Principal Drivers of Fungal Communities Associated with Needles, Shoots, Roots and Adjacent Soil of Pinus sylvestris
Abstract
1. Introduction
2. Materials and Methods
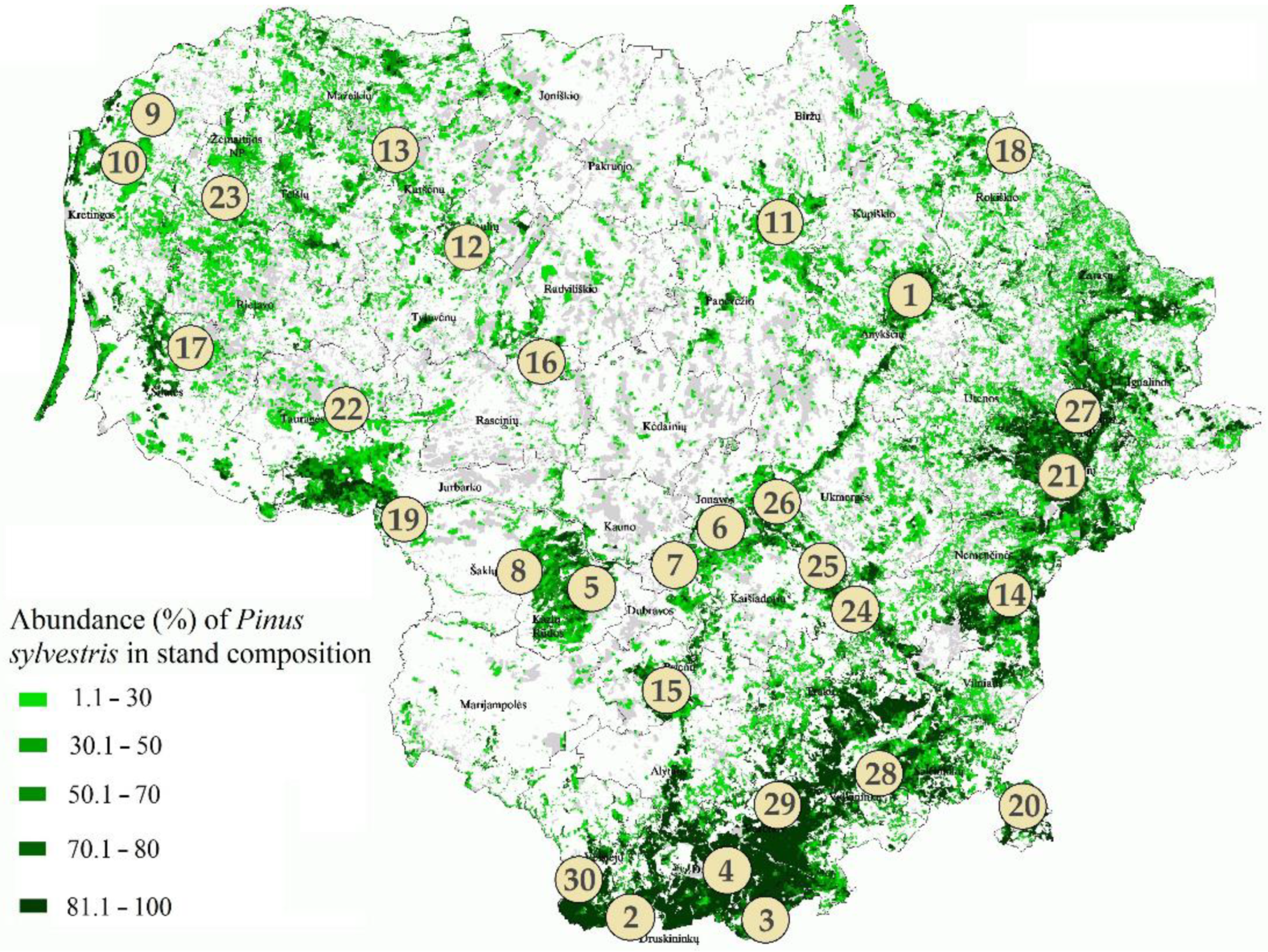
2.1. Study Sites and Sampling
2.2. DNA Work
2.3. Bioinformatics
2.4. Statistical Analyses
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Oleksyn, J.; Wyka, T.P.; Żytkowiak, R.; Zadworny, M.; Mucha, J.; Dering, M.; Ufnalski, K.; Nihlgård, B.; Reich, P.B. A fingerprint of climate change across pine forests of Sweden. Ecol. Lett. 2020, 23, 1739–1746. [Google Scholar] [CrossRef]
- Cavicchioli, R.; Ripple, W.J.; Timmis, K.N.; Azam, F.; Bakken, L.R.; Baylis, M.; Behrenfeld, M.J.; Boetius, A.; Boyd, P.W.; Classen, A.T.; et al. Scientists’ warning to humanity: Microorganisms and climate change. Nat. Rev. Microbiol. 2019, 17, 569–586. [Google Scholar] [CrossRef]
- Singh, D.P.; Prabha, R. Microbial Interventions in Agriculture and Environment: Volume 3: Soil and Crop Health Management; Springer: Berlin/Heidelberg, Germany, 2019; pp. 1–491. [Google Scholar] [CrossRef]
- de Habiyaremye, J.D.; Goldmann, K.; Reitz, T.; Herrmann, S.; Buscot, F. Tree Root Zone Microbiome: Exploring the Magnitude of Environmental Conditions and Host Tree Impact. Front. Microbiol. 2020, 11, 749. [Google Scholar] [CrossRef]
- Barnes, E.M.; Tringe, S.G. Exploring the roles of microbes in facilitating plant adaptation to climate change. Biochem. J. 2022, 479, 327–335. [Google Scholar] [CrossRef]
- Hardoim, P.R.; van Overbeek, L.S.; Berg, G.; Pirttilä, A.M.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The Hidden World within Plants: Ecological and Evolutionary Considerations for Defining Functioning of Microbial Endophytes. Microbiol. Mol. Biol. Rev. 2015, 79, 293–320. [Google Scholar] [CrossRef]
- Terhonen, E.; Blumenstein, K.; Kovalchuk, A.; Asiegbu, F.O. Forest tree microbiomes and associated fungal endophytes: Functional roles and impact on forest health. Forests 2019, 10, 42. [Google Scholar] [CrossRef]
- Kasanen, R. Abiotic factors affecting the composition of forest tree microbiomes. In Forest Microbiology; Academic Press: Cambridge, MA, USA, 2021; pp. 59–73. [Google Scholar]
- Wu, Y.; Qu, M.; Pu, X.; Lin, J.; Shu, B. Distinct microbial communities among different tissues of citrus tree Citrus reticulata cv. Chachiensis. Sci. Rep. 2020, 10, 6068. [Google Scholar] [CrossRef]
- Turner, T.R.; James, E.K.; Poole, P.S. The plant microbiome. Genome Biol. 2013, 14, 209. [Google Scholar] [CrossRef]
- Trivedi, P.; Leach, J.E.; Tringe, S.G.; Sa, T.; Singh, B.K. Plant–microbiome interactions: From community assembly to plant health. Nat. Rev. Microbiol. 2020, 18, 607–621. [Google Scholar] [CrossRef]
- Faticov, M.; Abdelfattah, A.; Roslin, T.; Vacher, C.; Hambäck, P.; Blanchet, F.G.; Lindahl, B.D.; Tack, A.J.M. Climate warming dominates over plant genotype in shaping the seasonal trajectory of foliar fungal communities on oak. New Phytol. 2021, 231, 1770–1783. [Google Scholar] [CrossRef]
- Lau, J.A.; Lennon, J.T.; Heath, K.D. Trees harness the power of microbes to survive climate change. Proc. Natl. Acad. Sci. USA 2017, 114, 11009–11011. [Google Scholar] [CrossRef]
- Bennett, A.E.; Classen, A.T. Climate change influences mycorrhizal fungal–plant interactions, but conclusions are limited by geographical study bias. Ecology 2020, 101, e02978. [Google Scholar] [CrossRef]
- Bender, S.F.; Wagg, C.; van der Heijden, M.G.A. An Underground Revolution: Biodiversity and Soil Ecological Engineering for Agricultural Sustainability. Trends Ecol. Evol. 2016, 31, 440–452. [Google Scholar] [CrossRef]
- Frac, M.; Hannula, S.E.; Belka, M.; Jȩdryczka, M. Fungal biodiversity and their role in soil health. Front. Microbiol. 2018, 9, 707. [Google Scholar] [CrossRef]
- Mercado-Blanco, J.; Abrantes, I.; Caracciolo, A.B.; Bevivino, A.; Ciancio, A.; Grenni, P.; Hrynkiewicz, K.; Kredics, L.; Proença, D.N. Belowground microbiota and the health of tree crops. Front. Microbiol. 2018, 9, 1006. [Google Scholar] [CrossRef]
- Zhou, Z.; Wang, C.; Luo, Y. Effects of forest degradation on microbial communities and soil carbon cycling: A global meta-analysis. Glob. Ecol. Biogeogr. 2018, 27, 110–124. [Google Scholar] [CrossRef]
- Rousk, J.; Brookes, P.C.; Bååth, E. Contrasting soil pH effects on fungal and bacterial growth suggest functional redundancy in carbon mineralization. Appl. Environ. Microbiol. 2009, 75, 1589–1596. [Google Scholar] [CrossRef]
- van der Heijden, M.G.A.; Martin, F.M.; Selosse, M.A.; Sanders, I.R. Mycorrhizal ecology and evolution: The past, the present, and the future. New Phytol. 2015, 205, 1406–1423. [Google Scholar] [CrossRef]
- Saravesi, K.; Aikio, S.; Wäli, P.R.; Ruotsalainen, A.L.; Kaukonen, M.; Huusko, K.; Suokas, M.; Brown, S.P.; Jumpponen, A.; Tuomi, J.; et al. Moth Outbreaks Alter Root-Associated Fungal Communities in Subarctic Mountain Birch Forests. Microb. Ecol. 2015, 69, 788–797. [Google Scholar] [CrossRef]
