Genome-Wide Identification of the Hsp70 Gene Family in Grape and Their Expression Profile during Abiotic Stress
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification of Hsp70 Proteins in Grape
2.2. Multiple Alignment and Phylogenetic Analysis
2.3. Analysis of Subcellular Localization, Gene Structure, and Conserved Motif
2.4. Gene Location and Duplication Analysis
2.5. Microarray Data Analysis
2.6. Plant Materials and Treatments
2.7. RNA Isolation and Quantitative Real-Time PCR (qRT-PCR) Analysis
3. Results
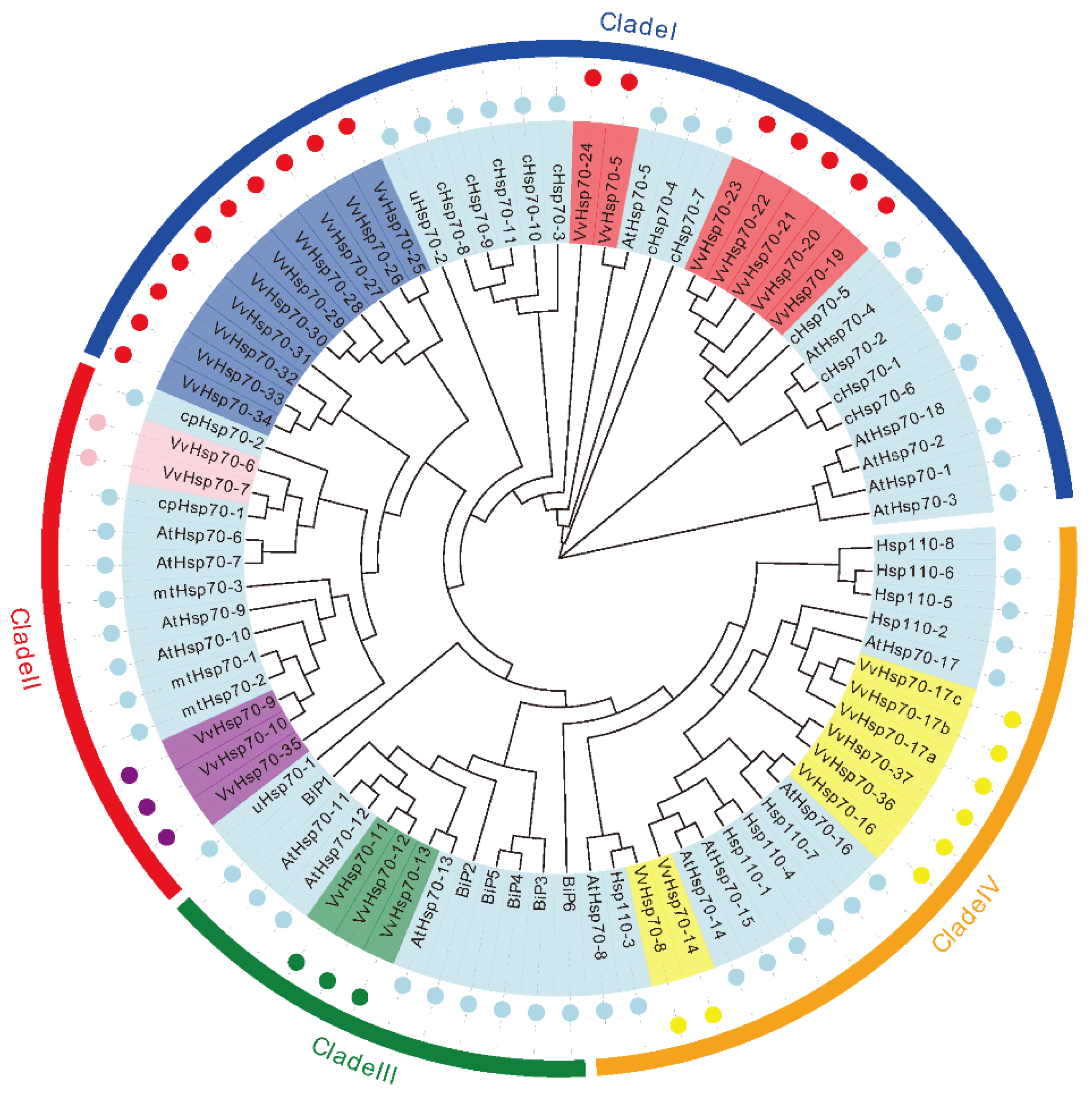
3.1. Identification and Phylogenetic Analysis of Hsp70 Genes in Grapevine
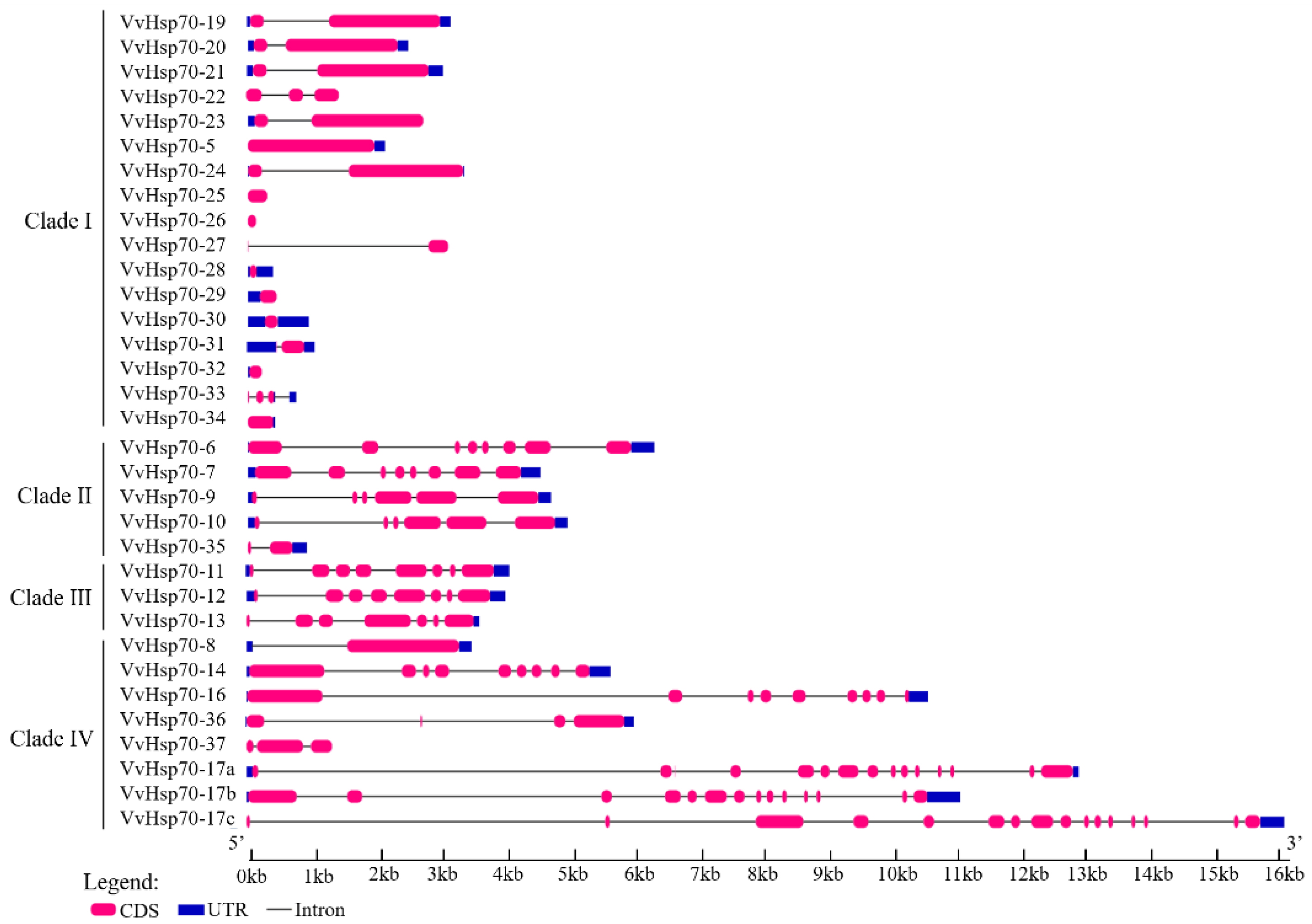
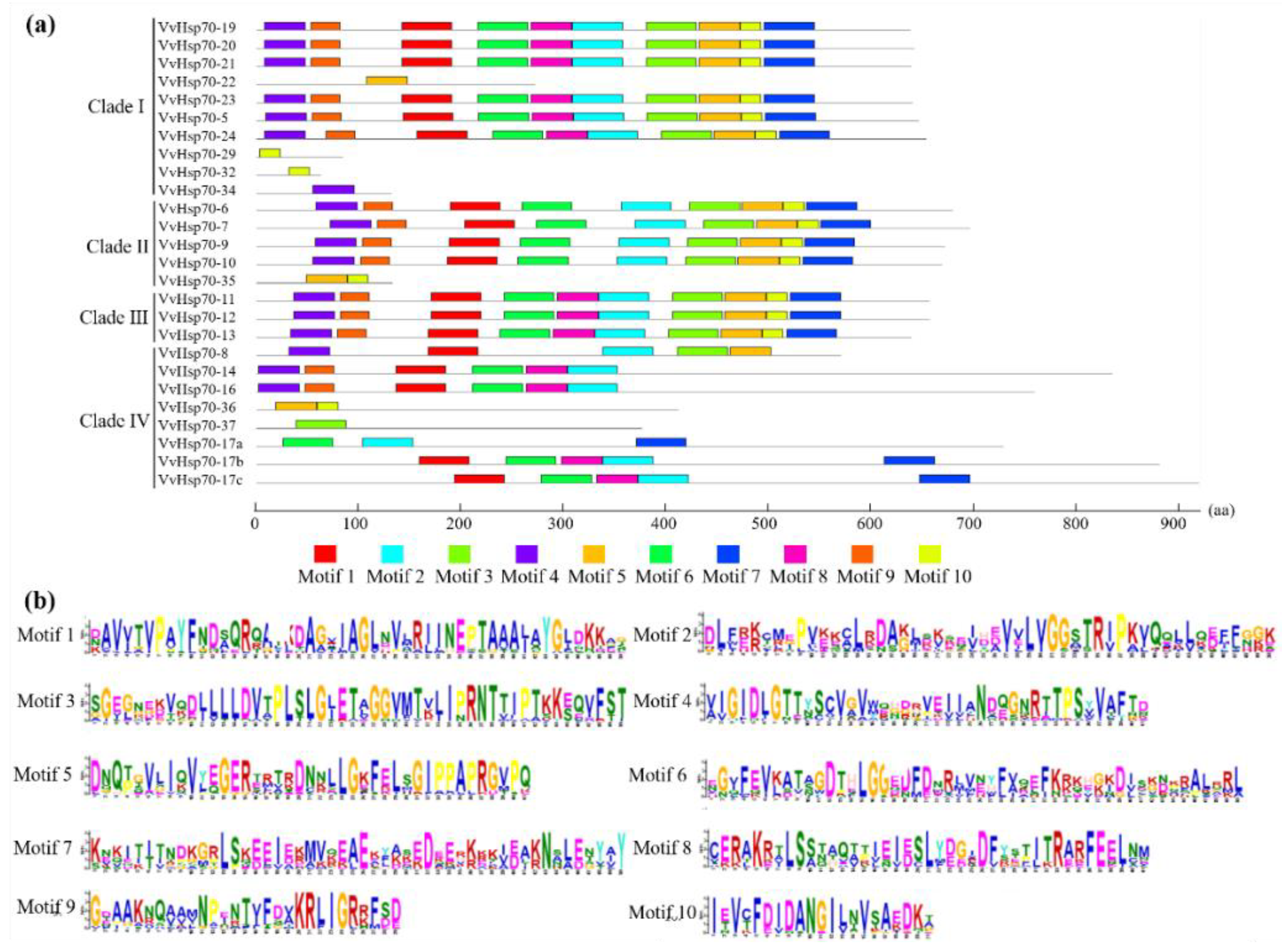
3.2. Subcellular Location, Gene Structure, and Conserved Motif Analysis
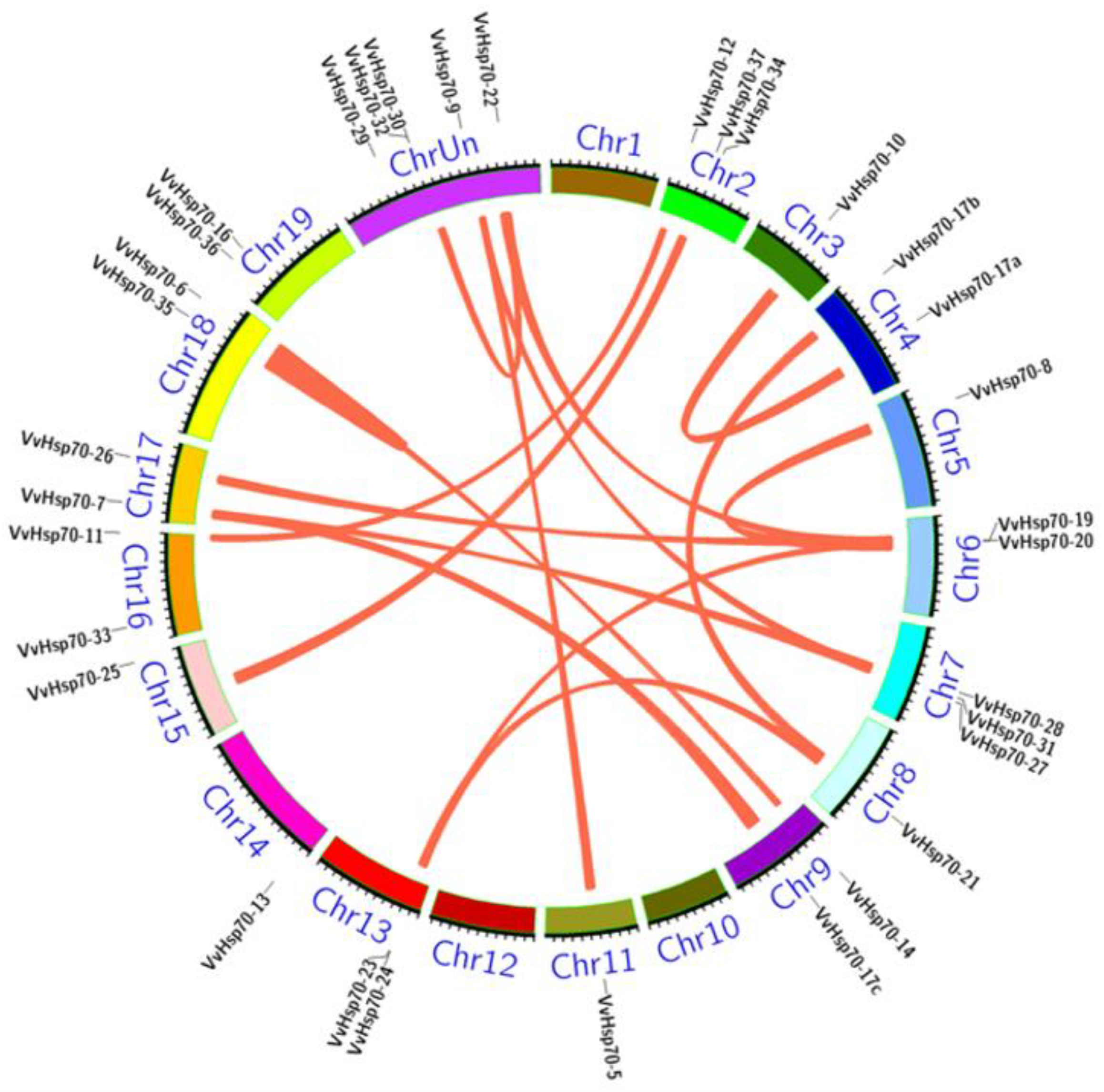
3.3. Gene Location and Gene Duplication
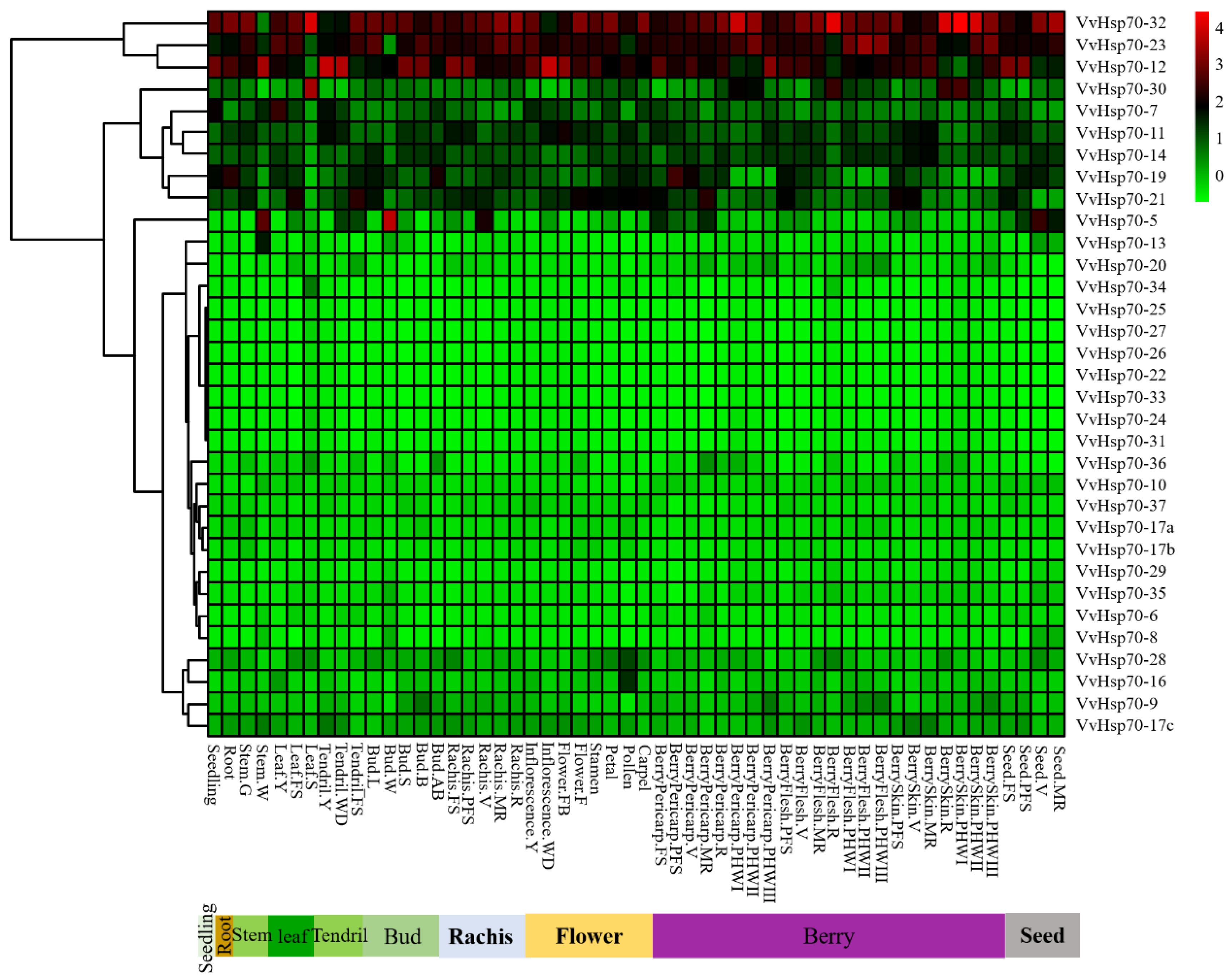
3.4. Expression Patterns of VvHsp70 Genes in Different Tissues and Development Stages
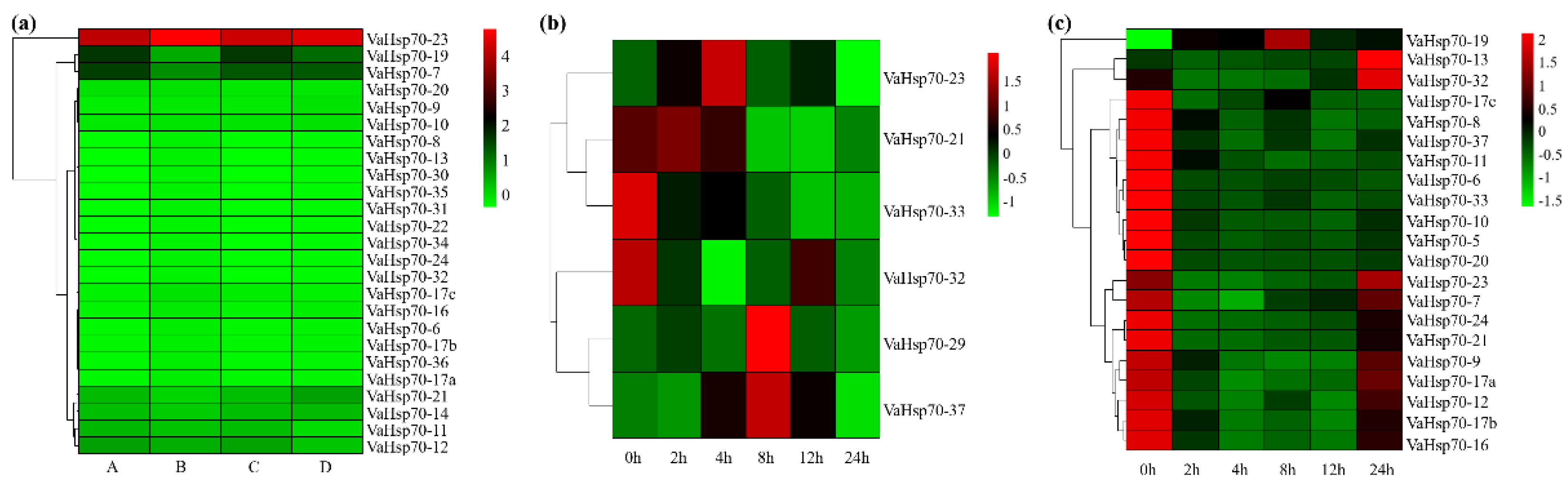
3.5. Expression Patterns of Grape Hsp70 Genes in Response to High Temperatures
3.6. VaHsp70 Genes in Response to Osmotic and Cold Stresses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, H.; Zhu, J.; Gong, Z.; Zhu, J. Abiotic stress responses in plants. Nat. Rev. Genet. 2022, 23, 104–119. [Google Scholar] [CrossRef] [PubMed]
- Nishad, A.; Nandi, A. Recent advances in plant thermomemory. Plant Cell Rep. 2021, 40, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Jacob, P.; Hirt, H.; Bendahmane, A. The heat-shock protein/chaperone network and multiple stress resistance. Plant Biotechnol. J. 2017, 15, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Swindell, W.; Huebner, M.; Weber, A. Transcriptional profiling of Arabidopsis heat shock proteins and transcription factors reveals extensive overlap between heat and non-heat stress response pathways. BMC Genom. 2007, 8, 125. [Google Scholar] [CrossRef]
- Sarkar, N.; Kim, Y.; Grover, A. Rice sHsp genes: Genomic organization and expression profiling under stress and development. BMC Genom. 2009, 10, 393. [Google Scholar] [CrossRef]
- Queitsch, C.; Hong, S.; Vierling, E.; Lindquist, S. Heat Shock Protein 101 Plays a Crucial Role in Thermotolerance in Arabidopsis. Plant Cell 2000, 12, 479–492. [Google Scholar] [CrossRef]
