Modeling and Optimization of the Culture Medium for Efficient 4′-N-Demethyl-Vicenistatin Production by Streptomyces parvus Using Response Surface Methodology and Artificial-Neural-Network-Genetic-Algorithm
Abstract
1. Introduction
2. Results
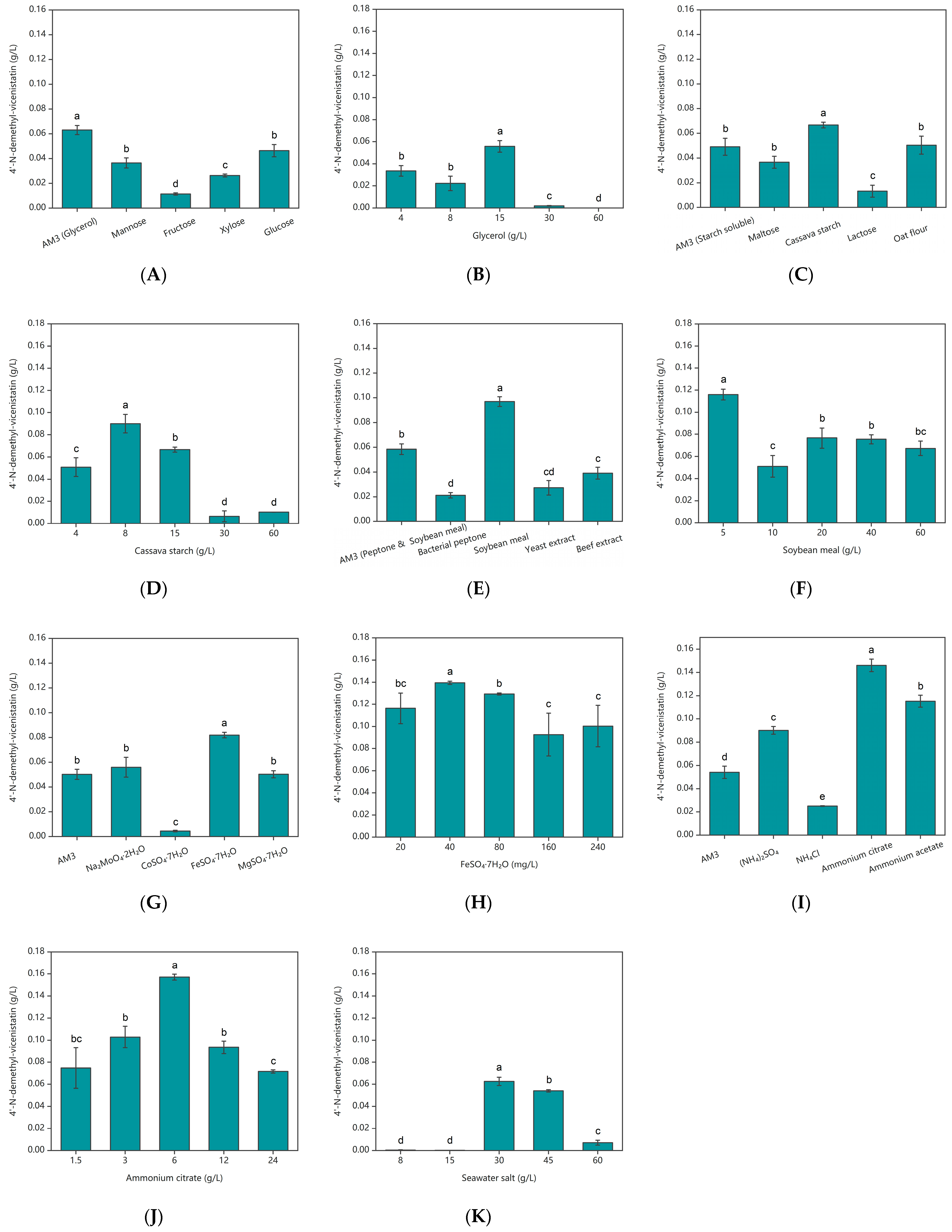
2.1. Screening and Optimizing the Medium Compositions by OFAT
2.2. Searching for the Most Significant Medium Components by Plackett–Burman Design (PBD)
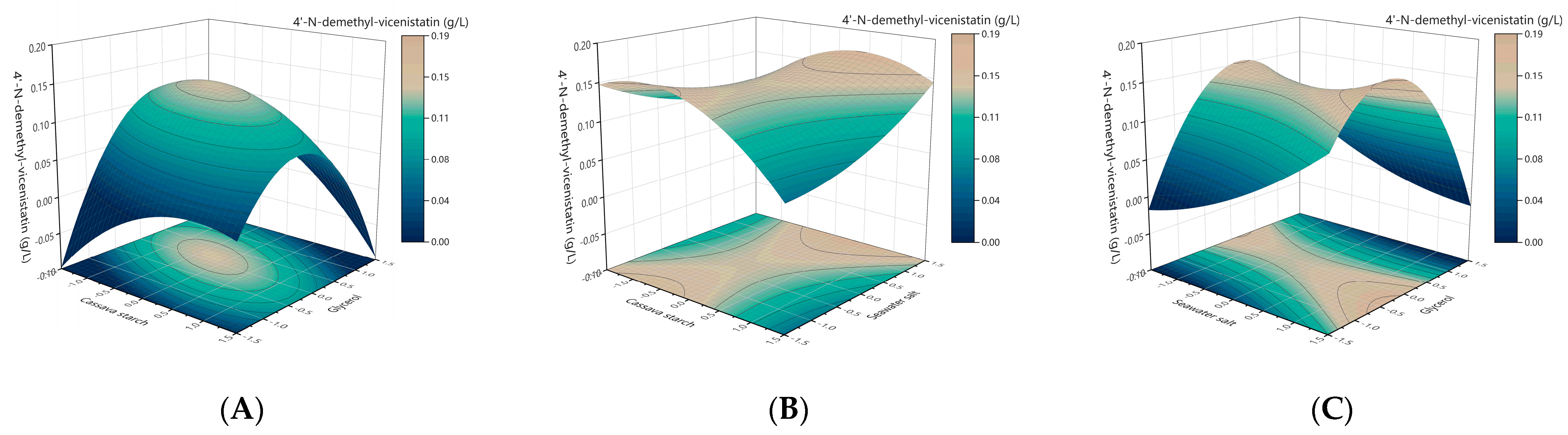
2.3. Modeling and Optimization of Medium Compositions by Response Surface Methodology (RSM)
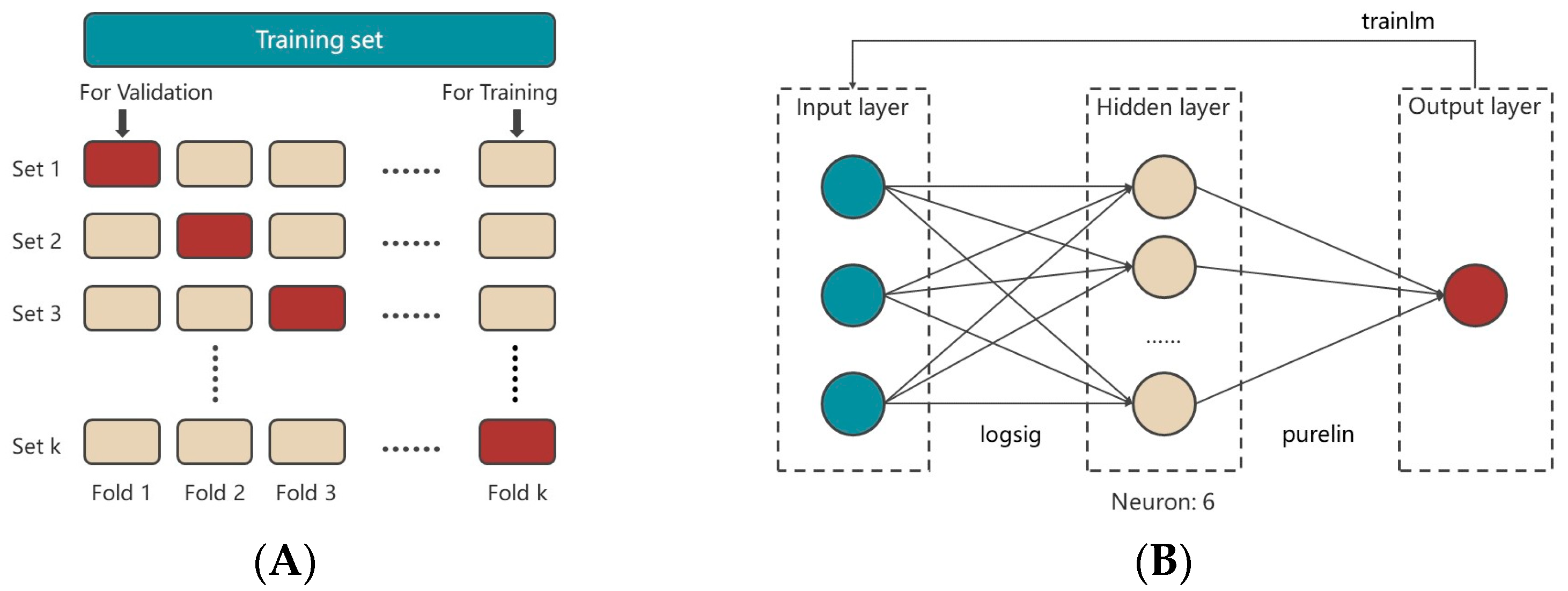
2.4. Modeling and Optimization of Medium Compositions by Artificial-Neural-Network-Genetic-Algorithm (ANN-GA)
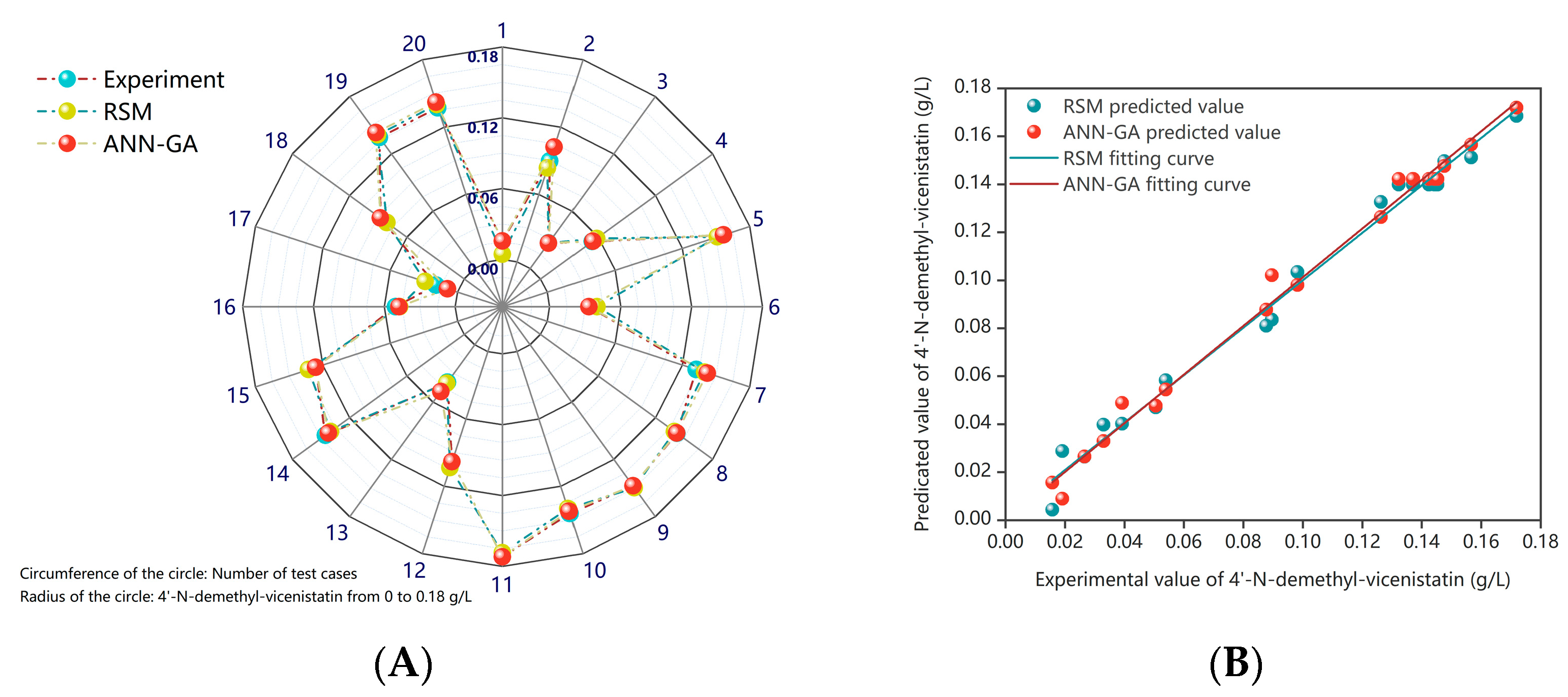
2.5. Comparison and Assessment of RSM and ANN-GA Models
3. Discussion
4. Materials and Methods
4.1. Bacterial Materials, Culture Medium, and Fermentation Conditions
4.2. Medium Optimization for Efficient 4′-N-Demethyl-Vicenistatin Production
4.2.1. Screening and Optimizing the Medium Compositions by One-Factor-at-a-Time (OFAT)
4.2.2. Searching for the Significant Medium Components by Plackett–Burman Design (PBD)
4.2.3. Modeling and Optimization of the Medium Compositions by Response Surface Methodology (RSM)
4.2.4. Modeling and Optimization of the Medium Compositions by Artificial-Neural-Network-Genetic-Algorithm (ANN-GA)
4.3. Analytical Method
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sánchez, S.; Chávez, A.; Forero, A.; García-Huante, Y.; Romero, A.; Sánchez, M.; Rocha, D.; Sánchez, B.; Ávalos, M.; Guzmán-Trampe, S.; et al. Carbon source regulation of antibiotic production. J. Antibiot. 2010, 63, 442–459. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Lu, Y.Y.; Cao, S.G. Antimicrobial compounds from marine actinomycetes. Arch. Pharm. Res. 2020, 43, 677–704. [Google Scholar] [CrossRef]
- Hutchings, M.I.; Truman, A.W.; Wilkinson, B. Antibiotics: Past, present and future. Curr. Opin. Microbiol. 2019, 51, 72–80. [Google Scholar] [CrossRef]
- Zhang, W.; Ma, S.; An, C.; Xue, W. Antibacterial Activity and Composition of the Fermentation Broth of Streptomyces parvus 33. Jundishapur J. Microbiol. 2017, 10, e12677. [Google Scholar] [CrossRef]
- El-Naggar, N.E.A. Extracellular Production of the Oncolytic Enzyme, L-Asparaginase, by Newly Isolated Streptomyces sp. Strain NEAE-95 as Potential Microbial Cell Factories: Optimization of Culture Conditions Using Response Surface Methodology. Curr. Pharm. Biotechnol. 2015, 16, 162–178. [Google Scholar] [CrossRef]
- Yang, Y.-K.; Shen, D.-D.; He, P.; Du, L.-D.; Wan, D.-J.; Wang, P.; Wang, T.; Feng, M.-Q. Chemically synthesized LYRM03 could inhibit the metastasis of human breast cancer MDA-MB-231 cells in vitro and in vivo. Bioorg. Med. Chem. Lett. 2019, 29, 1719–1726. [Google Scholar] [CrossRef]
- Bayram, S.; Dengiz, C.; Gercek, Y.C.; Cetin, I.; Topcul, M.R. Bioproduction, structure elucidation and in vitro antiproliferative effect of eumelanin pigment from Streptomyces parvus BSB49. Arch. Microbiol. 2020, 202, 2401–2409. [Google Scholar] [CrossRef]
