Genetic Identity and Diversity of Apple Accessions within a Candidate Collection for the Norwegian National Clonal Germplasm Repository
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Managements
2.2. Genetic Analyses
2.3. Biostatistical Analysis
3. Results and Discussion
3.1. Allele Polymorphism
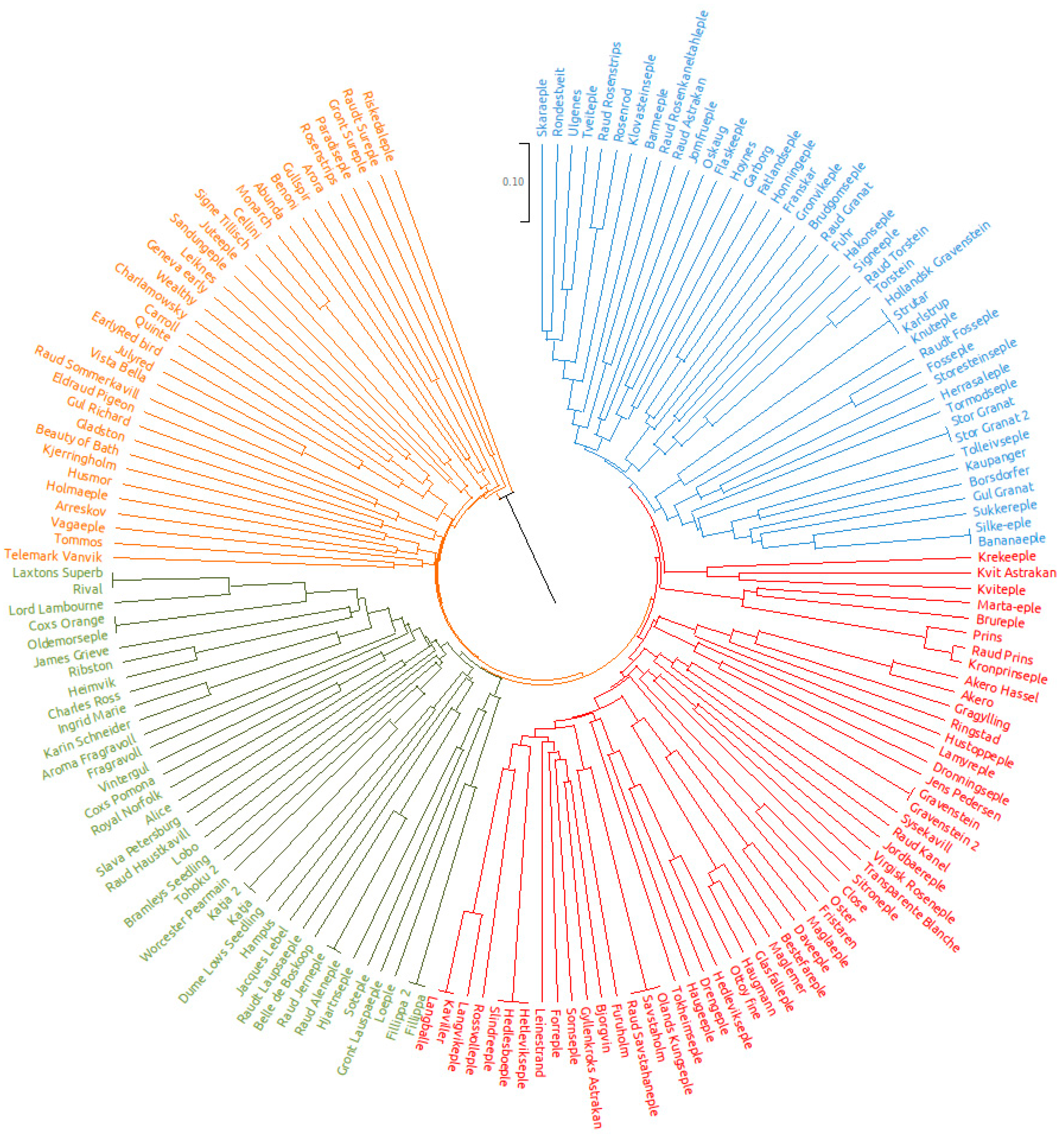
3.2. Genetic Identity and Relationships
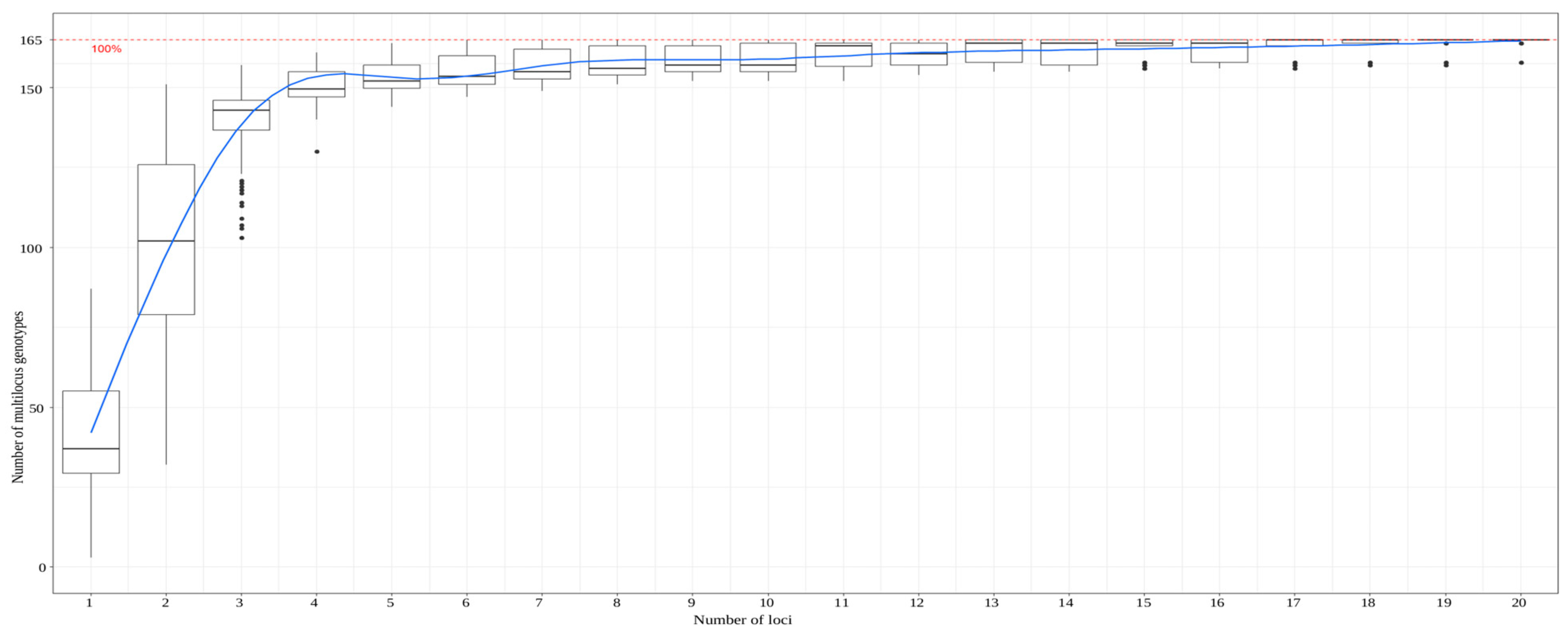
3.3. Accuracy of a Larger Marker Set
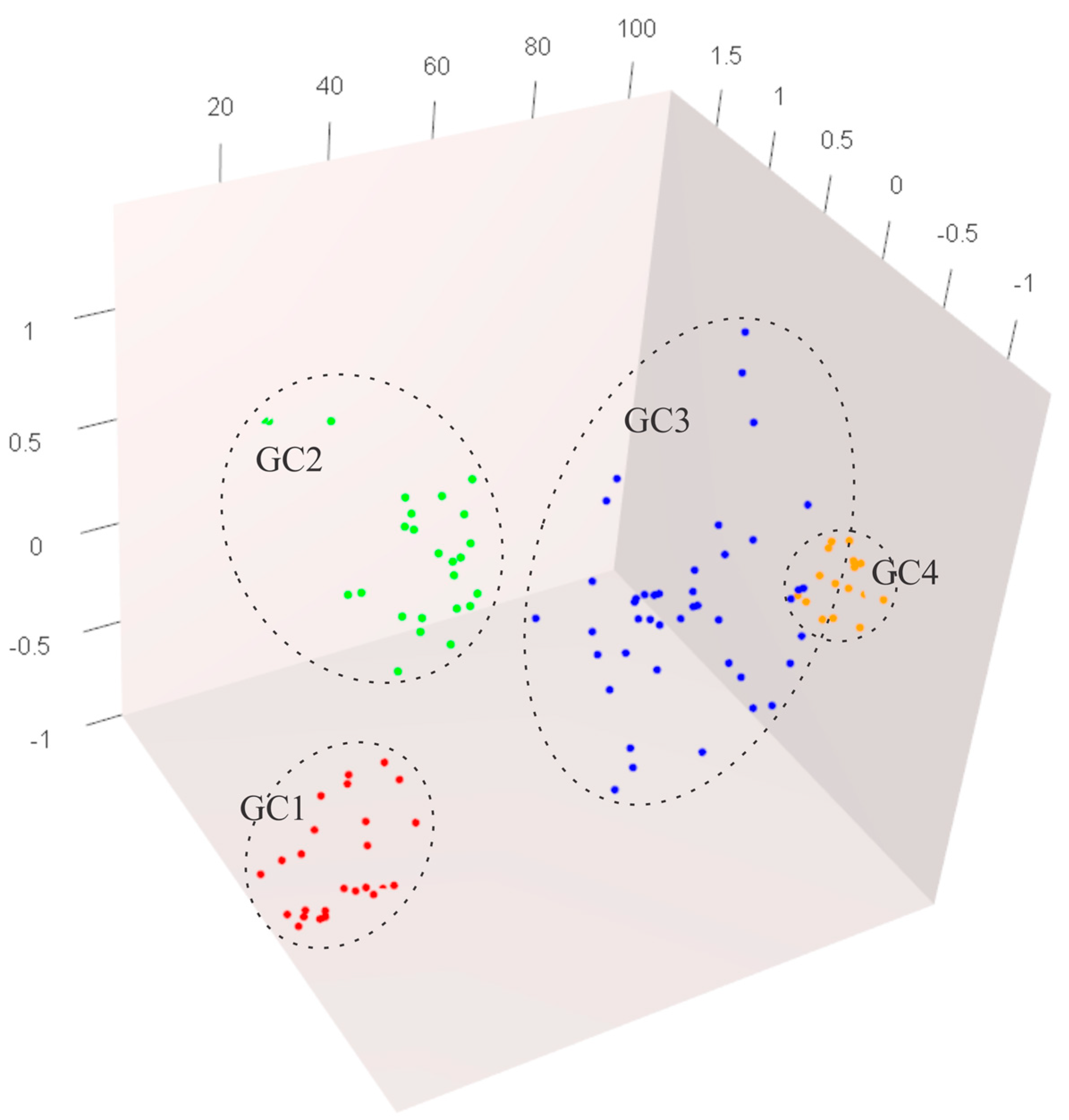
3.4. Genetic Structure
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fotiric Akšić, M.D.; Dabić Zagorac, U.; Gašić, T.; Tosti, M.; Natić, M.; Meland, M. Analysis of apple fruit (Malus × domestica Borkh.) quality attributes obtained from organic and integrated production systems. Sustainability 2022, 14, 5300. [Google Scholar] [CrossRef]
- Lacis, G.; Trajkovski, V.; Rashal, I. Phenotypical Variability and Genetic Diversity within Accessions of the Swedish Sour Cherry (Prunus cerasus L.) Genetic Resources Collection. Biologija 2010, 56, 1–8. [Google Scholar] [CrossRef]
- Jolivet, C.; Höltken, A.M.; Liesebach, H.; Steiner, W.; Degen, B. Spatial Genetic Structure in Wild Cherry (Prunus avium L.): I. Variation among Natural Populations of Different Density. Tree Genet. Genomes 2011, 7, 271–283. [Google Scholar] [CrossRef]
- Kellerhals, M.; Bertschinger, L.; Gessler, S. Use of genetic resources in apple breeding and for sustainable fruit production. J. Fruit Ornam. Plant Res. 2004, 12, 53–62. [Google Scholar]
- Gaši, F.; Kanlić, K.; Stroil, B.K.; Pojskić, N.; Asdal, Å.; Rasmussen, M.; Kaiser, C.; Meland, M. Redundancies and Genetic Structure among ex situ Apple Collections in Norway Examined with Microsatellite Markers. HortScience 2016, 51, 1458–1462. [Google Scholar] [CrossRef] [Green Version]
- Lassois, L.; Denancé, C.; Ravon, E.; Guyader, A.; Guisnel, R.; Hibrand-Saint-Oyant, L.; Poncet, C.; Lasserre-Zuber, P.; Feugey, L.; Durel, C.E. Genetic diversity, population structure, parentage analysis, and construction of core collections in the French apple germplasm based on SSR markers. Plant Mol. Biol. Report. 2016, 34, 827–844. [Google Scholar] [CrossRef] [Green Version]
