The Evolution of the Satratoxin and Atranone Gene Clusters of Stachybotrys chartarum
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Cultures and Culture Conditions
2.2. Sequence Analysis
2.3. PCR
2.4. Deep Re-Sequencing Analysis
2.5. Mass Spectrometry (LC-MS/MS)
3. Results
3.1. LC-MS/MS Analysis
3.2. Deep Re-Sequencing Analysis
3.3. G+C Content and Predicted Proteins
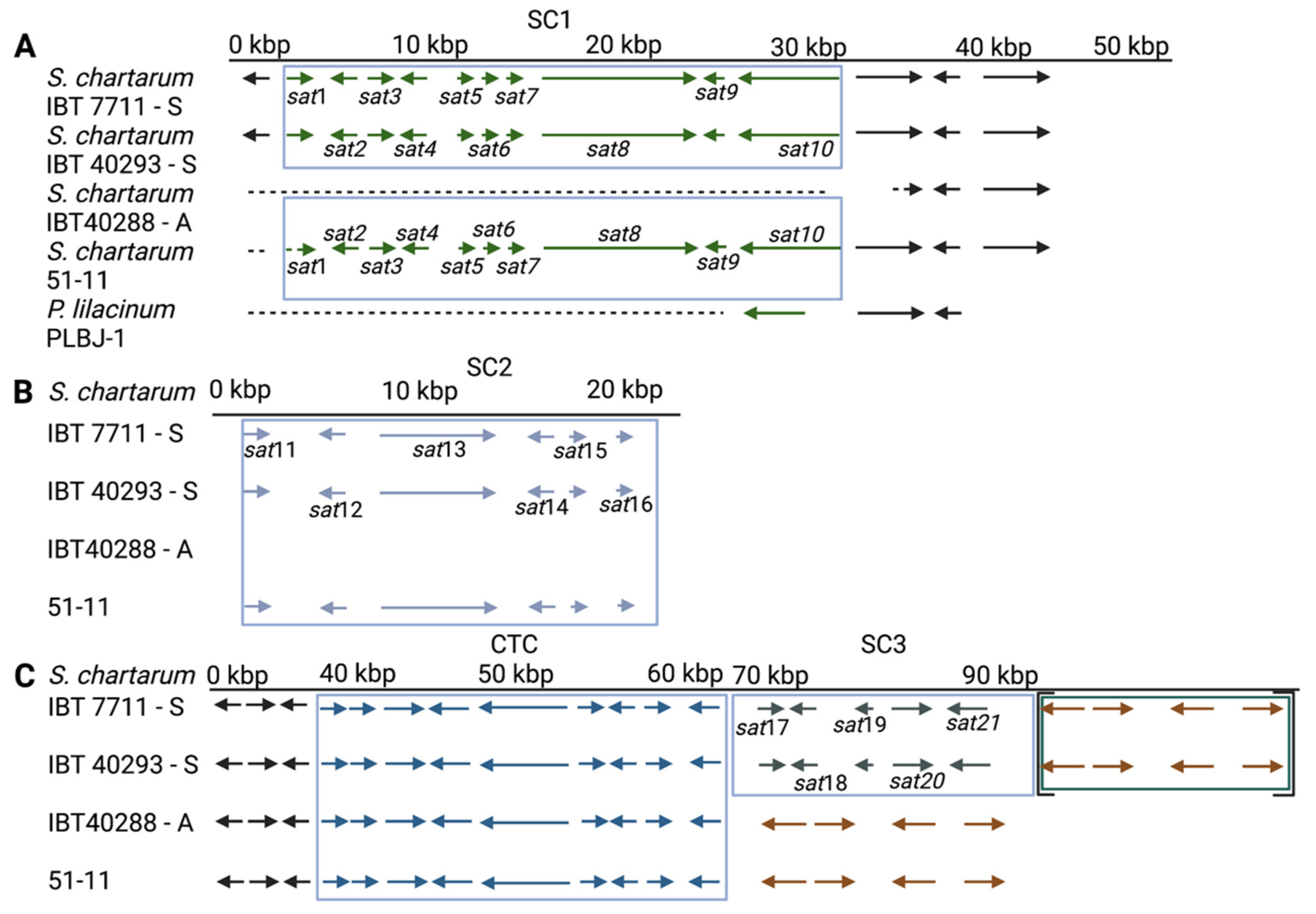
3.4. Satratoxin Cluster Alignment
3.4.1. Satratoxin Cluster 1 (SC1)
