A microRNA Arising from the Negative Strand of SARS-CoV-2 Genome Targets FOS to Reduce AP-1 Activity
Abstract
1. Introduction
2. Results
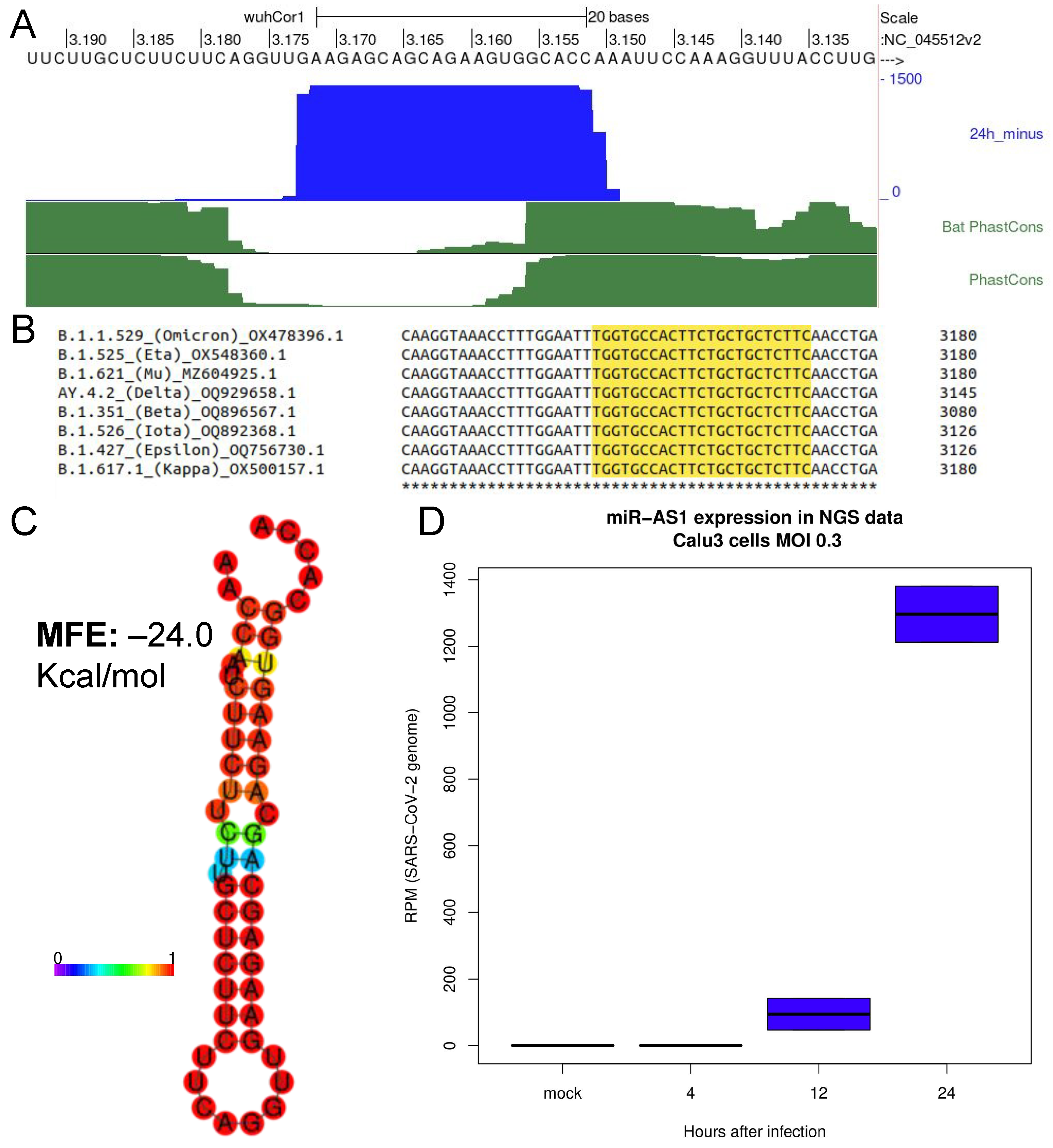
2.1. Identification of a Novel microRNA Encoded by SARS-CoV-2 Negative Strand Genomic RNA
2.2. Promoting miRNA Processing Increases the Expression of SARS-CoV-2 miR-AS1
2.3. A Set of Genes Translationally Repressed during SARS-CoV-2 Infection Is Significantly Enriched for miR-AS1 Seed Matches
2.4. miR-AS1 Is a Negative Modulator of AP-1 Transcription Factor via Targeting FOS
3. Methods
3.1. NGS Data Analysis
3.2. Cell Lines and Reagents
3.3. qRT-PCR
3.4. miR-AS1 Target Prediction
3.5. Statistical Analysis
3.6. Western Blot
3.7. Luciferase Assays
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pfeffer, S.; Zavolan, M.; Grässer, F.A.; Chien, M.; Russo, J.J.; Ju, J.; John, B.; Enright, A.J.; Marks, D.; Sander, C.; et al. Identification of Virus-Encoded MicroRNAs. Science 2004, 304, 734–736. [Google Scholar] [CrossRef] [PubMed]
- Kincaid, R.P.; Sullivan, C.S. Virus-Encoded MicroRNAs: An Overview and a Look to the Future. PLoS Pathog. 2012, 8, e1003018. [Google Scholar] [CrossRef] [PubMed]
- Grundhoff, A.; Sullivan, C.S.; Ganem, D. A Combined Computational and Microarray-Based Approach Identifies Novel MicroRNAs Encoded by Human Gamma-Herpesviruses. RNA 2006, 12, 733–750. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, C.S.; Grundhoff, A.T.; Tevethia, S.; Pipas, J.M.; Ganem, D. SV40-Encoded MicroRNAs Regulate Viral Gene Expression and Reduce Susceptibility to Cytotoxic T Cells. Nature 2005, 435, 682–686. [Google Scholar] [CrossRef]
- Grundhoff, A.; Sullivan, C.S. Virus-Encoded MicroRNAs. Virology 2011, 411, 325–343. [Google Scholar] [CrossRef]
- Nanbo, A.; Furuyama, W.; Lin, Z. RNA Virus-Encoded MiRNAs: Current Insights and Future Challenges. Front. Microbiol. 2021, 12, 679210. [Google Scholar] [CrossRef]
- Cullen, B.R. Five Questions about Viruses and MicroRNAs. PLoS Pathog. 2010, 6, e1000787. [Google Scholar] [CrossRef]
- Shapiro, J.S.; Schmid, S.; Aguado, L.C.; Sabin, L.R.; Yasunaga, A.; Shim, J.V.; Sachs, D.; Cherry, S.; tenOever, B.R. Drosha as an Interferon-Independent Antiviral Factor. Proc. Natl. Acad. Sci. USA 2014, 111, 7108–7113. [Google Scholar] [CrossRef]
- Shapiro, J.S.; Langlois, R.A.; Pham, A.M.; tenOever, B.R. Evidence for a Cytoplasmic Microprocessor of Pri-MiRNAs. RNA 2012, 18, 1338–1346. [Google Scholar] [CrossRef]
- Kozomara, A.; Griffiths-Jones, S. MiRBase: Integrating MicroRNA Annotation and Deep-Sequencing Data. Nucleic Acids Res. 2011, 39, D152–D157. [Google Scholar] [CrossRef]
- Long, S. SARS-CoV-2 Subgenomic RNAs: Characterization, Utility, and Perspectives. Viruses 2021, 13, 1923. [Google Scholar] [CrossRef] [PubMed]