- Hartmann, M.; Niklaus, P.A.; Zimmermann, S.; Schmutz, S.; Kremer, J.; Abarenkov, K.; Lüscher, P.; Widmer, F.; Frey, B. Resistance and resilience of the forest soil microbiome to logging-associated compaction. ISME J. 2014, 8, 226–244. [Google Scholar] [CrossRef]
- Treu, R.; Karst, J.; Randall, M.; Pec, G.J.; Cigan, P.W.; Simard, S.W.; Cooke, J.E.K.; Erbilgin, N.; Cahill, J.F. Decline of ectomycorrhizal fungi following a mountain pine beetle epidemic. Ecology 2014, 95, 1096–1103. [Google Scholar] [CrossRef] [PubMed]
- Karst, J.; Erbilgin, N.; Pec, G.J.; Cigan, P.W.; Najar, A.; Simard, S.W.; Cahill, J.F. Ectomycorrhizal fungi mediate indirect effects of a bark beetle outbreak on secondary chemistry and establishment of pine seedlings. New Phytol. 2015, 208, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Bödeker, I.T.M.; Lindahl, B.D.; Olson, Å.; Clemmensen, K.E. Mycorrhizal and saprotrophic fungal guilds compete for the same organic substrates but affect decomposition differently. Funct. Ecol. 2016, 30, 1967–1978. [Google Scholar] [CrossRef]
- Crowther, T.W.; van den Hoogen, J.; Wan, J.; Mayes, M.A.; Keiser, A.D.; Mo, L.; Averill, C.; Maynard, D.S. The global soil community and its influence on biogeochemistry. Science 2019, 365, eaav0550. [Google Scholar] [CrossRef]
- Usman, M.; Ho-Plágaro, T.; Frank, H.E.R.; Calvo-Polanco, M.; Gaillard, I.; Garcia, K.; Zimmermann, S.D. Mycorrhizal Symbiosis for Better Adaptation of Trees to Abiotic Stress Caused by Climate Change in Temperate and Boreal Forests. Front. For. Glob. Chang. 2021, 4, 141. [Google Scholar] [CrossRef]
- Steidinger, B.S.; Crowther, T.W.; Liang, J.; Van Nuland, M.E.; Werner, G.D.A.; Reich, P.B.; Nabuurs, G.; de-Miguel, S.; Zhou, M.; Picard, N.; et al. Climatic controls of decomposition drive the global biogeography of forest-tree symbioses. Nature 2019, 569, 404–408. [Google Scholar] [CrossRef]
- Park, K.H.; Oh, S.Y.; Yoo, S.; Park, M.S.; Fong, J.J.; Lim, Y.W. Successional Change of the Fungal Microbiome Pine Seedling Roots Inoculated With Tricholoma matsutake. Front. Microbiol. 2020, 11, 574146. [Google Scholar] [CrossRef]
- Lebeis, S.L. Greater than the sum of their parts: Characterizing plant microbiomes at the community-level. Curr. Opin. Plant Biol. 2015, 24, 82–86. [Google Scholar] [CrossRef]
- Wallander, H.; Johansson, U.; Sterkenburg, E.; Brandström Durling, M.; Lindahl, B.D. Production of ectomycorrhizal mycelium peaks during canopy closure in Norway spruce forests. New Phytol. 2010, 187, 1124–1134. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Rott, M.; Schlaeppi, K.; Ver Loren van Themaat, E.; Ahmadinejad, N.; Assenza, F.; Rauf, P.; Huettel, B.; Reinhardt, R.; Schmelzer, E.; et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 2012, 488, 91–95. [Google Scholar] [CrossRef]
- Lundberg, D.S.; Lebeis, S.L.; Paredes, S.H.; Yourstone, S.; Gehring, J.; Malfatti, S.; Tremblay, J.; Engelbrektson, A.; Kunin, V.; del Rio, T.G.; et al. Defining the core Arabidopsis thaliana root microbiome. Nature 2012, 488, 86–90. [Google Scholar] [CrossRef] [PubMed]
- Kohout, P.; Sudová, R.; Brabcová, V.; Vosolsobě, S.; Baldrian, P.; Albrechtová, J. Forest Microhabitat Affects Succession of Fungal Communities on Decomposing Fine Tree Roots. Front. Microbiol. 2021, 12, 541583. [Google Scholar] [CrossRef] [PubMed]
- Lladó, S.; López-Mondéjar, R.; Baldrian, P. Drivers of microbial community structure in forest soils. Appl. Microbiol. Biotechnol. 2018, 102, 4331–4338. [Google Scholar] [CrossRef] [PubMed]
- Lareen, A.; Burton, F.; Schäfer, P. Plant root-microbe communication in shaping root microbiomes. Plant Mol. Biol. 2016, 90, 575–587. [Google Scholar] [CrossRef]
- Peay, K.G.; Kennedy, P.G.; Talbot, J.M. Dimensions of biodiversity in the Earth mycobiome. Nat. Rev. Microbiol. 2016, 14, 434–447. [Google Scholar] [CrossRef]
- Mäkipää, R.; Rajala, T.; Schigel, D.; Rinne, K.T.; Pennanen, T.; Abrego, N.; Ovaskainen, O. Interactions between soil- and dead wood-inhabiting fungal communities during the decay of Norway spruce logs. ISME J. 2017, 11, 1964–1974. [Google Scholar] [CrossRef]
- Bagchi, R.; Gallery, R.E.; Gripenberg, S.; Gurr, S.J.; Narayan, L.; Addis, C.E.; Freckleton, R.P.; Lewis, O.T. Pathogens and insect herbivores drive rainforest plant diversity and composition. Nature 2014, 506, 85–88. [Google Scholar] [CrossRef]
- Millberg, H.; Boberg, J.; Stenlid, J. Changes in fungal community of Scots pine (Pinus sylvestris) needles along a latitudinal gradient in Sweden. Fungal Ecol. 2015, 17, 126–139. [Google Scholar] [CrossRef]
- Hyde, K.D.; Soytong, K. Understanding microfungal diversity: A critique. Cryptogam. Mycol. 2007, 28, 281–289. [Google Scholar]
- Osorio, M.; Stephan, B.R. Life cycle of Lophodermium piceae in Norway spruce needles. Eur. J. For. Pathol. 1991, 21, 152–163. [Google Scholar] [CrossRef]
- Larkin, B.G.; Hunt, L.S.; Ramsey, P.W. Foliar nutrients shape fungal endophyte communities in Western white pine (Pinus monticola) with implications for white-tailed deer herbivory. Fungal Ecol. 2012, 5, 252–260. [Google Scholar] [CrossRef]