3.4.2. Satratoxin Cluster 2 (SC2)
3.4.3. The Satratoxin Cluster 3 (SC3)
3.5. Atranone Cluster 1 and 2
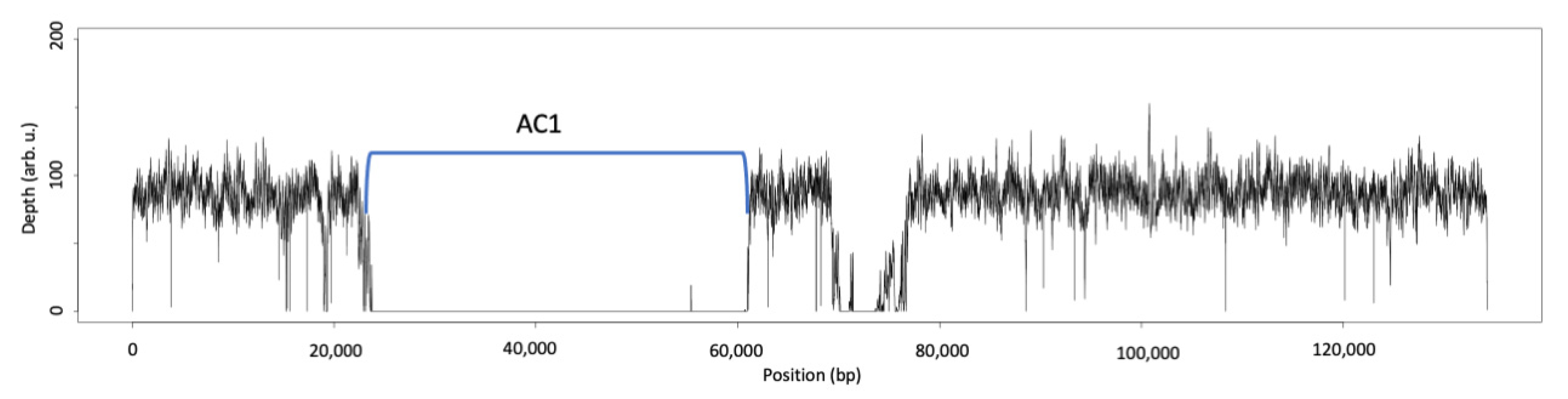
3.5.1. Atranone Cluster 1 (AC1)
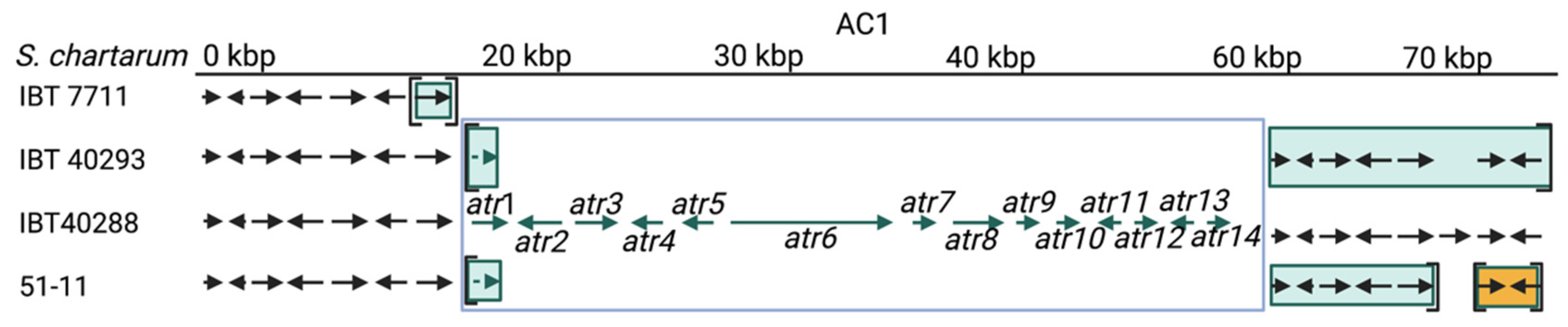
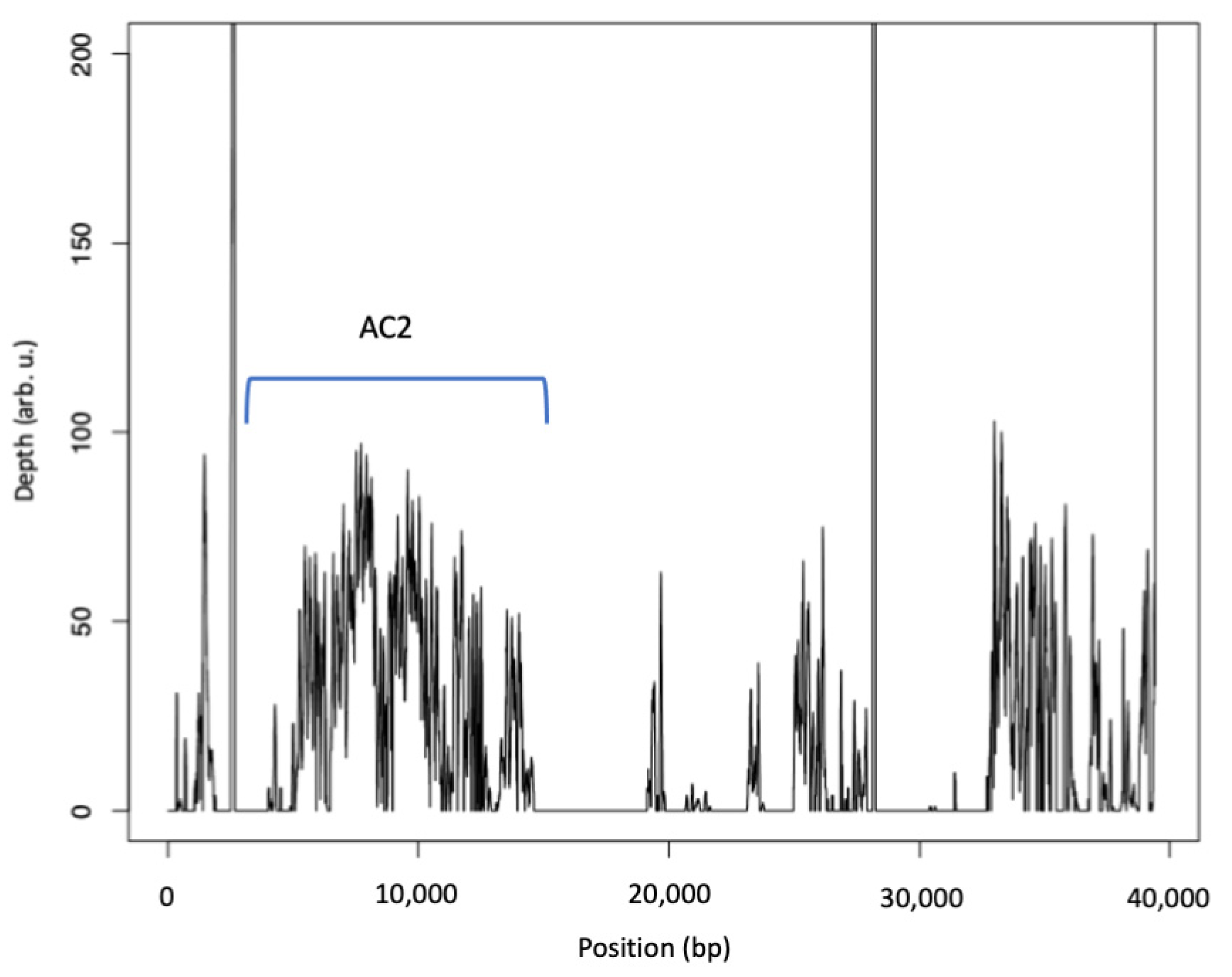
3.5.2. Atranone Cluster 2 (AC2)
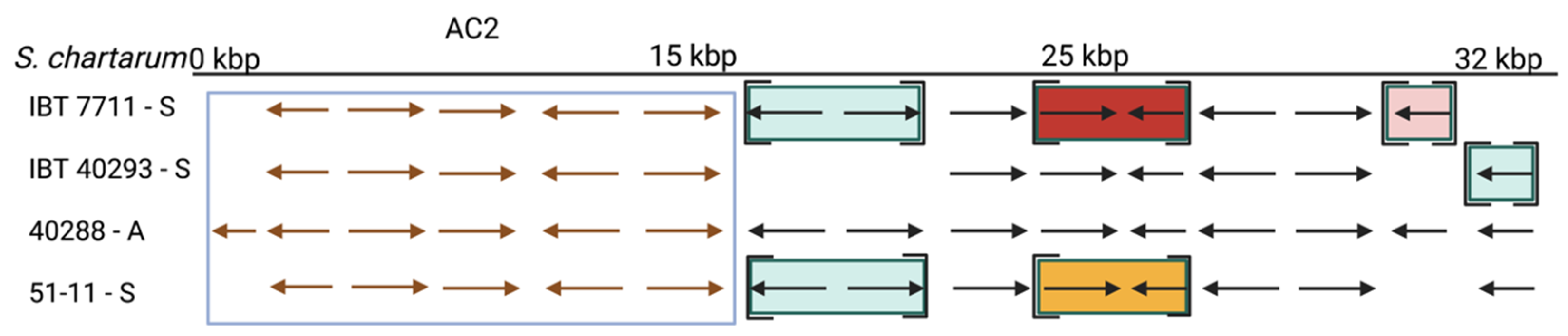
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lombard, L.; Houbraken, J.; Decock, C.; Samson, R.A.; Meijer, M.; Réblová, M.; Groenewald, J.Z.; Crous, P.W. Generic hyper-diversity in Stachybotriaceae. Pers. Mol. Phylogeny Evol. Fungi 2016, 36, 156–246. [Google Scholar] [CrossRef]
- Andersen, B.; Nielsen, K.F.; Jarvis, B.B. Characterization of Stachybotrys from water-damaged buildings based on morphology, growth, and metabolite production. Mycologia 2002, 94, 392–403. [Google Scholar] [CrossRef] [PubMed]
- Institute, W.F.B. Mycobank. Available online: https://www.mycobank.org/ (accessed on 30 January 2021).