- Pereira-Lorenzo, S.; Ramos-Cabrer, A.M.; Ferreira, V.; Díaz-Hernández, M.B.; Carnide, V.; Pinto-Carnide, O.; Rodrigues, R.; Velázquez Barrera, M.E.; Rios-Mesa, D.; Ascasíbar-Errasti, J.; et al. Genetic diversity and core collection of Malus × domestica in northwestern Spain, Portugal and the Canary Islands by SSRs. Sci. Hortic. 2018, 240, 49–56. [Google Scholar] [CrossRef]
- Bakɪr, M.; Dumanoglu, H.; Aygun, A.; Erdogan, V.; Efe Dost, S.; Gülsen, O.; Serdar, U.; Kalkisim, O.; Bastas, K. Genetic diversity and population structure of apple germplasm from Eastern Black Sea region of Turkey by SSRs. Sci. Hortic. 2022, 294, 110793. [Google Scholar] [CrossRef]
- Baric, S.; Storti, A.; Hofer, M.; Guerra, W.; Dalla Via, J. Molecular Genetic Identification of Apple Cultivars Based on Microsatellite DNA Analysis. I. The Database of 600 Validated Profiles. Erwerbs-Obstbau 2020, 62, 117–154. [Google Scholar] [CrossRef]
- Larsen, B.; Toldam-Andersen, T.B.; Pedersen, C.; Ørgaard, M. Unravelling genetic diversity and cultivar parentage in the Danish apple gene bank collection. Tree Genet. Genomes 2017, 13, 1–12. [Google Scholar] [CrossRef]
- Testolin, R.; Foria, S.; Baccichet, I.; Messina, R.; Danuso, F.; Losa, A.; Scarbolo, E.; Stocco, M.; Cipriani, G. Genotyping apple (Malus × domestica Borkh.) heirloom germplasm collected and maintained by the Regional Administration of Friuli Venezia Giulia (Italy). Sci. Hortic. 2019, 252, 229–237. [Google Scholar] [CrossRef]
- Urrestarazu, J.; Denancé, C.; Ravon, E.; Guyader, A.; Guisnel, R.; Feugey, L.; Poncet, C.; Lateur, M.; Houben, P.; Ordidge, M.; et al. Analysis of the genetic diversity and structure across a wide range of germplasm reveals prominent gene flow in apple at the European level. BMC Plant Biol. 2016, 16, 130. [Google Scholar] [CrossRef] [PubMed]
