Modelling of Catechin Extraction from Red Grape Solids under Conditions That Simulate Red Wine Fermentation
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental
2.1.1. Simulated Catechin Extraction—Experimental Design
2.1.2. Preparation of Grape Solids
2.1.3. Solid–Liquid Extractions
2.1.4. Catechin Quantification
2.2. Extraction Model Development
2.3. Model Fitting and Statistical Analysis
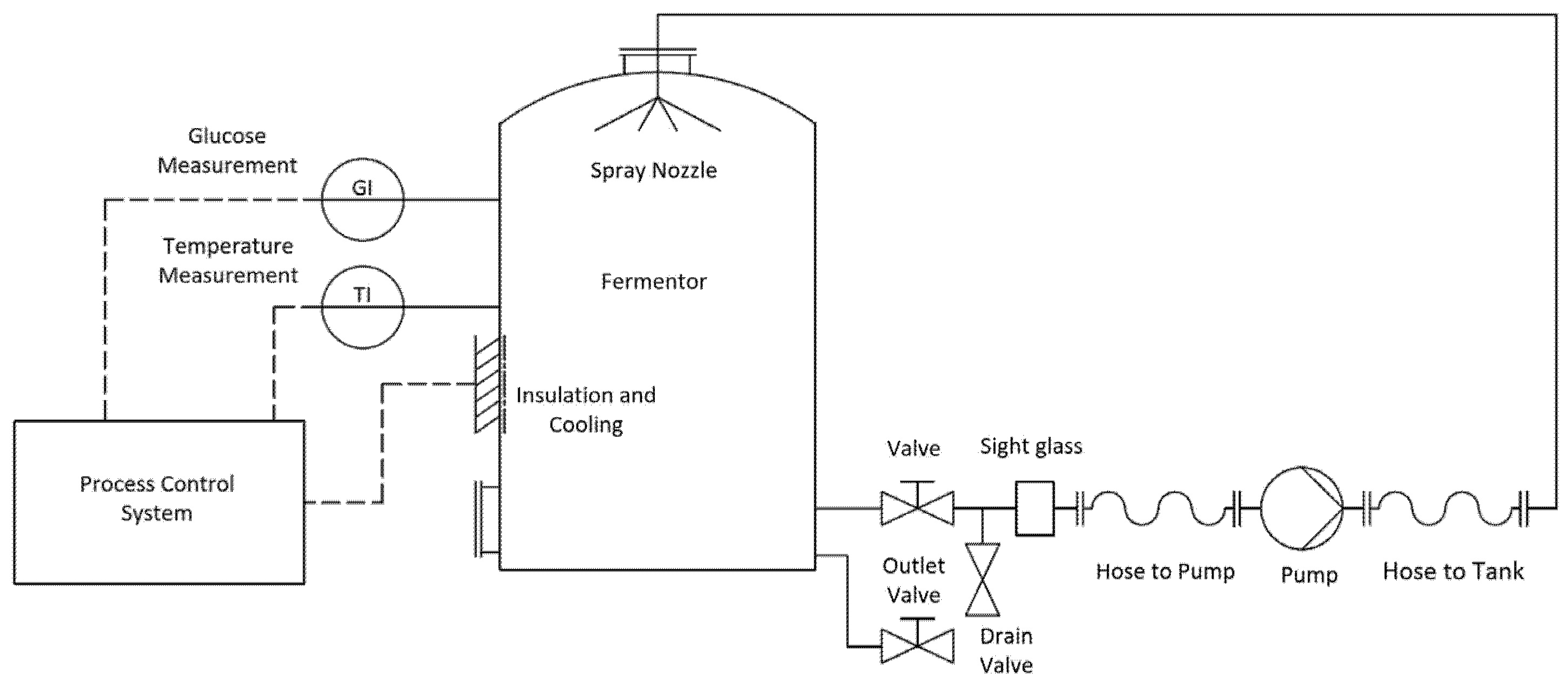
2.4. Control System Design
3. Results and Discussion
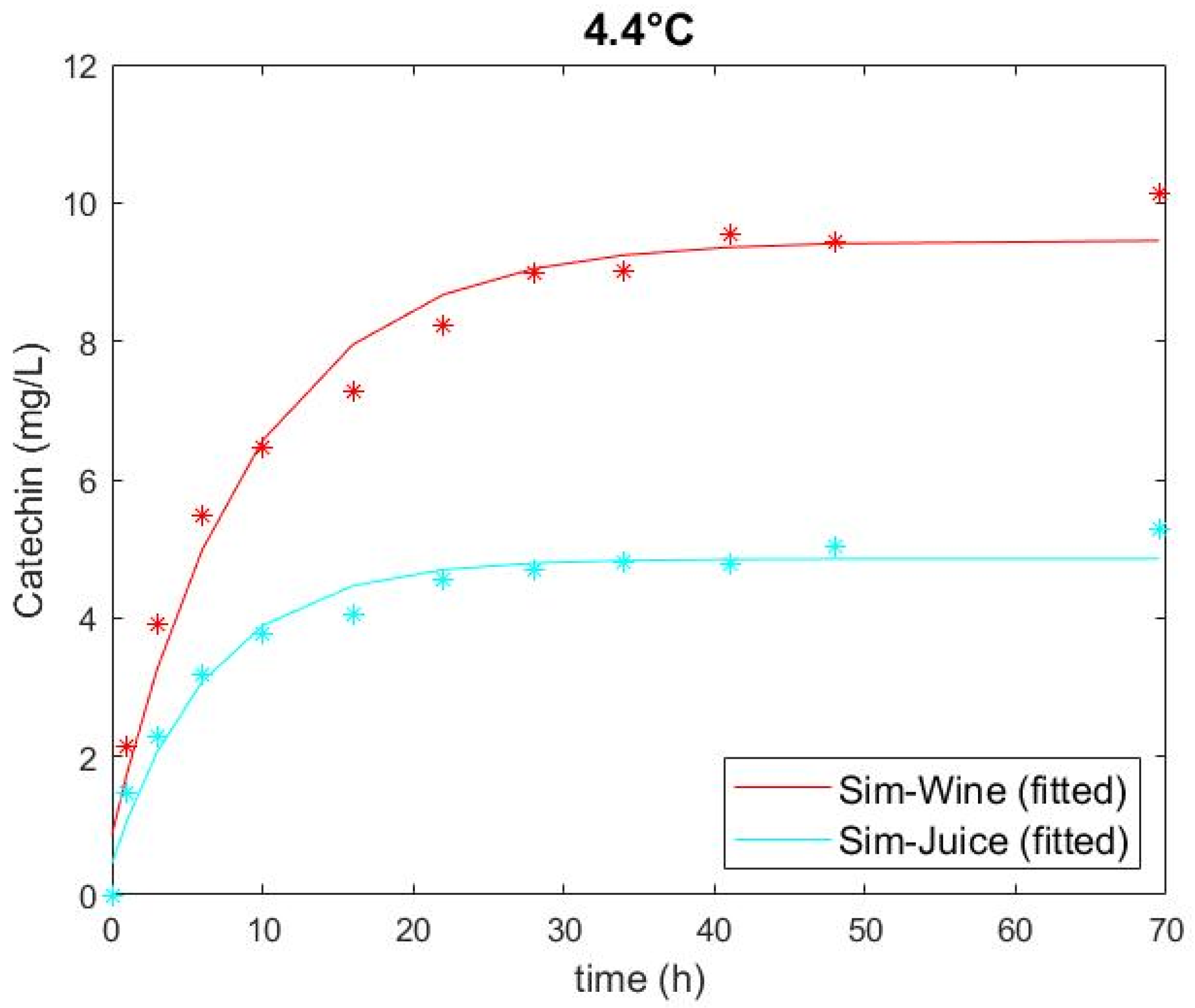
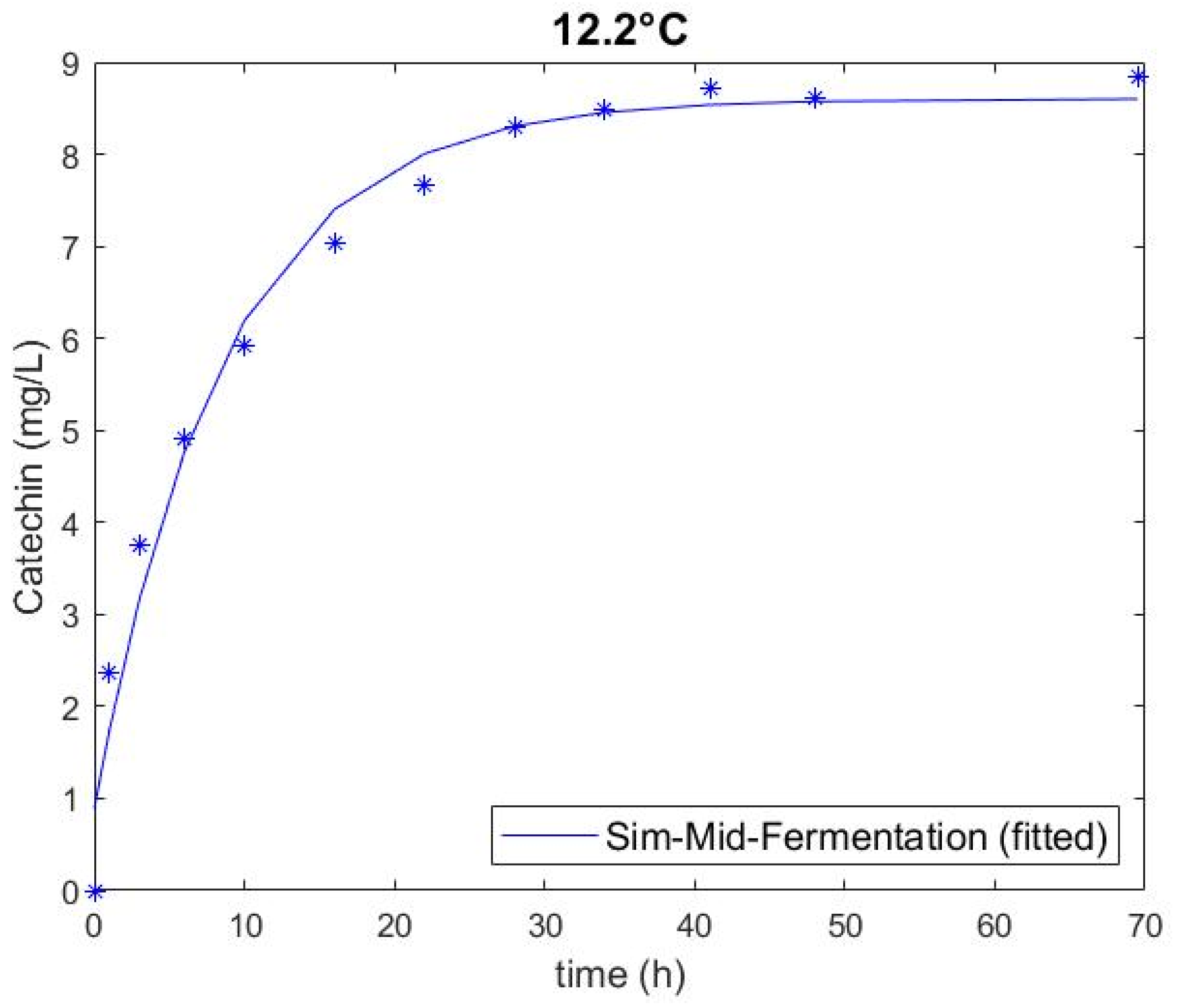
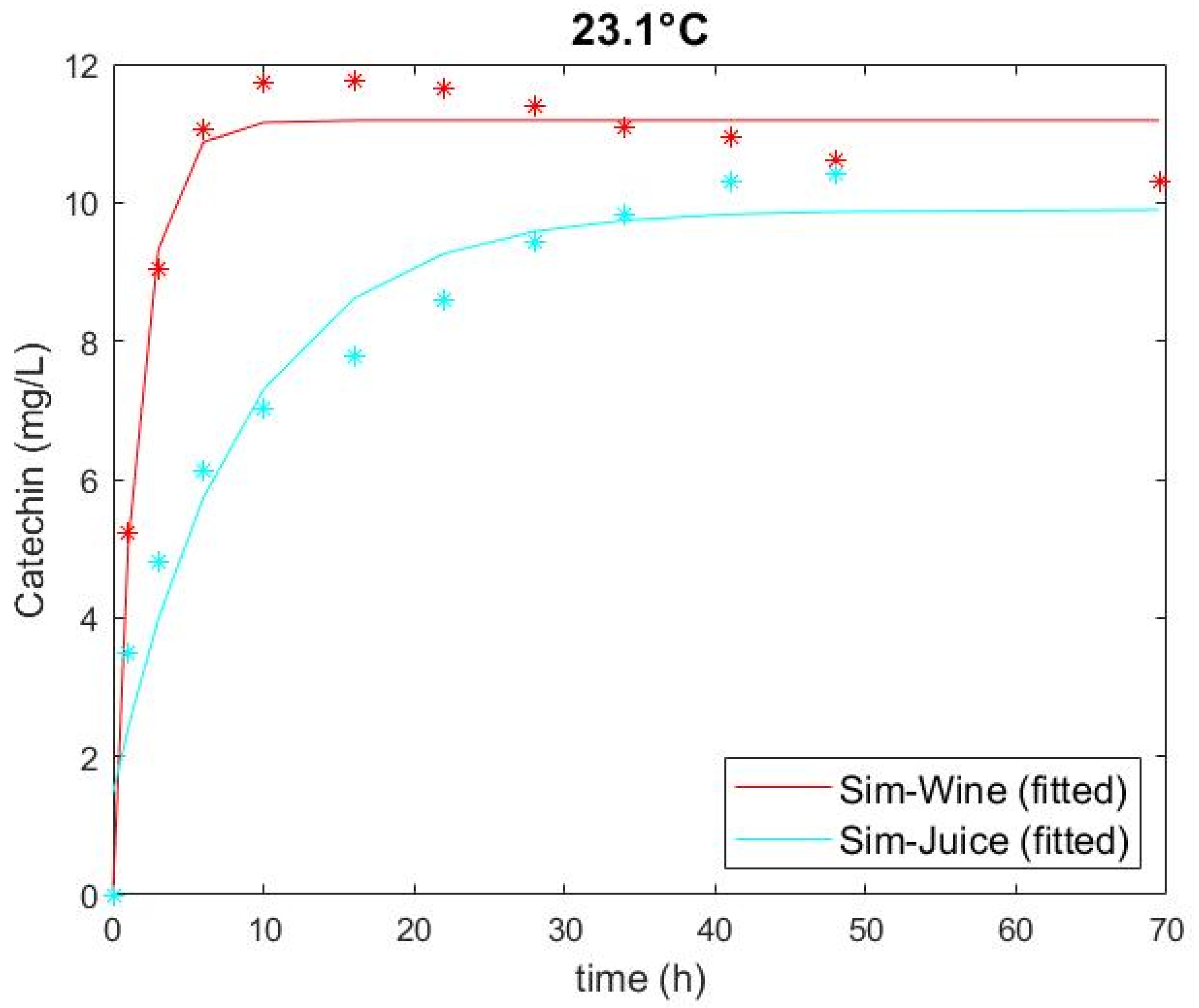
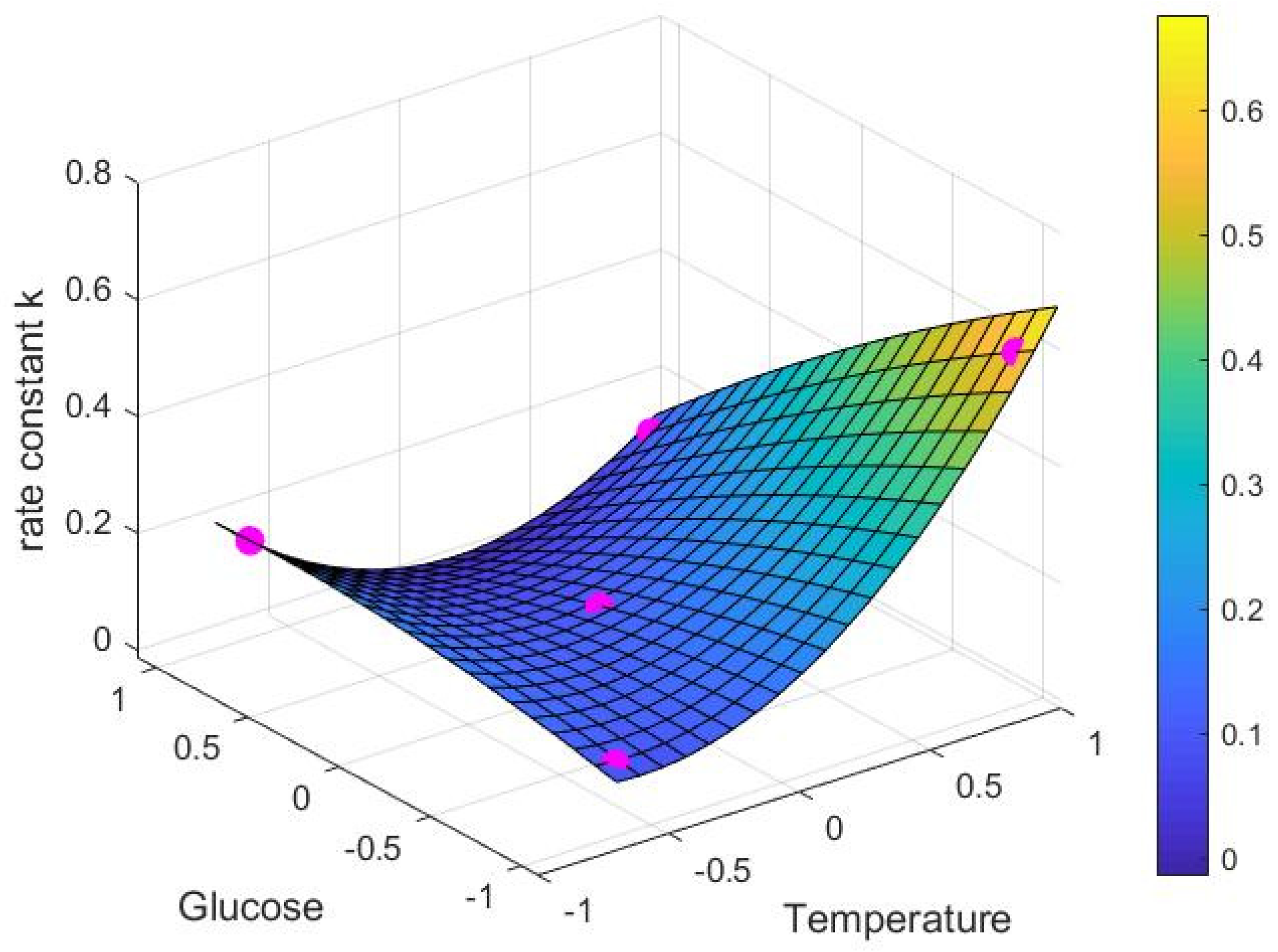
3.1. Experimental Catechin Extrction and Model Performance
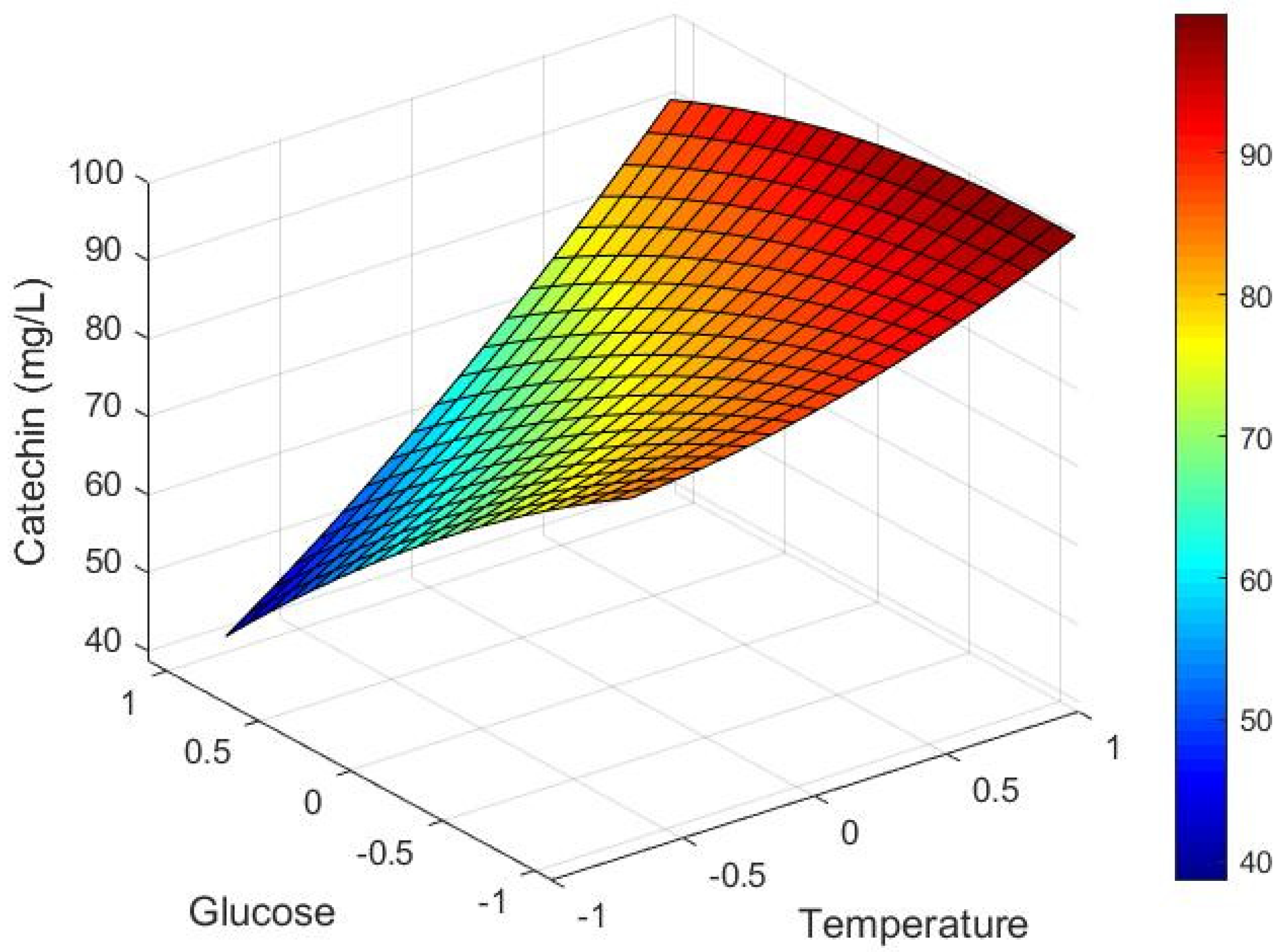
3.2. Industrial Application: Future Implementation of Models for Process Control
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Nomenclature
| C | Catechin amount (mg) |
| C0 | Initial catechin amount (mg) |
| Catechin concentration (mg/L) | |
| Maximal extractable catechin concentration (mg/L) | |
| Maximal extractable catechin concentration in undiluted industrial red wine fermentation (mg/L) | |
| Catechin concentration predicted by model (mg/L) | |