- Zimmerman, N.B.; Vitousek, P.M. Fungal endophyte communities reflect environmental structuring across a Hawaiian landscape. Proc. Natl. Acad. Sci. USA 2012, 109, 13022–13027. [Google Scholar] [CrossRef] [PubMed]
- Romeralo, C.; Martín-García, J.; Martínez-Álvarez, P.; Muñoz-Adalia, E.J.; Gonçalves, D.R.; Torres, E.; Witzell, J.; Diez, J.J. Pine species determine fungal microbiome composition in a common garden experiment. Fungal Ecol. 2022, 56, 101137. [Google Scholar] [CrossRef]
- Rabiey, M.; Hailey, L.E.; Roy, S.R.; Grenz, K.; Al-Zadjali, M.A.S.; Barrett, G.A.; Jackson, R.W. Endophytes vs. tree pathogens and pests: Can they be used as biological control agents to improve tree health? Eur. J. Plant Pathol. 2019, 155, 711–729. [Google Scholar] [CrossRef]
- Sieber, T.N. Endophytic fungi in forest trees: Are they mutualists? Fungal Biol. Rev. 2007, 21, 75–89. [Google Scholar] [CrossRef]
- Mengistu, A.A. Endophytes: Colonization, Behaviour, and Their Role in Defense Mechanism. Int. J. Microbiol. 2020, 2020, 6927219. [Google Scholar] [CrossRef]
- Rajala, T.; Peltoniemi, M.; Pennanen, T.; Mäkipää, R. Fungal community dynamics in relation to substrate quality of decaying Norway spruce (Picea abies [L.] Karst.) logs in boreal forests. FEMS Microbiol. Ecol. 2012, 81, 494–505. [Google Scholar] [CrossRef]
- Purahong, W.; Wubet, T.; Krüger, D.; Buscot, F. Molecular evidence strongly supports deadwood-inhabiting fungi exhibiting unexpected tree species preferences in temperate forests. ISME J. 2018, 12, 289–295. [Google Scholar] [CrossRef]
- Olsson, J. Colonization Patterns of Wood-Inhabiting Fungi in Boreal Forest; Ekologi, Miljö Och Geovetenskap: Umeå, Sweden, 2008; p. 35. [Google Scholar]
- Bahram, M.; Netherway, T. Fungi as mediators linking organisms and ecosystems. FEMS Microbiol. Rev. 2022, 46, fuab058. [Google Scholar] [CrossRef]
- Hallingbäck, H.R.; Burton, V.; Vizcaíno-Palomar, N.; Trotter, F.; Liziniewicz, M.; Marchi, M.; Berlin, M.; Ray, D.; Benito Garzón, M. Managing Uncertainty in Scots Pine Range-Wide Adaptation Under Climate Change. Front. Ecol. Evol. 2021, 9, 724051. [Google Scholar] [CrossRef]
- Reich, P.B.; Oleksyn, J. Climate warming will reduce growth and survival of Scots pine except in the far north. Ecol. Lett. 2008, 11, 588–597. [Google Scholar] [CrossRef] [PubMed]
- Vorontsov, A.I.; Mozolevskaya, E.G.; Sokolova, E.S. Tekhnologiya Zashchity Lesa—M; Ekologiya: Moscow, Russia, 1991. [Google Scholar]
- Ozolincius, R.; Stakenas, V. Tree crown defoliation: Influencing factors. Balt. For. 1996, 2, 48–55. [Google Scholar]
- Ihrmark, K.; Bödeker, I.T.M.; Cruz-Martinez, K.; Friberg, H.; Kubartova, A.; Schenck, J.; Strid, Y.; Stenlid, J.; Brandström-Durling, M.; Clemmensen, K.E.; et al. New primers to amplify the fungal ITS2 region—Evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 2012, 82, 666–677. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and Direct Sequencing of Fungal Ribosomal Rna Genes for Phylogenetics. PCR Protoc. 1990, 18, 315–322. [Google Scholar] [CrossRef]
- Clemmensen, K.E.; Ihrmark, K.; Brandström-Durling, M.; Lindahl, B.D. Sample preparation for fungal community analysis by high-throughput sequencing of barcode amplicons. In Microbial Environmental Genomics (MEG); Methods in Molecular Biology; Martin, F., Uroz, S., Eds.; Humana Press: New York, NY, USA, 2016; Volume 1399, pp. 61–88. [Google Scholar] [CrossRef]
- Nguyen, N.H.; Song, Z.; Bates, S.T.; Branco, S.; Tedersoo, L.; Menke, J.; Schilling, J.S.; Kennedy, P.G. FUNGuild: An open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 2016, 20, 241–248. [Google Scholar] [CrossRef]
- Tedersoo, L.; Bahram, M.; Põlme, S.; Kõljalg, U.; Yorou, N.S.; Wijesundera, R.; Ruiz, L.V.; Vasco-Palacios, A.M.; Thu, P.Q.; Suija, A.; et al. Global diversity and geography of soil fungi. Science 2014, 346, 1256688. [Google Scholar] [CrossRef]
- Brooks, M.E.; Kristensen, K.; van Benthem, K.J.; Magnusson, A.; Berg, C.W.; Nielsen, A.; Skaug, H.J.; Mächler, M.; Bolker, B.M. glmmTMB balances speed and flexibility among packages for zero-inflated generalized linear mixed modeling. R J. 2017, 9, 378–400. [Google Scholar] [CrossRef]
- R: A Language and Environment for Statistical Computing. In R Foundation for Statistical Computing; R Core Team: Vienna, Austria, 2020.
- Lüdecke, D. ggeffects: Tidy Data Frames of Marginal Effects from Regression Models. J. Open Source Softw. 2018, 3, 772. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.; O’Hara, B.; Simpson, G.; Solymos, P.; Stevens, H.; Wagner, H. Vegan: Community Ecology Package, R Package Version 2.5–3. 2018. Available online: https://cran.r-project.org (accessed on 28 November 2021).
- Hervé, M. RVAideMemoire: Testing and Plotting Procedures for Biostatistics, R Packag. Version 0.9-81-2. 2022. Available online: https://cran.r-project.org/web/packages/RVAideMemoire/RVAideMemoire.pdf (accessed on 15 June 2022).