- Usman, M.; Rafii, M.; Martini, M.; Yusuff, O.; Ismail, M.; Miah, G. Molecular analysis of Hsp70 mechanisms in plants and their function in response to stress. Biotechnol. Genet. Eng. Rev. 2017, 33, 26–39. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Wang, J.; Liu, H.; Chen, R.; Meyer, Y.; Barakat, A.; Delseny, M. Genomic analysis of the Hsp70 superfamily in Arabidopsis thaliana. Cell Stress Chaperones 2001, 6, 201–208. [Google Scholar] [CrossRef]
- Sarkar, N.; Kundnani, P.; Grover, A. Functional analysis of Hsp70 superfamily proteins of rice (Oryza sativa). Cell Stress Chaperones 2013, 18, 427–437. [Google Scholar] [CrossRef]
- Song, Z.; Pan, F.; Lou, X.; Wang, D.; Yang, C.; Zhang, B.; Zhang, H. Genome-Wide identification and characterization of Hsp70 gene family in Nicotiana tabacum. Mol. Biol. Rep. 2019, 46, 1941–1954. [Google Scholar] [CrossRef]
- Jiang, L.; Hu, W.; Qian, Y.; Ren, Q.; Zhang, J. Genome-Wide identification, classification and expression analysis of the Hsf and Hsp70 gene families in maize. Gene 2021, 770, 145348. [Google Scholar] [CrossRef] [PubMed]
- Song, P.; Jia, Q.; Xiao, X.; Tang, Y.; Liu, C.; Li, W.; Li, T.; Li, L.; Chen, H.; Zhang, W.; et al. HSP70-3 Interacts with Phospholipase Ddelta and Participates in Heat Stress Defense. Plant Physiol. 2021, 185, 1148–1165. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Xia, X.; Su, J.; Wei, M.; Wu, Y.; Tao, J. Overexpression of herbaceous peony HSP70 confers high temperature tolerance. BMC Genom. 2019, 20, 70. [Google Scholar] [CrossRef] [PubMed]
- Montero-Barrientos, M.; Hermosa, R.; Cardoza, R.; Gutierrez, S.; Nicolas, C.; Monte, E. Transgenic expression of the Trichoderma harzianum hsp70 gene increases Arabidopsis resistance to heat and other abiotic stresses. J. Plant Physiol. 2010, 167, 659–665. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.; Choi, Y. A nuclear-Localized HSP70 confers thermoprotective activity and drought-stress tolerance on plants. Biotechnol. Lett. 2009, 31, 597–606. [Google Scholar] [CrossRef]
- Yer, E.; Baloglu, M.; Ziplar, U.; Ayan, S.; Unver, T. Drought-Responsive Hsp70 Gene Analysis in Populus at Genome-Wide Level. Plant Mol. Biol. Report. 2015, 34, 483–500. [Google Scholar] [CrossRef]
