A Novel Long Non-Coding RNA in the hTERT Promoter Region Regulates hTERT Expression
Abstract
1. Introduction
2. Results
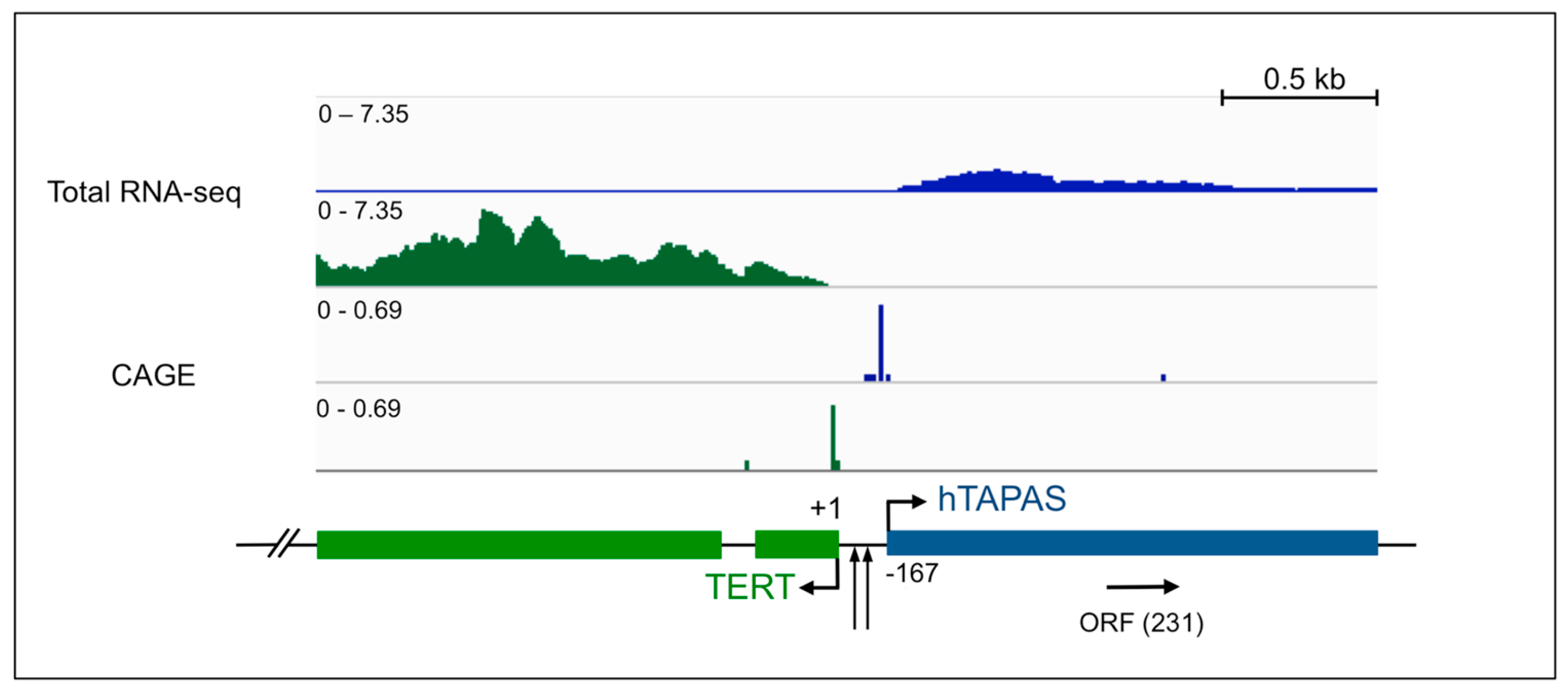
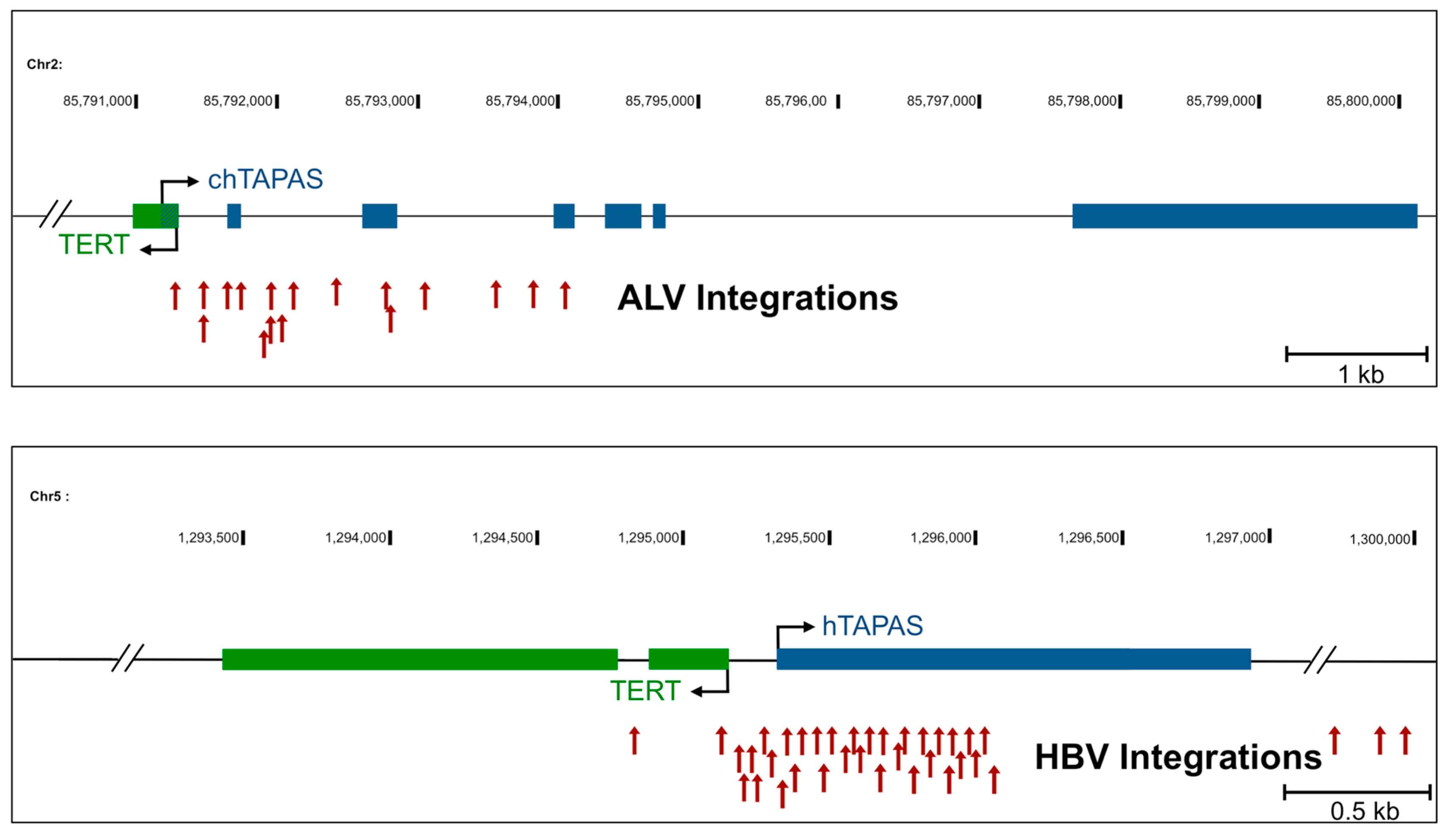
2.1. An Antisense Transcript Is Expressed Upstream of hTERT
2.2. hTAPAS Exhibits Features of a Long Non-Coding RNA
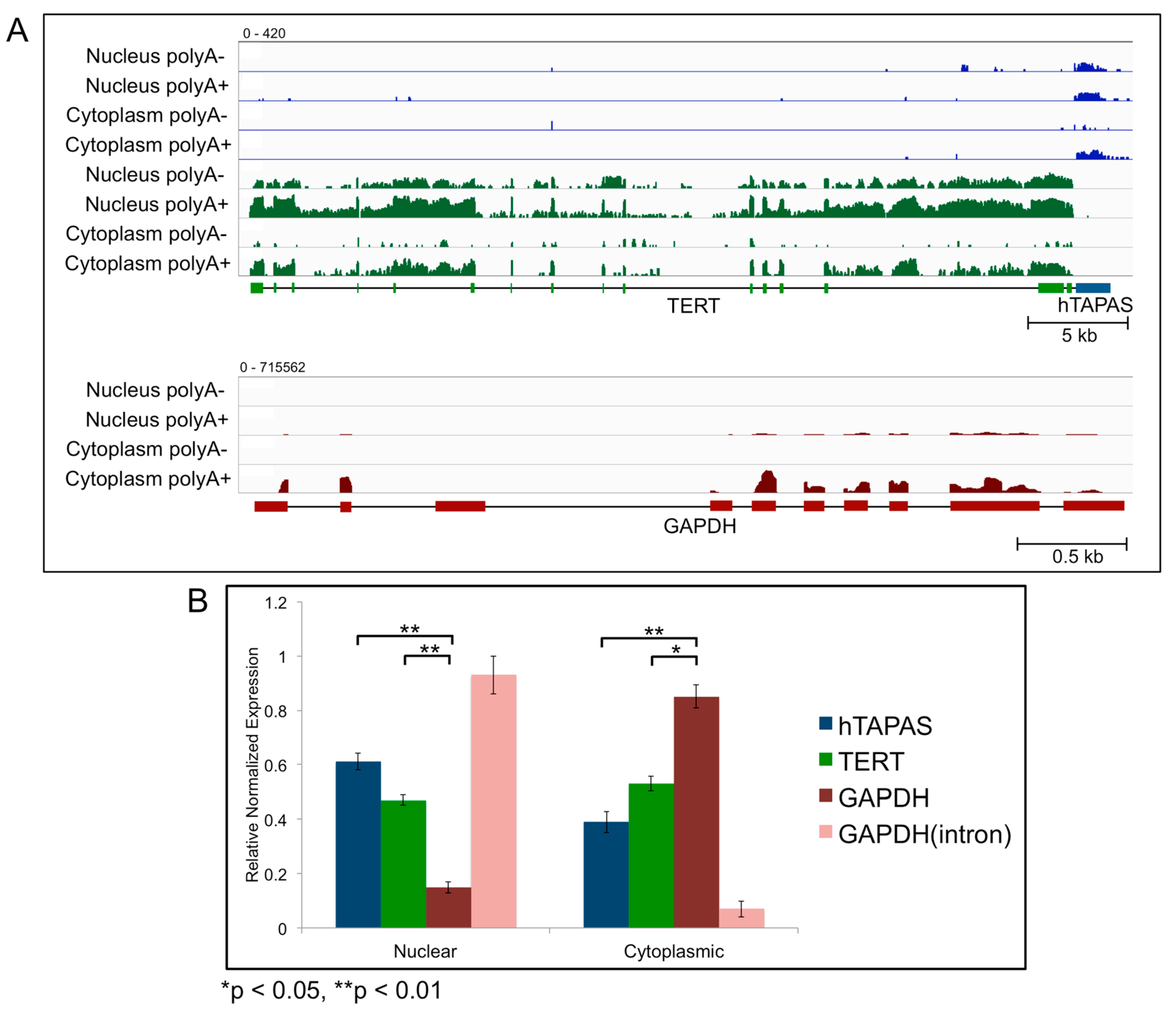
2.3. Sub-Cellular Localization of hTAPAS and hTERT RNAs
2.4. hTAPAS Expression Negatively Correlates with hTERT Expression in Cancer Patients
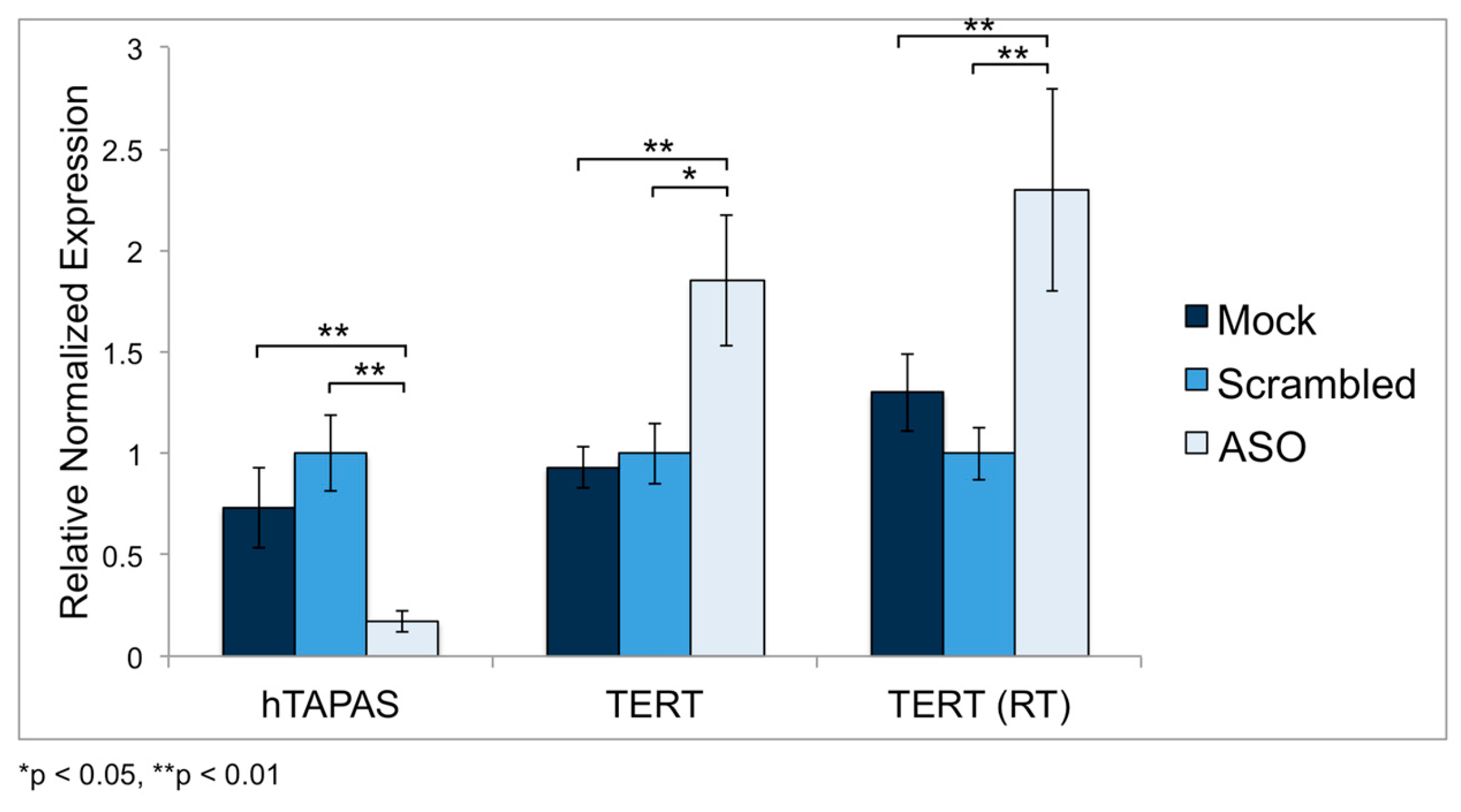
2.5. Knockdown of hTAPAS up Regulates hTERT Expression in HEK-293 Cells
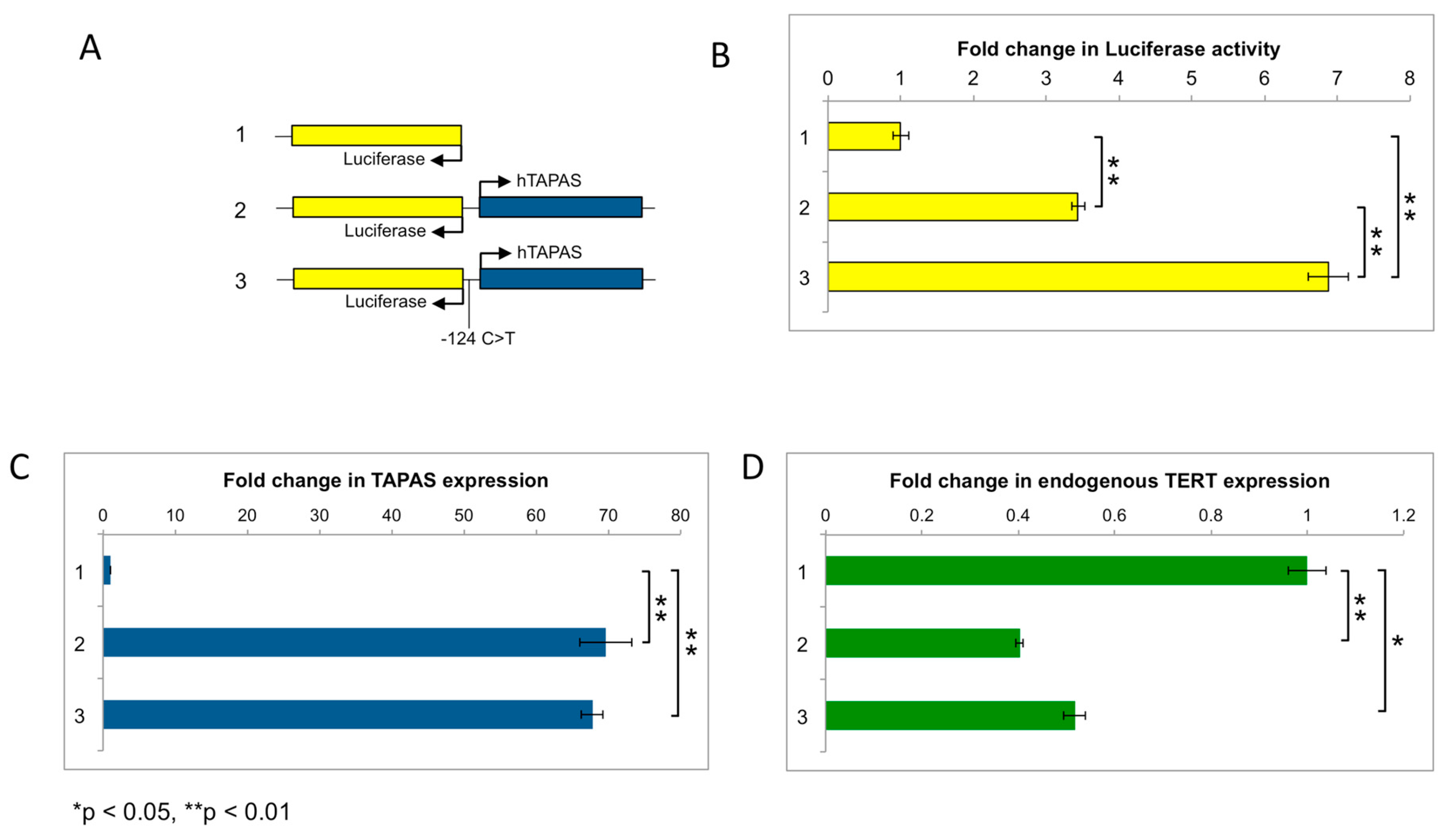
2.6. hTAPAS Expression Is Not Affected by the hTERT Promoter Mutation
3. Discussion
4. Materials and Methods
4.1. Analysis of RNA Sequencing and CAGE Data
4.2. Evolutionary Conservation and Coding Potential Analysis
4.3. Fractionation Experiments
4.4. RNA Isolation and Quantitative Reverse Transcription PCR
4.5. hTAPAS Knockdown Experiments Using Antisense Oligonucleotides
4.6. hTAPAS Constructs and Luciferase Assays
4.7. Nucleotide Sequence Accession Number
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Ethical Statement
References
- Blasco, M.A. Telomeres and human disease: Ageing, cancer and beyond. Nat. Rev. Genet. 2005, 6, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.W.; Piatyszek, M.A.; Prowse, K.R.; Harley, C.B.; West, M.D.; Ho, P.L.; Coviello, G.M.; Wright, W.E.; Weinrich, S.L.; Shay, J.W. Specific association of human telomerase activity with immortal cells and cancer. Science 1994, 266, 2011–2015. [Google Scholar] [CrossRef] [PubMed]
