Novel Roles of Non-Coding RNAs in Opioid Signaling and Cardioprotection
Abstract
1. Introduction
2. The Opioid System
3. Mechanisms of Opioid Conditioning in the Heart
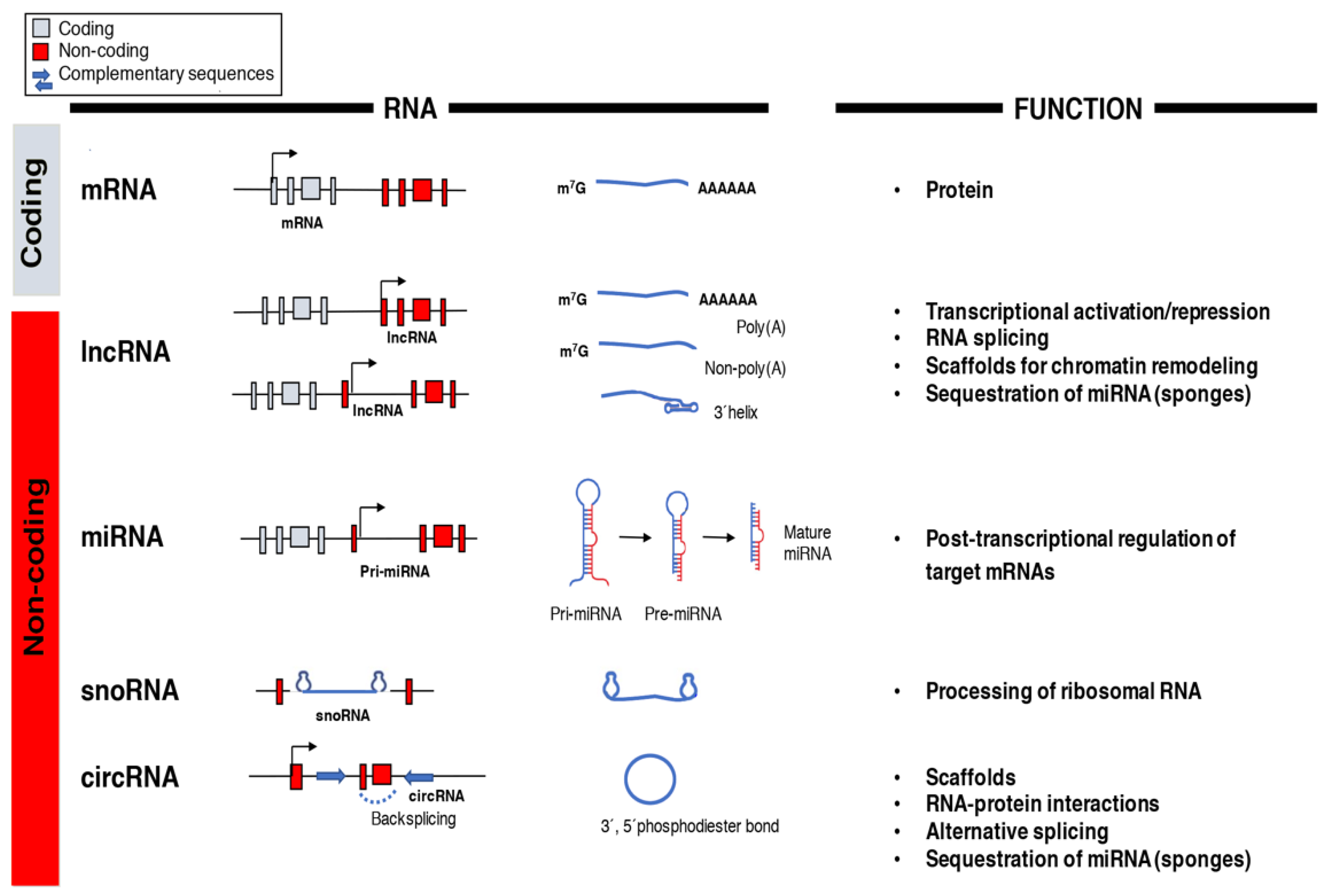
4. Non-Coding RNAs
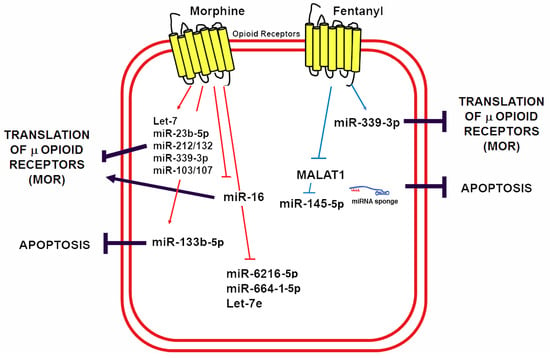
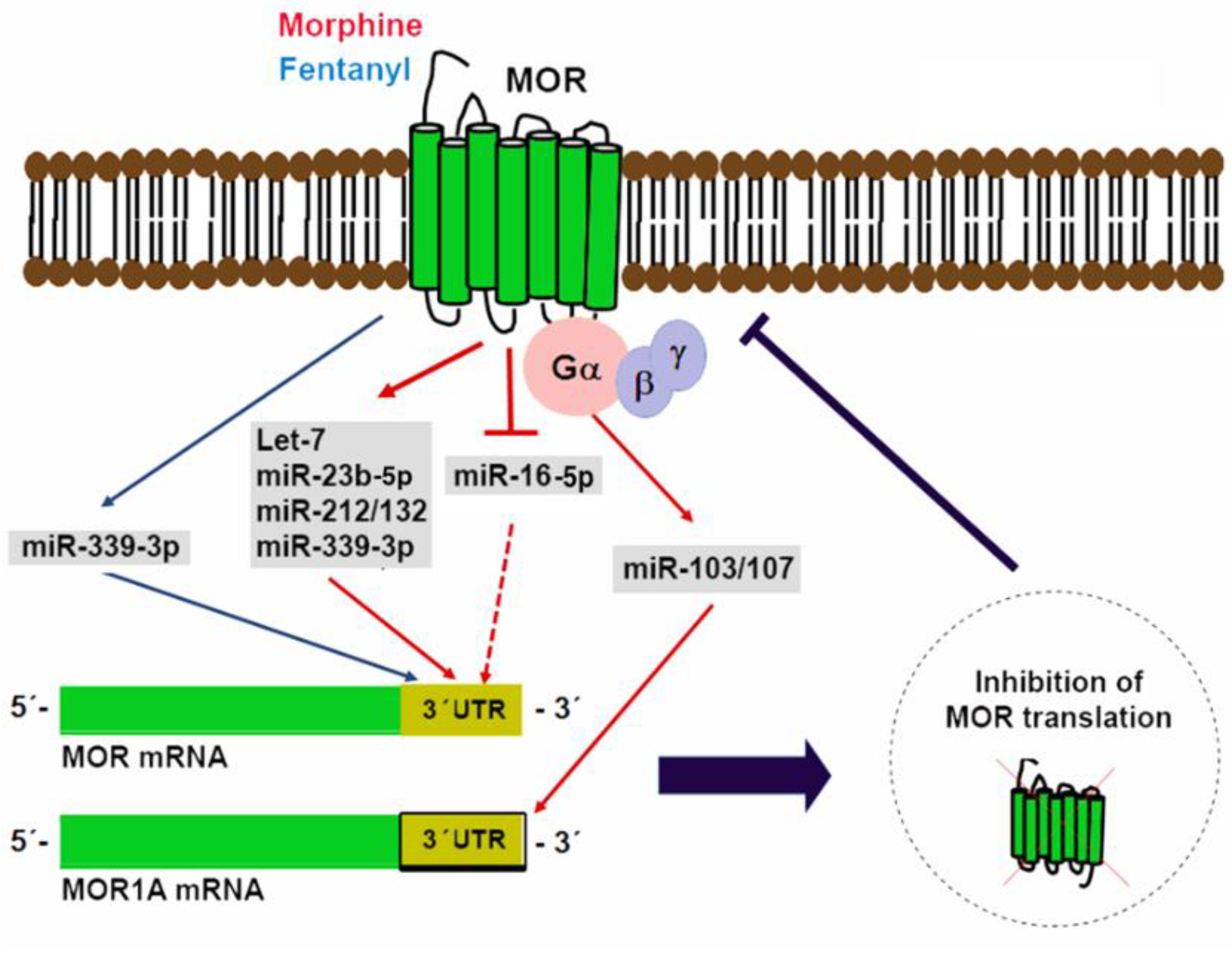
5. miRNAs in Control of Opioid Signaling
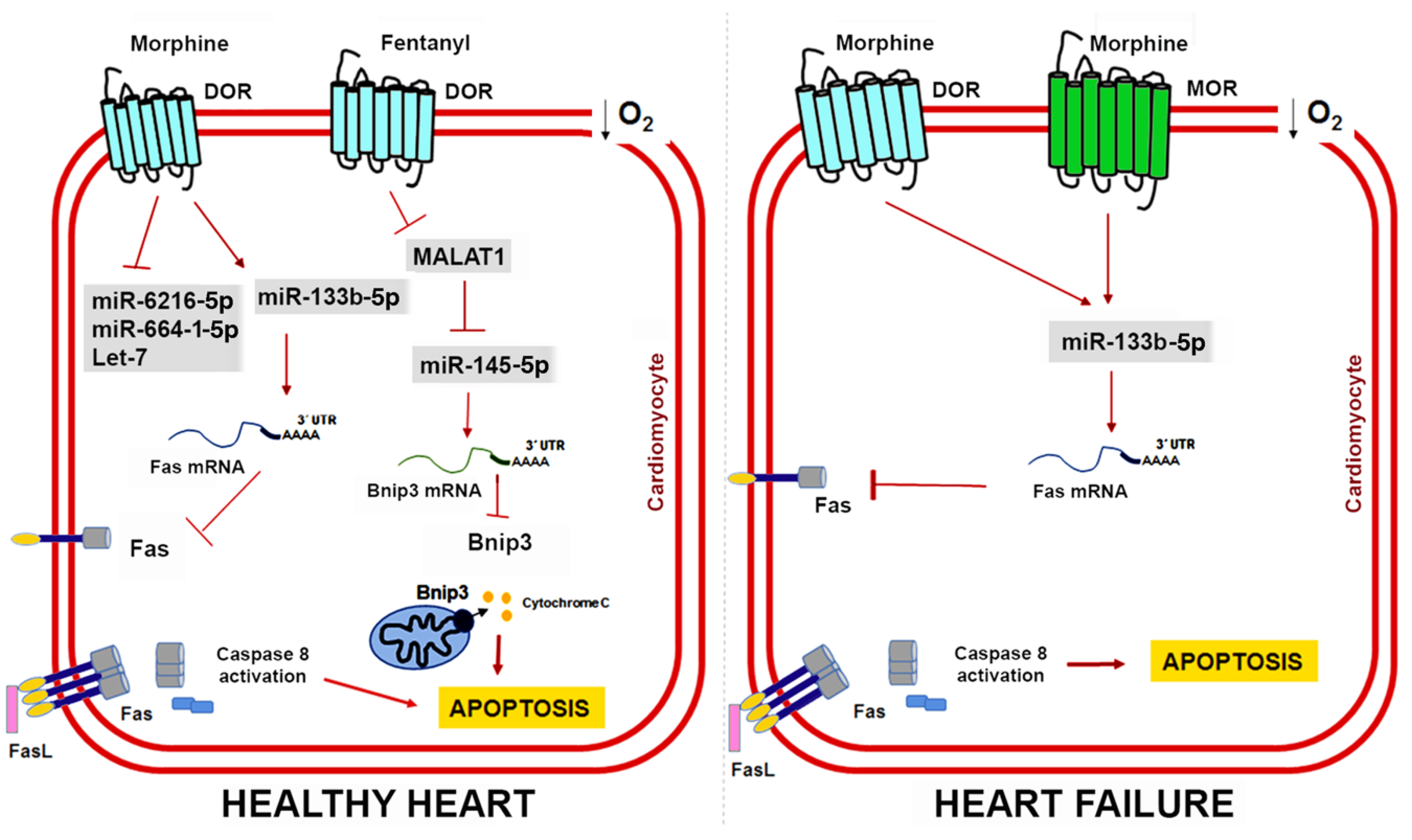
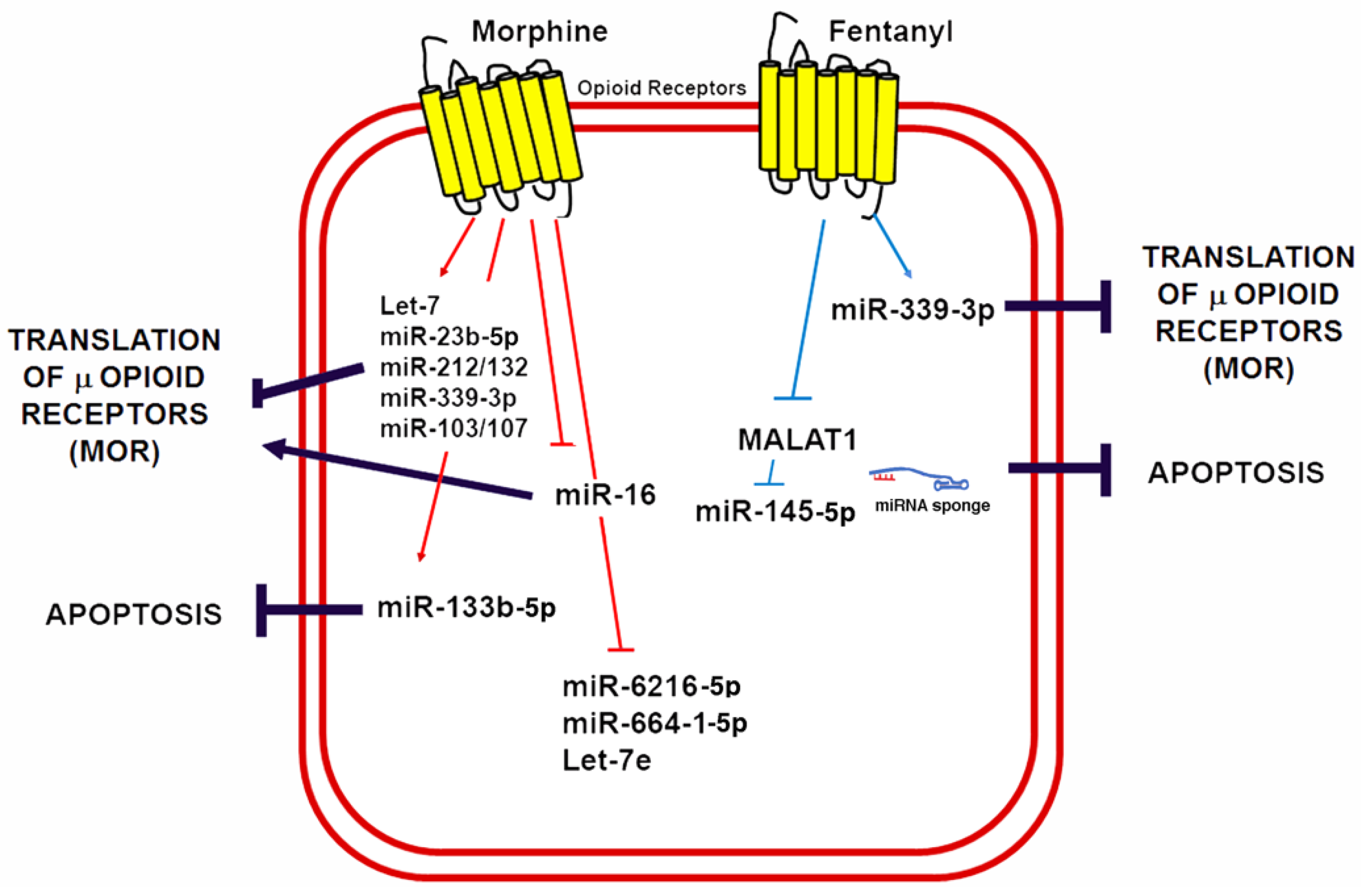
6. Non-Coding RNAs in Opioid-Induced Cardioprotection
7. Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Roth, G.A.; Johnson, C.; Abajobir, A.; Abd-Allah, F.; Abera, S.F.; Abyu, G.; Ahmed, M.; Aksut, B.; Alam, T.; Alam, K.; et al. Global, regional, and national burden of cardiovascular diseases for 10 causes, 1990 to 2015. J. Am. Coll. Cardiol. 2017, 70, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Libby, P.; Theroux, P. Pathophysiology of coronary artery disease. Circulation 2005, 111, 3481–3488. [Google Scholar] [CrossRef] [PubMed]
- Hausenloy, D.J.; Yellon, D.M. Myocardial ischemia-reperfusion injury: A neglected therapeutic target. J. Clin. Investig. 2013, 123, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Headrick, J.P.; See Hoe, L.E.; Du Toit, E.F.; Peart, J.N. Opioid receptors and cardioprotection-‘Opioidergic conditioning’ of the heart. Br. J. Pharmacol. 2015, 172, 2026–2050. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.E.; Gross, G.J. Opioids and cardioprotection. Pharmacol. Ther. 2001, 89, 123–137. [Google Scholar] [CrossRef]
- Schultz, J.E.; Hsu, A.K.; Gross, G.J. Ischemic preconditioning in the intact rat heart is mediated by delta1- but not mu- or kappa-opioid receptors. Circulation 1998, 97, 1282–1289. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.J.; Hsu, A.K.; Gross, G.J. Ischemic preconditioning and morphine-induced cardioprotection involve the delta (δ)-opioid receptor in the intact rat heart. J. Mol. Cell. Cardiol. 1997, 29, 2187–2195. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.E.; Hsu, A.K.; Gross, G.J. Morphine mimics the cardioprotective effect of ischemic preconditioning via a glibenclamide-sensitive mechanism in the rat heart. Circ. Res. 1996, 78, 1100–1104. [Google Scholar] [CrossRef] [PubMed]
- Uszczynska-Ratajczak, B.; Lagarde, J.; Frankish, A.; Guigó, R.; Johnson, R. Towards a complete map of the human long non-coding RNA transcriptome. Nat. Rev. Genet. 2018, 19, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Abascal, F.; Juan, D.; Jungreis, I.; Martinez, L.; Rigau, M.; Rodriguez, J.M.; Vazquez, J.; Tress, M.L. Loose ends: Almost one in five human genes still have unresolved coding status. Nucleic Acids Res. 2018, 46, 7070–7084. [Google Scholar] [CrossRef] [PubMed]
- Pennisi, E. Genomics. ENCODE project writes eulogy for junk DNA. Science 2012, 337, 1159–1161. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R.; Steitz, J.A. The noncoding RNA revolution-trashing old rules to forge new ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Frías-Lasserre, D.; Villagra, C.A. The importance of ncRNAs as epigenetic mechanisms in phenotypic variation and organic evolution. Front. Microbiol. 2017, 8, 2483. [Google Scholar] [CrossRef] [PubMed]
- Poller, W.; Dimmeler, S.; Heymans, S.; Zeller, T.; Haas, J.; Karakas, M.; Leistner, D.M.; Jakob, P.; Nakagawa, S.; Blankenberg, S.; et al. Non-coding RNAs in cardiovascular diseases: Diagnostic and therapeutic perspectives. Eur. Heart J. 2017. [Google Scholar] [CrossRef] [PubMed]
- Toyama, K.; Kiyosawa, N.; Watanabe, K.; Ishizuka, H. Identification of circulating miRNAs differentially regulated by opioid treatment. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Law, P.Y.; Loh, H.H. Non-coding RNAs regulating morphine function: With emphasis on the in vivo and in vitro functions of miR-190. Front. Genet. 2012, 3, 113. [Google Scholar] [CrossRef] [PubMed]
- Volkow, N.; Benveniste, H.; McLellan, A.T. Use and misuse of opioids in chronic pain. Annu. Rev. Med. 2018, 69, 451–465. [Google Scholar] [CrossRef] [PubMed]
- Al-Hasani, R.; Bruchas, M.R. Molecular mechanisms of opioid receptor-dependent signaling and behavior. Anesthesiology 2011, 115, 1363–1381. [Google Scholar] [CrossRef] [PubMed]
- Bodnar, R.J. Endogenous opiates and behavior: 2009. Peptides 2010, 31, 2325–2359. [Google Scholar] [CrossRef] [PubMed]
- Seyfried, O.; Hester, J. Opioids and endocrine dysfunction. Br. J. Pain 2012, 6, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Le Merrer, J.; Becker, J.A.; Befort, K.; Kieffer, B.L. Reward processing by the opioid system in the brain. Physiol. Rev. 2009, 89, 1379–1412. [Google Scholar] [CrossRef] [PubMed]
- Pathan, H.; Williams, J. Basic opioid pharmacology: An update. Br. J. Pain 2012, 6, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Pert, C.B.; Snyder, S.H. Properties of opiate-receptor binding in rat brain. Proc. Natl. Acad. Sci. USA 1973, 70, 2243–2247. [Google Scholar] [CrossRef] [PubMed]
- Simon, E.J.; Hiller, J.M.; Edelman, I. Stereospecific binding of the potent narcotic analgesic (3H) Etorphine to rat-brain homogenate. Proc. Natl. Acad. Sci. USA 1973, 70, 1947–1949. [Google Scholar] [CrossRef] [PubMed]