- Haynes, W. Benjamini–Hochberg Method BT—Encyclopedia of Systems Biology; Dubitzky, W., Wolkenhauer, O., Cho, K.-H., Yokota, H., Eds.; Springer: New York, NY, USA, 2013; p. 78. ISBN 978-1-4419-9863-7. [Google Scholar]
- Shannon, C.E. A Mathematical Theory of Communication. Bell Syst. Technol. J. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Magurran, A.E. Ecological Diversity and Its Measurement. J. Appl. Ecol. 1989, 26, 1101. [Google Scholar] [CrossRef]
- Marčiulynas, A.; Marčiulynienė, D.; Mishcherikova, V.; Franić, I.; Lynikienė, J.; Gedminas, A.; Menkis, A. High Variability of Fungal Communities Associated with the Functional Tissues and Rhizosphere Soil of Picea abies in the Southern Baltics. Forests 2022, 13, 1103. [Google Scholar] [CrossRef]
- Hoeber, S.; Baum, C.; Weih, M.; Manzoni, S.; Fransson, P. Site-Dependent Relationships between Fungal Community Composition, Plant Genotypic Diversity and Environmental Drivers in a Salix Biomass System. Front. Fungal Biol. 2021, 2, 31. [Google Scholar] [CrossRef]
- Alzarhani, A.K.; Clark, D.R.; Underwood, G.J.C.; Ford, H.; Cotton, T.E.A.; Dumbrell, A.J. Are drivers of root-associated fungal community structure context specific? ISME J. 2019, 13, 1330–1344. [Google Scholar] [CrossRef] [PubMed]
- Adamo, I.; Ortiz-Malavasi, E.; Chazdon, R.; Chaverri, P.; Ter Steege, H.; Geml, J. Soil fungal community composition correlates with site-specific abiotic factors, tree community structure, and forest age in regenerating tropical rainforests. Biology 2021, 10, 1120. [Google Scholar] [CrossRef]
- Lumibao, C.Y.; Borer, E.T.; Condon, B.; Kinkel, L.; May, G.; Seabloom, E.W. Site-specific responses of foliar fungal microbiomes to nutrient addition and herbivory at different spatial scales. Ecol. Evol. 2019, 9, 12231–12244. [Google Scholar] [CrossRef]
- Smith, J.E.; Molina, R.; Huso, M.M.P.; Luoma, D.L.; McKay, D.; Castellano, M.A.; Lebel, T.; Valachovic, Y. Species richness, abundance, and composition of hypogeous and epigeous ectomycorrhizal fungal sporocarps in young, rotation-age, and old-growth stands of Douglas-fir (Pseudotsuga menziesii) in the Cascade Range of Oregon, U.S.A. Can. J. Bot. 2002, 80, 186–204. [Google Scholar] [CrossRef]
- Dang, P.; Yu, X.; Le, H.; Liu, J.; Shen, Z.; Zhao, Z. Effects of stand age and soil properties on soil bacterial and fungal community composition in Chinese pine plantations on the Loess Plateau. PLoS ONE 2017, 12, e0186501. [Google Scholar] [CrossRef]
- Odriozola, I.; Martinovic, T.; Bahnmann, B.D.; Ryšánek, D.; Mašínová, T.; Sedlák, P.; Merunková, K.; Kohout, P.; Tomšovský, M.; Baldrian, P. Stand age affects fungal community composition in a Central European temperate forest. Fungal Ecol. 2020, 48, 100985. [Google Scholar] [CrossRef]
- Unuk, T.; Martinović, T.; Finžgar, D.; Šibanc, N.; Grebenc, T.; Kraigher, H. Root-Associated Fungal Communities from Two Phenologically Contrasting Silver Fir (Abies alba Mill.) Groups of Trees. Front. Plant Sci. 2019, 10, 214. [Google Scholar] [CrossRef]
- Izzo, A.; Agbowo, J.; Bruns, T.D. Detection of plot-level changes in ectomycorrhizal communities across years in an old-growth mixed-conifer forest. New Phytol. 2005, 166, 619–630. [Google Scholar] [CrossRef]
- Barnes, C.J.; van der Gast, C.J.; McNamara, N.P.; Rowe, R.; Bending, G.D. Extreme rainfall affects assembly of the root-associated fungal community. New Phytol. 2018, 220, 1172–1184. [Google Scholar] [CrossRef] [PubMed]
- Bui, A.; Orr, D.; Lepori-Bui, M.; Konicek, K.; Young, H.S.; Moeller, H.V. Soil fungal community composition and functional similarity shift across distinct climatic conditions. FEMS Microbiol. Ecol. 2020, 96, fiaa193. [Google Scholar] [CrossRef] [PubMed]
- Talley, S.M.; Coley, P.D.; Kursar, T.A. The effects of weather on fungal abundance and richness among 25 communities in the Intermountain West. BMC Ecol. 2002, 2, 7–10. [Google Scholar] [CrossRef] [PubMed]
- Swaty, R.L.; Gehring, C.A.; Van Ert, M.; Theimer, T.A.D.C.; Keim, P.; Whitham, T.G. Temporal variation in temperature and rainfall differentially affects ectomycorrhizal colonization at two contrasting sites. New Phytol. 1998, 139, 733–739. [Google Scholar] [CrossRef]
- Järvan, M.; Edesi, L.; Adamson, A.; Võsa, T. Soil microbial communities and dehydrogenase activity depending on farming systems. Plant Soil Environ. 2014, 60, 459–463. [Google Scholar] [CrossRef]
- Schindlbacher, A.; Rodler, A.; Kuffner, M.; Kitzler, B.; Sessitsch, A.; Zechmeister-Boltenstern, S. Experimental warming effects on the microbial community of a temperate mountain forest soil. Soil Biol. Biochem. 2011, 43, 1417–1425. [Google Scholar] [CrossRef]
- Davidson, E.A.; Janssens, I.A. Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature 2006, 440, 165–173. [Google Scholar] [CrossRef]
- Kirschbaum, M.U.F. The temperature dependence of soil organic matter decomposition, and the effect of global warming on soil organic C storage. Soil Biol. Biochem. 1995, 27, 753–760. [Google Scholar] [CrossRef]
- Martin, M.P.; Wattenbach, M.; Smith, P.; Meersmans, J.; Jolivet, C.; Boulonne, L.; Arrouays, D. Spatial distribution of soil organic carbon stocks in France. Biogeosciences 2011, 8, 1053–1065. [Google Scholar] [CrossRef]