- Garcia, C.K.; Wright, W.E.; Shay, J.W. Human diseases of telomerase dysfunction: Insights into tissue aging. Nucleic Acids Res. 2007, 35, 7406–7416. [Google Scholar] [CrossRef] [PubMed]
- Shay, J.W.; Wright, W.E. Role of telomeres and telomerase in cancer. Semin. Cancer Biol. 2011, 21, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Zhao, Y.; Wang, S. Chromatin and epigenetic regulation of the telomerase reverse transcriptase gene. Protein Cell 2010, 1, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Cevik, D.; Yildiz, G.; Ozturk, M. Common telomerase reverse transcriptase promoter mutations in hepatocellular carcinomas from different geographical locations. World J. Gastroenterol. 2015, 21, 311–317. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.W.; Hodis, E.; Xu, M.J.; Kryukov, G.V.; Chin, L.; Garraway, L.A. Highly recurrent TERT promoter mutations in human melanoma. Science 2013, 339, 957–959. [Google Scholar] [CrossRef] [PubMed]
- Withers, J.B.; Ashvetiya, T.; Beemon, K.L. Exclusion of Exon 2 Is a Common mRNA Splice Variant of Primate Telomerase Reverse Transcriptases. PLoS ONE 2012, 7, e48016. [Google Scholar] [CrossRef] [PubMed]
- Cong, Y.-S.; Wright, W.E.; Shay, J.W. Human telomerase and its regulation. Microbiol. Mol. Biol. Rev. 2002, 66, 407–425. [Google Scholar] [CrossRef] [PubMed]
- Horn, S.; Figl, A.; Rachakonda, P.S.; Fischer, C.; Sucker, A.; Gast, A.; Kadel, S.; Moll, I.; Nagore, E.; Hemminki, K.; et al. TERT Promoter Mutations in Familial and Sporadic Melanoma. Science 2013, 339, 959–961. [Google Scholar] [CrossRef] [PubMed]