- Terenius, L. Stereospecific interaction between narcotic analgesics and a synaptic plasm a membrane fraction of rat cerebral cortex. Acta Pharmacol. Toxicol. 1973, 32, 317–320. [Google Scholar] [CrossRef]
- Evans, C.J.; Keith, D.E.; Morrison, H.; Magendzo, K.; Edwards, R.H. Cloning of a delta opioid receptor by functional expression. Science 1992, 258, 1952–1955. [Google Scholar] [CrossRef] [PubMed]
- Kieffer, B.L.; Befort, K.; Gaveriaux-Ruff, C.; Hirth, C.G. The delta-opioid receptor: Isolation of a cDNA by expression cloning and pharmacological characterization. Proc. Natl. Acad. Sci. USA 1992, 89, 12048–12052. [Google Scholar] [CrossRef] [PubMed]
- Minami, M.; Toya, T.; Katao, Y.; Maekawa, K.; Nakamura, S.; Onogi, T.; Kaneko, S.; Satoh, M. Cloning and expression of a cDNA for the rat kappa-opioid receptor. FEBS Lett. 1993, 329, 291–295. [Google Scholar] [CrossRef]
- Chen, Y.; Mestek, A.; Liu, J.; Hurley, J.A.; Yu, L. Molecular cloning and functional expression of a mu-opioid receptor from rat brain. Mol. Pharmacol. 1993, 44, 8–12. [Google Scholar] [CrossRef]
- Bagley, J.R.; Kudzma, L.V.; Lalinde, N.L.; Colapret, J.A.; Huang, B.S.; Lin, B.S.; Jerussi, T.P.; Benvenga, M.J.; Doorley, B.M.; Ossipov, M.H. Evolution of the 4-anilidopiperidine class of opioid analgesics. Med. Res. Rev. 1991, 11, 403–436. [Google Scholar] [CrossRef] [PubMed]
- Zylbergold, P.; Ramakrishnan, N.; Hebert, T. The role of G proteins in assembly and function of Kir3 inwardly rectifying potassium channels. Channels 2010, 4, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Rusin, K.I.; Giovannucci, D.R.; Stuenkel, E.L.; Moises, H.C. Kappa-opioid receptor activation modulates Ca2+ currents and secretion in isolated neuroendocrine nerve terminals. J. Neurosci. 1997, 17, 6565–6574. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Loh, H.H.; Law, P.Y. A novel noncanonical signaling pathway for the μ-opioid receptor. Mol. Pharmacol. 2013, 84, 844–853. [Google Scholar] [CrossRef] [PubMed]
- Lefkowitz, R.J.; Shenoy, S.K. Transduction of receptor signals by beta-arrestins. Science 2005, 308, 512–517. [Google Scholar] [CrossRef] [PubMed]
- Luttrell, L.M.; Daaka, Y.; Lefkowitz, R.J. Regulation of tyrosine kinase cascades by G-protein-coupled receptors. Curr. Opin. Cell Biol. 1999, 11, 177–183. [Google Scholar] [CrossRef]
- Urban, J.D.; Clarke, W.P.; von Zastrow, M.; Nichols, D.E.; Kobilka, B.; Weinstein, H.; Javitch, J.A.; Roth, B.L.; Christopoulos, A.; Sexton, P.M.; et al. Functional selectivity and classical concepts of quantitative pharmacology. J. Pharmacol. Exp. Ther. 2007, 320, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Zeng, Y.; Zhang, X.; Chu, J.; Loh, H.H.; Law, P.Y. mu-opioid receptor agonists differentially regulate the expression of miR-190 and NeuroD. Mol. Pharmacol. 2010, 77, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Loh, H.H.; Law, P.Y. Beta-arrestin-dependent mu-opioid receptor-activated extracellular signal-regulated kinases (ERKs) translocate to nucleus in contrast to G protein-dependent ERK activation. Mol. Pharmacol. 2008, 73, 178–190. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Chu, J.; Qiu, Y.; Loh, H.H.; Law, P.Y. Agonist-selective signaling is determined by the receptor location within the membrane domains. Proc. Natl. Acad. Sci. USA 2008, 105, 9421–9426. [Google Scholar] [CrossRef] [PubMed]
- Takenouchi, O.; Yoshimura, H.; Ozawa, T. Unique roles of β-arrestin in GPCR trafficking revealed by photoinducible dimerizers. Sci. Rep. 2018, 8, 677. [Google Scholar] [CrossRef] [PubMed]
- DeWire, S.M.; Ahn, S.; Lefkowitz, R.J.; Shenoy, S.K. Beta-arrestins and cell signaling. Annu. Rev. Physiol. 2007, 69, 483–510. [Google Scholar] [CrossRef] [PubMed]
- Violin, J.D.; Lefkowitz, R.J. Beta-arrestin-biased ligands at seven-transmembrane receptors. Trends Pharmacol. Sci. 2007, 28, 416–422. [Google Scholar] [CrossRef] [PubMed]
- Gesty-Palmer, D.; Chen, M.; Reiter, E.; Ahn, S.; Nelson, C.D.; Wang, S.; Eckhardt, A.E.; Cowan, C.L.; Spurney, R.F.; Luttrell, L.M.; et al. Distinct beta-arrestin- and G protein-dependent pathways for parathyroid hormone receptor-stimulated ERK1/2 activation. J. Biol. Chem. 2006, 281, 10856–10864. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Chu, J.; Zhang, Y.; Loh, H.H.; Law, P.Y. Modulating micro-opioid receptor phosphorylation switches agonist-dependent signaling as reflected in PKCepsilon activation and dendritic spine stability. J. Biol. Chem. 2011, 286, 12724–12733. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.G. AMP-activated protein kinase: The guardian of cardiac energy status. J. Clin. Investig. 2004, 114, 465–468. [Google Scholar] [CrossRef] [PubMed]
- Gross, G.J.; Auchampach, J.A. Role of ATP dependent potassium channels in myocardial ischaemia. Cardiovasc. Res. 1992, 26, 1011–1016. [Google Scholar] [CrossRef] [PubMed]
- Turer, A.T.; Hill, J.A. Pathogenesis of myocardial ischemia-reperfusion injury and rationale for therapy. Am. J. Cardiol. 2010, 106, 360–368. [Google Scholar] [CrossRef] [PubMed]
- Reimer, K.A.; Lowe, J.E.; Rasmussen, M.M.; Jennings, R.B. The wavefront phenomenon of ischemic cell death. 1. Myocardial infarct size vs duration of coronary occlusion in dogs. Circulation 1977, 56, 786–794. [Google Scholar] [CrossRef] [PubMed]