- Al-Dhabi, N.A.; Ghilan, A.-K.M.; Esmail, G.A.; Arasu, M.V.; Duraipandiyan, V.; Ponmurugan, K. Environmental friendly synthesis of silver nanomaterials from the promising Streptomyces parvus strain Al-Dhabi-91 recovered from the Saudi Arabian marine regions for antimicrobial and antioxidant properties. J. Photochem. Photobiol. B-Biol. 2019, 197, 111529. [Google Scholar] [CrossRef]
- Shindo, K.; Kamishohara, M.; Odagawa, A.; Matsuoka, M.; Kawai, H. Vicenistatin, a novel 20-membered macrocyclic lactam antitumor antibiotic. J. Antibiot. 1993, 46, 1076–1081. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yang, Z.; Shi, C.; Wu, X.; Zhou, L.; Liang, Y.; Li, Q.; Ju, J. Semi-synthesis and structure-activity relationship study yield antibacterial vicenistatin derivatives with low cytotoxicity. J. Antibiot. 2024, 77, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.C.; Li, J.; Ling, C.Y.; Xu, R.; Yi, X.X.; Ju, J.H.; Li, Q.L. Characterization of the Aminosugar Biosynthetic and Regulatory Genes of Vicenistatin in Monodonata labio-Associated Streptomyces parvus SCSIO Mla-L010. J. Nat. Prod. 2022, 85, 256–263. [Google Scholar] [CrossRef]
- Qi, W.; Yong-Chun, S.; Hao, X.; Ye, G.; Jing, L.; Ren-Xiang, T. Medium optimization for enhanced co-production of two bioactive metabolites in the same fermentation by a statistical approach. J. Asian Nat. Prod. Res. 2011, 13, 1110–1121. [Google Scholar]
- Shivamathi, C.S.; Gunaseelan, S.; Soosai, M.R.; Vignesh, N.S.; Varalakshmi, P.; Kumar, R.S.; Karthikumar, S.; Kumar, R.V.; Baskar, R.; Rigby, S.P.; et al. Process optimization and characterization of pectin derived from underexploited pineapple peel biowaste as a value-added product. Food Hydrocoll. 2022, 123, 107141. [Google Scholar] [CrossRef]
- Hu, X.; Xu, F.; Li, J.; Li, J.; Mo, C.; Zhao, M.; Wang, L. Ultrasonic-assisted extraction of polysaccharides from coix seeds: Optimization, purification, and in vitro digestibility. Food Chem. 2022, 374, 131636. [Google Scholar] [CrossRef]
- Kowalczyk-Jusko, A.; Pochwatka, P.; Zaborowicz, M.; Czekala, W.; Mazurkiewicz, J.; Mazur, A.; Janczak, D.; Marczuk, A.; Dach, J. Energy value estimation of silages for substrate in biogas plants using an artificial neural network. Energy 2020, 202, 117729. [Google Scholar] [CrossRef]
- Yang, J.; Huang, Y.; Xu, H.; Gu, D.; Xu, F.; Tang, J.; Fang, C.; Yang, Y. Optimization of fungi co-fermentation for improving anthraquinone contents and antioxidant activity using artificial neural networks. Food Chem. 2020, 313, 126138. [Google Scholar] [CrossRef]
- Shakeel, Q.; Lyu, A.; Zhang, J.; Wu, M.D.; Chen, S.W.; Chen, W.D.; Li, G.Q.; Yang, L. Optimization of the cultural medium and conditions for production of antifungal substances by Streptomyces platensis 3–10 and evaluation of its efficacy in suppression of clubroot disease (Plasmodiophora brassicae) of oilseed rape. Biol. Control 2016, 101, 59–68. [Google Scholar] [CrossRef]
- Ismet, A.; Vikineswary, S.; Paramaswari, S.; Wong, W.H.; Ward, A.; Seki, T.; Fiedler, H.P.; Goodfellow, M. Production and chemical characterization of antifungal metabolites from Micromonospora sp. M39 isolated from mangrove rhizosphere soil. World J. Microbiol. Biotechnol. 2004, 20, 523–528. [Google Scholar] [CrossRef]