- Marconi, G.; Ferradini, N.; Russi, L.; Concezzi, L.; Veronesi, F.; Albertini, E. Genetic Characterization of the Apple Germplasm Collection in Central Italy: The Value of Local Varieties. Front. Plant Sci. 2018, 9, 1460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hokanson, S.C.; Lamboy, W.F.; Szewc-McFadden, A.K.; McFerson, J.R. Microsatellite (SSR) variation in a collection of Malus (apple) species and hybrids. Euphytica 2001, 118, 281–294. [Google Scholar] [CrossRef]
- Garkava-Gustavsson, L.; Mujaju, C.; Sehic, J.; Zborowska, A.; Backes, G.M.; Hietaranta, T.; Antonius, K. Genetic diversity in Swedish and Finnish heirloom apple cultivars revealed with SSR markers. Sci. Hortic. 2013, 162, 43–48. [Google Scholar] [CrossRef]
- Cmejlova, J.; Rejlova, M.; Paprstein, F.; Cmejla, R. A new one-tube reaction kit for the SSR genotyping of apple (Malus × domestica Borkh.). Plant Sci. 2021, 303, 110768. [Google Scholar] [CrossRef]
- Skytte af Sätra, J.; Troggio, M.; Odilbekov, F.; Sehic, J.; Mattisson, H.; Hjalmarsson, I.; Ingvarsson, P.K.; Garkava-Gustavsson, L. Genetic Status of the Swedish Central collection of heirloom apple cultivars. Sci. Hortic. 2020, 272, 109599. [Google Scholar] [CrossRef]
- Leforestier, D.; Ravon, E.; Muranty, H.; Cornille, A.; Lemaire, C.; Giraud, T.; Durel, C.E.; Branca, A. Genomic basis of the differences between cider and dessert apple varieties. Evolut. Appl. 2015, 8, 650–661. [Google Scholar] [CrossRef]
- Muranty, H.; Denancé, C.; Feugey, L.; Crépin, J.L.; Barbier, Y.; Tartarini, S.; Ordidge, M.; Troggio, M.; Lateur, M.; Nybom, H.; et al. Using whole-genome SNP data to reconstruct a large multi-generation pedigree in apple germplasm. BMC Plant Biol. 2020, 20, 2. [Google Scholar] [CrossRef] [Green Version]
- Ordidge, M.; Kirdwichai, P.; Baksh, M.F.; Venison, E.P.; Gibbings, J.G.; Dunwell, J.M. Genetic analysis of a major international collection of cultivated apple varieties reveals previously unknown historic heteroploid and inbred relationships. PLoS ONE 2018, 13, e0202405. [Google Scholar] [CrossRef] [Green Version]