| Catechin concentration observed experimentally (mg/L) | |
| Mean of the catechin concentration observed experimentally (mg/L) | |
| c1-c6 | Constants describing catechin extraction rate |
| d1-d6 | Constants describing maximum extracted catechin |
| G | Glucose concentration (g/L) |
| G0 | Glucose concentration at centre point of the system (g/L) |
| Dimensionless glucose | |
| k | Catechin extraction rate (1/h) |
| N | Number of replicates |
| P(t) | Extractable amount of catechin in grape pomace at time t (mg) |
| R2 | Coefficient of determination |
| RMSE | Root mean square error |
| t | Time (h) |
| T | Temperature (°C) |
| T0 | Temperature at centre point of the system (°C) |
| Dimensionless temperature |
References
- Australia, W. Australian Wine: Production, Sales and Inventory 2021–22; Wine Australia: Adelaide, Australia, 2022. [Google Scholar]
- Unterkofler, J.; Muhlack, R.A.; Jeffery, D.W. Processes and purposes of extraction of grape components during winemaking: Current state and perspectives. Appl. Microbiol. Biotechnol. 2020, 104, 4737–4755. [Google Scholar] [CrossRef] [PubMed]
- Corriou, J.-P. Process Control, 2nd ed.; Springer: London, UK, 2009; ISBN 9783319611426. [Google Scholar]