- Raytapadar, S.; Paul, A.K. Production of an antifungal antibiotic by Streptomyces aburaviensis 1DA-28. Microbiol. Res. 2001, 155, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.Y.; Li, J.J.; Li, L.Z.; Zhang, H.Y.; Ma, J.; Chen, Z.; Hu, C.Y.; Ju, X.; Fu, J.L. Improvement of FK506 Production by Selection of 4-Aminobutyric Acid-tolerant Mutant and Optimization of its Fermentation Using Response Surface Methodology. J. Korean Soc. Appl. Biol. Chem. 2014, 57, 715–722. [Google Scholar] [CrossRef]
- Chen, J.; Lan, X.; Jia, R.; Hu, L.; Wang, Y. Response Surface Methodology (RSM) Mediated Optimization of Medium Components for Mycelial Growth and Metabolites Production of Streptomyces alfalfae XN-04. Microorganisms 2022, 10, 1854. [Google Scholar] [CrossRef]
- Yu, F.; Zhang, M.; Sun, J.; Wang, F.; Li, X.; Liu, Y.; Wang, Z.; Zhao, X.; Li, J.; Chen, J.; et al. Improved Neomycin Sulfate Potency in Streptomyces fradiae Using Atmospheric and Room Temperature Plasma (ARTP) Mutagenesis and Fermentation Medium Optimization. Microorganisms 2022, 10, 94. [Google Scholar] [CrossRef]
- Zhang, Y.; Bai, J.; Wu, C.; Wang, Y.; Ju, X.; Qi, X.; Li, L.; Ji, L.; Fu, J. Efficient production of epsilon-poly-L-lysine using cassava starch and fish meal by Streptomyces albulus FQC-24. Prep. Biochem. Biotechnol. 2022, 52, 525–533. [Google Scholar] [CrossRef]
- Hawary, H.; Rasmey, A.H.M.; Aboseidah, A.A.; El-Morsi, E.; Hafez, M. Enhancement of glycerol production by UV-mutagenesis of the marine yeast Wickerhamomyces anomalus HH16: Kinetics and optimization of the fermentation process. 3 Biotech 2019, 9, 446. [Google Scholar] [CrossRef]
- Lai, L.S.T.; Pan, C.C.; Tzeng, B.K. Medium optimization for lovastatin production by Aspergillus terreus in submerged cultures. J. Chin. Inst. Chem. Eng. 2002, 33, 517–527. [Google Scholar]
- El-Naggar, N.E.-A.; Moawad, H.; Abdelwahed, N.A.M. Optimization of fermentation conditions for enhancing extracellular production of L-asparaginase, an anti-leukemic agent, by newly isolated Streptomyces brollosae NEAE-115 using solid state fermentation. Ann. Microbiol. 2017, 67, 1–15. [Google Scholar] [CrossRef]
- Mayer, A.F.; Deckwer, W.D. Simultaneous production and decomposition of clavulanic acid during Streptomyces clavuligerus cultivations. Appl. Microbiol. Biotechnol. 1996, 45, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Bao, Y.; Ma, Z.; Zhou, J.; Chen, H.; Lu, Y.; Zhu, L.; Chen, X. Optimization of fermentation medium and conditions for enhancing valinomycin production by Streptomyces sp. ZJUT-IFE-354. Prep. Biochem. Biotechnol. 2022, 53, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Xia, B.; Zou, H.; Li, L.; Zhang, B.; Xiang, Y.; Zou, Y.; Shen, Z.; Xue, S.; Han, Y. Screening and fermentation medium optimization of a strain favorable to Rice-fish Coculture. Front. Microbiol. 2022, 13, 1054797. [Google Scholar] [CrossRef] [PubMed]
- Rehder. Bioinorganic Chemistry; Oxford University Press: Oxford, UK, 2014. [Google Scholar]
- Ge, Y.; Zhang, K. Fermentation Medium Useful for Culturing and Fermenting Pseudomonas fluorescens to Prepare Mupirocin Comprises Sodium Molybdate and Urea. CN111996136-A, 24 July 2020.