- Hodel, R.G.; Segovia-Salcedo, M.C.; Landis, J.B.; Crowl, A.A.; Sun, M.; Liu, X.; Gitzendanner, M.A.; Douglas, N.A.; Germain-Aubrey, C.C.; Chen, S.; et al. The report of my death was an exaggeration: A review for researchers using microsatellites in the 21st century. APPS 2016, 4, apps.1600025. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fernandez, F.F. Common Set of ECPGR SSR Markers for Malus Characterization. In Report of a Working Group on Malus/Pyrus, Fourth Meeting, Weggis, Switzerland, 7–9 March 2012; Lateur, M., Ordidge, M., Engels, J., Lipman, E., Eds.; Bioversity International: Rome, Italy, 2013; pp. 26–27, (abstract of presentation). [Google Scholar]
- Akšić, M.F.; Lazarević, K.; Šegan, S.; Natić, M.; Tosti, T.; Ćirić, I.; Meland, M. Assessing the Fatty Acid, Carotenoid, and Tocopherol Compositions of Seeds from Apple Cultivars (Malus domestica Borkh.) Grown Norway. Foods 2021, 10, 1956. [Google Scholar] [CrossRef] [PubMed]
- Gianfranceschi, L.; Seglias, N.; Tarchini, R.; Komjanc, M.; Gessler, C. Simple sequence repeats for the genetic analysis of apple. Theor. Appl. Genet. 1998, 96, 1069–1076. [Google Scholar] [CrossRef]
- Hokanson, S.C.; Szewc-McFadden, A.K.; Lamboy, W.F.; McFerson, J.R. Microsatellite (SSR) markers reveal genetic identities, genetic diversity and relationships in a Malus × domestica borkh. core subset collection. Theor. Appl. Genet. 1998, 97, 671–683. [Google Scholar] [CrossRef]
- Liebhard, R.; Gianfranceschi, L.; Koller, B.; Ryder, C.D.; Tarchini, R.; Van de Weg, E.; Gessler, C. Development and characterisation of 140 new microsatellites in apple (Malus x domestica Borkh.). Mol. Breed. 2002, 10, 217–241. [Google Scholar] [CrossRef]
- Fernandez-Fernandez, F.; Harvey, N.G.; James, C.M. Isolation and characterization of polymorphic microsatellite markers from european pear (Pyrus Communis L.). Mol. Ecol. 2006, 6, 1039–1041. [Google Scholar] [CrossRef]