- Crous, P.W.; Gams, W.; Stalpers, J.A.; Robert, V.; Stegehuis, G. MycoBank: An online initiative to launch mycology into the 21st century. Stud. Mycol. 2004, 50, 19–22. [Google Scholar]
- Nielsen, K.F.; Huttunen, K.; Hyvarinen, A.; Andersen, B.; Jarvis, B.B.; Hirvonen, M.R. Metabolite profiles of Stachybotrys isolates from water-damaged buildings and their induction of inflammatory mediators and cytotoxicity in macrophages. Mycopathologia 2002, 154, 201–205. [Google Scholar] [CrossRef]
- Wang, Y.; Hyde, K.D.; McKenzie, E.H.C.; Jiang, Y.-L.; Li, D.-W.; Zhao, D.-G. Overview of Stachybotrys (Memnoniella) and current species status. Fungal Divers. 2015, 71, 17–83. [Google Scholar] [CrossRef]
- Biermaier, B.; Gottschalk, C.; Schwaiger, K.; Gareis, M. Occurrence of Stachybotrys chartarum chemotype S in dried culinary herbs. Mycotoxin Res. 2015, 31, 23–32. [Google Scholar] [CrossRef]
- El-Kady, I.A.; Moubasher, M.H. Toxigenicity and toxins of Stachybotrys isolates from wheat straw samples in Egypt. Exp. Mycol. 1982, 6, 25–30. [Google Scholar] [CrossRef]
- Ulrich, S.; Gottschalk, C.; Biermaier, B.; Bahlinger, E.; Twarużek, M.; Asmussen, S.; Schollenberger, M.; Valenta, H.; Ebel, F.; Dänicke, S. Occurrence of type A, B and D trichothecenes, zearalenone and stachybotrylactam in straw. Arch. Anim. Nutr. 2021, 75, 105–120. [Google Scholar] [CrossRef]
- Andersen, B.; Nielsen, K.F.; Thrane, U.; Szaro, T.; Taylor, J.W.; Jarvis, B.B. Molecular and phenotypic descriptions of Stachybotrys chlorohalonata sp. nov. and two chemotypes of Stachybotrys chartarum found in water-damaged buildings. Mycologia 2003, 95, 1227–1258. [Google Scholar] [CrossRef]
- Ulrich, S.; Niessen, L.; Ekruth, J.; Schäfer, C.; Kaltner, F.; Gottschalk, C. Truncated satratoxin gene clusters in selected isolates of the atranone chemotype of Stachybotrys chartarum (Ehrenb.) S. Hughes. Mycotoxin Res. 2020, 36, 83–91. [Google Scholar] [CrossRef]
- Jarvis, B.B.; Sorenson, W.G.; Hintikka, E.L.; Nikulin, M.; Zhou, Y.; Jiang, J.; Wang, S.; Hinkley, S.; Etzel, R.A.; Dearborn, D. Study of toxin production by isolates of Stachybotrys chartarum and Memnoniella echinata isolated during a study of pulmonary hemosiderosis in infants. Appl. Environ. Microbiol. 1998, 64, 3620–3625. [Google Scholar] [CrossRef] [PubMed]
- Hinkley, S.F.; Moore, J.A.; Squillari, J.; Tak, H.; Oleszewski, R.; Mazzola, E.P.; Jarvis, B.B. New atranones from the fungus Stachybotrys chartarum. Magn. Reson. Chem. 2003, 41, 337–343. [Google Scholar] [CrossRef]
- Hinkley, S.F.; Jiang, J.; Mazzola, E.P.; Jarvis, B.B. Atranones: Novel diterpenoids from the toxigenic mold Stachybotrys atra. Tetrahedron Lett. 1999, 40, 2725–2728. [Google Scholar] [CrossRef]
- Hanelt, M.; Gareis, M.; Kollarczik, B. Cytotoxicity of mycotoxins evaluated by the MTT-cell culture assay. Mycopathologia 1994, 128, 167–174. [Google Scholar] [CrossRef]
- Hernández, F.; Cannon, M. Inhibition of protein synthesis in Saccharomyces cerevisiae by the 12,13-epoxytrichothecenes trichodermol, diacetoxyscirpenol and verrucarin A. Reversibility of the effects. J. Antibiot. 1982, 35, 875–881. [Google Scholar] [CrossRef]
- Rocha, O.; Ansari, K.; Doohan, F.M. Effects of trichothecene mycotoxins on eukaryotic cells: A review. Food Addit. Contam. 2005, 22, 369–378. [Google Scholar] [CrossRef]