- DU Wen, C.C. Effects of Co2+ on the erythromycin biosynthesis. Front. Chem. Sci. Eng. 2007, 1, 246–250. [Google Scholar] [CrossRef]
- Zhu, T.; Wang, L.; Wang, W.; Hu, Z.; Yu, M.; Wang, K.; Cui, Z. Enhanced production of lipstatin from Streptomyces toxytricini by optimizing fermentation conditions and medium. J. Gen. Appl. Microbiol. 2014, 60, 106–111. [Google Scholar] [CrossRef]
- Ju, Y.; Son, K.-H.; Jin, C.; Hwang, B.S.; Park, D.-J.; Kim, C.-J. Statistical optimization of culture medium for improved production of antimicrobial compound by Streptomyces rimosus AG-P1441. Food Sci. Biotechnol. 2018, 27, 581–590. [Google Scholar] [CrossRef] [PubMed]
- Mojicevic, M.; D’Agostino, P.M.; Pavic, A.; Vojnovic, S.; Senthamaraikannan, R.; Vasiljevic, B.; Gulder, T.A.M.; Nikodinovic-Runic, J. Streptomyces sp. BV410 isolate from chamomile rhizosphere soil efficiently produces staurosporine with antifungal and antiangiogenic properties. Microbiologyopen 2020, 9, e986. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Gu, Q.; Zhou, T. Statistical Optimization of Novel Medium to Maximize the Yield of Exopolysaccharide from Lacticaseibacillus rhamnosus ZFM216 and Its Immunomodulatory Activity. Front. Nutr. 2022, 9, 924495. [Google Scholar] [CrossRef] [PubMed]
- Aruwajoye, G.S.; Faloye, F.D.; Kassim, A.; Saha, A.K.; Kana, E.G. Intelligent modelling of fermentable sugar concentration and combined severity factor (CSF) index from pretreated starch-based lignocellulosic biomass. Biomass Convers. Biorefinery 2022. [Google Scholar] [CrossRef]
- Suganthi, V.; Mohanasrinivasan, V. Optimization studies for enhanced bacteriocin production by Pediococcus pentosaceus KC692718 using response surface methodology. J. Food Sci. Technol. 2015, 52, 3773–3783. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, Q.; Li, Y.; Zhang, H. Modeling and optimization of photo-fermentation biohydrogen production from co-substrates basing on response surface methodology and artificial neural network integrated genetic algorithm. Bioresour. Technol. 2023, 374, 128789. [Google Scholar] [CrossRef] [PubMed]
- Mondal, P.; Sadhukhan, A.K.; Ganguly, A.; Gupta, P. Optimization of process parameters for bio-enzymatic and enzymatic saccharification of waste broken rice for ethanol production using response surface methodology and artificial neural network-genetic algorithm. 3 Biotech 2021, 11, 28. [Google Scholar] [CrossRef]
- Pirmoradi, N.; Ghaneian, M.T.; Ehrampoush, M.H.; Salmani, M.H.; Hatami, B. The conversion of poultry slaughterhouse wastewater sludge into biodiesel: Process modeling and optimization. Renew. Energy 2021, 178, 1236–1249. [Google Scholar] [CrossRef]
- Alizadeh, M.J.; Shabani, A.; Kavianpour, M.R. Predicting longitudinal dispersion coefficient using ANN with metaheuristic training algorithms. Int. J. Environ. Sci. Technol. 2017, 14, 2399–2410. [Google Scholar] [CrossRef]
- Wei, P.L.; Si, Z.J.; Lu, Y.; Yu, Q.F.; Huang, L.; Xu, Z.N. Medium optimization for pyrroloquinoline quinone (PQQ) production by Methylobacillus sp. zju323 using response surface methodology and artificial neural network-genetic algorithm. Prep. Biochem. Biotechnol. 2017, 47, 709–719. [Google Scholar] [CrossRef]
- Liu, Y.; Gong, H.; Shi, C.; Yuan, H.; Zuo, X.; Chang, Y.; Li, X. Modeling and optimization of the hydrolysis and acidification via liquid fraction of digestate from corn straw by response surface methodology and artificial neural network. J. Clean. Prod. 2022, 361, 132241. [Google Scholar] [CrossRef]
- Cai, H.; Li, X.; Li, D.; Liu, W.; Han, Y.; Xu, X.; Yang, P.; Meng, K. Optimization of Gamma-Aminobutyric Acid Production by Lactiplantibacillus plantarum FRT7 from Chinese Paocai. Foods 2023, 12, 3034. [Google Scholar] [CrossRef]
- Ibrahim, A.G.; Baazeem, A.; Al-Zaban, M.I.; Fawzy, M.A.; Hassan, S.H.A.; Koutb, M. Sustainable Biodiesel Production from a New Oleaginous Fungus, Aspergillus carneus Strain OQ275240: Biomass and Lipid Production Optimization Using Box–Behnken Design. Sustainability 2023, 15, 6836. [Google Scholar]
- Pan, M.; Zhu, M.; Jiang, H.; Zhang, X.; Tan, J. The Characterization of the Inhibitory Substances Produced by Bacillus pumilus LYMC-3 and the Optimization of Fermentation Conditions. Fermentation 2023, 9, 966. [Google Scholar] [CrossRef]
- Ahsan, T.; Liang, C.; Yu, S.; Pei, X.; Xie, J.; Lin, Y.; Liu, X.; Umair, M.; Zang, C. Screening and Optimization of Fermentation Medium for Bacillus velezensis BP-1 and Its Biocontrol Effects against Peyronellaea arachidicola. Appl. Sci. 2023, 13, 4653. [Google Scholar] [CrossRef]
- Djinni, I.; Djoudi, W.; Boumezoued, C.; Barchiche, H.; Souagui, S.; Kecha, M.; Mancini, I. Statistical Medium Optimization for the Production of Anti-Methicillin-Resistant Staphylococcus aureus Metabolites from a Coal-Mining-Soil-Derived Streptomyces rochei CMB47. Fermentation 2023, 9, 381. [Google Scholar] [CrossRef]
- He, Z.; Sun, Y.; Chu, M.; Zhu, J.; Zhang, Y.; Tang, Q.; Osman, G.; Jiang, L.; Zhang, Z. Screening of a Novel Fibrinolytic Enzyme-Producing Streptomyces from a Hyper-Arid Area and Optimization of Its Fibrinolytic Enzyme Production. Fermentation 2023, 9, 410. [Google Scholar] [CrossRef]