- Zhang, L.; Zhao, H.; Dong, Q.; Zhang, Y.; Wang, Y.; Li, H.; Xing, G.; Li, Q.; Dong, Y. Genome-Wide analysis and expression profiling under heat and drought treatments of HSP70 gene family in soybean (Glycine max L.). Front. Plant Sci. 2015, 6, 773. [Google Scholar] [CrossRef]
- Augustine, S.; Narayan, J.; Syamaladevi, D.; Appunu, C.; Chakravarthi, M.; Ravichandran, V.; Subramonian, N. Erianthus arundinaceus HSP70 (EaHSP70) overexpression increases drought and salinity tolerance in sugarcane (Saccharum spp. hybrid). Plant Sci. 2015, 232, 23–34. [Google Scholar] [CrossRef]
- Liu, J.; Pang, X.; Cheng, Y.; Yin, Y.; Zhang, Q.; Su, W.; Hu, B.; Guo, Q.; Ha, S.; Zhang, J.; et al. The Hsp70 Gene Family in Solanum tuberosum: Genome-Wide Identification, Phylogeny, and Expression Patterns. Sci. Rep. 2018, 8, 16628. [Google Scholar] [CrossRef]
- Fei, J.; Wang, Y.; Zhou, Q.; Gu, J. Cloning and expression analysis of HSP70 gene from mangrove plant Kandelia obovata under cold stress. Ecotoxicology 2015, 24, 1677–1685. [Google Scholar] [CrossRef]
- Xu, M.; Tong, Q.; Wang, Y.; Wang, Z.; Xu, G.; Elias, G.; Li, S.; Liang, Z. Transcriptomic analysis of the grapevine LEA gene family in response to osmotic and cold stress reveals a key role for VamDHN3. Plant Cell Physiol. 2020, 61, 775–786. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wong, D.; Wang, Y.; Xu, G.; Ren, C.; Liu, Y.; Kuang, Y.; Fan, P.; Li, S.; Xin, H.; et al. GRAS-Domain transcription factor PAT1 regulates jasmonic acid biosynthesis in grape cold stress response. Plant Physiol. 2021, 186, 1660–1678. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Liu, G.; Yan, B.; Duan, W.; Wang, L.; Li, S. Comparison of investigation methods of heatinjury in grapevine (Vitis) and assessment to heat tolerance in different cultivars and species. BMC Plant Biol. 2014, 14, 156. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Chai, F.; Wang, Y.; Jiang, J.; Duan, W.; Wang, Y.; Wang, F.; Li, S.; Wang, L. Genome-Wide Identification and Classification of HSF Family in Grape, and Their Transcriptional Analysis under Heat Acclimation and Heat Stress. Hortic. Plant J. 2018, 4, 133–143. [Google Scholar] [CrossRef]