- Killela, P.J.; Reitman, Z.J.; Jiao, Y.; Bettegowda, C.; Agrawal, N.; Diaz, L.A., Jr.; Friedman, A.H.; Friedman, H.; Gallia, G.L.; Giovanella, B.C.; et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc. Natl. Acad. Sci. USA 2013, 110, 6021–6026. [Google Scholar] [CrossRef] [PubMed]
- Renaud, S.; Bosman, F.T.; Benhattar, J. Implication of the exon region in the regulation of the human telomerase reverse transcriptase gene promoter. Biochem. Biophys. Res. Commun. 2003, 300, 47–54. [Google Scholar] [CrossRef]
- Collins, P.J.; Kobayashi, Y.; Nguyen, L.; Trinklein, N.D.; Myers, R.M. The ets-related transcription factor GABP directs bidirectional transcription. PLoS Genet. 2007, 3, 2247–2255. [Google Scholar] [CrossRef] [PubMed]
- Orekhova, A.S.; Rubtsov, P.M. Bidirectional promoters in the transcription of mammalian genomes. Biochemistry 2013, 78, 335–341. [Google Scholar] [CrossRef] [PubMed]
- Bell, R.J.A.; Rube, H.T.; Kreig, A.; Mancini, A.; Fouse, S.F.; Nagarajan, R.P.; Choi, S.; Hong, C.; He, D.; Pekmezci, M.; et al. The transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 2015, 348, 1036–1039. [Google Scholar] [CrossRef] [PubMed]
- Nehyba, J.; Malhotra, S.; Winans, S.; O’Hare, T.H.; Justice, J.; Beemon, K. Avian Leukosis Virus Activation of an Antisense RNA Upstream of TERT in B-Cell Lymphomas. J. Virol. 2016, 90, 9509–9517. [Google Scholar] [CrossRef] [PubMed]
- ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Siepel, A.; Bejerano, G.; Pedersen, J.S.; Hinrichs, A.S.; Hou, M.; Rosenbloom, K.; Clawson, H.; Spieth, J.; Hillier, L.W.; Richards, S.; et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 2005, 15, 1034–1050. [Google Scholar] [CrossRef] [PubMed]