- Ruiz, B.; Chávez, A.; Forero, A.; García-Huante, Y.; Romero, A.; Snchez, M.; Rocha, D.; Snchez, B.; Rodríguez-Sanoja, R.; Sánchez, S.; et al. Production of microbial secondary metabolites: Regulation by the carbon source. Crit. Rev. Microbiol. 2010, 36, 146–167. [Google Scholar] [CrossRef]
- Lebrihi, A.; Lefebvre, G.; Germain, P. Carbon catabolite regulation of cephamycin C and expandase biosynthesis in Streptomyces clavuligerus. Appl. Microbiol. Biotechnol. 1988, 28, 44–51. [Google Scholar] [CrossRef]
- Kim, E.S.; Hong, H.J.; Choi, C.Y.; Cohen, S.N. Modulation of actinorhodin biosynthesis in Streptomyces lividans by glucose repression of afsR2 gene transcription. J. Bacteriol. 2001, 183, 2198–2203. [Google Scholar] [CrossRef]
- Lounès, A.; Lebrihi, A.; Benslimane, C.; Lefebvre, G.; Germain, P. Regulation of spiramycin synthesis in Streptomyces ambofaciens: Effects of glucose and inorganic phosphate. Appl. Microbiol. Biotechnol. 1996, 45, 204–211. [Google Scholar] [CrossRef]
- Wei, Z.H.; Chen, N.; Li, Y.J.; Fan, Q.L.; Yu, T.F.; Wang, K.X.; Dong, B.T.; Fan, E.Y.; Yuan, P.L.; Hu, G.W.; et al. Glucose fed-batch integrated dissolved oxygen control strategy enhanced polysaccharide, total triterpenoids and inotodiol production in fermentation of a newly isolated Inonotus obliquus strain. Process Biochem. 2018, 66, 1–6. [Google Scholar] [CrossRef]
- Ma, Y.F.; Shen, W.; Chen, X.Z.; Liu, L.; Zhou, Z.M.; Xu, F.; Yang, H.Q. Significantly enhancing recombinant alkaline amylase production in Bacillus subtilis by integration of a novel mutagenesis-screening strategy with systems-level fermentation optimization. J. Biol. Eng. 2016, 10, 13. [Google Scholar] [CrossRef]
- Li, S.; Li, F.; Chen, X.S.; Wang, L.; Xu, J.; Tang, L.; Mao, Z.G. Genome Shuffling Enhanced ε-Poly-L-Lysine Production by Improving Glucose Tolerance of Streptomyces graminearus. Appl. Biochem. Biotechnol. 2012, 166, 414–423. [Google Scholar] [CrossRef]
- van Wezel, G.P.; McDowall, K.J. The regulation of the secondary metabolism of Streptomyces: New links and experimental advances. Nat. Prod. Rep. 2011, 28, 1311–1333. [Google Scholar] [CrossRef]
- Romero-Rodríguez, A.; Rocha, D.; Ruiz-Villafán, B.; Guzmán-Trampe, S.; Maldonado-Carmona, N.; Vázquez-Hernández, M.; Zelarayán, A.; Rodríguez-Sanoja, R.; Sánchez, S. Carbon catabolite regulation in Streptomyces: New insights and lessons learned. World J. Microbiol. Biotechnol. 2017, 33, 162. [Google Scholar] [CrossRef]
- Romero-Rodríguez, A.; Maldonado-Carmona, N.; Ruiz-Villafán, B.; Koirala, N.; Rocha, D.; Sánchez, S. Interplay between carbon, nitrogen and phosphate utilization in the control of secondary metabolite production in Streptomyces. Antonie Leeuwenhoek 2018, 111, 761–781. [Google Scholar] [CrossRef]
- Cotârlet, M.; Bahrim, G.E. Optimization of cold-adapted amylases and protease production by psychrotrophic Streptomyces 4 Alga using response surface methodology. Turk. J. Biochem. 2011, 36, 83–92. [Google Scholar]
- Cotârlet, M. Medium Optimization for the Production of Cold-Active Beta Amylase by Psychrotrophic Streptomyces MIUG 4 Alga Using Response Surface Methodology. Microbiology 2013, 82, 147–154. [Google Scholar] [CrossRef]
- Smaoui, S.; Ennouri, K.; Chakchouk-Mtibaa, A.; Sellem, I.; Bouchaala, K.; Karray-Rebai, I.; Mellouli, L. Statistical versus artificial intelligence -based modeling for the optimization of antifungal activity against Fusarium oxysporum using Streptomyces sp. strain TN71. J. Mycol. Med. 2018, 28, 551–560. [Google Scholar] [CrossRef] [PubMed]
- Al-Ansari, M.; Kalaiyarasi, M.; Almalki, M.A.; Vijayaraghavan, P. Optimization of medium components for the production of antimicrobial and anticancer secondary metabolites from Streptomyces sp. AS11 isolated from the marine environment. J. King Saud Univ. Sci. 2020, 32, 1993–1998. [Google Scholar] [CrossRef]
- Ni, H.J.; Lv, S.Y.; Sheng, Y.T.; Wang, H.; Chu, X.H.; Zhang, H.W. Optimization of fermentation conditions and medium compositions for the production of chrysomycin a by a marine-derived strain Streptomyces sp. 891. Prep. Biochem. Biotechnol. 2021, 51, 998–1003. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Zhao, J.; Chen, X.; Mao, Z.; Miao, W. Insights into the simultaneous utilization of glucose and glycerol by Streptomyces albulus M-Z18 for high ε-poly-l-lysine productivity. Bioprocess Biosyst. Eng. 2017, 40, 1775–1785. [Google Scholar] [CrossRef] [PubMed]
- Grahovac, J.; Grahovac, M.; Dodić, J.; Bajić, B.; Balaž, J. Optimization of cultivation medium for enhanced production of antifungal metabolites by Streptomyces hygroscopicus. Crop Prot. 2014, 65, 143–152. [Google Scholar] [CrossRef]
- Singh, N.; Rai, V. Optimization of cultural parameters for antifungal and antibacterial metabolite from microbial isolate; Streptomyces rimosus MTCC 10792 from soil of Chhattisgarh. Int. J. Pharmcy Pharm. Sci. 2012, 4, 94–101. [Google Scholar]
- Roberts, M.F. Osmoadaptation and osmoregulation in archaea: Update 2004. Front. Biosci. 2004, 9, 1999–2019. [Google Scholar] [CrossRef] [PubMed]