- Morales, L.; Oliveros, J.C.; Fernandez-Delgado, R.; tenOever, B.R.; Enjuanes, L.; Sola, I. SARS-CoV-Encoded Small RNAs Contribute to Infection-Associated Lung Pathology. Cell Host Microbe. 2017, 21, 344–355. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Siu, G.K.-H.; Mok, B.W.-Y.; Sun, J.; Fung, K.S.C.; Lam, J.Y.-W.; Wong, N.K.; Gedefaw, L.; Luo, S.; Lee, T.M.H.; et al. Viral MicroRNAs Encoded by Nucleocapsid Gene of SARS-CoV-2 Are Detected during Infection, and Targeting Metabolic Pathways in Host Cells. Cells 2021, 10, 1762. [Google Scholar] [CrossRef]
- Singh, M.; Chazal, M.; Quarato, P.; Bourdon, L.; Malabat, C.; Vallet, T.; Vignuzzi, M.; van der Werf, S.; Behillil, S.; Donati, F.; et al. A Virus-Derived MicroRNA Targets Immune Response Genes during SARS-CoV-2 Infection. EMBO Rep. 2022, 23, e54341. [Google Scholar] [CrossRef]
- Pawlica, P.; Yario, T.A.; White, S.; Wang, J.; Moss, W.N.; Hui, P.; Vinetz, J.M.; Steitz, J.A. SARS-CoV-2 Expresses a MicroRNA-like Small RNA Able to Selectively Repress Host Genes. Proc. Natl. Acad. Sci. USA 2021, 118, e2116668118. [Google Scholar] [CrossRef]
- Wyler, E.; Mösbauer, K.; Franke, V.; Diag, A.; Gottula, L.T.; Arsiè, R.; Klironomos, F.; Koppstein, D.; Hönzke, K.; Ayoub, S.; et al. Transcriptomic Profiling of SARS-CoV-2 Infected Human Cell Lines Identifies HSP90 as Target for COVID-19 Therapy. iScience 2021, 24, 102151. [Google Scholar] [CrossRef]
- Lorenz, R.; Bernhart, S.H.; Höner zu Siederdissen, C.; Tafer, H.; Flamm, C.; Stadler, P.F.; Hofacker, I.L. ViennaRNA Package 2.0. Algorithms Mol. Biol. 2011, 6, 26. [Google Scholar] [CrossRef]
- Shan, G.; Li, Y.; Zhang, J.; Li, W.; Szulwach, K.E.; Duan, R.; Faghihi, M.A.; Khalil, A.M.; Lu, L.; Paroo, Z.; et al. A Small Molecule Enhances RNA Interference and Promotes MicroRNA Processing. Nat. Biotechnol. 2008, 26, 933–940. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.; Villanueva, A.; Moutinho, C.; Davalos, V.; Spizzo, R.; Ivan, C.; Rossi, S.; Setien, F.; Casanovas, O.; Simo-Riudalbas, L.; et al. Small Molecule Enoxacin Is a Cancer-Specific Growth Inhibitor That Acts by Enhancing TAR RNA-Binding Protein 2-Mediated MicroRNA Processing. Proc. Natl. Acad. Sci. USA 2011, 108, 4394–4399. [Google Scholar] [CrossRef]
- Gioia, U.; Francia, S.; Cabrini, M.; Brambillasca, S.; Michelini, F.; Jones-Weinert, C.W.; d’Adda di Fagagna, F. Pharmacological Boost of DNA Damage Response and Repair by Enhanced Biogenesis of DNA Damage Response RNAs. Sci. Rep. 2019, 9, 6460. [Google Scholar] [CrossRef]