- Cooper, G.M.; Stone, E.A.; Asimenos, G.; Green, E.D.; Batzoglou, S.; Sidow, A. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 2005, 15, 901–913. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.F.; Jungreis, I.; Kellis, M. PhyloCSF: A comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 2011, 27, i275–i282. [Google Scholar] [CrossRef] [PubMed]
- Wakano, C.; Byun, J.S.; Di, L.J.; Gardner, K. The dual lives of bidirectional promoters. Biochim. Biophys. Acta Gene Regul. Mech. 2012, 1819, 688–693. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Pelechano, V.; Järvelin, A.I.; Steinmetz, L.M. Functional consequences of bidirectional promoters. Trends Genet. 2011, 27, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.; Creighton, C.J.; Davis, C.; Donehower, L.; Drummond, J.; Wheeler, D.; Ally, A.; Balasundaram, M.; Birol, I.; Butterfield, Y.S.N.; et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Masutomi, K.; Yu, E.Y.; Khurts, S.; Ben-Porath, I.; Currier, J.L.; Metz, G.B.; Brooks, M.W.; Kaneko, S.; Murakami, S.; DeCaprio, J.A.; et al. Telomerase maintains telomere structure in normal human cells. Cell 2003, 114, 241–253. [Google Scholar] [CrossRef]
- Heidenreich, B.; Kumar, R. TERT promoter mutations in telomere biology. Mutat. Res. Mutat. Res. 2017, 771, 15–31. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.-C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Sun, B.K.; Erwin, J.A.; Song, J.-J.; Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 2008, 322, 750–756. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.A.; Shah, N.; Wang, K.C.; Kim, J.; Horlings, H.M.; Wong, D.J.; Tsai, M.