- Schwenzer, M.; Ay, M.; Bergs, T.; Abel, D. Review on model predictive control: An engineering perspective. Int. J. Adv. Manuf. Technol. 2021, 117, 1327–1349. [Google Scholar] [CrossRef]
- Chai, W.Y.; Teo, K.T.K.; Tan, M.K.; Tham, H.J. Model Predictive Control in Fermentation Process—A Review. AIP Conf. Proc. 2022, 2610, 070008. [Google Scholar] [CrossRef]
- Fabri, S.G.; Agius, J.; Ghirlando, R.; Axisa, R. Modelling and temperature control of a wine fermentation process with solar cooling. In Proceedings of the 2016 24th Mediterranean Conference on Control and Automation (MED), Athens, Greece, 21–24 June 2016; pp. 985–990. [Google Scholar] [CrossRef]
- Jang, J.R. ANFIS: Adaptive-network-based fuzzy inference system. IEEE Trans. Syst. Man. Cybern. 1993, 23, 665–685. [Google Scholar] [CrossRef]
- Waterhouse, A.L.; Sacks, G.L.; Jeffery, D.W. Understanding Wine Chemistry; John Wiley & Sons Ltd.: Hoboken, NJ, USA, 2016; ISBN 9781118627808. [Google Scholar]
- Cheynier, V.; Dueñas-Paton, M.; Salas, E.; Maury, C.; Souquet, J.M.; Sarni-Manchado, P.; Fulcrand, H. Structure and Properties of Wine Pigments and Tannins. Am. J. Enol. Vitic. 2006, 57, 298–305. [Google Scholar] [CrossRef]