- Chiong, M.; Wang, Z.V.; Pedrozo, Z.; Cao, D.J.; Troncoso, R.; Ibacache, M.; Criollo, A.; Nemchenko, A.; Hill, J.A.; Lavandero, S. Cardiomyocyte death: Mechanisms and translational implications. Cell Death Dis. 2011, 2, e244. [Google Scholar] [CrossRef] [PubMed]
- Yellon, D.M.; Hausenloy, D.J. Myocardial reperfusion injury. N. Engl. J. Med. 2007, 357, 1121–1135. [Google Scholar] [CrossRef] [PubMed]
- Braunwald, E.; Kloner, R.A. Myocardial reperfusion: A double-edged sword? J. Clin. Investig. 1985, 76, 1713–1719. [Google Scholar] [CrossRef] [PubMed]
- Murry, C.E.; Jennings, R.B.; Reimer, K.A. Preconditioning with ischemia: A delay of lethal cell injury in ischemic myocardium. Circulation 1986, 74, 1124–1136. [Google Scholar] [CrossRef] [PubMed]
- Peart, J.N.; Patel, H.H.; Gross, G.J. Delta-opioid receptor activation mimics ischemic preconditioning in the canine heart. J. Cardiovasc. Pharmacol. 2003, 42, 78–81. [Google Scholar] [CrossRef] [PubMed]
- Mayfield, K.P.; D’Alecy, L.G. Role of endogenous opioid peptides in the acute adaptation to hypoxia. Brain Res. 1992, 582, 226–231. [Google Scholar] [CrossRef]
- Korthuis, R.J. Filling GAPs in the understanding of cardioprotection induced by GPCR activation: RGS proteins modulate ischaemic injury. Cardiovasc. Res. 2011, 91, 5–6. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.R.; Hsu, A.K.; Gross, G.J. Acute methadone treatment reduces myocardial infarct size via the delta-opioid receptor in rats during reperfusion. Anesth. Analg. 2009, 109, 1395–1402. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Sun, Y.; Li, J.; Xing, W.; Zhang, S.; Gu, X.; Feng, N.; Zhao, L.; Fan, R.; Wang, Y.; et al. Quaternary ammonium salt of U50488H, a new κ-opioid receptor agonist, protects rat heart against ischemia/reperfusion injury. Eur. J. Pharmacol. 2014, 737, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Schultz, J.E.; Rose, E.; Yao, Z.; Gross, G.J. Evidence for involvement of opioid receptors in ischemic preconditioning in rat hearts. Am. J. Physiol. 1995, 268, H2157–H2161. [Google Scholar] [CrossRef] [PubMed]
- Miki, T.; Cohen, M.V.; Downey, J.M. Opioid receptor contributes to ischemic preconditioning through protein kinase C activation in rabbits. Mol. Cell. Biochem. 1998, 186, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Fryer, R.M.; Wang, Y.; Hsu, A.K.; Gross, G.J. Essential activation of PKC-delta in opioid-initiated cardioprotection. Am. J. Physiol. Heart Circ. Physiol. 2001, 280, H1346–H1353. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.; Liu, F.; Ji, F.; Liang, J.; Liu, L.; Wu, Q.; Wang, T. Effect of c-Jun N-terminal kinase (JNK)/p38 mitogen-activated protein kinase (p38 MAPK) in morphine-induced tau protein hyperphosphorylation. Behav. Brain Res. 2013, 237, 249–255. [Google Scholar] [CrossRef] [PubMed]
- Fryer, R.M.; Hsu, A.K.; Gross, G.J. ERK and p38 MAP kinase activation are components of opioid-induced delayed cardioprotection. Basic. Res. Cardiol. 2001, 96, 136–142. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.R.; Hsu, A.K.; Gross, G.J. The JAK/STAT pathway is essential for opioid-induced cardioprotection: JAK2 as a mediator of STAT3, Akt, and GSK-3 beta. Am. J. Physiol. Heart Circ. Physiol. 2006, 291, H827–H834. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.R.; Hsu, A.K.; Gross, G.J. Opioid-induced cardioprotection occurs via glycogen synthase kinase beta inhibition during reperfusion in intact rat hearts. Circ. Res. 2004, 94, 960–966. [Google Scholar] [CrossRef] [PubMed]
- Kodani, E.; Xuan, Y.T.; Shinmura, K.; Takano, H.; Tang, X.L.; Bolli, R. Delta-opioid receptor-induced late preconditioning is mediated by cyclooxygenase-2 in conscious rabbits. Am. J. Physiol. Heart Circ. Physiol. 2002, 283, H1943–H1957. [Google Scholar] [CrossRef] [PubMed]
- Peart, J.N.; Gross, G.J. Adenosine and opioid receptor-mediated cardioprotection in the rat: Evidence for cross-talk between receptors. Am. J. Physiol. Heart Circ. Physiol. 2003, 285, H81–H89. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.V.; Yang, X.M.; Liu, G.S.; Heusch, G.; Downey, J.M. Acetylcholine, bradykinin, opioids, and phenylephrine, but not adenosine, trigger preconditioning by generating free radicals and opening mitochondrial K(ATP) channels. Circ. Res. 2001, 89, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Fagbemi, O.; Kane, K.A.; Lepran, I. Anti-arrhythmic actions of meptazinol, a partial agonist at opiate receptors, in acute myocardial ischemia. Br. J. Pharmacol. 1983, 78, 455–460. [Google Scholar] [CrossRef] [PubMed]
- Saini, V.; Carr, D.B.; Verrier, R.L. Comparative effects of the opioids fentanyl and buprenorphine on ventricular vulnerability during acute coronary artery occlusion. Cardiovasc. Res. 1989, 23, 1001–1006. [Google Scholar] [CrossRef] [PubMed]
- Hess, L.; Vrana, M.; Vranova, Z.; Fejfar, Z. The antifibrillatory effect of fentanyl, sufentanil and carfentanil in acute phase of local myocardial ischemia in the dog. Acta Cardiol. 1989, 44, 303–311. [Google Scholar] [PubMed]