- de Graaff, M.-A.; Classen, A.T.; Castro, H.F.; Schadt, C.W. Labile soil carbon inputs mediate the soil microbial community composition and plant residue decomposition rates. New Phytol. 2010, 188, 1055–1064. [Google Scholar] [CrossRef]
- Bradford, M.A.; Wieder, W.R.; Bonan, G.B.; Fierer, N.; Raymond, P.A.; Crowther, T.W. Managing uncertainty in soil carbon feedbacks to climate change. Nat. Clim. Chang. 2016, 6, 751–758. [Google Scholar] [CrossRef]
- Shi, L.; Dossa, G.G.O.; Paudel, E.; Zang, H.; Xu, J.; Harrison, R.D. Changes in fungal communities across a forest disturbance gradient. Appl. Environ. Microbiol. 2019, 85, e00080-19. [Google Scholar] [CrossRef] [PubMed]
- Pennanen, T.; Fitze, H.; Vanhala, P.; Kiikkilä, O.; Neuvonen, S.; Bååth, E. Structure of a microbial community in soil after prolonged addition of low levels of simulated acid rain. Appl. Environ. Microbiol. 1998, 64, 2173–2180. [Google Scholar] [CrossRef] [PubMed]
- Albright, M.B.; Johansen, R.; Thompson, J.; Lopez, D.; Gallegos-Graves, L.V.; Kroeger, M.E.; Runde, A.; Mueller, R.C.; Washburne, A.; Munsky, B.; et al. Soil bacterial and fungal richness forecast patterns of early pine litter decomposition. Front. Microbiol. 2020, 11, 542220. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, L.O.; Bååth, E.; Falkengren-Grerup, U.; Wallander, H. Growth of ectomycorrhizal mycelia and composition of soil microbial communities in oak forest soils along a nitrogen deposition gradient. Oecologia 2007, 153, 375–384. [Google Scholar] [CrossRef] [PubMed]
- Coleman-Derr, D.; Desgarennes, D.; Fonseca-Garcia, C.; Gross, S.; Clingenpeel, S.; Woyke, T.; North, G.; Visel, A.; Partida-Martinez, L.P.; Tringe, S.G. Plant compartment and biogeography affect microbiome composition in cultivated and native Agave species. New Phytol. 2016, 209, 798–811. [Google Scholar] [CrossRef]
- Fonseca-García, C.; Coleman-Derr, D.; Garrido, E.; Visel, A.; Tringe, S.G.; Partida-Martínez, L.P. The Cacti Microbiome: Interplay between habitat-filtering and host-specificity. Front. Microbiol. 2016, 7, 150. [Google Scholar] [CrossRef] [PubMed]
- Rosling, A.; Cox, F.; Cruz-Martinez, K.; Ihrmark, K.; Grelet, G.A.; Lindahl, B.D.; Menkis, A.; James, T.Y. Archaeorhizomycetes: Unearthing an ancient class of ubiquitous soil fungi. Science 2011, 333, 876–879. [Google Scholar] [CrossRef]
- Qadri, M.; Rajput, R.; Abdin, M.Z.; Vishwakarma, R.A.; Riyaz-Ul-Hassan, S. Diversity, Molecular Phylogeny, and Bioactive Potential of Fungal Endophytes Associated with the Himalayan Blue Pine (Pinus wallichiana). Microb. Ecol. 2014, 67, 877–887. [Google Scholar] [CrossRef]
- Jeewon, R.; Yeung, Q.S.Y.; Wannasinghe, D.N.; Rampadarath, S.; Puchooa, D.; Wang, H.K.; Hyde, K.D. Hidden mycota of pine needles: Molecular signatures from PCR-DGGE and Ribosomal DNA phylogenetic characterization of novel phylotypes. Sci. Rep. 2018, 8, 18053. [Google Scholar] [CrossRef]
- Lynikienė, J.; Marčiulynienė, D.; Marčiulynas, A.; Gedminas, A.; Vaičiukynė, M.; Menkis, A. Managed and unmanaged Pinus sylvestris forest stands harbour similar diversity and composition of the phyllosphere and soil fungi. Microorganisms 2020, 8, 259. [Google Scholar] [CrossRef] [PubMed]
- Bahram, M.; Netherway, T.; Hildebrand, F.; Pritsch, K.; Drenkhan, R.; Loit, K.; Anslan, S.; Bork, P.; Tedersoo, L. Plant nutrient—Acquisition strategies drive topsoil microbiome structure and function. New Phytol. 2020, 227, 1189–1199. [Google Scholar] [CrossRef]
- Hou, J.; Feng, H.; Wu, M. Incorporating Effect Factors into the Relationship between Biodiversity and Ecosystem Functioning (BEF). Diversity 2022, 14, 274. [Google Scholar] [CrossRef]
- de la Fuente Cantó, C.; Simonin, M.; King, E.; Moulin, L.; Bennett, M.J.; Castrillo, G.; Laplaze, L. An extended root phenotype: The rhizosphere, its formation and impacts on plant fitness. Plant J. 2020, 103, 951–964. [Google Scholar] [CrossRef] [PubMed]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; Van Themaat, E.V.L.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef] [PubMed]
- Hanso, M.; Drenkhan, R. Lophodermium needle cast, insect defoliation and growth responses of young Scots pines in Estonia. For. Pathol. 2012, 42, 124–135. [Google Scholar] [CrossRef]
- Millberg, H. Foliar Fungi of Scots Pine (Pinus sylvestris). Ph.D. Thesis, Department of Forest Mycology and Plant Pathology, Swedish University of Agricultural Sciences, Uppsala, Zweden, 2015; pp. 1652–6880. [Google Scholar]
- Haas, J.C.; Street, N.R.; Sjödin, A.; Lee, N.M.; Högberg, M.N.; Näsholm, T.; Hurry, V. Microbial community response to growing season and plant nutrient optimisation in a boreal Norway spruce forest. Soil Biol. Biochem. 2018, 125, 197–209. [Google Scholar] [CrossRef]
- Policelli, N.; Bruns, T.D.; Vilgalys, R.; Nuñez, M.A. Suilloid fungi as global drivers of pine invasions. New Phytol. 2019, 222, 714–725. [Google Scholar] [CrossRef]
- Peay, K.G.; Baraloto, C.; Fine, P.V.A. Strong coupling of plant and fungal community structure across western Amazonian rainforests. ISME J. 2013, 7, 1852–1861. [Google Scholar] [CrossRef]
- van der Linde, S.; Suz, L.M.; Orme, C.D.L.; Cox, F.; Andreae, H.; Asi, E.; Atkinson, B.; Benham, S.; Carroll, C.; Cools, N.; et al. Author Correction: Environment and host as large-scale controls of ectomycorrhizal fungi. Nature 2018, 561, E42. [Google Scholar] [CrossRef]