- Su, L.; Dai, Z.; Li, S.; Xin, H. A novel system for evaluating drought-cold tolerance of grapevines using chlorophyll fluorescence. BMC Plant Biol. 2015, 15, 82. [Google Scholar] [CrossRef]
- Cheng, C.; Wang, Y.; Chai, F.; Li, S.; Xin, H.; Liang, Z. Genome-Wide identification and characterization of the 14-3-3 family in Vitis vinifera L. during berry development and cold- and heat-stress response. BMC Genom. 2018, 19, 579. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.; Marra, M. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Jiang, J.; Liu, X.; Liu, C.; Liu, G.; Li, S.; Wang, L. Integrating Omics and Alternative Splicing Reveals Insights into Grape Response to High Temperature. Plant Physiol. 2017, 173, 1502–1518. [Google Scholar] [CrossRef]
- Fasoli, M.; Dal Santo, S.; Zenoni, S.; Tornielli, G.; Farina, L.; Zamboni, A.; Porceddu, A.; Venturini, L.; Bicego, M.; Murino, V.; et al. The grapevine expression atlas reveals a deep transcriptome shift driving the entire plant into a maturation program. Plant Cell 2012, 24, 3489–3505. [Google Scholar] [CrossRef]
- Tashiro, R.M.; Philips, J.G.; Winefield, C.S. Identification of suitable grapevine reference genes for qRT-PCR derived from heterologous species. Mol. Genet. Genom. 2016, 291, 483–492. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Chen, T.; Yin, X.; Xiang, G.; Peng, J.; Fu, Q.; Li, M.; Shang, B.; Ma, H.; Liu, G.; et al. A Plasmopara viticola RXLR effector targets a chloroplast protein PsbP to inhibit ROS production in grapevine. Plant J. 2021, 106, 1557–1570. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Li, M.; Zhang, N.; Shang, B.; Liu, G.; Xu, Y. Cloning and functional analysis of the CDS and promoter of VpPR4b gene response to downy mildew in Chinese wild grape. Acta Hortic. Sin. 2021, 48, 265–275. [Google Scholar] [CrossRef]
- Grimplet, J.; Adam-Blondon, A.; Bert, P.; Bitz, O.; Cantu, D.; Davies, C.; Pezzotti, M.; Rombauts, S.; Cramer, G. The grapevine gene nomenclature system. BMC Genom. 2014, 15, 1077. [Google Scholar] [CrossRef] [PubMed]
- Timperio, A.; Egidi, M.; Zolla, L. Proteomics applied on plant abiotic stresses: Role of heat shock proteins (HSP). J. Proteom. 2008, 71, 391–411. [Google Scholar] [CrossRef]