- Bhowmick, S.; Shenouda, M.L.; Tschowri, N. Osmotic stress responses and the biology of the second messenger c-di-AMP in Streptomyces. microLife 2023, 4, uqad020. [Google Scholar] [CrossRef]
- Sanjivkumar, M.; Vijayalakshmi, K.; Silambarasan, T.; Sholkamy, E.N.; Immanuel, G. Biosynthesis, statistical optimization and molecular modeling of chitinase from crab shell wastes by a mangrove associated actinobacterium Streptomyces olivaceus (MSU3) using Box-Behnken design and its antifungal effects. Bioresour. Technol. Rep. 2020, 11, 100493. [Google Scholar] [CrossRef]
- Selvaraj, J.N.; Ganapathi, U.; Vincent, S.G.P.; Ramamoorthy, S.; Thavasimuthu, C. Statistical optimization of media components for antibiotic production in Streptomyces sp. CMSTAAHAL-3. Electron. J. Biotechnol. 2023, 65, 1–13. [Google Scholar] [CrossRef]
- Manivasagan, P.; Venkatesan, J.; Kang, K.-H.; Sivakumar, K.; Park, S.-J.; Kim, S.-K. Production of α-amylase for the biosynthesis of gold nanoparticles using Streptomyces sp. MBRC-82. Int. J. Biol. Macromol. 2015, 72, 71–78. [Google Scholar] [CrossRef]
- Venkatachalam, M.; Shum-Chéong-Sing, A.; Caro, Y.; Dufossé, L.; Fouillaud, M. OVAT Analysis and Response Surface Methodology Based on Nutrient Sources for Optimization of Pigment Production in the Marine-Derived Fungus Talaromyces albobiverticillius 30548 Submerged Fermentation. Mar. Drugs 2021, 19, 248. [Google Scholar] [CrossRef]
- Manhas, R.S.; Kumar, A.; Chaubey, A. A biostatistical approach for augmenting rare bianthraquinone antibiotic production by Streptomyces sp. RA-WS2 using Taguchi design. AMB Express 2022, 12, 155. [Google Scholar] [CrossRef]
- Li, L.; Zhang, L.; Li, K.; Wang, Y.; Gao, C.; Han, B.; Ma, C.; Xu, P. A newly isolated Bacillus licheniformis strain thermophilically produces 2,3-butanediol, a platform and fuel bio-chemical. Biotechnol. Biofuels 2013, 6, 123. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Li, J.; Chen, X.; Wu, J.; Wang, P.; Ye, J.; Yao, J. Enzymatical hydrolysis of food waste and ethanol production from the hydrolysate. Renew. Energy 2011, 36, 1259–1265. [Google Scholar] [CrossRef]
- Du, L.; Wang, C.; Li, J.; Xiao, D.; Li, C.; Xu, Y. Optimization of Headspace Solid-Phase Microextraction Coupled with Gas Chromatography-Mass Spectrometry for Detecting Methoxyphenolic Compounds in Pu-erh Tea. J. Agric. Food Chem. 2013, 61, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Liu, Y.; Zhang, S.; Luo, L.; Zeng, L. Effect of brewing conditions on phytochemicals and sensory profiles of black tea infusions: A primary study on the effects of geraniol and beta-ionone on taste perception of black tea infusions. Food Chem. 2021, 354, 129504. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Wang, Y.; Li, S.; Zhu, G. Air-assisted liquid-liquid microextraction based on the solidification of floating deep eutectic solvents for the simultaneous determination of bisphenols and polycyclic aromatic hydrocarbons in tea infusions via HPLC. Food Chem. 2021, 348, 129106. [Google Scholar] [CrossRef] [PubMed]
- Karimifard, S.; Moghaddam, M.R.A. Application of response surface methodology in physicochemical removal of dyes from wastewater: A critical review. Sci. Total Environ. 2018, 640, 772–797. [Google Scholar] [CrossRef]
- Sun, Y.C.; Peng, Y.X.; Chen, Y.X.; Shukla, A.J. Application of artificial neural networks in the design of controlled release drug delivery systems. Adv. Drug Deliv. Rev. 2003, 55, 1201–1215. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.-l.; Zhang, Y.-b.; Lu, J.-h.; Jiang, L.-y.; Teng, L.-R.; Wang, Y.; Liang, Y.-c. Optimization of fermentation medium for nisin production from Lactococcus lactis subsp lactis using response surface methodology (RSM) combined with artificial neural network-genetic algorithm (ANN-GA). Afr. J. Biotechnol. 2010, 9, 6264–6272. [Google Scholar]
- Yang, Y.; Gao, M.; Yu, X.; Zhang, Y.; Lyu, S. Optimization of medium composition for two-step fermentation of vitamin C based on artificial neural network-genetic algorithm techniques. Biotechnol. Biotechnol. Equip. 2015, 29, 1128–1134. [Google Scholar] [CrossRef]
- Chen, H.; Wang, B.; Li, J.; Xu, J.; Zeng, J.; Gao, W.; Chen, K. Comparative study on the extraction efficiency, characterization, and bioactivities of Bletilla striata polysaccharides using response surface methodology (RSM) and genetic algorithm-artificial neural network (GA-ANN). Int. J. Biol. Macromol. 2023, 226, 982–995. [Google Scholar] [CrossRef]
- Witek-Krowiak, A.; Chojnacka, K.; Podstawczyk, D.; Dawiec, A.; Pokomeda, K. Application of response surface methodology and artificial neural network methods in modelling and optimization of biosorption process. Bioresour. Technol. 2014, 160, 150–160. [Google Scholar] [CrossRef]
- Zhang, C.; Guo, Y.; Li, M. Review of Development and Application of Artificial Neural Network Models. Comput. Eng. Appl. 2021, 57, 57–69. [Google Scholar] [CrossRef]
- Wu, W.; Dandy, G.C.; Maier, H.R. Protocol for developing ANN models and its application to the assessment of the quality of the ANN model development process in drinking water quality modelling. Environ. Model. Softw. 2014, 54, 108–127. [Google Scholar] [CrossRef]