- Baar, J.; Horton, T.R.; Kretzer, A.; Bruns, T.D. From Resistant Propagules After a Stand- Replacing Wildfire. New Phytol. 1999, 143, 409–418. [Google Scholar] [CrossRef]
- Qu, Z.L.; Santalahti, M.; Köster, K.; Berninger, F.; Pumpanen, J.; Heinonsalo, J.; Sun, H. Soil Fungal Community Structure in Boreal Pine Forests: From Southern to Subarctic Areas of Finland. Front. Microbiol. 2021, 12, 653896. [Google Scholar] [CrossRef] [PubMed]
- Hoff, J.A.; Klopfenstein, N.B.; Mcdonald, G.I.; Tonn, J.R.; Kim, M.S.; Zambino, P.J.; Hessburg, P.F.; Rogers, J.D.; Peever, T.L.; Carris, L.M. Fungal endophytes in woody roots of Douglas-fir (Pseudotsuga menziesii) and ponderosa pine (Pinus ponderosa). For. Pathol. 2004, 34, 255–271. [Google Scholar] [CrossRef]
- Summerbell, R.C. Root endophyte and mycorrhizosphere fungi of black spruce, Picea mariana, in a boreal forest habitat: Influence of site factors on fungal distributions. Stud. Mycol. 2005, 53, 121–145. [Google Scholar] [CrossRef]
- Bååth, E. Microfungi in a clear-cut pine forest soil in central Sweden. Can. J. Bot. 1981, 59, 1331–1337. [Google Scholar] [CrossRef]
- Kohout, P.; Charvátová, M.; Štursová, M.; Mašínová, T.; Tomšovský, M.; Baldrian, P. Clearcutting alters decomposition processes and initiates complex restructuring of fungal communities in soil and tree roots. ISME J. 2018, 12, 692–703. [Google Scholar] [CrossRef]
- Halmschlager, E.; Kowalski, T. The mycobiota in nonmycorrhizal roots of healthy and declining oaks. Can. J. Bot. 2004, 82, 1446–1458. [Google Scholar] [CrossRef]
- Sheng, Y.; Cong, J.; Lu, H.; Yang, L.; Liu, Q.; Li, D.; Zhang, Y. Broad-leaved forest types affect soil fungal community structure and soil organic carbon contents. Microbiologyopen 2019, 8, e874. [Google Scholar] [CrossRef]
- Crous, P.W.; Quaedvlieg, W.; Hansen, K.; Hawksworth, D.L.; Groenewald, J.Z. Phacidium and Ceuthospora (Phacidiaceae) are congeneric: Taxonomic and nomenclatural implications. IMA Fungus 2014, 5, 173–193. [Google Scholar] [CrossRef]
- Nawrot-Chorabik, K.; Grad, B.; Kowalski, T. Interactions between callus cultures of Pinus sylvestris and pine fungi with different trophic properties. For. Pathol. 2016, 46, 179–186. [Google Scholar] [CrossRef]
- Ozturk, I.K.; Amiri, A. Pathogenicity and Control of Phacidium lacerum, an Emerging Pome Fruit Pathogen in Washington State. Plant Dis. 2020, 104, 3124–3130. [Google Scholar] [CrossRef] [PubMed]
- Silva, A.C.; Henriques, J.; Diogo, E.; Ramos, A.P.; Bragança, H. First report of Sydowia polyspora causing disease on Pinus pinea shoots. For. Pathol. 2020, 50, 27–30. [Google Scholar] [CrossRef]
- Cleary, M.; Oskay, F.; Doğmuş, H.T.; Lehtijärvi, A.; Woodward, S.; Vettraino, A.M. Cryptic risks to forest biosecurity associated with the global movement of commercial seed. Forests 2019, 10, 459. [Google Scholar] [CrossRef]
- Davydenko, K.; Vasaitis, R.; Meshkova, V.; Menkis, A. Fungi associated with the red-haired bark beetle, Hylurgus ligniperda (Coleoptera: Curculionidae) in the forest-steppe zone in eastern Ukraine. Eur. J. Entomol. 2014, 111, 561–565. [Google Scholar] [CrossRef]
- Lygis, V.; Vasiliauskaite, I.; Matelis, A.; Pliūra, A.; Vasaitis, R. Fungi in living and dead stems and stumps of pinus mugo on coastal dunes of the baltic sea. Plant Prot. Sci. 2014, 50, 221–226. [Google Scholar] [CrossRef]
- Behnke, J.; Hanna, B. Fungi associated with Cyclaneusma needle cast in Scots pine in the west of Poland. For. Pathol. 2019, 49, e12487. [Google Scholar] [CrossRef]


| Site | Position | Tree Species | Age | Soil Chemical Parameters | Climate Data | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. | Longitude (N) | Latitude (E) | Composition, % * | (y) | pH, mol/L (KCl) | P2O5, mg/kg | K2O, mg/kg | Ca, mg/kg | Mg, mg/kg | Cl, mg/kg | Salts, ms/cm | Average Annual Temp. °C | Precipitation, mm/year |
| 1 | 55°37′14″ | 25°11′25″ | 100P | 60 | 4.0 | 345 | 25 | 155 | 48 | 3.6 | 2.55 | 8.0 | 508.8 |
| 2 | 53°58′52″ | 23°56′56″ | 100P | 98 | 4.2 | 93 | 31 | 212 | 36 | 3.6 | 1.61 | 8.0 | 484.6 |
| 3 | 53°58′27″ | 24°25′48″ | 100P | 53 | 4.2 | 123 | 25 | 256 | 74 | 3.6 | 1.89 | 8.0 | 484.6 |
| 4 | 54°8′20″ | 24°11′51″ | 100P | 53 | 4.4 | 64 | 23 | 260 | 60 | 3.6 | 1.44 | 8.0 | 484.6 |
| 5 | 54°52′40″ | 23°42′17″ | 80P,20B | 67 | 4.1 | 60 | 19 | 205 | 50 | 3.6 | 2.18 | 8.3 | 614.9 |
| 6 | 55°10′27″ | 24°26′43″ | 100P | 57 | 4.0 | 139 | 19 | 234 | 55 | 5.3 | 2.41 | 8.3 | 614.9 |
| 7 | 55°1′10″ | 24°12′56″ | 100P | 71 | 3.5 | 19 | 57 | 181 | 45 | 5.3 | 2.71 | 8.6 | 519.8 |
| 8 | 54°52′15″ | 23°26′41″ | 100P | 49 | 3.9 | 18 | 61 | 298 | 62 | 3.6 | 3.45 | 8.9 | 491.7 |
| 9 | 56°11′40″ | 21°28′13″ | 100P | 59 | 3.8 | 34 | 65 | 643 | 115 | 5.3 | 4.45 | 8.9 | 491.7 |
| 10 | 56°1′1″ | 21°6′45″ | 90P,10S | 47 | 4.0 | 92 | 31 | 246 | 46 | 3.6 | 1.74 | 8.3 | 643.6 |
| 11 | 55°45′36″ | 24°41′38″ | 70P,20S,10B | 72 | 3.4 | 13 | 36 | 191 | 50 | 3.6 | 3.07 | 8.1 | 479.8 |
| 12 | 55°45′36″ | 24°41′38″ | 80P,20B | 68 | 5.9 | 71 | 60 | 4100 | 670 | 6.9 | 20.2 | 8.1 | 505.5 |
| 13 | 56°11′12″ | 22°23′4″ | 100P | 58 | 4.2 | 25 | 86 | 2081 | 272 | 5.3 | 5.18 | 8.0 | 480.4 |
