.B.; Gotway, M.B.; Liang, J. Convolutional Neural Networks for Medical Image Analysis: Full Training or Fine Tuning? IEEE Trans. Med. Imaging 2016, 35, 1299–1312. [Google Scholar] [CrossRef] [PubMed]
- Sethi, A.; Sha, L.; Vahadane, A.; Deaton, R.; Kumar, N.; Macias, V.; Gann, P. Empirical comparison of color normalization methods for epithelial-stromal classification in H and E images. J. Pathol. Inform. 2016, 7, 17. [Google Scholar] [CrossRef] [PubMed]
- Galdran, A.; Alvarez-Gila, A.; Meyer, M.I.; Saratxaga, C.L.; Araújo, T.; Garrote, E.; Aresta, G.; Costa, P.; Mendonça, A.M.; Campilho, A. Data-Driven Color Augmentation Techniques for Deep Skin Image Analysis. arXiv, 2017; arXiv:1703.03702. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. arXiv, 2014; arXiv:1409.4842v1. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. arXiv, 2015; arXiv:1512.03385v1. [Google Scholar]
| Author | Species | Stain | Number of Whole Slide Images (WSIs) | Method | Performance | ||
|---|---|---|---|---|---|---|---|
| Recall | Precision | F-Measure | |||||
| Kato et al. [21] | Rat | Desmin | 20 | R-HOG + SVM | 0.911 | 0.777 | 0.838 |
| S-HOG + SVM | 0.897 | 0.874 | 0.897 | ||||
| Temerinac-Ott et al. [23] | Human | HE/PAS/CD10/SR | 80 | R-HOG + SVM | N/A | N/A | 0.405–0.551 |
| CNN | N/A | N/A | 0.522–0.716 | ||||
| Gallego et al. [24] | Human | PAS | 108 | CNN | 1.000 | 0.881 | 0.937 |
| Simon et al. [22] | Mouse | HE | 15 | mrcLBP + SVM | 0.800 | 0.900 | 0.850 |
| Rat | HE/PAS/JS/TRI/CR | 25 | 0.560–0.730 | 0.750–0.914 | 0.680–0.801 | ||
| Human | PAS | 25 | 0.761 | 0.917 | 0.832 | ||
| Stain | WSIs | Total Number of Glomeruli | Number of Glomeruli per WSI (Min–Max) |
|---|---|---|---|
| PAS | 200 | 8058 | 40.3 (2–166) |
| PAM | 200 | 8459 | 42.3 (4–173) |
| MT | 200 | 8569 | 42.8 (3–187) |
| Azan | 200 | 8203 | 41.0 (2–195) |
| Average | 8323 | 41.6 (2–195) |
| Author | Species | Stain | Number of WSIs | Method | Performance | ||
|---|---|---|---|---|---|---|---|
| Recall | Precision | F-Measure | |||||
| Kato et al. [21] | Rat | Desmin | 20 | R-HOG + SVM | 0.911 | 0.777 | 0.838 |
| S-HOG + SVM | 0.897 | 0.874 | 0.897 | ||||
| Temerinac-Ott et al. [23] | Human | HE/PAS/CD10/SR | 80 | R-HOG + SVM | N/A | N/A | 0.405–0.551 |
| CNN | N/A | N/A | 0.522–0.716 | ||||
| Gallego et al. [24] | Human | PAS | 108 | CNN | 1.000 | 0.881 | 0.937 |
| Simon et al. [22] | Mouse | HE | 15 | mrcLBP + SVM | 0.800 | 0.900 | 0.850 |
| Rat | HE/PAS/JS/TRI/CR | 25 | 0.560–0.730 | 0.750–0.914 | 0.680–0.801 | ||
| Human | PAS | 25 | 0.761 | 0.917 | 0.832 | ||
| Proposed | Human | PAS | 200 | Faster R-CNN | 0.919 | 0.931 | 0.925 |
| PAM | 200 | 0.918 | 0.939 | 0.928 | |||
| MT | 200 | 0.878 | 0.915 | 0.896 | |||
| Azan | 200 | 0.849 | 0.904 | 0.876 | |||
| Stain | 140 WSIs (Training 60, Validation 40, Testing 40) | 170 WSIs (Training 90, Validation 40, Testing 40) | 200 WSIs (Training 120, Validation 40, Testing 40) |
|---|---|---|---|
| PAS | 0.907 | 0.905 | 0.925 |
| PAM | 0.927 | 0.926 | 0.928 |
| MT | 0.898 | 0.892 | 0.896 |
| Azan | 0.877 | 0.876 | 0.876 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kawazoe, Y.; Shimamoto, K.; Yamaguchi, R.; Shintani-Domoto, Y.; Uozaki, H.; Fukayama, M.; Ohe, K. Faster R-CNN-Based Glomerular Detection in Multistained Human Whole Slide Images. J. Imaging 2018, 4, 91. https://doi.org/10.3390/jimaging4070091
Kawazoe Y, Shimamoto K, Yamaguchi R, Shintani-Domoto Y, Uozaki H, Fukayama M, Ohe K. Faster R-CNN-Based Glomerular Detection in Multistained Human Whole Slide Images. Journal of Imaging. 2018; 4(7):91. https://doi.org/10.3390/jimaging4070091
Chicago/Turabian StyleKawazoe, Yoshimasa, Kiminori Shimamoto, Ryohei Yamaguchi, Yukako Shintani-Domoto, Hiroshi Uozaki, Masashi Fukayama, and Kazuhiko Ohe. 2018. "Faster R-CNN-Based Glomerular Detection in Multistained Human Whole Slide Images" Journal of Imaging 4, no. 7: 91. https://doi.org/10.3390/jimaging4070091
APA StyleKawazoe, Y., Shimamoto, K., Yamaguchi, R., Shintani-Domoto, Y., Uozaki, H., Fukayama, M., & Ohe, K. (2018). Faster R-CNN-Based Glomerular Detection in Multistained Human Whole Slide Images. Journal of Imaging, 4(7), 91. https://doi.org/10.3390/jimaging4070091