- Ribéreau-Gayon, P.; Glories, Y.; Maujean, A.; Dubourdieu, D. Handbook of Enology, Volume 2: The Chemistry of Wine Stabilization and Treatments, 2nd ed.; John Wiley & Sons, Ltd.: Chichester, UK, 2006; Volume 2, ISBN 9780470010396. [Google Scholar]
- Setford, P.C.; Jeffery, D.W.; Grbin, P.R.; Muhlack, R.A. Mass transfer of anthocyanins during extraction from pre-fermentative grape solids under simulated fermentation conditions: Effect of convective conditions. Molecules 2019, 24, 73. [Google Scholar] [CrossRef]
- Setford, P.C.; Jeffery, D.W.; Grbin, P.R.; Muhlack, R.A. Modelling the mass transfer process of malvidin-3-glucoside during simulated extraction from fresh grape solids under wine-like conditions. Molecules 2018, 23, 2159. [Google Scholar] [CrossRef]
- Setford, P.C.; Jeffery, D.W.; Grbin, P.R.; Muhlack, R.A. Mathematical modelling of anthocyanin mass transfer to predict extraction in simulated red wine fermentation scenarios. Food Res. Int. 2019, 121, 705–713. [Google Scholar] [CrossRef]
- Setford, P.C.; Jeffery, D.W.; Grbin, P.R.; Muhlack, R.A. A new approach to predicting the extraction of malvidin-3-glucoside during red wine fermentation at industrial-scale. Food Bioprod. Process. 2022, 131, 217–223. [Google Scholar] [CrossRef]