-C.; Hung, T.; Argani, P.; Rinn, J.L.; et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Kotake, Y.; Nakagawa, T.; Kitagawa, K.; Suzuki, S.; Liu, N.; Kitagawa, M.; Xiong, Y. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15INK4B tumor suppressor gene. Oncogene 2011, 30, 1956–1962. [Google Scholar] [CrossRef] [PubMed]
- Huarte, M. The emerging role of lncRNAs in cancer. Nat. Med. 2015, 21, 1253–1261. [Google Scholar] [CrossRef] [PubMed]
- Ingolia, N.T.; Brar, G.A.; Stern-Ginossar, N.; Harris, M.S.; Talhouarne, G.J.S.; Jackson, S.E.; Wills, M.R.; Weissman, J.S. Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep. 2014, 8, 1365–1379. [Google Scholar] [CrossRef] [PubMed]
- Bodnar, A.G.; Ouellette, M.; Frolkis, M.; Holt, S.E.; Chiu, C.P.; Morin, G.B.; Harley, C.B.; Shay, J.W.; Lichtsteiner, S.; Wright, W.E. Extension of life-span by introduction of telomerase into normal human cells. Science 1998, 279, 349–352. [Google Scholar] [CrossRef] [PubMed]
- Harley, C.B.; Futcher, A.B.; Greider, C.W. Telomeres shorten during ageing of human fibroblasts. Nature 1990, 345, 458–460. [Google Scholar] [CrossRef] [PubMed]
- Ulaner, G.A.; Giudice, L.C. Developmental regulation of telomerase activity in human fetal tissues during gestation. Mol. Hum. Reprod. 1997, 3, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, H.; Benchimol, S. Reconstitution of telomerase activity in normal human cells leads to elongation of telomeres and extended replicative life span. Curr. Biol. 1998, 8, 279–282. [Google Scholar] [CrossRef]