- Ueno, Y. Trichothecenes-Chemical, Biological and Toxicological Aspects; Elsevier Science Publisher B.V.: Amsterdam, The Netherlands, 1983. [Google Scholar]
- Karunasena, E.; Larranaga, M.D.; Simoni, J.S.; Douglas, D.R.; Straus, D.C. Building-associated neurological damage modeled in human cells: A mechanism of neurotoxic effects by exposure to mycotoxins in the indoor environment. Mycopathologia 2010, 170, 377–390. [Google Scholar] [CrossRef]
- Dearborn, D.G.; Smith, P.G.; Dahms, B.B.; Allan, T.M.; Sorenson, W.G.; Montana, E.; Etzel, R.A. Clinical profile of 30 infants with acute pulmonary hemorrhage in Cleveland. Pediatrics 2002, 110, 627–637. [Google Scholar] [CrossRef]
- Miller, J.D.; Rand, T.G.; Jarvis, B.B. Stachybotrys chartarum: Cause of human disease or media darling? Med. Mycol. 2003, 41, 271–291. [Google Scholar] [CrossRef]
- CDC. Pulmonary hemorrhage/hemosiderosis among infants Cleveland, Ohio, 1993/1996. Morb. Mortal. Wkly. Rep. 2002, 49, 180–184. [Google Scholar]
- Johanning, E.; Biagini, R.; Hull, D.; Morey, P.; Jarvis, B.; Landsbergis, P. Health and immunology study following exposure to toxigenic fungi (Stachybotrys chartarum) in a water-damaged office environment. Int. Arch. Occup. Environ. Health 1996, 68, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Hintikka, E.-L. The Role of Stachybotrys in the Phenomenon Known as Sick Building Syndrome. In Advances in Applied Microbiology; Academic Press: Cambridge, MA, USA, 2004; Volume 55, pp. 155–173. [Google Scholar]
- Nikulin, M.; Reijula, K.; Jarvis, B.B.; Hintikka, E.L. Experimental lung mycotoxicosis in mice induced by Stachybotrys atra. Int. J. Exp. Pathol. 1996, 77, 213–218. [Google Scholar] [CrossRef] [PubMed]
- Vesper, S.; Dearborn, D.G.; Yike, I.; Allan, T.; Sobolewski, J.; Hinkley, S.F.; Jarvis, B.B.; Haugland, R.A. Evaluation of Stachybotrys chartarum in the house of an infant with pulmonary hemorrhage: Quantitative assessment before, during, and after remediation. J. Urban Health 2000, 77, 68–85. [Google Scholar] [CrossRef] [PubMed]
- Forgacs, J.; Carll, W.T.; Herring, A.S.; Hinshaw, W.R. Toxicity of Stachybotrys atra for animals. Trans. N. Y. Acad. Sci. 1958, 20, 787–808. [Google Scholar] [CrossRef] [PubMed]
- Schneider, D.J.; Marasas, W.F.; Dale Kuys, J.C.; Kriek, N.P.; Van Schalkwyk, G.C. A field outbreak of suspected stachybotryotoxicosis in sheep. J. S. Afr. Vet. Assoc. 1979, 50, 73–81. [Google Scholar] [PubMed]
- Kriek, N.P.J.; Marasas, W.F.O. Field outbreak of ovine stachybotryotoxicosis in South Africa. In Thrichothecenes-Chemical, Biological and Toxicological Aspects; Ueno, Y., Ed.; Elsevier Science Publisher B.V.: Amsterdam, The Netherlands, 1983; pp. 279–284. [Google Scholar]
- Semis, M.; Dadwal, S.S.; Tegtmeier, B.R.; Wilczynski, S.P.; Ito, J.I.; Kalkum, M. First Case of Invasive Stachybotrys Sinusitis. Clin. Infect. Dis. 2020, 72, 1386–1391. [Google Scholar] [CrossRef]
- Vesper, S.J.; Magnuson, M.L.; Dearborn, D.G.; Yike, I.; Haugland, R.A. Initial characterization of the hemolysin stachylysin from Stachybotrys chartarum. Infect. Immun. 2001, 69, 912–916. [Google Scholar] [CrossRef]