- Boulton, R.B.; Singleton, V.L.; Bisson, L.F.; Kunkee, R.E. Principles and Practices of Winemaking; Springer: Boston, MA, USA, 1999; ISBN 978-1-4419-5190-8. [Google Scholar]
- Miller, K.; Noguera, R.; Beaver, J.; Medina-Plaza, C.; Oberholster, A.; Block, D. A mechanistic model for the extraction of phenolics from grapes during red wine fermentation. Molecules 2019, 24, 1275. [Google Scholar] [CrossRef]
- Abi-Habib, E.; Vernhet, A.; Roi, S.; Carrillo, S.; Veran, F.; Ducasse, M.-A.; Poncet-Legrand, C. Diffusion of phenolic compounds during a model maceration. J. Sci. Food Agric. 2022, 103, 2004–2013. [Google Scholar] [CrossRef]
- Luna, R.; Lima, B.M.; Cuevas-valenzuela, J.; Normey-rico, J.E.; Pérez-correa, J.R. Optimal Control Applied to Oenological Management of Red Wine Fermentative Macerations. Fermentation 2021, 7, 94. [Google Scholar] [CrossRef]
- Zanoni, B.; Siliani, S.; Canuti, V.; Rosi, I.; Bertuccioli, M. A kinetic study on extraction and transformation phenomena of phenolic compounds during red wine fermentation. Int. J. Food Sci. Technol. 2010, 45, 2080–2088. [Google Scholar] [CrossRef]
- Hobbie, R.K.; Roth, B.J. Intermediate Physics for Medicine and Biology, 4th ed.; Springer: New York, NY, USA, 2007; ISBN 978-0-387-30942-2. [Google Scholar]