- Wright, W.E.; Piatyszek, M.A.; Rainey, W.E.; Byrd, W.; Shay, J.W. Telomerase activity in human germline and embryonic tissues and cells. Dev. Genet. 1996, 18, 173–179. [Google Scholar] [CrossRef]
- Cristofari, G.; Lingner, J. Telomere length homeostasis requires that telomerase levels are limiting. EMBO J. 2006, 25, 565–574. [Google Scholar] [CrossRef] [PubMed]
- Xi, L.; Cech, T.R. Inventory of telomerase components in human cells reveals multiple subpopulations of hTR and hTERT. Nucleic Acids Res. 2014, 42, 8565–8577. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Xian, R.R.; Li, Y.; Polony, T.S.; Beemon, K.L. Telomerase reverse transcriptase expression elevated by avian leukosis virus integration in B cell lymphomas. Proc. Natl. Acad. Sci. USA 2007, 104, 18952–18957. [Google Scholar] [CrossRef] [PubMed]
- Justice, J.F.; Morgan, R.W.; Beemon, K.L. Common viral integration sites identified in avian leukosis virus- induced B-cell lymphomas. MBio 2015, 6, e01863-15. [Google Scholar] [CrossRef] [PubMed]
- Buendia, M.-A.; Neuveut, C. Hepatocellular Carcinoma. Cold Spring Harb. Perspect. Med. 2015, 5, a021444. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.-H.; Liu, X.; Yan, H.-X.; Li, W.-Y.; Zeng, X.; Yang, Y.; Zhao, J.; Liu, S.-P.; Zhuang, X.-H.; Lin, C.; et al. Genomic and oncogenic preference of HBV integration in hepatocellular carcinoma. Nat. Commun. 2016, 7, 12992. [Google Scholar] [CrossRef] [PubMed]
- Toh, S.T.; Jin, Y.; Liu, L.; Wang, J.; Babrzadeh, F.; Gharizadeh, B.; Ronaghi, M.; Toh, H.C.; Chow, P.K.-H.; Chung, A.Y.-F.; et al. Deep sequencing of the hepatitis B virus in hepatocellular carcinoma patients reveals enriched integration events, structural alterations and sequence variations. Carcinogenesis 2013, 34, 787–798. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Lou, X.; Hua, D.; Yu, W.; Li, L.; Wang, J.; Gao, F.; Zhao, N.; Ren, G.; Li, L.; et al. Recurrent Targeted Genes of Hepatitis B Virus in the Liver Cancer Genomes Identified by a Next-Generation Sequencing-Based Approach. PLoS Genet. 2012, 8, e1003065. [Google Scholar] [CrossRef] [PubMed]
- Delany, M.E.; Daniels, L.M. The chicken telomerase reverse transcriptase (chTERT): Molecular and cytogenetic characterization with a comparative analysis. Gene 2004, 339, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Delany, M.E.; Krupkin, A.B.; Miller, M.M. Organization of telomere sequences in birds: Evidence for arrays of extreme length and for in vivo shortening. Cytogenet. Cell Genet. 2000, 90, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Hackett, J.A.; Greider, C.W. Balancing instability: Dual roles for telomerase and telomere dysfunction in tumorigenesis. Oncogene 2002, 21, 619–626. [Google Scholar] [CrossRef] [PubMed]
- Goodspeed, A.; Heiser, L.M.; Gray, J.W.; Costello, J.C. Tumor-Derived Cell Lines as Molecular Models of Cancer Pharmacogenomics. Mol. Cancer Res. 2016, 14, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Domcke, S.; Sinha, R.; Levine, D.A.; Sander, C.; Schultz, N. Evaluating cell lines as tumour models by comparison of genomic profiles. Nat. Commun. 2013, 4, 2126. [Google Scholar] [CrossRef] [PubMed]
- Everaert, C.; Luypaert, M.; Maag, J.L.V.; Cheng, Q.X.; Dinger, M.E.; Hellemans, J.; Mestdagh, P. Benchmarking of RNA-sequencing analysis workflows using whole-transcriptome RT-qPCR expression data. Sci. Rep. 2017, 7, 1559. [Google Scholar] [CrossRef] [PubMed]