- Gaši, F.; Žulj-Mihaljević, M.; Šimon, S.; Grahić, J.; Pojskić, N.; Kurtović, M.; Nikolić, D.; Pejić, I. Genetic structure of apple accessions maintained ex situ in Bosnia and Herzegovina examined by microsatellite markers. Genetika 2013, 45, 467–478. [Google Scholar] [CrossRef]
- Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 1978, 89, 583–590. [Google Scholar] [CrossRef]
- Hardy, O.J.; Vekemans, X. A versatile computer program to analyse spatial genetic structure at the individual or population level. Mol. Ecol. 2002, 2, 618–620. [Google Scholar] [CrossRef] [Green Version]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef] [Green Version]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; Von Holdt, B.M. Structure harvester: A Website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2011, 4, 359–361. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [Green Version]
- Vigouroux, Y.; Glaubitz, J.C.; Matsuoka, Y.; Goodman, M.M.; Sánchez, G.J.; Doebley, J. Population structure and genetic diversity of New World maize races assessed by DNA microsatellites. Am. J. Bot. 2008, 95, 1240–1253. [Google Scholar] [CrossRef] [PubMed]
- Urrestarazu, J.; Miranda, C.; Santesteban, L.G.; Royo, J.B. Genetic diversity and structure of local apple cultivars from northeastern Spain assessed by microsatellite markers. Tree Genet. Genomes 2012, 8, 1163–1180. [Google Scholar] [CrossRef]
- Excoffier, L.; Smouse, P.E.; Quattro, J.M. Analysis of molecular variance interfered from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 1992, 131, 479–491. [Google Scholar] [CrossRef]
- Ohta, T.; Kimura, M. A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in a finite population. Genet. Resour. 1973, 22, 201–204. [Google Scholar] [CrossRef]
- Meirmans, P.; Van Tienderen, P. Genotype and genodive: Two programs for the analysis of genetic diversity of asexual organisms. Mol. Ecol. Notes 2004, 4, 792–794. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2022; Available online: http://www.R-project.org/ (accessed on 15 March 2022).