- Maslov, L.N.; Oeltgen, P.R.; Lishmanov, Y.B.; Brown, S.A.; Barzakh, E.I.; Krylatov, A.V.; Pei, J.M. Activation of peripheral delta opioid receptors increases cardiac tolerance to arrhythmogenic effect of ischemia/reperfusion. Acad. Emerg. Med. 2014, 21, 31–39. [Google Scholar] [CrossRef] [PubMed]
- Wittert, G.; Hope, P.; Pyle, D. Tissue distribution of opioid receptor gene expression in the rat. Biochem. Biophys. Res. Commun. 1996, 218, 877–881. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, L.O.; Bergh, N.; Li, L.; Bissessar, E.; Bobrova, I.; Gross, G.J.; Akyürek, L.M.; Grip, L. Dose-dependent cardioprotection of enkephalin analogue Eribis peptide 94 and cardiac expression of opioid receptors in a porcine model of ischaemia and reperfusion. Eur. J. Pharmacol. 2012, 674, 378–383. [Google Scholar] [CrossRef] [PubMed]
- Theisen, M.M.; Schlottmann, S.; August, C.; Herzog, C.; Theilmeier, G.; Maas, M.; Blumenstiel, J.M.; Weber, T.P.; Van Aken, H.K.; Kaerlein, K.T. Detection and distribution of opioid peptide receptors in porcine myocardial tissue. Pharmacol. Res. 2014, 84, 45–49. [Google Scholar] [CrossRef] [PubMed]
- Head, B.P.; Patel, H.H.; Roth, D.M.; Lai, N.C.; Niesman, I.R.; Farquhar, M.G.; Insel, P.A. G-protein-coupled receptor signaling components localize in both sarcolemmal and intracellular caveolin-3-associated microdomains in adult cardiac myocytes. J. Biol. Chem. 2005, 280, 31036–31044. [Google Scholar] [CrossRef] [PubMed]
- He, S.F.; Jin, S.Y.; Yang, W.; Pan, Y.L.; Huang, J.; Zhang, S.J.; Zhang, L.; Zhang, Y. Cardiac μ-opioid receptor contributes to opioid-induced cardioprotection in chronic heart failure. Br. J. Anaesth. 2018, 121, 26–37. [Google Scholar] [CrossRef] [PubMed]
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561–563. [Google Scholar] [CrossRef] [PubMed]
- Saleembhasha, A.; Mishra, S. Novel molecules lncRNAs, tRFs and circRNAs deciphered from next-generation sequencing/RNA sequencing: Computational databases and tools. Brief. Funct. Genom. 2018, 17, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.L. Linking long noncoding RNA localization and function. Trends Biochem. Sci. 2016, 41, 761–772. [Google Scholar] [CrossRef] [PubMed]
- DiStefano, J.K. The emerging role of long noncoding RNAs in human disease. Methods Mol. Biol. 2018, 1706, 91–110. [Google Scholar] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Kim, D.; Chang, H.R.; Baek, D. Rules for functional microRNA targeting. BMB Rep. 2017, 50, 554–559. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.; Ajay, S.S.; Yook, J.I.; Kim, H.S.; Hong, S.H.; Kim, N.H.; Dhanasekaran, S.M.; Chinnaiyan, A.M.; Athey, B.D. New class of microRNA targets containing simultaneous 5′-UTR and 3′-UTR interaction sites. Genome Res. 2009, 19, 1175–1183. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Xu, Y.; Xie, X.; Wang, T.; Ko, J.H.; Zhou, T. The role of RNA structure at 5’ untranslated region in microRNA-mediated gene regulation. RNA 2014, 20, 1369–1375. [Google Scholar] [CrossRef] [PubMed]
- Hausser, J.; Syed, A.P.; Bilen, B.; Zavolan, M. Analysis of CDS-located miRNA target sites suggests that they can effectively inhibit translation. Genome Res. 2013, 23, 604–615. [Google Scholar] [CrossRef] [PubMed]
- Lytle, J.R.; Yario, T.A.; Steitz, J.A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5’ UTR as in the 3’ UTR. Proc. Natl. Acad. Sci. USA 2007, 104, 9667–9672. [Google Scholar] [CrossRef] [PubMed]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Schirle, N.T.; Sheu-Gruttadauria, J.; MacRae, I.J. Structural basis for microRNA targeting. Science 2014, 31, 608–613. [Google Scholar] [CrossRef] [PubMed]
- Pratt, A.J.; MacRae, I.J. The RNA-induced silencing complex: A versatile gene-silencing machine. J. Biol. Chem. 2009, 284, 17897–17901. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef] [PubMed]
- Borchert, G.M.; Lanier, W.; Davidson, B.L. RNA polymerase III transcribes human microRNAs. Nat. Struct. Mol. Biol. 2006, 13, 1097–1101. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Rådmark, O.; Kim, S.; et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: MicroRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Huntzinger, E.; Izaurralde, E. Gene silencing by microRNAs: Contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011, 12, 99–110. [Google Scholar] [CrossRef] [PubMed]
- Ruby, J.G.; Jan, C.H.; Bartel, D.P. Intronic microRNA precursors that bypass Drosha processing. Nature 2007, 448, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, D.S.; Hutvágner, G.; Du, T.; Xu, Z.; Aronin, N.; Zamore, P.D. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003, 115, 199–208. [Google Scholar] [CrossRef]