| 14 | 54°52′16″ | 25°41′56″ | 100P | 65 | 4.1 | 41 | 57 | 339 | 68 | 3.6 | 2.81 | 8.0 | 755.4 |
| 15 | 54°34′28″ | 23°57′2″ | 100P | 47 | 4.0 | 115 | 41 | 243 | 57 | 3.6 | 2.21 | 8.6 | 595.2 |
| 16 | 55°28′16″ | 23°26′39″ | 100P | 55 | 4.2 | 70 | 39 | 373 | 66 | 5.3 | 2.08 | 7.8 | 458.0 |
| 17 | 55°29′54″ | 21°57′21″ | 70P,20S,10B | 65 | 2.8 | 138 | 381 | 1171 | 271 | 7.1 | 15.9 | 7.7 | 514.6 |
| 18 | 56°3′26″ | 25°42′28″ | 90P,10S | 59 | 3.9 | 10 | 51 | 208 | 41 | 3.6 | 3.02 | 8.4 | 617.0 |
| 19 | 55°1′46″ | 22°42′14″ | 70P,20J,10B | 36 | 3.9 | 78 | 30 | 178 | 53 | 3.6 | 1.78 | 8.6 | 519.8 |
| 20 | 54°17′17″ | 25°39′41″ | 90P,10S | 50 | 3.9 | 44 | 44 | 191 | 54 | 5.3 | 3.29 | 8.0 | 530.6 |
| 21 | 55°10′12″ | 25°41′56″ | 100P | 65 | 4.4 | 57 | 13 | 97 | 28 | 3.6 | 2.62 | 7.4 | 669.2 |
| 22 | 55°19′38″ | 22°27′14″ | 70P,20B,10S | 60 | 3.5 | 16 | 174 | 834 | 281 | 3.6 | 5 | 8.6 | 591.3 |
| 23 | 55°45′34″ | 21°43′12″ | 70P,20S,10B | 65 | 3.4 | 30 | 83 | 383 | 81 | 5.3 | 4.29 | 8.0 | 598.7 |
| 24 | 54°44′58″ | 24°41′59″ | 100P | 58 | 5.9 | 271 | 67 | 1624 | 215 | 3.6 | 8.49 | 7.3 | 614.9 |
| 25 | 54°56′54″ | 24°41′46″ | 100P | 55 | 4.9 | 104 | 29 | 399 | 68 | 5.3 | 3.19 | 8.0 | 535.8 |
| 26 | 55°10′47″ | 24°42′10″ | 100P | 50 | 3.9 | 130 | 36 | 183 | 52 | 3.6 | 2.72 | 8.0 | 506.9 |
| 27 | 55°19′1″ | 25°42′54″ | 100P | 65 | 4.5 | 37 | 48 | 675 | 97 | 5.3 | 4.86 | 7.7 | 505.4 |
| 28 | 54°25′23″ | 24°57′33″ | 90P,10B | 48 | 4.1 | 14 | 33 | 180 | 44 | 5.3 | 2.5 | 8.0 | 593.2 |
| 29 | 54°25′39″ | 24°27′15″ | 90P,10B | 70 | 4.3 | 38 | 48 | 323 | 66 | 3.6 | 3.88 | 8.0 | 540.9 |
| 30 | 54°7′57″ | 23°41′50″ | 100P | 69 | 5.3 | 62 | 40 | 1053 | 115 | 3.6 | 4.03 | 8.4 | 565.2 |
| Site | No. of High-Quality Sequences/OTUs | Shannon Diversity Index (H) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. | Roots | Shoots | Needles | Soil O * | Soil M * | Soil O + M | Roots | Shoots | Needles | Soil O * | Soil M * | Soil O + M |
| 1 | 6569/74 | 271/43 | 1117/136 | 3165/283 | 209/77 | 3374/311 | 2.28 | 2.78 | 3.78 | 4.20 | 3.84 | 4.29 |
| 2 | 2558/117 | 224/59 | 1529/134 | 414/85 | 639/148 | 1053/193 | 2.83 | 3.34 | 3.26 | 3.35 | 4.06 | 4.19 |
| 3 | 2189/91 | 597/61 | 2518/125 | 1092/145 | 863/193 | 1955/279 | 3.21 | 2.61 | 2.99 | 3.58 | 4.57 | 4.34 |
| 4 | 1107/50 | 3779/134 | 870/61 | 3205/237 | 1962/131 | 5167/312 | 2.60 | 2.91 | 2.82 | 3.64 | 3.53 | 3.91 |
| 5 | 907/67 | 458/93 | 770/85 | 3180/273 | 742/87 | 3922/305 | 3.04 | 3.88 | 3.06 | 3.20 | 3.05 | 3.40 |
| 6 | 139/33 | 49/30 | 1846/182 | 236/101 | 390/136 | 626/191 | 3.06 | 3.25 | 4.03 | 4.05 | 4.00 | 4.27 |
| 7 | 3026/108 | 517/104 | 1974/174 | 1/1 | 21/8 | 22/9 | 2.93 | 3.81 | 3.54 | - | 1.59 | 1.70 |
| 8 | 2390/97 | 1006/125 | 479/76 | 708/142 | 1609/157 | 2317/246 | 3.30 | 3.53 | 3.35 | 4.34 | 3.70 | 4.31 |
| 9 | 969/88 | 708/119 | 2286/180 | 1427/234 | 1888/208 | 3315/322 | 3.18 | 3.84 | 3.75 | 4.47 | 4.02 | 4.42 |
| 10 | 1309/126 | 1715/222 | 4841/325 | 39/29 | 2407/184 | 2446/198 | 3.04 | 4.19 | 4.24 | 3.24 | 3.80 | 3.85 |
| 11 | 411/104 | 3032/248 | 1058/128 | 382/97 | 191/57 | 573/125 | 3.76 | 4.06 | 3.78 | 3.82 | 3.50 | 4.05 |
| 12 | 885/89 | 3134/237 | 689/97 | 1074/223 | 731/80 | 1805/253 | 3.18 | 4.03 | 2.58 | 4.66 | 2.93 | 4.37 |
| 13 | 75/36 | 2018/169 | 3590/188 | 6088/242 | 3955/262 | 10043/371 | 3.30 | 3.80 | 3.01 | 3.81 | 4.08 | 4.09 |
| 14 | 5076/103 | 1726/156 | 677/102 | 1275/267 | 1018/202 | 2293/367 | 2.47 | 3.68 | 3.23 | 4.86 | 4.65 | 5.11 |
| 15 | 223/56 | 1252/115 | 4097/219 | 1508/152 | 1339/137 | 2847/245 | 3.28 | 3.56 | 3.83 | 3.71 | 3.20 | 4.02 |
| 16 | 79/14 | 4344/221 | 3936/232 | 168/75 | 87/46 | 255/107 | 1.97 | 3.67 | 3.77 | 3.71 | 3.48 | 4.02 |
| 17 | 75/30 | 3770/194 | 3037/178 | 883/123 | 640/138 | 1523/231 | 3.04 | 3.85 | 3.29 | 3.73 | 4.10 | 4.42 |
| 18 | 841/51 | 5417/202 | 3649/209 | 113/48 | 285/78 | 398/106 | 1.97 | 3.68 | 3.87 | 3.15 | 3.54 | 3.75 |
| 19 | 174/50 | 5273/186 | 5146/204 | 27/21 | 152/79 | 179/93 | 3.21 | 3.36 | 3.37 | 2.97 | 3.97 | 4.12 |
| 20 | 139/38 | 2440/150 | 3122/140 | 1355/242 | 574/106 | 1929/289 | 3.00 | 3.43 | 3.28 | 4.81 | 3.86 | 4.84 |
| 21 | 121/42 | 913/86 | 2819/189 | 701/142 | 234/92 | 935/210 | 3.18 | 3.18 | 3.19 | 4.18 | 4.17 | 4.61 |
| 22 | -/- | 2368/153 | 4445/216 | 87/54 | 61/31 | 148/83 | - | 3.37 | 3.71 | 3.70 | 3.35 | 4.04 |
| 23 | 169/33 | 4620/233 | 1885/137 | 1047/128 | 1752/200 | 2799/286 | 2.56 | 3.67 | 2.86 | 3.47 | 4.40 | 4.54 |
| 24 | 219/41 | 2890/158 | 2572/148 | 68/16 | 136/35 | 204/45 | 2.65 | 3.89 | 3.39 | 1.84 | 1.65 | 1.88 |
| 25 | 2193/61 | 1490/155 | 682/89 | 4009/345 | 1721/159 | 5730/396 | 1.62 | 3.93 | 3.26 | 4.39 | 4.06 | 4.57 |
| 26 | 149/35 | 4207/239 | 397/89 | 1575/183 | 553/112 | 2128/245 | 2.52 | 3.95 | 3.66 | 3.57 | 3.37 | 3.74 |
| 27 | 1395/88 | 4081/159 | 417/71 | 3611/321 | 2120/148 | 5731/392 | 2.71 | 3.36 | 3.08 | 4.40 | 3.48 | 4.44 |
| 28 | 1868/109 | 652/85 | 1102/119 | 1257/166 | 1188/195 | 2445/298 | 3.12 | 3.32 | 3.18 | 3.62 | 4.34 | 4.39 |
| 29 | 699/58 | 5143/217 | 710/91 | 1960/242 | 2598/218 | 4558/364 | 2.58 | 3.47 | 3.48 | 4.56 | 4.21 | 4.67 |
| 30 | -/- | 10667/233 | 776/100 | 1502/166 | 1234/183 | 2736/276 | - | 2.78 | 3.42 | 3.83 | 4.13 | 4.27 |