- Muller, L.A.H.; McCusker, J.H. Microsatellite analysis of genetic diversity among clinical and nonclinical Saccharomyces cerevisiae isolates suggests heterozygote advantage in clinical environments. Mol. Ecol. 2009, 18, 2779–2786. [Google Scholar] [CrossRef] [Green Version]
- Adler, D.; Murdoch, D. RGL: 3D Visualization Device System (Open GL), R Package Version 0.93.945; 2013. Available online: http://rgl.neoscientists.org (accessed on 15 March 2022).
- Drost, H.G. Philentropy: Information Theory and Distance Quantification with R. J. Open Source Softw. 2018, 3, 765. [Google Scholar] [CrossRef]
- Bittinger, K. Usedist: Functions to Re-Arrange, Extract, and Work with Distances, R package version 0.4.0. 2020. Available online: https://CRAN.R-project.org/package=usedist (accessed on 15 March 2022).
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grahić, A.; Grahić, J. MADC—Marker Analysis Data Compiler User’s Manual. 2017. Available online: www.divisionagro.ba/apps/docs/madc-marker-analysis-data-compiler/usermanual (accessed on 15 March 2022).
- Van Treuren, R.; Kemp, H.; Ernsting, G.; Jongejans, B.; Houtman, H.; Visser, L. Microsatellite genotyping of apple (Malus · domestica Borkh.) genetic resources in the Netherlands: Application in collection management and variety identification. Genet. Resour. Crop Evolut. 2010, 57, 853–865. [Google Scholar] [CrossRef] [Green Version]
- Liang, W.; Dondini, L.; De Franceschi, P.; Paris, R.; Sansavini, S.; Tartarini, S. Genetic diversity, population structure and construction of a core collection of apple cultivars from Italian germplasm. Plant Mol. Biol. Rep. 2015, 33, 458–473. [Google Scholar] [CrossRef]
- Pereira-Lorenzo, S.; Ramos-Cabrer, A.M.; Díaz-Hernández, M.B. Evaluation of genetic identity and variation of local apple cultivars (Malus × domestica Borkh.) from Spain using microsatellite markers. Genet. Resour. Crop. Evolut. 2007, 54, 405–420. [Google Scholar] [CrossRef]
- Gasi, F.; Simon, S.; Pojskic, N.; Kurtovic, M.; Pejic, I. Genetic assessment apple germplasm in Bosnia and Herzegovina using microsatellite and morphologic markers. Sci. Hortic. 2010, 126, 164–171. [Google Scholar] [CrossRef]
- Kanlić, K.; Kalamujic-Stroil, B.; Grahic, J.; Asdal, Å.; Meland, M.; Kurtovic, M.; Gasi, F. Influence of selection pressure on the frequency of triploid genotypes among different traditional apple germplasms. Work. Fac. Agric. Food Sci. Univ. Sarajevo 2016, 66, 287–290. [Google Scholar]
- Wiehle, M.; Nawaz, M.A.; Dahlem, R.; Alam, I.; Khan, A.A.; Gailing, O.; Mueller, M.; Buerkert, A. Pheno-genetic studies of apple varieties in northern Pakistan: A hidden pool of diversity. Sci. Hortic. 2021, 281, 109950. [Google Scholar] [CrossRef]
| Apple Accessions | GC | Apple Accessions | GC | Apple Accessions | GC |
|---|---|---|---|---|---|
| Abunda | 3 | Håkonseple * | 2 | Quinte | 3 |
| Åkerø * | 1 | Hampus * | 3 | Raudt Fosseple | 2 |