- Johanning, E.; Landsbergis, P.; Gareis, M.; Yang, C.S.; Olmsted, E. Clinical experience and results of a Sentinel Health Investigation related to indoor fungal exposure. Environ. Health Perspect. 1999, 107, 489–494. [Google Scholar] [CrossRef]
- Rand, T.G.; Flemming, J.; David Miller, J.; Womiloju, T.O. Comparison of inflammatory responses in mouse lungs exposed to atranones A and C from Stachybotrys chartarum. J. Toxicol. Environ. Health 2006, 69, 1239–1251. [Google Scholar] [CrossRef]
- Semeiks, J.; Borek, D.; Otwinowski, Z.; Grishin, N.V. Comparative genome sequencing reveals chemotype-specific gene clusters in the toxigenic black mold Stachybotrys. BMC Genom. 2014, 15, 590. [Google Scholar] [CrossRef]
- Niessen, L.; Vogel, R.F. Detection of Fusarium graminearum DNA using a loop-mediated isothermal amplification (LAMP) assay. Int. J. Food Microbiol. 2010, 140, 183–191. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Ulrich, S.; Schäfer, C. Toxin Production by Stachybotrys chartarum Genotype S on Different Culture Media. J. Fungi 2020, 6, 159. [Google Scholar]
- Vesper, S.J.; Dearborn, D.G.; Yike, I.; Sorenson, W.G.; Haugland, R.A. Hemolysis, toxicity, and randomly amplified polymorphic DNA analysis of Stachybotrys chartarum strains. Appl. Environ. Microbiol. 1999, 65, 3175–3181. [Google Scholar] [CrossRef]
- Vining, L.C. Secondary Metabolism; VCH: Weinheim, Germany, 1986. [Google Scholar]
- Medema, M.H.; Kottmann, R.; Yilmaz, P.; Cummings, M.; Biggins, J.B.; Blin, K.; de Bruijn, I.; Chooi, Y.H.; Claesen, J.; Coates, R.C.; et al. Minimum Information about a Biosynthetic Gene cluster. Nat. Chem. Biol. 2015, 11, 625–631. [Google Scholar] [CrossRef]
- Degenkolb, T.; Dieckmann, R.; Nielsen, K.F.; Gräfenhan, T.; Theis, C.; Zafari, D.; Chaverri, P.; Ismaiel, A.; Brückner, H.; von Döhren, H.; et al. The Trichoderma brevicompactum clade: A separate lineage with new species, new peptaibiotics, and mycotoxins. Mycol. Prog. 2008, 7, 177–219. [Google Scholar] [CrossRef]
- Rokas, A.; Mead, M.E.; Steenwyk, J.L.; Raja, H.A.; Oberlies, N.H. Biosynthetic gene clusters and the evolution of fungal chemodiversity. Nat. Prod. Rep. 2020, 37, 868–878. [Google Scholar] [CrossRef]
- Palmer, M.A.; Menninger, H.L.; Bernhardt, E. River restoration, habitat heterogeneity and biodiversity: A failure of theory or practice? Freshw. Biol. 2010, 55, 205–222. [Google Scholar] [CrossRef]
- Hoogendoorn, K.; Barra, L.; Waalwijk, C.; Dickschat, J.S.; van der Lee, T.A.J.; Medema, M.H. Evolution and Diversity of Biosynthetic Gene Clusters in Fusarium. Front. Microbiol. 2018, 9, 1158. [Google Scholar] [CrossRef]
- Alexander, W.G.; Wisecaver, J.H.; Rokas, A.; Hittinger, C.T. Horizontally acquired genes in early-diverging pathogenic fungi enable the use of host nucleosides and nucleotides. Proc. Natl. Acad. Sci. USA 2016, 113, 4116. [Google Scholar] [CrossRef] [PubMed]
- Kock, J.; Gottschalk, C.; Ulrich, S.; Schwaiger, K.; Gareis, M.; Niessen, L. Rapid and selective detection of macrocyclic trichothecene producing Stachybotrys chartarum strains by loop-mediated isothermal amplification (LAMP). Anal. Bioanal. Chem. 2021, 413, 4801–4813. [Google Scholar] [CrossRef] [PubMed]