| Total | 35954/734 | 78760/907 | 63031/953 | 42117/1440 | 31254/1233 | 73371/1854 | ||||||
| Fungal OTU | Phylum * | Genbank/UNITE Reference | Sequence Similarity, % | Needles, % | Shoots, % | Roots, % | Soil O **, % | Soil M **, % | Soil all, % | All, % |
|---|---|---|---|---|---|---|---|---|---|---|
| Dothideomycetes sp. 5208_5 | A | KX908472 | 99 | 20.445 | 3.389 | 0.039 | 3.217 | 1.709 | 2.575 | 6.953 |
| Unidentified sp. 5208_1 | A | KP891398 | 100 | 4.314 | 12.786 | 0.017 | 1.363 | 1.654 | 1.487 | 5.530 |
| Archaeorhizomyces sp. 5208_0 | A | MH248043 | 100 | 1.231 | 0.382 | 8.352 | 8.393 | 9.839 | 9.009 | 4.257 |
| Unidentified sp. 5208_2 | A | MN902367 | 100 | 4.406 | 7.096 | 0.011 | 2.275 | 1.814 | 2.078 | 3.940 |
| Helotiales sp. 5208_17 | A | KY742593 | 100 | 1.577 | 6.136 | 0.011 | 0.617 | 0.573 | 0.598 | 2.497 |
| Rhinocladiella sp. 5208_3 | A | KM056296 | 98 | 0.398 | 6.435 | 0.008 | 0.769 | 0.518 | 0.662 | 2.313 |
| Phacidium lacerum | A | MN588163 | 100 | 4.515 | 0.113 | 0.036 | 3.039 | 1.248 | 2.276 | 1.839 |
| Phialocephala fortinii | A | MN947395 | 100 | 0.160 | 0.041 | 11.056 | 0.468 | 0.458 | 0.463 | 1.771 |
| Scoliciosporum umbrinum | A | KX133008 | 100 | 0.887 | 4.247 | 0.003 | 0.477 | 0.694 | 0.570 | 1.722 |
| Sydowia polyspora | A | MN900630 | 100 | 3.428 | 1.894 | 0.031 | 0.567 | 0.467 | 0.525 | 1.612 |
| Mycena cinerella | B | KT900146 | 100 | 0.024 | 0.009 | 8.847 | 1.135 | 0.205 | 0.739 | 1.491 |
| Malassezia restricta | B | LT854697 | 100 | 1.228 | 0.670 | 0.456 | 1.854 | 3.180 | 2.419 | 1.291 |
| Microsphaeropsis olivacea | A | MT561396 | 100 | 2.588 | 0.819 | 0.111 | 0.803 | 1.641 | 1.160 | 1.261 |
| Penicillium camemberti | A | MT355566 | 100 | 0.159 | 0.024 | 5.326 | 1.648 | 1.037 | 1.387 | 1.215 |
| Phaeomoniella pinifoliorum | A | MK762595 | 100 | 0.982 | 1.769 | 0.006 | 0.558 | 0.483 | 0.526 | 0.956 |
| Unidentified sp. 5208_36 | A | MG828311 | 100 | 0.695 | 2.051 | 0.008 | 0.480 | 0.368 | 0.432 | 0.945 |
| Cladosporium herbarum | A | MT635288 | 100 | 1.826 | 0.432 | 0.631 | 0.784 | 0.848 | 0.811 | 0.921 |
| Chaetothyriomycetidae sp. 5208_56 | A | KX589170 | 98 | 0.227 | 2.400 | - | 0.078 | 0.483 | 0.251 | 0.883 |
| Leptosphaeria sp. 5208_64 | A | JQ044439 | 97 | 0.252 | 2.417 | - | 0.024 | 0.022 | 0.023 | 0.828 |
| Unidentified sp. 5208_39 | A | MT242010 | 100 | 1.599 | 1.023 | 0.006 | 0.380 | 0.262 | 0.330 | 0.820 |
| Hyaloscypha variabilis | A | MT469925 | 100 | 0.057 | - | 3.221 | 1.099 | 1.142 | 1.118 | 0.802 |
| Umbelopsis dimorpha | Z | MT138616 | 100 | 0.114 | 0.004 | 0.028 | 1.885 | 3.209 | 2.449 | 0.749 |
| Chaetothyriales sp. 5208_15 | A | KP400572 | 100 | 0.836 | 1.346 | 0.019 | 0.297 | 0.381 | 0.333 | 0.732 |
| Trechispora sp. 5208_19 | B | JX392812 | 99 | 0.002 | - | 3.919 | 0.582 | 0.477 | 0.537 | 0.718 |
| Unidentified sp. 5208_12 | A | FJ553582 | 100 | 0.062 | 0.004 | 0.006 | 2.123 | 2.726 | 2.380 | 0.713 |
| Unidentified sp. 5208_32 | B | MN902363 | 100 | 2.189 | 0.325 | - | 0.252 | 0.070 | 0.174 | 0.702 |
| Cenococcum geophilum | A | HM189724 | 100 | 0.010 | - | 2.517 | 0.560 | 1.817 | 1.096 | 0.683 |
| Unidentified sp. 5208_72 | A | MN902396 | 100 | 1.637 | 0.458 | 0.014 | 0.525 | 0.224 | 0.397 | 0.673 |
| Unidentified sp. 5208_23 | A | MT237078 | 100 | 1.109 | 0.684 | 0.006 | 0.565 | 0.461 | 0.521 | 0.646 |
| Unidentified sp. 5208_69 | A | MN902387 | 100 | 1.474 | 0.543 | - | 0.216 | 0.083 | 0.159 | 0.587 |
| Substrate | Needles | Roots | Shoots | SoilM | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| R2 | F | p | R2 | F | p | R2 | F | p | R2 | F | p | |
| Roots | 0.37 | 24.19 | 0.0011 | - | - | - | - | - | - | - | - | - |
| Shoots | 0.23 | 14.86 | 0.0011 | 0.36 | 22.06 | 0.0011 | - | - | - | - | - | - |
| SoilM | 0.26 | 15.64 | 0.0011 | 0.20 | 8.25 | 0.0011 | 0.25 | 14.50 | 0.0011 | - | - | - |
| SoilO | 0.21 | 11.88 | 0.0011 | 0.18 | 7.68 | 0.0011 | 0.20 | 11.29 | 0.0011 | 0.02 | 0.90 | 0.591 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marčiulynienė, D.; Marčiulynas, A.; Mishcherikova, V.; Lynikienė, J.; Gedminas, A.; Franic, I.; Menkis, A. Principal Drivers of Fungal Communities Associated with Needles, Shoots, Roots and Adjacent Soil of Pinus sylvestris. J. Fungi 2022, 8, 1112. https://doi.org/10.3390/jof8101112
Marčiulynienė D, Marčiulynas A, Mishcherikova V, Lynikienė J, Gedminas A, Franic I, Menkis A. Principal Drivers of Fungal Communities Associated with Needles, Shoots, Roots and Adjacent Soil of Pinus sylvestris. Journal of Fungi. 2022; 8(10):1112. https://doi.org/10.3390/jof8101112
Chicago/Turabian StyleMarčiulynienė, Diana, Adas Marčiulynas, Valeriia Mishcherikova, Jūratė Lynikienė, Artūras Gedminas, Iva Franic, and Audrius Menkis. 2022. "Principal Drivers of Fungal Communities Associated with Needles, Shoots, Roots and Adjacent Soil of Pinus sylvestris" Journal of Fungi 8, no. 10: 1112. https://doi.org/10.3390/jof8101112
APA StyleMarčiulynienė, D., Marčiulynas, A., Mishcherikova, V., Lynikienė, J., Gedminas, A., Franic, I., & Menkis, A. (2022). Principal Drivers of Fungal Communities Associated with Needles, Shoots, Roots and Adjacent Soil of Pinus sylvestris. Journal of Fungi, 8(10), 1112. https://doi.org/10.3390/jof8101112