- Strauss, W.A. Partial Differential Equations: An Introduction, 2nd ed.; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2007; Volume 5, ISBN 9781119130536. [Google Scholar]
- Cussler, E.L. Diffusion: Mass Transfer in Fluid Systems, 3rd ed.; Cambridge University Press: Cambridge, UK, 2009; ISBN 9780521871211. [Google Scholar]
- Feynman, R.P.; Leighton, R.B.; Sands, M. The Feynman Lectures on Physics; California Institute of Technology: Pasadena, CA, USA, 1963; Volume 1, ISBN 978-0201021165. [Google Scholar]
- Haynes, W.M.; Lide, D.R.; Bruno, T.J. CRC Handbook of Chemistry and Physics, 97th ed.; CRC Press: Boca Raton, FL, USA, 2016; ISBN 9781498754293. [Google Scholar]
- Shi, J.; Yu, J.; Pohorly, J.; Young, J.C.; Bryan, M.; Wu, Y. Optimization of the extraction of polyphenols from grape seed meal by aqueous ethanol solution. J. Food Agric. Environ. 2003, 1, 42–47. [Google Scholar]
- Hernández-Jiménez, A.; Kennedy, J.A.; Bautista-Ortín, A.B.; Gómez-Plaza, E. Effect of ethanol on grape seed proanthocyanidin extraction. Am. J. Enol. Vitic. 2012, 63, 57–61. [Google Scholar] [CrossRef]
- The MathWorks Inc. Deep Learning Toolbox: User’s Guide (R2018a). Available online: https://au.mathworks.com/help/optim/ug/least-squares-model-fitting-algorithms.html (accessed on 13 December 2022).