- Martyn, R.D. Monosporascus Root Rot and Vine Decline of Melons (MRR/VD). Also referred to as sudden wilt, sudden death, melon collapse, Monosporascus wilt, and black pepper root rot. Plant Health Instr. 2002, 10, 0612-01. [Google Scholar] [CrossRef]
- Zhu, M.; Cen, Y.; Ye, W.; Li, S.; Zhang, W. Recent Advances on Macrocyclic Trichothecenes, Their Bioactivities and Biosynthetic Pathway. Toxins 2020, 12, 417. [Google Scholar] [CrossRef]
- Sidhu, G.S. Mycotoxin Genetics and Gene Clusters. Eur. J. Plant Pathol. 2002, 108, 705–711. [Google Scholar] [CrossRef]
- Alexander, N.J.; Proctor, R.H.; McCormick, S.P. Genes, gene clusters, and biosynthesis of trichothecenes and fumonisins in Fusarium. Toxin Rev. 2009, 28, 198–215. [Google Scholar] [CrossRef]
- Proctor, R.H.; McCormick, S.P.; Alexander, N.J.; Desjardins, A.E. Evidence that a secondary metabolic biosynthetic gene cluster has grown by gene relocation during evolution of the filamentous fungus Fusarium. Mol. Microbiol. 2009, 74, 1128–1142. [Google Scholar] [CrossRef]
- Rokas, A.; Wisecaver, J.H.; Lind, A.L. The birth, evolution and death of metabolic gene clusters in fungi. Nat. Rev. Microbiol. 2018, 16, 731–744. [Google Scholar] [CrossRef]
- Coleman, J.J.; Mylonakis, E. Efflux in fungi: La piece de resistance. PLoS Pathog. 2009, 5, e1000486. [Google Scholar] [CrossRef]
- Slot, J.C.; Rokas, A. Horizontal transfer of a large and highly toxic secondary metabolic gene cluster between fungi. Curr. Biol. 2011, 21, 134–139. [Google Scholar] [CrossRef]
- Shen, L.; Porée, F.H.; Gaslonde, T.; Lalucque, H.; Chapeland-Leclerc, F.; Ruprich-Robert, G. Functional characterization of the sterigmatocystin secondary metabolite gene cluster in the filamentous fungus Podospora anserina: Involvement in oxidative stress response, sexual development, pigmentation and interspecific competitions. Environ. Microbiol. 2019, 21, 3011–3026. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.F.; Xiao, J.H.; He, S.; Liu, L.; Murphy, R.W.; Huang, D.W. Multiple interkingdom horizontal gene transfers in Pyrenophora and closely related species and their contributions to phytopathogenic lifestyles. PLoS ONE 2013, 8, e60029. [Google Scholar] [CrossRef]
- Fitzpatrick, D.A. Horizontal gene transfer in fungi. FEMS Microbiol. Lett. 2012, 329, 1–8. [Google Scholar] [CrossRef] [PubMed]
| Compound | Q1 (m/z) a | Q3 (m/z) a | Retention Time (min) b | DP (V) | CE (V) | CXP (V) |
|---|---|---|---|---|---|---|
| 6-hydroxydolabella-3,7,12-trien-14-one | 303.3 | 205.1 | 12.0 | 86 | 15 | 12 |
| 303.3 | 243.0 | 12.0 | 86 | 11 | 14 | |
| 3,4-epoxy 6-hydroxy-dolabella-7,12-diene-one | 319.2 | 221.1 | 10.5 | 101 | 11 | 12 |
| 319.2 | 259.0 | 10.5 | 101 | 9 | 14 | |
| atranone A | 417.3 | 205.2 | 11.4 | 200 | 33 | 12 |
| 417.3 | 357.2 | 11.4 | 200 | 15 | 11 |
| Sample Name | 6-hydroxydolabella-3,7,12-trien-14-one | 3,4-epoxy-6-hydroxy-dolabella-7,12-diene-one | Atranone A | |||
|---|---|---|---|---|---|---|
| Quantifier | Qualifier | Quantifier | Qualifier | Quantifier | Qualifier | |
| 303.2/205.1 Da | 303.2/243.0 Da | 319.2/221.1 Da | 319.2/259.0 Da | 417.3/357.2 Da | 417.3/205.2 Da | |
| IBT 40288 | 230,000 * | 54,900 * | 349,000 * | 107,000 * | 491,000 * | 379,000 * |
| IBT 40293 | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
| A: Satratoxin Clusters | |||||||||
| Strain ID | SC1 scaffolds | SC2 scaffolds | SC3 scaffolds | ||||||