- Goff, L.A.; Rinn, J.L. Linking RNA biology to lncRNAs. Genome Res. 2015, 25, 1456–1465. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, J.E. Long noncoding RNAs: Re-writing dogmas of RNA processing and stability. Biochim. Biophys. Acta 2016, 1859, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Yang, L.; Chen, L.L. The diversity of long noncoding RNAs and their generation. Trends Genet. 2017, 33, 540–552. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.E.; Alvarez-Dominguez, J.R.; Kline, N.; Huynh, N.J.; Geisler, S.; Hu, W.; Coller, J.; Baker, K.E. Translation of small open reading frames within unannotated RNA transcripts in Saccharomyces cerevisiae. Cell Rep. 2014, 7, 1858–1866. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Mattick, J.S. Structure and function of long noncoding RNAs in epigenetic regulation. Nat. Struct. Mol. Biol. 2013, 20, 300–307. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Kutter, C.; Watt, S.; Stefflova, K.; Wilson, M.D.; Goncalves, A.; Ponting, C.P.; Odom, D.T.; Marques, A.C. Rapid turnover of long noncoding RNAs and the evolution of gene expression. PLoS Genet. 2012, 8. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Martin, L.; Chang, H.Y. Uncovering the role of genomic “dark matter” in human disease. J. Clin. Investig. 2012, 122, 1589–1595. [Google Scholar] [CrossRef] [PubMed]
- Barbierato, M.; Zusso, M.; Skaper, S.D.; Giusti, P. MicroRNAs: Emerging role in the endogenous μ opioid system. CNS Neurol. Disord. Drug. Targets 2015, 14, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Dave, R.S.; Khalili, K. Morphine treatment of human monocyte-derived macrophages induces differential miRNA and protein expression: Impact on inflammation and oxidative stress in the central nervous system. J. Cell Biochem. 2010, 110, 834–845. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Yang, C.; Kirkmire, C.M.; Wang, Z.J. Regulation of opioid tolerance by let-7 family microRNA targeting the mu opioid receptor. J. Neurosci. 2010, 30, 10251–10258. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Zhang, L.; Law, P.Y.; Wei, L.N.; Loh, H.H. Long-term morphine treatment decreases the association of mu-opioid receptor (MOR1) mRNA with polysomes through miRNA23b. Mol. Pharmacol. 2009, 75, 744–750. [Google Scholar] [CrossRef] [PubMed]
- Pillai, R.S.; Bhattacharyya, S.N.; Artus, C.G.; Zoller, T.; Cougot, N.; Basyuk, E.; Bertrand, E.; Filipowicz, W. Inhibition of translational initiation by let-7 MicroRNA in human cells. Science 2005, 309, 1573–1576. [Google Scholar] [CrossRef] [PubMed]
- Wanet, A.; Tacheny, A.; Arnould, T.; Renard, P. miR-212/132 expression and functions: Within and beyond the neuronal compartment. Nucleic Acids Res. 2012, 40, 4742–4753. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Gao, Y.; Gong, S.; Guo, S.; Hisamitsu, T.; Jiang, X. Regulation of μ-opioid type 1 receptors by microRNA134 in dorsal root ganglion neurons following peripheral inflammation. Eur. J. Pain 2013, 17, 313–323. [Google Scholar] [CrossRef] [PubMed]
- Aqeilan, R.I.; Calin, G.A.; Croce, C.M. miR-15a and miR-16-1 in cancer: Discovery, function and future perspectives. Cell Death Differ. 2010, 17, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wei, Y.; Li, X.; Liang, X.; Wang, L.; Song, J.; Zhang, X.; Zhang, C.; Niu, J.; Zhang, P.; et al. microRNA-874 suppresses tumor proliferation and metastasis in hepatocellular carcinoma by targeting the DOR/EGFR/ERK pathway. Cell Death Dis. 2018, 9, 130. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Li, H.; Jiang, W.; Zhang, C.; McNutt, M.A.; Li, G. Simian immunodeficiency virus impacts microRNA-16 mediated post-transcriptional regulation of mu opioid receptor in CEM × 174 cells. J. Cell. Biochem. 2016, 117, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Ide, S.; Han, W.; Kasai, S.; Hata, H.; Sora, I.; Ikeda, K. Characterization of the 3’ untranslated region of the human mu-opioid receptor (MOR-1) mRNA. Gene 2005, 364, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Han, W.; Kasai, S.; Hata, H.; Takahashi, T.; Takamatsu, Y.; Yamamoto, H.; Uhl, G.R.; Sora, I.; Ikeda, K. Intracisternal A-particle element in the 3’ noncoding region of the mu-opioid receptor gene in CXBK mice: A new genetic mechanism underlying differences in opioid sensitivity. Pharm. Genom. 2006, 16, 451–460. [Google Scholar] [CrossRef] [PubMed]
- Regan, P.M.; Sariyer, I.K.; Langford, T.D.; Datta, P.K.; Khalili, K. Morphine-induced MOR-1X and ASF/SF2 expressions are independent of transcriptional regulation: Implications for MOR-1X signaling. J. Cell. Physiol. 2016, 231, 1542–1553. [Google Scholar] [CrossRef] [PubMed]
- Brodsky, M.; Elliott, K.; Hynansky, A.; Inturrisi, C.E. CNS levels of mu opioid receptor (MOR-1) mRNA during chronic treatment with morphine or naltrexone. Brain Res. Bull. 1995, 38, 135–141. [Google Scholar] [CrossRef]
- Lee, H.; Han, S.; Kwon, C.S.; Lee, D. Biogenesis and regulation of the let-7 miRNAs and their functional implications. Protein Cell. 2016, 7, 100–113. [Google Scholar] [CrossRef] [PubMed]
- Grimson, A.; Farh, K.K.; Johnston, W.K.; Garrett-Engele, P.; Lim, L.P.; Bartel, D.P. MicroRNA targeting specificity in mammals: Determinants beyond seed pairing. Mol. Cell 2007, 27, 91–105. [Google Scholar] [CrossRef] [PubMed]
- Roush, S.; Slack, F.J. The let-7 family of microRNAs. Trends Cell Biol. 2008, 18, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007, 21, 1025–1030. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.S.; Erkeland, S.J.; Pester, R.E.; Chen, C.Y.; Ebert, M.S.; Sharp, P.A.; Jacks, T. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc. Natl. Acad. Sci. USA 2008, 105, 3903–3908. [Google Scholar] [CrossRef] [PubMed]