- Scroggs, S.L.P.; Offerdahl, D.K.; Flather, D.P.; Morris, C.N.; Kendall, B.L.; Broeckel, R.M.; Beare, P.A.; Bloom, M.E. Fluoroquinolone Antibiotics Exhibit Low Antiviral Activity against SARS-CoV-2 and MERS-CoV. Viruses 2021, 13, 8. [Google Scholar] [CrossRef] [PubMed]
- Puray-Chavez, M.; Lee, N.; Tenneti, K.; Wang, Y.; Vuong, H.R.; Liu, Y.; Horani, A.; Huang, T.; Gunsten, S.P.; Case, J.B.; et al. The Translational Landscape of SARS-CoV-2-Infected Cells Reveals Suppression of Innate Immune Genes. mBio 2022, 13, e0081522. [Google Scholar] [CrossRef] [PubMed]
- Rehmsmeier, M.; Steffen, P.; Höchsmann, M.; Giegerich, R. Fast and Effective Prediction of MicroRNA/Target Duplexes. RNA 2004, 10, 1507–1517. [Google Scholar] [CrossRef] [PubMed]
- Varshney, B.; Lal, S.K. SARS-CoV Accessory Protein 3b Induces AP-1 Transcriptional Activity through Activation of JNK and ERK Pathways. Biochemistry 2011, 50, 5419–5425. [Google Scholar] [CrossRef]
- Wu, Y.; Shi, Z.; Chen, J.; Zhang, H.; Li, M.; Zhao, Y.; Shi, H.; Shi, D.; Guo, L.; Feng, L. Porcine Deltacoronavirus E Protein Induces Interleukin-8 Production via NF-ΚB and AP-1 Activation. Vet. Microbiol. 2022, 274, 109553. [Google Scholar] [CrossRef]
- Yuan, L.X.; Liang, J.Q.; Zhu, Q.C.; Dai, G.; Li, S.; Fung, T.S.; Liu, D.X. Gammacoronavirus Avian Infectious Bronchitis Virus and Alphacoronavirus Porcine Epidemic Diarrhea Virus Exploit a Cell-Survival Strategy via Upregulation of CFOS to Promote Viral Replication. J. Virol. 2021, 95, e02107-20. [Google Scholar] [CrossRef]
- Patra, T.; Meyer, K.; Geerling, L.; Isbell, T.S.; Hoft, D.F.; Brien, J.; Pinto, A.K.; Ray, R.B.; Ray, R. SARS-CoV-2 Spike Protein Promotes IL-6 Trans-Signaling by Activation of Angiotensin II Receptor Signaling in Epithelial Cells. PLoS Pathog. 2020, 16, e10091282020. [Google Scholar] [CrossRef]
- Vasanwala, F.H.; Kusam, S.; Toney, L.M.; Dent, A.L. Repression of AP-1 Function: A Mechanism for the Regulation of Blimp-1 Expression and B Lymphocyte Differentiation by the B Cell Lymphoma-6 Protooncogene. J. Immunol. 2002, 169, 1922–1929. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve Years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Sola, I.; Almazán, F.; Zúñiga, S.; Enjuanes, L. Continuous and Discontinuous RNA Synthesis in Coronaviruses. Annu. Rev. Virol. 2015, 2, 265–288. [Google Scholar] [CrossRef] [PubMed]
- Quinlan, A.R.; Hall, I.M. BEDTools: A Flexible Suite of Utilities for Comparing Genomic Features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.D.; Hinrichs, A.S.; Clawson, H.; Gonzalez, J.N.; Lee, B.T.; Nassar, L.R.; Raney, B.J.; Rosenbloom, K.R.; Nerli, S.; Rao, A.; et al. The UCSC SARS-CoV-2 Genome Browser. Nat. Genet. 2020, 52, 991–998. [Google Scholar] [CrossRef] [PubMed]