- Strang, G. Introduction to Linear Algebra, 4th ed.; Wellesley-Cambridge Press: Wellesley, MA, USA, 2009; ISBN 0980232716. [Google Scholar]
- Marquardt, D.W. An algorithm for least-squares estimation of nonlinear parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Phipps, M.C.; Quine, M.P. A Primer of Statistics: Data Analysis, Probability, Inference, 3rd ed.; Prentice Hall: Sydney, Australia, 1998; ISBN 0724809619. [Google Scholar]
- Montgomery, D.C. Design and Analysis of Experiments, 8th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2013; ISBN 9781118146927. [Google Scholar]
- Franklin, G.F.; Powell, J.D.; Emami-Naeini, A. Feedback Control of Dynamic Systems, 7th ed.; Pearson Education: London, UK, 2015; ISBN 1292068906. [Google Scholar]
- Lennart, L. System Identification, 2nd ed.; Pearson: London, UK, 1998; ISBN 0136566952. [Google Scholar]
- Ibrahim, J.; Chen, M.-H.; Sinha, D. The Elements of Statistical Learning, 2nd ed.; Springer: Berlin/Heidelberg, Germany, 2009; Volume 27, ISBN 9781441968241. [Google Scholar]
- Cozzolino, D.; Dambergs, R.G.; Janik, L.; Cynkar, W.U.; Gishen, M. Analysis of grapes and wine by near infrared spectroscopy. J. Near Infrared Spectrosc. 2006, 14, 279–289. [Google Scholar] [CrossRef]
- Tindal, R.A.; Jeffery, D.W.; Muhlack, R.A. Mathematical modelling to enhance winemaking efficiency: A review of red wine colour and polyphenol extraction and evolution. Aust. J. Grape Wine Res. 2021, 27, 219–233. [Google Scholar] [CrossRef]
- Wang, X.; Lim, L.-T. Modeling study of coffee extraction at different temperature and grind size conditions to better understand the cold and hot brewing process. J. Food Process Eng. 2021, 44, e13748. [Google Scholar] [CrossRef]
- Meziane, I.A.A.; Bali, N.; Belblidia, N.-B.; Abatzoglou, N.; Benyoussef, E.H. The first-order model in the simulation of essential oil extraction kinetics. J. Appl. Res. Med. Aromat. Plants 2019, 15, 100226. [Google Scholar] [CrossRef]
- Simonin, J.P. On the comparison of pseudo-first order and pseudo-second order rate laws in the modeling of adsorption kinetics. Chem. Eng. J. 2016, 300, 254–263. [Google Scholar] [CrossRef]
- Sacchi, K.L.; Bisson, L.F.; Adams, D.O. A review of the effect of winemaking techniques on phenolic extraction in red wines. Am. J. Enol. Vitic. 2005, 56, 197–206. [Google Scholar] [CrossRef]
- Nagel, C.W.; Wulf, L.W. Changes in the Anthocyanins, Flavonoids and Hydroxycinnamic Acid Esters during Fermentation and Aging of Merlot and Cabernet Sauvignon. Am. J. Enol. Vitic. 1979, 30, 111–116. [Google Scholar] [CrossRef]
- Landrault, N.; Poucheret, P.; Ravel, P.; Gasc, F.; Cros, G.; Teissedre, P.L. Antioxidant capacities and phenolics levels of French wines from different varieties and vintages. J. Agric. Food Chem. 2001, 49, 3341–3348. [Google Scholar] [CrossRef] [PubMed]
- Faustino, R.S.; Sobrattee, S.; Edel, A.L.; Pierce, G.N. Comparative analysis of the phenolic content of selected Chilean, Canadian and American Merlot red wines. Mol. Cell. Biochem. 2003, 249, 11–19. [Google Scholar] [CrossRef]
- Bindon, K.A.; Madani, S.H.; Pendleton, P.; Smith, P.A.; Kennedy, J.A. Factors affecting skin tannin extractability in ripening grapes. J. Agric. Food Chem. 2014, 62, 1130–1141. [Google Scholar] [CrossRef]
- Rousserie, P.; Lacampagne, S.; Vanbrabant, S.; Rabot, A.; Geny-Denis, L. Influence of berry ripeness on seed tannins extraction in wine. Food Chem. 2020, 315, 126307. [Google Scholar] [CrossRef]
- VanderWeide, J.; Del Zozzo, F.; Nasrollahiazar, E.; Kennedy, J.A.; Peterlunger, E.; Rustioni, L.; Sabbatini, P. Influence of freezing and heating conditions on grape seed flavan-3-ol extractability, oxidation, and galloylation pattern. Sci. Rep. 2022, 12, 3838. [Google Scholar] [CrossRef]