- Chafin, C.B.; Regna, N.L.; Caudell, D.L.; Reilly, C.M. MicroRNA-let-7a promotes E2F-mediated cell proliferation and NFκB activation in vitro. Cell. Mol. Immunol. 2014, 11, 79–83. [Google Scholar] [CrossRef] [PubMed]
- Büssing, I.; Slack, F.J.; Grosshans, H. Let-7 microRNAs in development, stem cells and cancer. Trends Mol. Med. 2008, 14, 400–409. [Google Scholar] [CrossRef] [PubMed]
- Klee, W.A.; Streaty, R.A. Narcotic receptor sites in morphine-dependent rats. Nature 1974, 248, 61–63. [Google Scholar] [CrossRef] [PubMed]
- Holaday, J.W.; Hitzemann, R.J.; Curell, J.; Tortella, F.C.; Belenky, G.L. Repeated electroconvulsive shock or chronic morphine treatment increases the number of 3H-D-Ala2, D-Leu5-enkephalin binding sites in rat brain membranes. Life Sci. 1982, 31, 2359–2362. [Google Scholar] [CrossRef]
- Lewis, J.W.; Lewis, M.E.; Loomus, D.J.; Akil, H. Acute systemic administration of morphine selectively increases mu opioid receptor binding in the rat brain. Neuropeptides 1984, 5, 117–120. [Google Scholar] [CrossRef]
- Rothman, R.B.; Danks, J.A.; Jacobson, A.E.; Burke, T.R.; Rice, K.C.; Tortella, F.C.; Holaday, J.W. Morphine tolerance increases mu-noncompetitive delta binding sites. Eur. J. Pharmacol. 1986, 124, 113–119. [Google Scholar] [CrossRef]
- Koch, T.; Höllt, V. Role of receptor internalization in opioid tolerance and dependence. Pharmacol. Ther. 2008, 117, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Law, P.Y.; Wei, L.N.; Loh, H.H. Post-transcriptional regulation of mouse mu opioid receptor (MOR1) via its 3’ untranslated region: A role for microRNA23b. FASEB J. 2008, 22, 4085–4095. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Concejo, A.; Jimenez-Gonzalez, A.; Rodríguez, R.E. μ opioid receptor expression after morphine administration is regulated by miR-212/132 cluster. PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.E.; Lioy, D.T.; Ma, L.; Impey, S.; Mandel, G.; Goodman, R.H. Homeostatic regulation of MeCP2 expression by a CREB-induced microRNA. Nat. Neurosci. 2007, 10, 1513–1514. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Hwang, C.K.; Zheng, H.; Wagley, Y.; Lin, H.Y.; Kim, D.K.; Law, P.Y.; Loh, H.H.; Wei, L.N. MicroRNA 339 down-regulates μ-opioid receptor at the post-transcriptional level in response to opioid treatment. FASEB J. 2013, 27, 522–535. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Xu, J.; Xu, M.; Pasternak, G.W.; Pan, Y.X. Morphine regulates expression of μ-opioid receptor MOR-1A, an intron-retention carboxyl terminal splice variant of the μ-opioid receptor (OPRM1) gene via miR-103/miR-107. Mol. Pharmacol. 2014, 85, 368–380. [Google Scholar] [CrossRef] [PubMed]
- Bare, L.A.; Mansson, E.; Yang, D. Expression of two variants of the human mu opioid receptor mRNA in SK-N-SH cells and human brain. FEBS Lett. 1994, 354, 213–216. [Google Scholar] [CrossRef]
- Trajkovski, M.; Hausser, J.; Soutschek, J.; Bhat, B.; Akin, A.; Zavolan, M.; Heim, M.H.; Stoffel, M. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 2011, 474, 649–653. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.Y.; Surapureddi, S.; Coulter, S.; Ferguson, S.S.; Goldstein, J.A. Human CYP2C8 is post-transcriptionally regulated by microRNAs 103 and 107 in human liver. Mol. Pharmacol. 2012, 82, 529–540. [Google Scholar] [CrossRef] [PubMed]
- Polster, B.J.; Westaway, S.K.; Nguyen, T.M.; Yoon, M.Y.; Hayflick, S.J. Discordant expression of miR-103/7 and pantothenate kinase host genes in mouse. Mol. Genet. Metab. 2010, 101, 292–295. [Google Scholar] [CrossRef] [PubMed]
- Wilfred, B.R.; Wang, W.X.; Nelson, P.T. Energizing miRNA research: A review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways. Mol. Genet. Metab. 2007, 91, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xu, W.; Zhong, T.; Song, Z.; Zou, Y.; Ding, Z.; Guo, Q.; Dong, X.; Zou, W. miR-365 targets β-arrestin 2 to reverse morphine tolerance in rats. Sci. Rep. 2016, 6, 38285. [Google Scholar] [CrossRef] [PubMed]
- Bohn, L.M.; Gainetdinov, R.R.; Lin, F.T.; Lefkowitz, R.J.; Caron, M.G. Mu-opioid receptor desensitization by beta-arrestin-2 determines morphine tolerance but not dependence. Nature 2000, 408, 720–723. [Google Scholar] [CrossRef] [PubMed]
- Zhi, F.; Xue, L.; Shao, N.; Deng, D.; Kang, X.; Chao, D.; Xu, Y.; Wang, R.; Yang, Y.; Xia, Y. δ-Opioid receptor activation and microRNA expression in the rat heart under prolonged hypoxia. Cell Physiol. Biochem. 2016, 39, 1118–1128. [Google Scholar] [CrossRef] [PubMed]