- Banilas, G.; Korkas, E.; Englezos, V.; Nisiotou, A.; Hatzopoulos, P.; Research, W. Genome-Wide analysis of the heat shock protein 90 gene family in grapevine (Vitis vinifera L.). Aust. J. Grape Wine Res. 2012, 18, 29–38. [Google Scholar] [CrossRef]
- Chen, T.; Xu, T.; Zhang, T.; Liu, T.; Shen, L.; Chen, Z.; Wu, Y.; Yang, J. Genome-Wide Identification and Characterization of DnaJ Gene Family in Grape (Vitis vinifera L.). Horticulturae 2021, 7, 589. [Google Scholar] [CrossRef]
- Ji, X.; Yu, Y.; Ni, P.; Zhang, G.; Guo, D. Genome-Wide identification of small heat-shock protein (HSP20) gene family in grape and expression profile during berry development. BMC Plant Biol. 2019, 19, 433. [Google Scholar] [CrossRef]
- Magadum, S.; Banerjee, U.; Murugan, P.; Gangapur, D.; Ravikesavan, R. Gene duplication as a major force in evolution. J. Genet. 2013, 92, 155–161. [Google Scholar] [CrossRef]
- Wei, Y.; Mu, H.; Xu, G.; Wang, Y.; Li, Y.; Li, S.; Wang, L. Genome-Wide Analysis and Functional Characterization of the UDP-Glycosyltransferase Family in Grapes. Horticulturae 2021, 7, 204. [Google Scholar] [CrossRef]
- Bond, U.; Schlesinger, M. Heat-Shock Proteins and Development. Adv. Genet. 1987, 24, 1–29. [Google Scholar] [PubMed]
- Chaudhry, S.; Sidhu, G. Climate change regulated abiotic stress mechanisms in plants: A comprehensive review. Plant Cell Rep. 2022, 41, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Schöffl, F. AnHsp70 antisense gene affects the expression of HSP70/HSC70, the regulation of HSF, and the acquisition of thermotolerance in transgenic Arabidopsis thaliana. Mol. Gen. Genet. 1996, 252, 11–19. [Google Scholar] [PubMed]
- Fernández-Bautista, N.; Fernández-Calvino, L.; Muñoz, A.; Toribio, R.; Mock, H.; Castellano, M. HOP family plays a major role in long-term acquired thermotolerance in Arabidopsis. Plant Cell Environ. 2018, 41, 1852–1869. [Google Scholar] [CrossRef]
- Alvim, F.; Carolino, S.; Cascardo, J.; Nunes, C.; Martinez, C.; Otoni, W.; Fontes, E. Enhanced accumulation of BiP in transgenic plants confers tolerance to water stress. Plant Physiol. 2001, 126, 1042–1054. [Google Scholar] [CrossRef]
- Chaudhary, R.; Baranwal, V.; Kumar, R.; Sircar, D.; Chauhan, H. Genome-Wide identification and expression analysis of Hsp70, Hsp90, and Hsp100 heat shock protein genes in barley under stress conditions and reproductive development. Funct. Integr. Genom. 2019, 19, 1007–1022. [Google Scholar] [CrossRef]