| Åkerø Hassel * | 1 | Haugeeple * | admx | Raud Åleneple | 3 |
| Alice * | admx | Haugmann * | 1 | Raud Granat | 2 |
| Arora | admx | Hedlesbøeple | 1 | Raud Haustkavill | admx |
| Arreskov * | 3 | Hedlevikseple | admx | Raud Jerneple | 3 |
| Gyllenkroks Astrakan * | 1 | Heimvik * | 4 | Raud Kanel | 3 |
| Kvit Astrakan * | 3 | Herrasaleple | 2 | Raud Rosenkaneleple | admx |
| Raud Astrakan * | admx | Hetlevikseple | 1 | Raud Rosenstrips | 2 |
| Bananeple | 2 | Hjartnseple * | admx | Raud Sävstaholml | 1 |
| Barmeeple | admx | Hollandsk Gravenstein * | admx | Raud Sommerkavill | 3 |
| Beauty of Bath * | 3 | Holmaeple | admx | Raudt Laupsaeple | 3 |
| Belle de Boskoop | 3 | Honningeple | admx | Raudt Sureple | 3 |
| Benoni * | 3 | Høynes * | admx | Ribston * | 4 |
| Bestefareple | 1 | Husmor * | 3 | Ringstad | admx |
| Bjørgvin | admx | Hustoppeple | admx | Riskedaleple * | 3 |
| Borsdorfer * | 2 | Ingrid Marie * | 4 | Rival * | 4 |
| Bramleys Seedling * | 3 | Jacques Lebel * | 3 | Rondestveit * | 2 |
| Brudgomseple* | admx | James Grieve * | 4 | Rosenrød * | 2 |
| Brureple * | 2 | Jens Pedersen * | 3 | Rosenstrips * | 3 |
| Carroll | 3 | Jomfrueple | 3 | Rossvolleple | 1 |
| Cellini * | 3 | Jordbæreple * | 3 | Royal Norfolk | 4 |
| Charlamowsky * | 3 | Julyred * | admx | Sandungeple | admx |
| Charles Ross * | 4 | Juteeple | 3 | Sävstaholm * | 1 |
| Close | admx | Karin Schneider | 4 | Signe Tillisch * | admx |
| Cox’s Pomona * | admx | Karlstrup | admx | Signeeple | 2 |
| Cox’s Orange * | 4 | Katja * | 4 | Silke-eple * | 2 |
| Daveeple | 1 | Katja 2 * | 4 | Sitroneple * | admx |
| Drengeple | 1 | Kaupanger * | 2 | Skaraeple | 2 |
| Dronningseple | admx | Kaviller | 1 | Slava Petersburg | admx |
| Dume Low’s Seedling | admx | Kjerringholm * | 3 | Slindreeple | 1 |
| EarlyRed bird * | 3 | Klovasteinseple | 2 | Sørnseple * | 1 |
| Eldraud Pigeon * | 3 | Knuteple * | 2 | Søteple | admx |
| Fatlandseple | admx | Krekeeple * | 3 | Stor Granat | admx |
| Filippa * | 3 | Kviteple * | admx | Stor Granat | admx |
| Fillippa 2 * | 3 | Lamyreple * | admx | Storesteinseple * | admx |
| Flaskeeple | 3 | Langballe * | 1 | Strutar * | admx |
| Førreple | admx | Langvikeple | 1 | Sukkereple * | 2 |
| Fosseple * | admx | Laxtons Superb * | 4 | Grønt Sureple | admx |
| Aroma Fragravoll | admx | Leiknes * | admx | Sysekavill * | admx |
| Fragravoll | admx | Leinestrand * | 1 | Telemark Vanvik | admx |
| Franskar * | 1 | Lobo * | 3 | Tohoku2 | admx |
| Fristaren * | admx | Løeple * | admx | Tokheimseple | 1 |
| Fuhr * | admx | Lord Lambourne * | 4 | Tolleivseple * | admx |
| Furuholm * | admx | Maglaeple | 1 | Tommos * | 3 |
| Garborg * | admx | Maglemer * | 1 | Tormodseple | admx |
| Geneva Early * | 3 | Marta-eple * | admx | Torstein | 2 |
| Gladstone | admx | Monarch | 3 | Raud Torstein * | 2 |
| Glasfalleple | 1 | Ølands Kungseple | 1 | Transparente Blanche | admx |
| Grågylling | 3 | Oldemorseple | 4 | Tveiteple * | 2 |
