Genome-Scale Modeling-Guided Metabolic Engineering Enables Heterologous Production of 3-Amino-4-hydroxybenzoic Acid in Streptomyces thermoviolaceus
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains, Plasmids, and Enzymes
2.2. Plasmid Construction
2.3. Preparation of Transformant Strains
2.4. Fermentation and 3,4-AHBA Analysis
2.5. Model Simulations
2.5.1. Flux Balance Analysis (FBA)
2.5.2. Construction of the Metabolic Model
2.5.3. Simulation Settings and Parameter Constraints
2.5.4. Introduction of the 3,4-AHBA Biosynthetic Pathway
3. Results and Discussion
3.1. In Silico Simulation of 3,4-AHBA Production in S. thermoviolaceus Strains
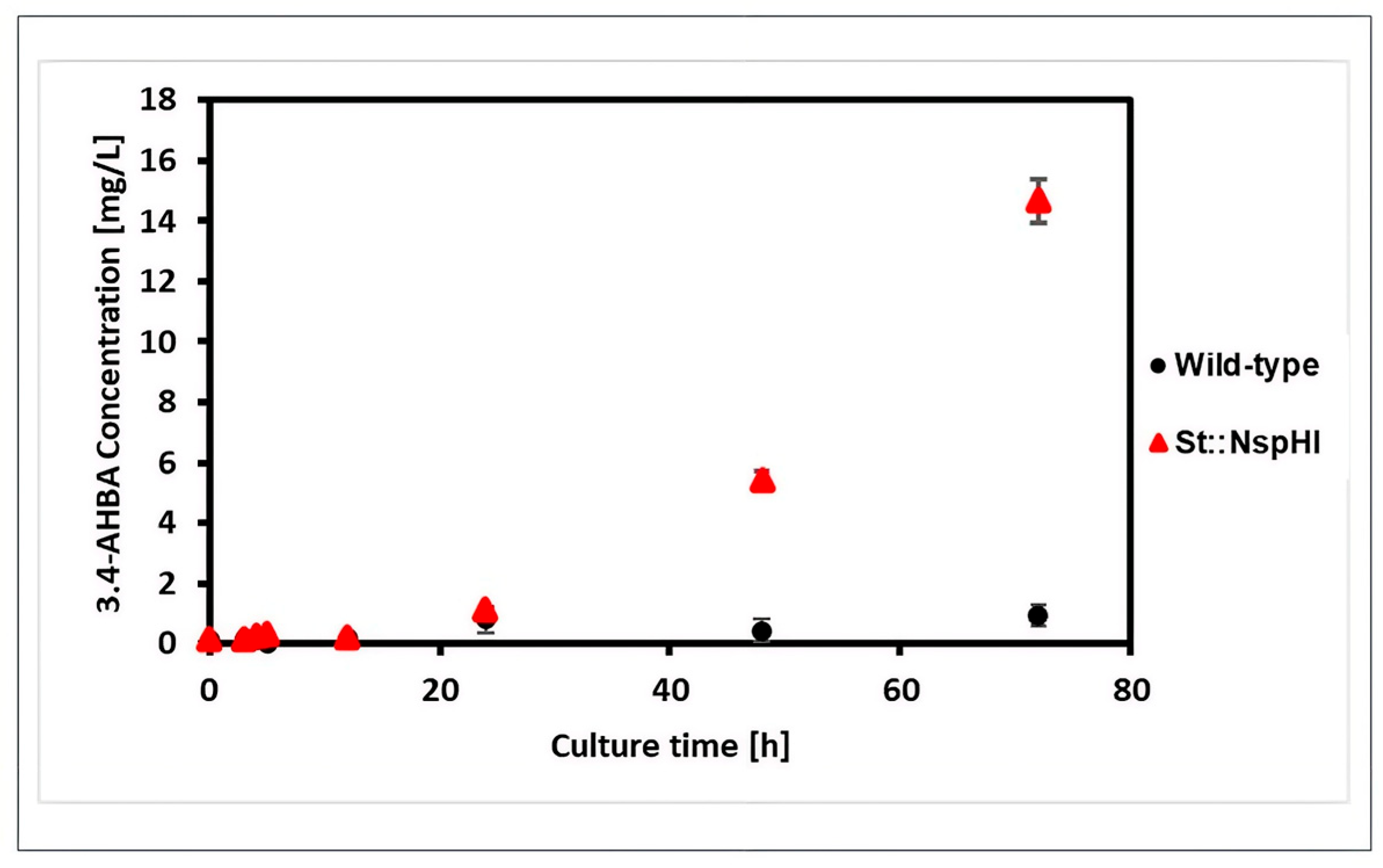
3.2. Effect of nspH–nspI Insertion on 3,4-AHBA Production



3.3. Comparison of FBA Predictions with Experimental Data
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| 3,4-AHBA | 3-amino-4-hydroxybenzoic acid |
| GSM | Genome-Scale Metabolic modeling |
| GEM | Genome-scale metabolic model |
| FBA | Flux Balance Analysis |
| S. thermoviolaceus | Streptomyces thermoviolaceus |
| GAM | Growth-associated maintenance |
| NGAM | Non-growth-associated maintenance |
| NBRC | NITE Biological Resource Center |
| NCBI | National Center for Biotechnology Information |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
References
- Osterlund, T.; Nookaew, I.; Bordel, S.; Nielsen, J. Mapping Condition-Dependent Regulation of Metabolism in Yeast through Genome-Scale Modeling. BMC Syst. Biol. 2013, 7, 36. [Google Scholar] [CrossRef]
- Vikromvarasiri, N.; Shirai, T.; Kondo, A. Metabolic Engineering Design to Enhance (R,R)-2,3-Butanediol Production from Glycerol in Bacillus Subtilis Based on Flux Balance Analysis. Microb. Cell Fact. 2021, 20, 196. [Google Scholar] [CrossRef]
- Feist, A.M.; Henry, C.S.; Reed, J.L.; Krummenacker, M.; Joyce, A.R.; Karp, P.D.; Broadbelt, L.J.; Hatzimanikatis, V.; Palsson, B.O. A Genome-scale Metabolic Reconstruction for Escherichia coli K-12 MG1655 That Accounts for 1260 ORFs and Thermodynamic Information. Mol. Syst. Biol. 2007, 3, 121. [Google Scholar] [CrossRef]
- Orth, J.D.; Thiele, I.; Palsson, B.Ø. What Is Flux Balance Analysis? Nat. Biotechnol. 2010, 28, 245–248. [Google Scholar] [CrossRef]
- Simensen, V.; Schulz, C.; Karlsen, E.; Bråtelund, S.; Burgos, I.; Thorfinnsdottir, L.B.; García-Calvo, L.; Bruheim, P.; Almaas, E. Experimental Determination of Escherichia coli Biomass Composition for Constraint-Based Metabolic Modeling. PLoS ONE 2022, 17, e0262450. [Google Scholar] [CrossRef]
- Tokuyama, K.; Ohno, S.; Yoshikawa, K.; Hirasawa, T.; Tanaka, S.; Furusawa, C.; Shimizu, H. Increased 3-Hydroxypropionic Acid Production from Glycerol, by Modification of Central Metabolism in Escherichia coli. Microb. Cell Fact. 2014, 13, 64. [Google Scholar] [CrossRef]