- Hlavackova, I.; Vitamvas, P.; Santrucek, J.; Kosova, K.; Zelenkova, S.; Prasil, I.; Ovesna, J.; Hynek, R.; Kodicek, M. Proteins involved in distinct phases of cold hardening process in frost resistant winter barley (Hordeum vulgare L.) cv Luxor. Int. J. Mol. Sci. 2013, 14, 8000–8024. [Google Scholar] [CrossRef]
- Jin, Y.; Zhang, C.; Yang, H.; Yang, Y.; Huang, C.; Tian, Y.; Lu, X. Proteomic analysis of cold stress responses in tobacco seedlings. Afr. J. Biotechnol. 2011, 10, 18991–19004. [Google Scholar]
- Kosova, K.; Vitamvas, P.; Planchon, S.; Renaut, J.; Vankova, R.; Prasil, I. Proteome analysis of cold response in spring and winter wheat (Triticum aestivum) crowns reveals similarities in stress adaptation and differences in regulatory processes between the growth habits. J. Proteome Res. 2013, 12, 4830–4845. [Google Scholar] [CrossRef]
- Maikova, A.; Zalutskaya, Z.; Lapina, T.; Ermilova, E. The HSP70 chaperone machines of Chlamydomonas are induced by cold stress. J. Plant Physiol. 2016, 204, 85–91. [Google Scholar] [CrossRef]
- Bae, M.; Cho, E.; Choi, E.; Park, O. Analysis of the Arabidopsis nuclear proteome and its response to cold stress. Plant J. 2003, 36, 652–663. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, M.; Mao, Q.; Hong, J.; Shi, L.; Ran, X.; Liaquat, F.; Uzair, M.; Liang, W.; Fernie, A.; Shi, J. HSP70-16 and VDAC3 jointly inhibit seed germination under cold stress in Arabidopsis. Plant Cell Environ. 2021, 44, 3616–3627. [Google Scholar] [CrossRef] [PubMed]
- Haider, M.; Zhang, C.; Kurjogi, M.; Pervaiz, T.; Zheng, T.; Zhang, C.; Lide, C.; Shangguan, L.; Fang, J. Insights into grapevine defense response against drought as revealed by biochemical, physiological and RNA-Seq analysis. Sci. Rep. 2017, 7, 13134. [Google Scholar] [CrossRef] [PubMed]
- Xin, H.; Zhu, W.; Wang, L.; Xiang, Y.; Fang, L.; Li, J.; Sun, X.; Wang, N.; Londo, J.P.; Li, S. Genome wide transcriptional profile analysis of Vitis amurensis and Vitis vinifera in response to cold stress. PLoS ONE 2013, 8, e58740. [Google Scholar] [CrossRef]