- Oberholster, A.; Elmendorf, B.L.; Lerno, L.A.; King, E.S.; Heymann, H.; Brenneman, C.E.; Boulton, R.B. Barrel maturation, oak alternatives and micro-oxygenation: Influence on red wine aging and quality. Food Chem. 2015, 173, 1250–1258. [Google Scholar] [CrossRef]
- Nordestgaard, S. Inspirations from the past and opportunities for the future part 2: In-tank fermentation monitoring and continuous processes. Aust. N. Z. Grapegrow. Winemak. 2020, 677, 50–56. [Google Scholar]
- Kondakci, T.; Zhou, W. Recent applications of advanced control techniques in food industry. Food Bioprocess Technol. 2017, 10, 522–542. [Google Scholar] [CrossRef]
- Mjalli, F.S.; Al-Asheh, S. Neural-networks-based feedback linearization versus model predictive control of continuous alcoholic fermentation process. Chem. Eng. Technol. 2005, 28, 1191–1200. [Google Scholar] [CrossRef]
- Scrimgeour, N. Evaluating the Viability of Process Sensor Technologies for Measurement of Sugar Levels during Fermentation; Australian Wine Research Institute: Adelaide, Australia, 2015. [Google Scholar]
- Jaywant, S.A.; Singh, H.; Arif, K.M. Sensors and instruments for brix measurement: A review. Sensors 2022, 22, 2290. [Google Scholar] [CrossRef] [PubMed]
- Nichols, J.A.; Herbert Chan, H.W.; Baker, M.A.B. Machine learning: Applications of artificial intelligence to imaging and diagnosis. Biophys. Rev. 2019, 11, 111–118. [Google Scholar] [CrossRef] [PubMed]

| (°C) | ||
|---|---|---|
| T(low) | 4.4 | −0.639 |
| T(med) | 12.2 | 0 |
| T(high) | 23.1 | 0.893 |
| (g/L) | ||
|---|---|---|
| G(low) | 0 | −1 |
| G(med) | 133 | 0 |
| G(high) | 266 | 1 |
| Trial Conditions | Model Parameters | Model Fit | ||||||
|---|---|---|---|---|---|---|---|---|
| Temp. (°C) | Glucose (g/L) | Ethanol (% v/v) | 0 (mg/L) | ∞ (mg/L) | k (1/h) | RMSE | R2 | |
| Sim-Juice | Low (4.4) | 266 | 0 | 0.452 | 4.856 | 0.152 | 0.268 | 0.971 |
| Sim-Wine | Low (4.4) | 0 | 14 | 0.867 | 9.457 | 0.109 | 0.480 | 0.976 |
| Sim-Mid-Ferment | Med (12.2) | 133 | 7 | 0.889 | 8.603 | 0.116 | 0.407 | 0.979 |
| Sim-Juice | High (23.1) | 266 | 0 | 1.473 | 9.898 | 0.118 | 0.737 | 0.944 |
| Sim-Wine | High (23.1) | 0 | 14 | 0.059 | 11.190 | 0.595 | 0.451 | 0.984 |
| Constant | Fitted Value (h−1) |
|---|---|
| c1 | 0.116 |
| c2 | 0.090 |
| c3 | −0.087 |
| c4 | 0.225 |
| c5 | −0.021 |
| c6 | −0.170 |
| Constant | Fitted Value (mg/L) |
|---|---|
| d1 | 8.603 |
| d2 | 2.072 |
| d3 | −1.610 |
| d4 | 0.544 |
| d5 | −0.344 |
| d6 | 1.079 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Unterkofler, J.; Jeffery, D.W.; Setford, P.C.; Macintyre, J.; Muhlack, R.A. Modelling of Catechin Extraction from Red Grape Solids under Conditions That Simulate Red Wine Fermentation. Fermentation 2023, 9, 394. https://doi.org/10.3390/fermentation9040394
Unterkofler J, Jeffery DW, Setford PC, Macintyre J, Muhlack RA. Modelling of Catechin Extraction from Red Grape Solids under Conditions That Simulate Red Wine Fermentation. Fermentation. 2023; 9(4):394. https://doi.org/10.3390/fermentation9040394
Chicago/Turabian StyleUnterkofler, Judith, David W. Jeffery, Patrick C. Setford, Jean Macintyre, and Richard A. Muhlack. 2023. "Modelling of Catechin Extraction from Red Grape Solids under Conditions That Simulate Red Wine Fermentation" Fermentation 9, no. 4: 394. https://doi.org/10.3390/fermentation9040394
APA StyleUnterkofler, J., Jeffery, D. W., Setford, P. C., Macintyre, J., & Muhlack, R. A. (2023). Modelling of Catechin Extraction from Red Grape Solids under Conditions That Simulate Red Wine Fermentation. Fermentation, 9(4), 394. https://doi.org/10.3390/fermentation9040394