- Mohanraj, M.; Jayaraj, S.; Muraleedharan, C. Applications of artificial neural networks for thermal analysis of heat exchangers—A review. Int. J. Therm. Sci. 2015, 90, 150–172. [Google Scholar] [CrossRef]
- Afram, A.; Janabi-Sharifi, F.; Fung, A.S.; Raahemifar, K. Artificial neural network (ANN) based model predictive control (MPC) and optimization of HVAC systems: A state of the art review and case study of a residential HVAC system. Energy Build. 2017, 141, 96–113. [Google Scholar] [CrossRef]
- Abiodun, O.I.; Jantan, A.; Omolara, A.E.; Dada, K.V.; Umar, A.M.; Linus, O.U.; Arshad, H.; Kazaure, A.A.; Gana, U.; Kiru, M.U. Comprehensive Review of Artificial Neural Network Applications to Pattern Recognition. IEEE Access 2019, 7, 158820–158846. [Google Scholar] [CrossRef]
- Li, B.; Delpha, C.; Diallo, D.; Migan-Dubois, A. Application of Artificial Neural Networks to photovoltaic fault detection and diagnosis: A review. Renew. Sustain. Energy Rev. 2021, 138, 110512. [Google Scholar] [CrossRef]
- Buasri, A.; Sirikoom, P.; Pattane, S.; Buachum, O.; Loryuenyong, V. Process Optimization of Biodiesel from Used Cooking Oil in a Microwave Reactor: A Case of Machine Learning and Box–Behnken Design. Chem. Eng. 2023, 7, 65. [Google Scholar]
- Singh, V.; Haque, S.; Niwas, R.; Srivastava, A.; Pasupuleti, M.; Tripathi, C.K.M. Strategies for Fermentation Medium Optimization: An In-Depth Review. Front. Microbiol. 2017, 7, 2087. [Google Scholar] [CrossRef] [PubMed]
- Vinayagam, R.; Dave, N.; Varadavenkatesan, T.; Rajamohan, N.; Sillanpaa, M.; Nadda, A.K.; Govarthanan, M.; Selvaraj, R. Artificial neural network and statistical modelling of biosorptive removal of hexavalent chromium using macroalgal spent biomass. Chemosphere 2022, 296, 133965. [Google Scholar] [CrossRef] [PubMed]
- Hadjisolomou, E.; Stefanidis, K.; Herodotou, H.; Michaelides, M.; Papatheodorou, G.; Papastergiadou, E. Modelling Freshwater Eutrophication with Limited Limnological Data Using Artificial Neural Networks. Water 2021, 13, 1590. [Google Scholar] [CrossRef]
- Vu, H.L.; Ng, K.T.W.; Richer, A.; An, C. Analysis of input set characteristics and variances on k-fold cross validation for a Recurrent Neural Network model on waste disposal rate estimation. J. Environ. Manag. 2022, 311, 114869. [Google Scholar] [CrossRef] [PubMed]
- Das, L.; Maity, U.; Basu, J.K. The photocatalytic degradation of carbamazepine and prediction by artificial neural networks. Process Saf. Environ. Prot. 2014, 92, 888–895. [Google Scholar] [CrossRef]
- Elmolla, E.S.; Chaudhuri, M.; Eltoukhy, M.M. The use of artificial neural network (ANN) for modeling of COD removal from antibiotic aqueous solution by the Fenton process. J. Hazard. Mater. 2010, 179, 127–134. [Google Scholar] [CrossRef]
- Abdolrasol, M.G.M.; Hussain, S.M.S.; Ustun, T.S.; Sarker, M.R.; Hannan, M.A.; Mohamed, R.; Ali, J.A.; Mekhilef, S.; Milad, A. Artificial Neural Networks Based Optimization Techniques: A Review. Electronics 2021, 10, 2689. [Google Scholar] [CrossRef]
- Ma, J.Y.; Huang, H.B.; Xie, Y.C.; Liu, Z.Y.; Zhao, J.; Zhang, C.Y.; Jia, Y.X.; Zhang, Y.; Zhang, H.; Zhang, T.Y.; et al. Biosynthesis of ilamycins featuring unusual building blocks and engineered production of enhanced anti-tuberculosis agents. Nat. Commun. 2017, 8, 10. [Google Scholar] [CrossRef]

| Run | X1 | X2 | X3 | X4 | X5 | X6 | Y |
|---|---|---|---|---|---|---|---|
| Cassava Starch | Glycerol | Soybean Meal | Ammonium Citrate | FeSO4·7H2O | Seawater Salt | 4′-N-Demethyl-Vicenistatin | |
| (g/L) | (g/L) | (g/L) | (g/L) | (mg/L) | (g/L) | (g/L) | |
| 1 | 6(−1) | 10(−1) | 2.5(−1) | 7.5(+1) | 60(+1) | 36(+1) | 0.1070 ± 0.0039 |
| 2 | 10(+1) | 20(+1) | 7.5(+1) | 4.5(−1) | 60(+1) | 36(+1) | 0.1061 ± 0.0684 |
| 3 | 10(+1) | 10(−1) | 7.5(+1) | 4.5(−1) | 30(−1) | 24(−1) | 0.1099 ± 0.0214 |
| 4 | 10(+1) | 10(−1) | 7.5(+1) | 7.5(+1) | 30(−1) | 36(+1) | 0.1042 ± 0.0203 |
| 5 | 6(−1) | 20(+1) | 7.5(+1) | 4.5(−1) | 60(+1) | 24(−1) | 0.1173 ± 0.0575 |
| 6 | 10(+1) | 20(+1) | 2.5(−1) | 7.5(+1) | 30(−1) | 24(−1) | 0.1609 ± 0.0195 |
| 7 | 8(0) | 15(0) | 5(0) | 6(0) | 40(0) | 30(0) | 0.0832 ± 0.0211 |
| 8 | 6(−1) | 10(−1) | 2.5(−1) | 4.5(−1) | 30(−1) | 24(−1) | 0.0606 ± 0.0066 |
| 9 | 10(+1) | 10(−1) | 2.5(−1) | 4.5(−1) | 60(+1) | 36(+1) | 0.0564 ± 0.0075 |
| 10 | 8(0) | 15(0) | 5(0) | 6(0) | 40(0) | 30(0) | 0.0922 ± 0.0162 |
| 11 | 6(−1) | 20(+1) | 2.5(−1) | 4.5(−1) | 30(−1) | 36(+1) | 0.0571 ± 0.0060 |
| 12 | 6(−1) | 10(−1) | 7.5(+1) | 7.5(+1) | 60(+1) | 24(−1) | 0.0241 ± 0.0008 |
| 13 | 6(−1) | 20(+1) | 7.5(+1) | 7.5(+1) | 30(−1) | 36(+1) | 0.0908 ± 0.0251 |
| 14 | 8(0) | 15(0) | 5(0) | 6(0) | 40(0) | 30(0) | 0.0960 ± 0.0213 |
| 15 | 10(+1) | 20(+1) | 2.5(−1) | 7.5(+1) | 60(+1) | 24(−1) | 0.1667 ± 0.0139 |
| Model | |||||||
| Adj SS | 0.008588 | 0.008028 | 0.000026 | 0.000436 | 0.000529 | 0.003064 | 0.020671 |
| Adj MS | 0.008588 | 0.008028 | 0.000026 | 0.000436 | 0.000529 | 0.003064 | 0.002953 |
| F-Value | 23.49 | 21.96 | 0.07 | 1.19 | 1.45 | 8.38 | 8.08 |