- Bentley, S.D.; Chater, K.F.; Cerdeño-Tárraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete Genome Sequence of the Model Actinomycete Streptomyces Coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef]
- Kashiwagi, N.; Ogino, C.; Kondo, A. Production of Chemicals and Proteins Using Biomass-Derived Substrates from a Streptomyces Host. Bioresour. Technol. 2017, 245, 1655–1663. [Google Scholar] [CrossRef]
- Lee, N.; Hwang, S.; Kim, W.; Lee, Y.; Kim, J.H.; Cho, S.; Kim, H.U.; Yoon, Y.J.; Oh, M.-K.; Palsson, B.O.; et al. Systems and Synthetic Biology to Elucidate Secondary Metabolite Biosynthetic Gene Clusters Encoded in Streptomyces Genomes. Nat. Prod. Rep. 2021, 38, 1330–1361. [Google Scholar] [CrossRef]
- Tsujibo, H.; Hatano, N.; Endo, H.; Miyamoto, K.; Inamori, Y. Purification and Characterization of a Thermostable Chitinase from Streptomyces thermoviolaceus OPC-520 and Cloning of the Encoding Gene. Biosci. Biotechnol. Biochem. 2000, 64, 96–102. [Google Scholar] [CrossRef]
- Onaka, H.; Taniguchi, S.-I.; Ikeda, H.; Igarashi, Y.; Furumai, T. PTOYAMAcos, PTYM18, and PTYM19, Actinomycete-Escherichia coli Integrating Vectors for Heterologous Gene Expression. J. Antibiot. 2003, 56, 950–956. [Google Scholar] [CrossRef]
- Kawaguchi, H.; Sasaki, K.; Uematsu, K.; Tsuge, Y.; Teramura, H.; Okai, N.; Nakamura-Tsuruta, S.; Katsuyama, Y.; Sugai, Y.; Ohnishi, Y.; et al. 3-Amino-4-Hydroxybenzoic Acid Production from Sweet Sorghum Juice by Recombinant Corynebacterium glutamicum. Bioresour. Technol. 2015, 198, 410–417. [Google Scholar] [CrossRef]
- Kim, M.; Sang Yi, J.; Kim, J.; Kim, J.; Kim, M.W.; Kim, B. Reconstruction of a High-quality Metabolic Model Enables the Identification of Gene Overexpression Targets for Enhanced Antibiotic Production in Streptomyces coelicolor A3(2). Biotechnol. J. 2014, 9, 1185–1194. [Google Scholar] [CrossRef]
- Orth, J.D.; Fleming, R.M.T.; Palsson, B.Ø. Reconstruction and Use of Microbial Metabolic Networks: The Core Escherichia Coli Metabolic Model as an Educational Guide. EcoSal Plus 2010, 4, 10-1128. [Google Scholar] [CrossRef]
- Becker, S.A.; Feist, A.M.; Mo, M.L.; Hannum, G.; Palsson, B.Ø.; Herrgard, M.J. Quantitative Prediction of Cellular Metabolism with Constraint-Based Models: The COBRA Toolbox. Nat. Protoc. 2007, 2, 727–738. [Google Scholar] [CrossRef]
- Kawaguchi, H.; Hasunuma, T.; Ohnishi, Y.; Sazuka, T.; Kondo, A.; Ogino, C. Enhanced Production of γ-Amino Acid 3-Amino-4-Hydroxybenzoic Acid by Recombinant Corynebacterium glutamicum under Oxygen Limitation. Microb. Cell Fact. 2021, 20, 228. [Google Scholar] [CrossRef]