- Meng, S.; Cao, J.; Wang, L.; Zhou, Q.; Li, Y.; Shen, C.; Zhang, X.; Wang, C. MicroRNA 107 partly inhibits endothelial progenitor cells differentiation via HIF-1β. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Chan, Y.C.; Khanna, S.; Roy, S.; Sen, C.K. miR-200b targets Ets-1 and is down-regulated by hypoxia to induce angiogenic response of endothelial cells. J. Biol. Chem. 2011, 286, 2047–2056. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Li, R.; Zhang, C.; Bian, H.J.; Wang, F.; Xiao, J.; Liu, S.W.; Yi, W.; Zhang, M.X.; Wang, S.X.; et al. MicroRNA-7a/b protects against cardiac myocyte injury in ischemia/reperfusion by targeting poly (ADP-ribose) polymerase. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- He, S.F.; Zhu, H.J.; Han, Z.Y.; Wu, H.; Jin, S.Y.; Irwin, M.G.; Zhang, Y. MicroRNA-133b-5p is involved in cardioprotection of morphine preconditioning in rat cardiomyocytes by targeting fas. Can. J. Cardiol. 2016, 32, 996–1007. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Gu, H.; Xu, W.; Zhou, X. Down-regulation of lncRNA MALAT1 reduces cardiomyocyte apoptosis and improves left ventricular function in diabetic rats. Int. J. Cardiol. 2016, 203, 214–216. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.J.; Han, Z.Y.; He, S.F.; Jin, S.Y.; Xu, S.J.; Fang, X.D.; Zhang, Y. Specific MicroRNAs comparisons in hypoxia and morphine preconditioning against hypoxia-reoxgenation injury with and without heart failure. Life Sci. 2017, 170, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Xia, L.; Chen, M.; Lin, C.; Wu, H.; Zhang, Y.; Pan, S.; Li, X. miR-133b, a particular member of myomiRs, coming into playing its unique pathological role in human cancer. Oncotarget 2017, 8, 50193–50208. [Google Scholar] [CrossRef] [PubMed]
- Miki, T.; Miura, T.; Tanno, M.; Sakamoto, J.; Kuno, A.; Genda, S.; Matsumoto, T.; Ichikawa, Y.; Shimamoto, K. Interruption of signal transduction between G protein and PKC-epsilon underlies the impaired myocardial response to ischemic preconditioning in postinfarct remodeled hearts. Mol. Cell. Biochem. 2003, 247, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Kim, S.Y.; Kwak, Y.L.; Hwang, K.C.; Shim, Y.H. Hyperglycemia attenuates myocardial preconditioning of remifentanil. J. Surg. Res. 2012, 174, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Andersen, A.; Povlsen, J.A.; Bøtker, H.E.; Nielsen-Kudsk, J.E. Right ventricular hypertrophy and failure abolish cardioprotection by ischaemic pre-conditioning. Eur. J. Heart Fail. 2013, 15, 1208–1214. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Simon, F.M.; Zhang, X.X.; Loh, H.H.; Law, P.Y.; Rodriguez, R.E. Morphine regulates dopaminergic neuron differentiation via miR-133b. Mol. Pharmacol. 2010, 78, 935–942. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.C.; Zheng, J.Y.; Tang, L.J.; Huang, B.S.; Li, K.; Tao, Y.; Yu, W.; Zhu, R.L.; Li, S.; Li, L.X. miR-133b promotes neurite outgrowth by targeting RhoA expression. Cell. Physiol. Biochem. 2015, 35, 246–258. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.M.; Gibbs, K.M.; Davila, J.; Campbell, N.; Sung, S.; Todorova, T.I.; Otsuka, S.; Sabaawy, H.E.; Hart, R.P.; Schachner, M. MicroRNA miR-133b is essential for functional recovery after spinal cord injury in adult zebrafish. Eur. J. Neurosci. 2011, 33, 1587–1597. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.; Wu, Z.; Tian, L.; Li, D.; Wang, X.; He, Y.; Jin, W.; Li, M.; Zhu, Q.; Shang, T.; et al. Long noncoding RNA AK12348 is involved in the regulation of myocardial ischaemia-reperfusion injury by targeting PARP and caspase-3. Heart Lung Circ. 2018, 27, e51–e58. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.B.; Katwadi, K.; Kwek, X.Y.; Ismail, N.I.; Chinda, K.; Ong, S.G.; Hausenloy, D.J. Non-coding RNAs as therapeutic targets for preventing myocardial ischemia-reperfusion injury. Expert Opin. Ther. Targets 2018, 22, 247–261. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Wang, Q.; Yuan, Z.; Chen, A.; Liu, Z.; Wang, Z.; Li, H. LncRNA-MALAT1 promotes CPC proliferation and migration in hypoxia by up-regulation of JMJD6 via sponging miR-125. Biochem. Biophys. Res. Commun. 2018, 499, 711–718. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.Y.; Dong, B.; Tang, L.; Zhou, S.H. LncRNA MALAT1 sponges miR-133 to promote NLRP3 inflammasome expression in ischemia-reperfusion injured heart. Int. J. Cardiol. 2018, 254. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.H.; Hao, W.; Meng, Q.T.; Du, X.B.; Lei, S.Q.; Xia, Z.Y. Long non-coding RNA MALAT1 functions as a mediator in cardioprotective effects of fentanyl in myocardial ischemia-reperfusion injury. Cell. Biol. Int. 2017, 41, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Li, J.; Xu, T.; Zhou, D.D.; Zhang, L.; Wang, X. MicroRNA-145 induces apoptosis of glioma cells by targeting BNIP3 and Notch signaling. Oncotarget 2017, 8, 61510–61527. [Google Scholar] [CrossRef] [PubMed]
- Viereck, J.; Thum, T. Circulating noncoding RNAs as biomarkers of cardiovascular disease and injury. Circ. Res. 2017, 120, 381–399. [Google Scholar] [CrossRef] [PubMed]
- Huang, W. MicroRNAs: Biomarkers, diagnostics, and therapeutics. Methods Mol. Biol. 2017, 1617, 57–67. [Google Scholar] [PubMed]
- E, S.; Costa, M.C.; Kurc, S.; Drożdż, A.; Cortez-Dias, N.; Enguita, F.J. The circulating non-coding RNA landscape for biomarker research: Lessons and prospects from cardiovascular diseases. Acta Pharmacol. Sin. 2018, 39, 1085–1099. [Google Scholar] [CrossRef] [PubMed]