| Gravenstein * | admx | Oskaug * | 3 | Ulgenes * | 2 |
| Gravenstein 2 * | admx | Oster * | admx | Vågaeple | 3 |
| Grønt Lauspaeple | 4 | Ottøy fine | admx | Vintergul | admx |
| Grønvikeple | admx | Paradiseple | 3 | Virgisk Roseneple | 1 |
| Gul Granat | 2 | Prins * | 2 | VistaBella * | admx |
| Gul Richard * | 3 | Kronprinseple * | 2 | Wealthy * | 3 |
| Gullspir * | admx | Raud Prins * | 2 | Worcester Pearmain * | 4 |
| Locus Code | No. of Alleles | No. of Rare Alleles | No. of Effective Alleles | Size/Range of Different Alleles | Gene Diversity He |
|---|---|---|---|---|---|
| CH02B10 B | 18 | 12 | 5.79 | 182/264 | 0.83 |
| CH05E03 G | 22 | 18 | 5.44 | 144/196 | 0.81 |
| CH02C02a Y | 11 | 7 | 2.78 | 120/158 | 0.64 |
| CH02C02b B | 5 | 3 | 1.88 | 107/121 | 0.47 |
| CH04E02 G | 12 | 7 | 4.50 | 134/166 | 0.78 |
| CH02D08 Y | 14 | 9 | 6.36 | 206/258 | 0.84 |
| CH03D12 G | 20 | 15 | 6.57 | 97/153 | 0.85 |
| CH02C11 G | 14 | 4 | 8.27 | 201/235 | 0.88 |
| CH01D03 Y | 12 | 7 | 4.79 | 132/158 | 0.79 |
| CH01H01 B | 14 | 8 | 6.01 | 109/139 | 0.83 |
| CH01H02 B | 11 | 7 | 3.36 | 228/252 | 0.70 |
| CH01H10 G | 16 | 10 | 3.34 | 82/130 | 0.70 |
| CH05E04 G | 13 | 6 | 7.10 | 146/184 | 0.86 |
| CH02C06 B | 24 | 17 | 12.75 | 204/264 | 0.92 |
| CH01F07a B | 14 | 6 | 8.25 | 168/200 | 0.88 |
| CH04E03 G | 17 | 11 | 5.65 | 176/230 | 0.82 |
| CH01D09 Y | 20 | 15 | 6.20 | 128/164 | 0.84 |
| EMPC117 G | 9 | 5 | 1.88 | 90/112 | 0.47 |
| CH02C09 G | 11 | 5 | 5.65 | 232/258 | 0.82 |
| GD12 B | 9 | 5 | 1.96 | 144/188 | 0.49 |
| Average | 14.3 | 8.85 | 5.43 | 0.75 | |
| Fluorescent dyes labels: 6-FAM-B, HEX-G, and TAMRA-Y. | |||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Meland, M.; Aksic, M.F.; Frøynes, O.; Konjic, A.; Lasic, L.; Pojskic, N.; Gasi, F. Genetic Identity and Diversity of Apple Accessions within a Candidate Collection for the Norwegian National Clonal Germplasm Repository. Horticulturae 2022, 8, 630. https://doi.org/10.3390/horticulturae8070630
Meland M, Aksic MF, Frøynes O, Konjic A, Lasic L, Pojskic N, Gasi F. Genetic Identity and Diversity of Apple Accessions within a Candidate Collection for the Norwegian National Clonal Germplasm Repository. Horticulturae. 2022; 8(7):630. https://doi.org/10.3390/horticulturae8070630
Chicago/Turabian StyleMeland, Mekjell, Milica Fotiric Aksic, Oddmund Frøynes, Almira Konjic, Lejla Lasic, Naris Pojskic, and Fuad Gasi. 2022. "Genetic Identity and Diversity of Apple Accessions within a Candidate Collection for the Norwegian National Clonal Germplasm Repository" Horticulturae 8, no. 7: 630. https://doi.org/10.3390/horticulturae8070630
APA StyleMeland, M., Aksic, M. F., Frøynes, O., Konjic, A., Lasic, L., Pojskic, N., & Gasi, F. (2022). Genetic Identity and Diversity of Apple Accessions within a Candidate Collection for the Norwegian National Clonal Germplasm Repository. Horticulturae, 8(7), 630. https://doi.org/10.3390/horticulturae8070630