- Huang, D.; Wen, J.; Wang, G.; Yu, G.; Jia, X.; Chen, Y. In Silico Aided Metabolic Engineering of Streptomyces roseosporus for Daptomycin Yield Improvement. Appl. Microbiol. Biotechnol. 2012, 94, 637–649. [Google Scholar] [CrossRef]
- Rubio, A.; Razmilic, V.; Brain-Isasi, S.; Andrews, B.; Asenjo, J.A. Metabolic Engineering of Streptomyces leeuwenhoekii C34 T to Increase Chaxamycin Production Based on the i VR1007 Genome-Scale Model. Biotechnol. Bioeng. 2025, 122, 3376–3392. [Google Scholar] [CrossRef]
- Niimi-Nakamura, S.; Kawaguchi, H.; Uematsu, K.; Teramura, H.; Nakamura-Tsuruta, S.; Kashiwagi, N.; Sugai, Y.; Katsuyama, Y.; Ohnishi, Y.; Ogino, C.; et al. 3-Amino-4-Hydroxybenzoic Acid Production from Glucose and/or Xylose via Recombinant Streptomyces lividans. J. Gen. Appl. Microbiol. 2022, 68, 109–116. [Google Scholar] [CrossRef]
- Suzuki, H.; Ohnishi, Y.; Furusho, Y.; Sakuda, S.; Horinouchi, S. Novel Benzene Ring Biosynthesis from C3 and C4 Primary Metabolites by Two Enzymes. J. Biol. Chem. 2006, 281, 36944–36951. [Google Scholar] [CrossRef]
- Ohnishi, Y.; Furusho, Y.; Higashi, T.; Chun, H.-K.; Furihata, K.; Sakuda, S.; Horinouchi, S. Structures of Grixazone A and B, A-Factor-Dependent Yellow Pigments Produced under Phosphate Depletion by Streptomyces griseus. J. Antibiot. 2004, 57, 218–223. [Google Scholar] [CrossRef][Green Version]
- Martinet, L.; Naômé, A.; Deflandre, B.; Maciejewska, M.; Tellatin, D.; Tenconi, E.; Smargiasso, N.; De Pauw, E.; Van Wezel, G.P.; Rigali, S. A Single Biosynthetic Gene Cluster Is Responsible for the Production of Bagremycin Antibiotics and Ferroverdin Iron Chelators. mBio 2019, 10, e01230-19. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Yan, Q.; Jones, J.A.; Tang, Y.J.; Fong, S.S.; Koffas, M.A.G. Metabolic Burden: Cornerstones in Synthetic Biology and Metabolic Engineering Applications. Trends Biotechnol. 2016, 34, 652–664. [Google Scholar] [CrossRef]
- Asah-Asante, R.; Tang, L.; Gong, X.; Fan, S.; Yan, C.; Asante, J.O.; Zeng, Q. Exploring Pigment-Producing Streptomyces as an Alternative Source to Synthetic Pigments: Diversity, Biosynthesis, and Biotechnological Applications. A Review. World J. Microbiol. Biotechnol. 2025, 41, 211. [Google Scholar] [CrossRef]

| Materials | Description | Source |
|---|---|---|
| Strains: | ||
| E. coli Nova Blue | For routine plasmid maintenance (endA1 hsdR17 (rK12– mK12+) supE44 thi-1 recA1 gyrA96 relA1 lac F′[proA+B+ lacIqZΔM15::Tn10] (TetR) | |
| E. coli HST04 | Cloning host. ((F−, ara, Δ(lacproAB) [Φ80dlacZΔM15], rpsL(str), thi, Δ(mrr-hsdRMS-mcrBC), ΔmcrA, dam, dcm)) | Takara |
| E. coli JM109 | For conjugation plasmid into Streptomyces (endA1 recA1 gyrA96 thi-1 hsdR17 (rK−, mK+) relA1 supE44 Δ(lac-proAB)/F’ [traD36 proA+B+ lacIq lacZΔM15]) | Takara |