- Raymond, C.K.; Roberts, B.S.; Garrett-Engele, P.; Lim, L.P.; Johnson, J.M. Simple, quantitative primer-extension PCR assay for direct monitoring of microRNAs and short-interfering RNAs. RNA 2005, 11, 1737–1744. [Google Scholar] [CrossRef] [PubMed]
- Fulci, V.; Chiaretti, S.; Goldoni, M.; Azzalin, G.; Carucci, N.; Tavolaro, S.; Castellano, L.; Magrelli, A.; Citarella, F.; Messina, M.; et al. Quantitative Technologies Establish a Novel MicroRNA Profile of Chronic Lymphocytic Leukemia. Blood 2007, 109, 4944–4951. [Google Scholar] [CrossRef] [PubMed]
- Chiang, H.R.; Schoenfeld, L.W.; Ruby, J.G.; Auyeung, V.C.; Spies, N.; Baek, D.; Johnston, W.K.; Russ, C.; Luo, S.; Babiarz, J.E.; et al. Mammalian MicroRNAs: Experimental Evaluation of Novel and Previously Annotated Genes. Genes Dev. 2010, 24, 992–1009. [Google Scholar] [CrossRef] [PubMed]
- Fromm, B.; Billipp, T.; Peck, L.E.; Johansen, M.; Tarver, J.E.; King, B.L.; Newcomb, J.M.; Sempere, L.F.; Flatmark, K.; Hovig, E.; et al. A Uniform System for the Annotation of Vertebrate MicroRNA Genes and the Evolution of the Human MicroRNAome. Annu. Rev. Genet. 2015, 49, 213–242. [Google Scholar] [CrossRef]
- R: The R Project for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 28 March 2023).
- Wu, B.; Ramaiah, A.; Garcia, G.; Hasiakos, S.; Arumugaswami, V.; Srikanth, S. ORAI1 Limits SARS-CoV-2 Infection by Regulating Tonic Type I IFN Signaling. J. Immunol. 2022, 208, 74–84. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Greco, F.; Lorefice, E.; Carissimi, C.; Laudadio, I.; Ciccosanti, F.; Di Rienzo, M.; Colavita, F.; Meschi, S.; Maggi, F.; Fimia, G.M.; et al. A microRNA Arising from the Negative Strand of SARS-CoV-2 Genome Targets FOS to Reduce AP-1 Activity. Non-Coding RNA 2023, 9, 33. https://doi.org/10.3390/ncrna9030033
Greco F, Lorefice E, Carissimi C, Laudadio I, Ciccosanti F, Di Rienzo M, Colavita F, Meschi S, Maggi F, Fimia GM, et al. A microRNA Arising from the Negative Strand of SARS-CoV-2 Genome Targets FOS to Reduce AP-1 Activity. Non-Coding RNA. 2023; 9(3):33. https://doi.org/10.3390/ncrna9030033
Chicago/Turabian StyleGreco, Francesco, Elisa Lorefice, Claudia Carissimi, Ilaria Laudadio, Fabiola Ciccosanti, Martina Di Rienzo, Francesca Colavita, Silvia Meschi, Fabrizio Maggi, Gian Maria Fimia, and et al. 2023. "A microRNA Arising from the Negative Strand of SARS-CoV-2 Genome Targets FOS to Reduce AP-1 Activity" Non-Coding RNA 9, no. 3: 33. https://doi.org/10.3390/ncrna9030033
APA StyleGreco, F., Lorefice, E., Carissimi, C., Laudadio, I., Ciccosanti, F., Di Rienzo, M., Colavita, F., Meschi, S., Maggi, F., Fimia, G. M., & Fulci, V. (2023). A microRNA Arising from the Negative Strand of SARS-CoV-2 Genome Targets FOS to Reduce AP-1 Activity. Non-Coding RNA, 9(3), 33. https://doi.org/10.3390/ncrna9030033