| Gene Locus ID | Gene Symbol | Exon | Intron | Localization | Clade | Chr | Start Site | End Site | Gene Length (bp) | CDS (bp) | Protein (aa) | Strand |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| VIT_06s0004g04510 | VvHsp70-19 | 2 | 1 | Cyto | I | 6 | 5470302 | 5473460 | 3159 | 1947 | 648 | − |
| VIT_06s0004g04470 | VvHsp70-20 | 2 | 1 | Cyto | Ⅰ | 6 | 5418191 | 5420677 | 2487 | 1959 | 652 | + |
| VIT_08s0007g00130 | VvHsp70-21 | 2 | 1 | Cyto | Ⅰ | 8 | 14518534 | 14521575 | 3042 | 1950 | 649 | + |
| VIT_00s0787g00020 | VvHsp70-22 | 3 | 2 | Chlo | Ⅰ | Un | 35359691 | 35361118 | 1428 | 831 | 276 | + |
| VIT_13s0019g01430 | VvHsp70-23 | 2 | 1 | Cyto | Ⅰ | 13 | 3014601 | 3017332 | 2732 | 1953 | 650 | + |
| VIT_11s0037g00510 | VvHsp70-5 | 1 | 0 | Cyto | Ⅰ | 11 | 8347523 | 8349655 | 2133 | 1971 | 656 | − |
| VIT_13s0019g00930 | VvHsp70-24 | 2 | 1 | Cyto | Ⅰ | 13 | 2720364 | 2723716 | 3353 | 1995 | 664 | + |
| VIT_15s0046g02740 | VvHsp70-25 | 1 | 0 | Cyto | Ⅰ | 15 | 19488552 | 19488854 | 303 | 303 | 100 | − |
| VIT_17s0000g09950 | VvHsp70-26 | 1 | 0 | Cyto | Ⅰ | 17 | 11927365 | 11927493 | 129 | 129 | 42 | − |
| VIT_07s0095g00430 | VvHsp70-27 | 2 | 1 | Chlo | Ⅰ | 7 | 12876279 | 12879396 | 3118 | 315 | 104 | − |
| VIT_07s0005g06130 | VvHsp70-28 | 1 | 0 | Nucl | Ⅰ | 7 | 10927794 | 10928172 | 379 | 108 | 35 | + |
| VIT_00s0198g00100 | VvHsp70-29 | 1 | 0 | Nucl | Ⅰ | Un | 10909128 | 10909575 | 448 | 261 | 86 | − |
| VIT_00s0254g00030 | VvHsp70-30 | 1 | 0 | Cyto | Ⅰ | Un | 18000270 | 18001211 | 942 | 195 | 64 | − |
| VIT_07s0005g06650 | VvHsp70-31 | 2 | 1 | Cyto | Ⅰ | 7 | 12031939 | 12032979 | 1041 | 363 | 120 | − |
| VIT_00s0254g00040 | VvHsp70-32 | 1 | 0 | Cyto | Ⅰ | Un | 18001212 | 18001429 | 218 | 195 | 64 | + |
| VIT_16s0115g00060 | VvHsp70-33 | 3 | 3 | Nucl | Ⅰ | 16 | 3576063 | 3576810 | 748 | 225 | 74 | − |
| VIT_02s0012g01830 | VvHsp70-34 | 1 | 0 | Nucl | Ⅰ | 2 | 8260058 | 8260480 | 423 | 405 | 134 | − |
| VIT_18s0041g01230 | VvHsp70-6 | 8 | 7 | Chlo | Ⅱ | 18 | 25847216 | 25853530 | 6315 | 2073 | 690 | + |
| VIT_17s0000g03310 | VvHsp70-7 | 8 | 7 | Chlo | Ⅱ | 17 | 3189808 | 3194355 | 4548 | 2124 | 707 | + |
| VIT_00s0415g00030 | VvHsp70-9 | 6 | 5 | Mito | Ⅱ | Un | 28290680 | 28295392 | 4713 | 2049 | 682 | + |
| VIT_03s0097g00530 | VvHsp70-10 | 6 | 5 | Mito | Ⅱ | 3 | 10899795 | 10904772 | 4978 | 2040 | 679 | + |