| Upstream | SC1 | Downstream | Upstream | SC2 | Downstream | Upstream | SC3 | Downstream | |
| 51-11 | 451638bp | 5329,998bp | 48140,301bp | 272133bp | 4719,876bp | 27140,301bp | 51849,200bp | - - - | - - - |
| IBT 40293 | 444155bp | 5328,909bp | 5110,613bp | 42886bp | 4719,873bp | 4410,613bp | 51105,159bp | 5310,500bp | 48112bp/42,552bp ** |
| IBT 7711 | 454564bp | 5329,996 | 5072,574bp | 45772bp | 4719,854bp | 4372,574bp | 5195,549bp | 5410,496bp | 33122bp |
| IBT 40288 | - - - | - - - | 5296,283bp | - - - | - - - | - - - | 51129,561bp | - - - | - - - |
| B: Atranone Clusters | |||||||||
| Strain ID | AC1 scaffolds | AC2 scaffolds | |||||||
| Upstream | AC1 | Downstream | AC2 | Downstream | |||||
| 51-11 | 4685,474bp | - - - | 3921,564bp/849,200bp ** | 5310,350bp | 53n.d. * | ||||
| IBT 40293 | 4942,470bp | - - - | 4954,221bp | 579340bp | 54294,349bp * | ||||
| IBT 7711 | 5033,211bp/12,367bp | - - - | - - - | 5310,380bp | 5364,167bp * | ||||
| IBT 40288 | 4524,509bp | 4935,610bp | 4874,142bp | 5111,699bp | 5124,996bp | ||||
| Strain ID | Upstream of SC1 | SC1 Genes (sat1-10) | Downstream of SC1 | |||
|---|---|---|---|---|---|---|
| Identity (%) | Gaps (%) | Identity (%) | Gaps (%) | Identity (%) | Gaps (%) | |
| 51-11 vs. IBT 40293 | 46.5 | 37.6 | 95.7 | 4.1 | 98.7 | 0.6 |
| 51-11 vs. IBT 7711 | 45.4 | 40.0 | 95.7 | 4.1 | 98.5 | 0.6 |
| IBT 40293 vs. IBT 7711 | 85.7 | 11.7 | 99.8 | 0.0 | 99.3 | 0.1 |
| Strain ID | Upstream of SC2 | SC2 Genes (sat11-16) | Downstream of SC2 | |||
|---|---|---|---|---|---|---|
| Identity (%) | Gaps (%) | Identity (%) | Gaps (%) | Identity (%) | Gaps (%) | |
| 51-11 vs. IBT 40293 | 99.0 | 0.3 | 99.5 | 0.1 | 98.5 | 0.2 |
| 51-11 vs. IBT 7711 | 99.0 | 0.6 | 99.5 | 0.1 | 98.6 | 0.1 |
| IBT 40293 vs. IBT 7711 | 99.1 | 0.8 | 99.6 | 0.1 | 98.8 | 0.2 |
| Strain ID | AC2 | Downstream of AC2 | ||
|---|---|---|---|---|
| Mean Identity (%) | Mean Coverage (%) | Mean Identity (%) | Mean Cover (%) | |
| 51-11 | 88.4 | 100 | 74.5 | 86.0 |
| IBT 40293 | 89.2 | 98.8 | 54.1 | 64.4 |
| IBT 7711 | 88.7 | 100 | 73.5 | 84.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ulrich, S.; Lang, K.; Niessen, L.; Baschien, C.; Kosicki, R.; Twarużek, M.; Straubinger, R.K.; Ebel, F. The Evolution of the Satratoxin and Atranone Gene Clusters of Stachybotrys chartarum. J. Fungi 2022, 8, 340. https://doi.org/10.3390/jof8040340
Ulrich S, Lang K, Niessen L, Baschien C, Kosicki R, Twarużek M, Straubinger RK, Ebel F. The Evolution of the Satratoxin and Atranone Gene Clusters of Stachybotrys chartarum. Journal of Fungi. 2022; 8(4):340. https://doi.org/10.3390/jof8040340
Chicago/Turabian StyleUlrich, Sebastian, Katharina Lang, Ludwig Niessen, Christiane Baschien, Robert Kosicki, Magdalena Twarużek, Reinhard K. Straubinger, and Frank Ebel. 2022. "The Evolution of the Satratoxin and Atranone Gene Clusters of Stachybotrys chartarum" Journal of Fungi 8, no. 4: 340. https://doi.org/10.3390/jof8040340
APA StyleUlrich, S., Lang, K., Niessen, L., Baschien, C., Kosicki, R., Twarużek, M., Straubinger, R. K., & Ebel, F. (2022). The Evolution of the Satratoxin and Atranone Gene Clusters of Stachybotrys chartarum. Journal of Fungi, 8(4), 340. https://doi.org/10.3390/jof8040340