| Strptomyces thermoviolaceus NBRC 13905 | Expression host | NBRC |
| St::NspHI | Strain of Streptomyces thermoviolaceus::nspH::nspI | This study |
| Plasmids: | ||
| pTYM18 | [11] | |
| pTYM18-ermE-nspHI | Plasmid for nspHI expression; apramycin resistance marker; kasO promoter; pld terminator; rep, replication gene | This study |
| PUB307 | PΔTnAc,IncPαrepliconKmrTCr |
| Primers | Description |
|---|---|
| ermEp-MCS_Fw (N.1) | ggccagtgccaagcttCACGCGGTCGATCTTGACGGCTG |
| ermEp-MCS-PLDt_Rv (N.2) | ggatccggtaccCGCTGGATCCTACCAACCGGCAC- |
| PLDt_Fw | GCGggtaccggatcctctagaTGAGACGACTGAGCGCCCGGAC |
| PLDt_Rv | acatgattacgaattcATTTCCTCGCTGGTCGGTTC |
| nspHI-tandem1-Hind3-Fw (N.3) | gccagtgccaagcttCACGCGGTCGATCTTGACG |
| nspHI-tandem2-Xba1-Rv (N.4) | GTCGTCTCAtctagactacttctccaggcagaactc |
| Wild-Type | St::NspHI | ||||
|---|---|---|---|---|---|
| Glucose Uptake Rate [mmol gDCW−1 h−1] | Experimental Growth [h−1] | In Silico Growth [h−1] | Glucose Uptake Rate [mmol gDCW−1 h−1] | Experimental Growth [h−1] | In Silico Growth [h−1] |
| −12.02 | 0.916 | 0.874 | −3.087 | 0.0736 | 0.336 |
| Parameter (mmol/gDCW/h) | S. thermoviolaceus | |
|---|---|---|
| Wild Type | St::NspHI | |
| * Glucose consumption | −12.02 | −3.087 |
| * Biomass (Growth rate) [h−1] | 0.916 | 0.0377 |
| Oxygen consumption | −10 | −10 |
| DHAP exchange | 11.98 | 3.084 |
| ASA exchange | 0.0257 | 0.726 |
| 3,4-AHBA exchange | 0 | 0.724 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Yamada, T.; Apriliana, P.; Kahar, P.; Kobayashi, T.; Mori, Y.; Ogino, C. Genome-Scale Modeling-Guided Metabolic Engineering Enables Heterologous Production of 3-Amino-4-hydroxybenzoic Acid in Streptomyces thermoviolaceus. Fermentation 2026, 12, 108. https://doi.org/10.3390/fermentation12020108
Yamada T, Apriliana P, Kahar P, Kobayashi T, Mori Y, Ogino C. Genome-Scale Modeling-Guided Metabolic Engineering Enables Heterologous Production of 3-Amino-4-hydroxybenzoic Acid in Streptomyces thermoviolaceus. Fermentation. 2026; 12(2):108. https://doi.org/10.3390/fermentation12020108
Chicago/Turabian StyleYamada, Togo, Pamella Apriliana, Prihardi Kahar, Tomoya Kobayashi, Yutaro Mori, and Chiaki Ogino. 2026. "Genome-Scale Modeling-Guided Metabolic Engineering Enables Heterologous Production of 3-Amino-4-hydroxybenzoic Acid in Streptomyces thermoviolaceus" Fermentation 12, no. 2: 108. https://doi.org/10.3390/fermentation12020108
APA StyleYamada, T., Apriliana, P., Kahar, P., Kobayashi, T., Mori, Y., & Ogino, C. (2026). Genome-Scale Modeling-Guided Metabolic Engineering Enables Heterologous Production of 3-Amino-4-hydroxybenzoic Acid in Streptomyces thermoviolaceus. Fermentation, 12(2), 108. https://doi.org/10.3390/fermentation12020108