- Zampetaki, A.; Willeit, P.; Tilling, L.; Drozdov, I.; Prokopi, M.; Renard, J.M.; Mayr, A.; Weger, S.; Schett, G.; Shah, A.; et al. Prospective study on circulating microRNAs and risk of myocardial infarction. J. Am. Coll. Cardiol. 2012, 60, 290–299. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.K.; Zhu, J.Q.; Zhang, J.T.; Li, Q.; Li, Y.; He, J.; Qin, Y.W.; Jing, Q. Circulating microRNA: A novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur. Heart J. 2010, 31, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Devaux, Y.; Vausort, M.; McCann, G.P.; Kelly, D.; Collignon, O.; Ng, L.L.; Wagner, D.R.; Squire, I.B. A panel of 4 microRNAs facilitates the prediction of left ventricular contractility after acute myocardial infarction. PLoS ONE 2013, 8. [Google Scholar] [CrossRef]
- Feistritzer, H.J.; Klug, G.; Reinstadler, S.J.; Reindl, M.; Mayr, A.; Mair, J.; Metzler, B. Novel biomarkers predicting cardiac function after acute myocardial infarction. Br. Med. Bull. 2016, 119, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Choong, O.K.; Lee, D.S.; Chen, C.Y.; Hsieh, P.C.H. The roles of non-coding RNAs in cardiac regenerative medicine. Noncoding RNA Res. 2017, 2, 100–110. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, J. Therapeutic potentials of noncoding RNAs: Targeted delivery of ncRNAs in cancer cells. Adv. Exp. Med. Biol. 2016, 927, 429–458. [Google Scholar] [PubMed]
- Somogyi, A.A.; Barratt, D.T.; Coller, J.K. Pharmacogenetics of opioids. Clin. Pharmacol. Ther. 2007, 81, 429–444. [Google Scholar] [CrossRef] [PubMed]
- Owusu Obeng, A.; Hamadeh, I.; Smith, M. Review of opioid pharmacogenetics and considerations for pain management. Pharmacotherapy 2017, 37, 1105–1121. [Google Scholar] [CrossRef] [PubMed]
| Opioid Agonist | Location | Species | miRNAs | Expression | Reference |
|---|---|---|---|---|---|
| Morphine | Macrophages | Human | miR-146a-5p miR-24-5p miR-1826-5p miR-16-5p miR-150-5p miR-30c-5p miR-155-5p miR-23a/b-5p miR-221-5p miR-15b-5p miR-132-5p | Upregulated | [109] |
| Morphine | Macrophages | Human | miR-26a-5p miR-191-5p miR-26b-5p miR-21-5p miR-320c-5p miR-423-5p miR-103-5p miR-107-5p miR-320a-5p miR-99b-5p miR-1915-5p miR-1469-5p miR-638-5p miR-181b-5p | Downregulated | [109] |
| Hydromorphone Oxycodone | Serum | Human | Let-7a miR-423-3p miR-199a-3p miR-146a/b-5p miR-23b-3p miR-24-3p miR-221-3p miR-223-3p | Upregulated | [15] |
| Hydromorphone Oxycodone | Serum | Human | miR-144-3p miR-451a-5p miR-192-5p miR-215-5p miR-363-3p miR-194-5p miR-140-3p miR-16-5p miR-15a-5p miR-92a-3p miR-101-3p miR-19b-3p miR-19a-3p miR-15b-3p miR-425-5p miR-93-5p miR-20a-5p | Downregulated | [15] |
| Morphine | Neuroblastoma derived cells | Mice | Let-7 | Upregulated | [110] |
| Morphine | Neuronal cells | Mice | miR-23b-5p | Upregulated | [111,113] |
| Morphine | Embryos and neurons | Zebrafish | miR-212-5p miR-132-5p | Upregulated | [114] |
| Morphine Fentanyl | Hippocampus | Mice | miR-339-3p | Upregulated | [115] |
| Morphine | Lymphocytes | Human | miR-16-5p | Downregulated | [112] |
| Morphine | HEK-293 cells, Be(2)C cells and striatum | Human Mice | miR-103-5p miR-107-5p | Upregulated | [116] |
| Morphine | Neurons | Rat | miR-365-5p | Downregulated | [117] |
| Opioid Agonist | Location | Species | miRNAs | Expression | Reference |
|---|---|---|---|---|---|
| UFP-512 * | Heart | Rat | miR-107-3p miR-141-3p miR-350-5p | Upregulated | [146] |
| UFP-512 * | Heart | Rat | miR-7a/b mi-107-3p miR-200b-5p miR-376a-3p miR-134-5p | Upregulated ** | [146] |
| Morphine | Ventricular myocytes | Rat | miR-423-5p miR-106b-5p miR-345-5p miR-3571-5p miR-130a-3p miR-133b-5p miR-3596d-5p miR-152-3p miR-15b-5p miR-3582-5p miR-374-5p miR-3596a-5p miR-3473-5p miR-3068-3p miR-16-5p miR-133a-5p miR-6215-5p | Upregulated | [132] |
| Morphine | Ventricular myocytes | Rat | miR-25-3p miR-466b-2-3p miR-214-3p miR-328a-5p miR-181a/b/c-5p miR-148b-3p miR-133a/b-3p miR-29a-3p let-7i miR-125a-5p miR-466b-1-3p miR-208a-3p miR-499-3p miR-466c-3p miR-483-5p miR-505-3p miR-1224-5p miR-3584-5p miR-6216-5p | Downregulated | [132] |
| Morphine | Cardiomyocytes *** | Rat | miR-130a-3p miR-133a/b-5p miR-16-5p miR-6215-5p miR-151-5p miR-143-3p miR-107-3p miR-125b-5p miR-150-5p miR-29a-3p miR-378b-5p | Upregulated | [133] |
| Morphine | Cardiomyocytes *** | Rat | Let-7i miR-1224-5p miR-125a-5p miR-133a/b-3p miR-181a-5p miR-29a-3p miR-6216-5p miR-30a/e-5p miR-6215-5p | Downregulated | [133] |
| Fentanyl | Cardiomyocytes | Mice | miRNA-145-5p | Upregulated | [152] |
| Fentanyl | Cardiomyocytes | Mice | lncRNA MALAT1 | Downregulated | [152] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Melo, Z.; Ishida, C.; Goldaraz, M.D.l.P.; Rojo, R.; Echavarria, R. Novel Roles of Non-Coding RNAs in Opioid Signaling and Cardioprotection. Non-Coding RNA 2018, 4, 22. https://doi.org/10.3390/ncrna4030022
Melo Z, Ishida C, Goldaraz MDlP, Rojo R, Echavarria R. Novel Roles of Non-Coding RNAs in Opioid Signaling and Cardioprotection. Non-Coding RNA. 2018; 4(3):22. https://doi.org/10.3390/ncrna4030022
Chicago/Turabian StyleMelo, Zesergio, Cecilia Ishida, Maria De la Paz Goldaraz, Rocio Rojo, and Raquel Echavarria. 2018. "Novel Roles of Non-Coding RNAs in Opioid Signaling and Cardioprotection" Non-Coding RNA 4, no. 3: 22. https://doi.org/10.3390/ncrna4030022
APA StyleMelo, Z., Ishida, C., Goldaraz, M. D. l. P., Rojo, R., & Echavarria, R. (2018). Novel Roles of Non-Coding RNAs in Opioid Signaling and Cardioprotection. Non-Coding RNA, 4(3), 22. https://doi.org/10.3390/ncrna4030022