| p-Value | 0.002 | 0.002 | 0.799 | 0.311 | 0.268 | 0.023 | 0.007 |
| Run | Cassava Starch | Glycerol | Seawater Salt | 4′-N-Demethyl-Vicenistatin |
|---|---|---|---|---|
| (g/L) | (g/L) | (g/L) | (g/L) | |
| Step size | +3 | +3 | −2.5 | |
| 1 | 8 | 15 | 30 | 0.1070 ± 0.0037 |
| 2 | 11 | 18 | 27.5 | 0.1399 ± 0.0036 |
| 3 | 14 | 21 | 25 | 0.0159 ± 0.0007 |
| 4 | 17 | 24 | 22.5 | 0.0022 ± 0.0003 |
| 5 | 20 | 27 | 20 | 0.0013 ± 0.0007 |
| Run | X1 | X2 | X3 | Y |
|---|---|---|---|---|
| Cassava Starch | Glycerol | Seawater Salt | 4′-N-Demethyl-Vicenistatin | |
| (g/L) | (g/L) | (g/L) | (g/L) | |
| 1 | 11(0) | 9(−1.5) | 28(0) | 0.0157 ± 0.0049 |
| 2 | 5(−1) | 24(+1) | 34(+1) | 0.0896 ± 0.0414 |
| 3 | 5(−1) | 12(−1) | 22(−1) | 0.0266 ± 0.0098 |
| 4 | 5(−1) | 12(−1) | 34(+1) | 0.0539 ± 0.0100 |
| 5 | 5(−1) | 24(+1) | 22(−1) | 0.1566 ± 0.0275 |
| 6 | 17(+1) | 12(−1) | 22(−1) | 0.0330 ± 0.0075 |
| 7 | 11(0) | 18(0) | 28(0) | 0.1323 ± 0.0274 |
| 8 | 11(0) | 18(0) | 28(0) | 0.1424 ± 0.0380 |
| 9 | 11(0) | 18(0) | 19(−1.5) | 0.1476 ± 0.1049 |
| 10 | 11(0) | 18(0) | 28(0) | 0.1443 ± 0.0048 |
| 11 | 11(0) | 18(0) | 37(+1.5) | 0.1719 ± 0.0388 |
| 12 | 2(−1.5) | 18(0) | 28(0) | 0.0982 ± 0.0011 |
| 13 | 17(+1) | 24(+1) | 34(+1) | 0.0392 ± 0.0028 |
| 14 | 11(0) | 18(0) | 28(0) | 0.1453 ± 0.0061 |
| 15 | 17(+1) | 12(−1) | 34(+1) | 0.1263 ± 0.0066 |
| 16 | 17(+1) | 24(+1) | 22(−1) | 0.0505 ± 0.0596 |
| 17 | 11(0) | 27(+1.5) | 28(0) | 0.0191 ± 0.0008 |
| 18 | 20(+1.5) | 18(0) | 28(0) | 0.0877 ± 0.0090 |
| 19 | 11(0) | 18(0) | 28(0) | 0.1372 ± 0.0037 |
| 20 | 11(0) | 18(0) | 28(0) | 0.1370 ± 0.0179 |
| Source | DF | Adj SS | Adj MS | F-Value | p-Value |
|---|---|---|---|---|---|
| Model | 9 | 0.052301 | 0.005811 | 93.65 | 0.183 × 10−7 |
| X1 | 1 | 0.000699 | 0.000699 | 11.26 | 0.007 |
| X2 | 1 | 0.000822 | 0.000822 | 13.25 | 0.005 |
| X3 | 1 | 0.000496 | 0.000496 | 7.99 | 0.018 |
| X12 | 1 | 0.004644 | 0.004644 | 74.84 | 0.590 × 10−5 |
| X22 | 1 | 0.030963 | 0.030963 | 498.96 | 0.727 × 10−9 |
| X32 | 1 | 0.000741 | 0.000741 | 11.94 | 0.006 |
| X1 × X2 | 1 | 0.006916 | 0.006916 | 111.45 | 0.965 × 10−6 |
| X1 × X3 | 1 | 0.001852 | 0.001852 | 29.84 | 0.276 × 10−3 |
| X2 × X3 | 1 | 0.004950 | 0.004950 | 79.76 | 0.443 × 10−5 |
| Error | 10 | 0.000621 | 0.000062 | ||
| Lack-of-Fit | 5 | 0.000493 | 0.000099 | 3.85 | 0.083 |
| Pure Error | 5 | 0.000128 | 0.000026 | ||
| Total | 19 | 0.052922 | |||
| R2 = 0.9883 | Adj R2 = 0.9777 | RMSE = 0.0056 | |||
| Parameters | MSE | |
|---|---|---|
| Number of neurons | 3 | 0.3331 |
| 4 | 0.3181 | |
| 5 | 0.3138 | |
| 6 | 0.2588 | |
| 7 | 0.2733 | |
| 8 | 0.2833 | |
| 9 | 0.2789 | |
| 10 | 0.2812 | |
| 11 | 0.3034 | |
| 12 | 0.2868 | |
| Transfer functions | logsig + purelin | 0.2177 |
| tansig + purelin | 0.2539 | |
| logsig + tansig | 0.2202 | |
| tansig + logsig | 0.3618 | |
| tansig + tansig | 0.2703 | |
| logsig + logsig | 0.3234 | |
| Backpropagation training algorithm | trainbr | 0.4448 |
| traincgb | 0.3799 | |
| traincgf | 0.3392 | |
| traincgp | 0.3746 | |
| traingd | 0.4474 | |
| Backpropagation training algorithm | traingda | 0.4341 |
| traingdm | 0.5184 | |
| traingdx | 0.5052 | |
| trainlm | 0.2453 | |
| trainrp | 0.3207 | |
| trainscg | 0.3811 | |
| Model | Process Parameter | Statistical Values | 4′-N-Demethyl-Vicenistatin (g/L) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Cassava Starch | Glycerol | Seawater Salt | Error | R2 | MSE | RSME | APD | Experimental Data | Predicted Data | |
| (g/L) | (g/L) | (g/L) | ||||||||
| AM3 | 15 | 15 | 30 | - | - | - | - | - | 0.0502 ± 0.0041 | - |
| RSM | 4 | 22 | 19 | 11.4% | 0.9883 | 3.11 × 10−5 | 0.0056 | 10.58 | 0.1637 ± 0.0036 | 0.1848 |
| ANN-GA | 12 | 17 | 34 | 1.9% | 0.9962 | 2.62 × 10−5 | 0.0051 | 5.88 | 0.1921 ± 0.0052 | 0.1885 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, Z.; Fu, H.; Wang, J. Modeling and Optimization of the Culture Medium for Efficient 4′-N-Demethyl-Vicenistatin Production by Streptomyces parvus Using Response Surface Methodology and Artificial-Neural-Network-Genetic-Algorithm. Fermentation 2024, 10, 154. https://doi.org/10.3390/fermentation10030154
Yu Z, Fu H, Wang J. Modeling and Optimization of the Culture Medium for Efficient 4′-N-Demethyl-Vicenistatin Production by Streptomyces parvus Using Response Surface Methodology and Artificial-Neural-Network-Genetic-Algorithm. Fermentation. 2024; 10(3):154. https://doi.org/10.3390/fermentation10030154
Chicago/Turabian StyleYu, Zhixin, Hongxin Fu, and Jufang Wang. 2024. "Modeling and Optimization of the Culture Medium for Efficient 4′-N-Demethyl-Vicenistatin Production by Streptomyces parvus Using Response Surface Methodology and Artificial-Neural-Network-Genetic-Algorithm" Fermentation 10, no. 3: 154. https://doi.org/10.3390/fermentation10030154
APA StyleYu, Z., Fu, H., & Wang, J. (2024). Modeling and Optimization of the Culture Medium for Efficient 4′-N-Demethyl-Vicenistatin Production by Streptomyces parvus Using Response Surface Methodology and Artificial-Neural-Network-Genetic-Algorithm. Fermentation, 10(3), 154. https://doi.org/10.3390/fermentation10030154