- Chiba, K.; Johnson, J.Z.; Vogan, J.M.; Wagner, T.; Boyle, J.M.; Hockemeyer, D. Cancer-associated TERT promoter mutations abrogate telomerase silencing. Elife 2015, 4, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Cossio, M.L.T.; Giesen, L.F.; Araya, G.; Pérez-Cotapos, M.L.S.; Vergara, R.L.; Manca, M.; Tohme, R.A.; Holmberg, S.D.; Bressmann, T.; Lirio, D.R.; et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and cufflinks. Nat. Protoc. 2012, 7, 562. [Google Scholar] [CrossRef]
- Liu, H.; Naismith, J.H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 2008, 8, 91. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malhotra, S.; Freeberg, M.A.; Winans, S.J.; Taylor, J.; Beemon, K.L. A Novel Long Non-Coding RNA in the hTERT Promoter Region Regulates hTERT Expression. Non-Coding RNA 2018, 4, 1. https://doi.org/10.3390/ncrna4010001
Malhotra S, Freeberg MA, Winans SJ, Taylor J, Beemon KL. A Novel Long Non-Coding RNA in the hTERT Promoter Region Regulates hTERT Expression. Non-Coding RNA. 2018; 4(1):1. https://doi.org/10.3390/ncrna4010001
Chicago/Turabian StyleMalhotra, Sanandan, Mallory A. Freeberg, Shelby J. Winans, James Taylor, and Karen L. Beemon. 2018. "A Novel Long Non-Coding RNA in the hTERT Promoter Region Regulates hTERT Expression" Non-Coding RNA 4, no. 1: 1. https://doi.org/10.3390/ncrna4010001
APA StyleMalhotra, S., Freeberg, M. A., Winans, S. J., Taylor, J., & Beemon, K. L. (2018). A Novel Long Non-Coding RNA in the hTERT Promoter Region Regulates hTERT Expression. Non-Coding RNA, 4(1), 1. https://doi.org/10.3390/ncrna4010001