| VIT_18s0075g00260 | VvHsp70-35 | 2 | 1 | Mito | Ⅱ | 18 | 21622733 | 21623656 | 924 | 408 | 135 | − |
| VIT_16s0098g01580 | VvHsp70-11 | 8 | 7 | ER | Ⅲ | 16 | 21738759 | 21742840 | 4082 | 2004 | 667 | − |
| VIT_02s0025g02140 | VvHsp70-12 | 8 | 7 | ER | Ⅲ | 2 | 1900784 | 1904806 | 4023 | 2004 | 667 | + |
| VIT_14s0060g02340 | VvHsp70-13 | 7 | 6 | ER | Ⅲ | 14 | 1920620 | 1924241 | 3622 | 1950 | 649 | + |
| VIT_05s0020g03330 | VvHsp70-8 | 1 | 1 | Cyto | Ⅳ | 5 | 5092723 | 5096216 | 3494 | 1740 | 579 | + |
| VIT_09s0002g04680 | VvHsp70-14 | 9 | 8 | Cyto | Ⅳ | 9 | 4243222 | 4248888 | 5667 | 2547 | 848 | + |
| VIT_19s0090g00340 | VvHsp70-16 | 9 | 8 | Nucl | Ⅳ | 19 | 6484998 | 6495602 | 10,605 | 2316 | 771 | + |
| VIT_19s0014g03420 | VvHsp70-36 | 4 | 3 | Cyto | Ⅳ | 19 | 3567945 | 3573976 | 6032 | 1257 | 418 | − |
| VIT_02s0012g00960 | VvHsp70-37 | 3 | 2 | Cyto | Ⅳ | 2 | 6881520 | 6882848 | 1329 | 1149 | 382 | + |
| VIT_04s0043g00710 | VvHsp70-17a | 15 | 13 | Cyto | Ⅳ | 4 | 14692195 | 14705141 | 12,947 | 2223 | 740 | − |
| VIT_04s0008g04410 | VvHsp70-17b | 14 | 13 | ER | Ⅳ | 4 | 3800929 | 3812026 | 11,098 | 2688 | 895 | − |
| VIT_09s0096g00090 | VvHsp70-17c | 16 | 15 | Chlo | Ⅳ | 9 | 11486035 | 11502175 | 16,141 | 2805 | 934 | + |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Chen, H.; Li, S.; Wang, L. Genome-Wide Identification of the Hsp70 Gene Family in Grape and Their Expression Profile during Abiotic Stress. Horticulturae 2022, 8, 743. https://doi.org/10.3390/horticulturae8080743
Liu X, Chen H, Li S, Wang L. Genome-Wide Identification of the Hsp70 Gene Family in Grape and Their Expression Profile during Abiotic Stress. Horticulturae. 2022; 8(8):743. https://doi.org/10.3390/horticulturae8080743
Chicago/Turabian StyleLiu, Xinna, Haiyang Chen, Shenchang Li, and Lijun Wang. 2022. "Genome-Wide Identification of the Hsp70 Gene Family in Grape and Their Expression Profile during Abiotic Stress" Horticulturae 8, no. 8: 743. https://doi.org/10.3390/horticulturae8080743
APA StyleLiu, X., Chen, H., Li, S., & Wang, L. (2022). Genome-Wide Identification of the Hsp70 Gene Family in Grape and Their Expression Profile during Abiotic Stress. Horticulturae, 8(8), 743. https://doi.org/10.3390/horticulturae8080743





