In Silico Predictions of Ecological Plasticity Mediated by Protein Family Expansions in Early-Diverging Fungi
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Acquisition, Proteome Mapping, and Protein Families Selection
2.2. Sequence Alignment and Phylogenetic Analysis
3. Results and Discussion
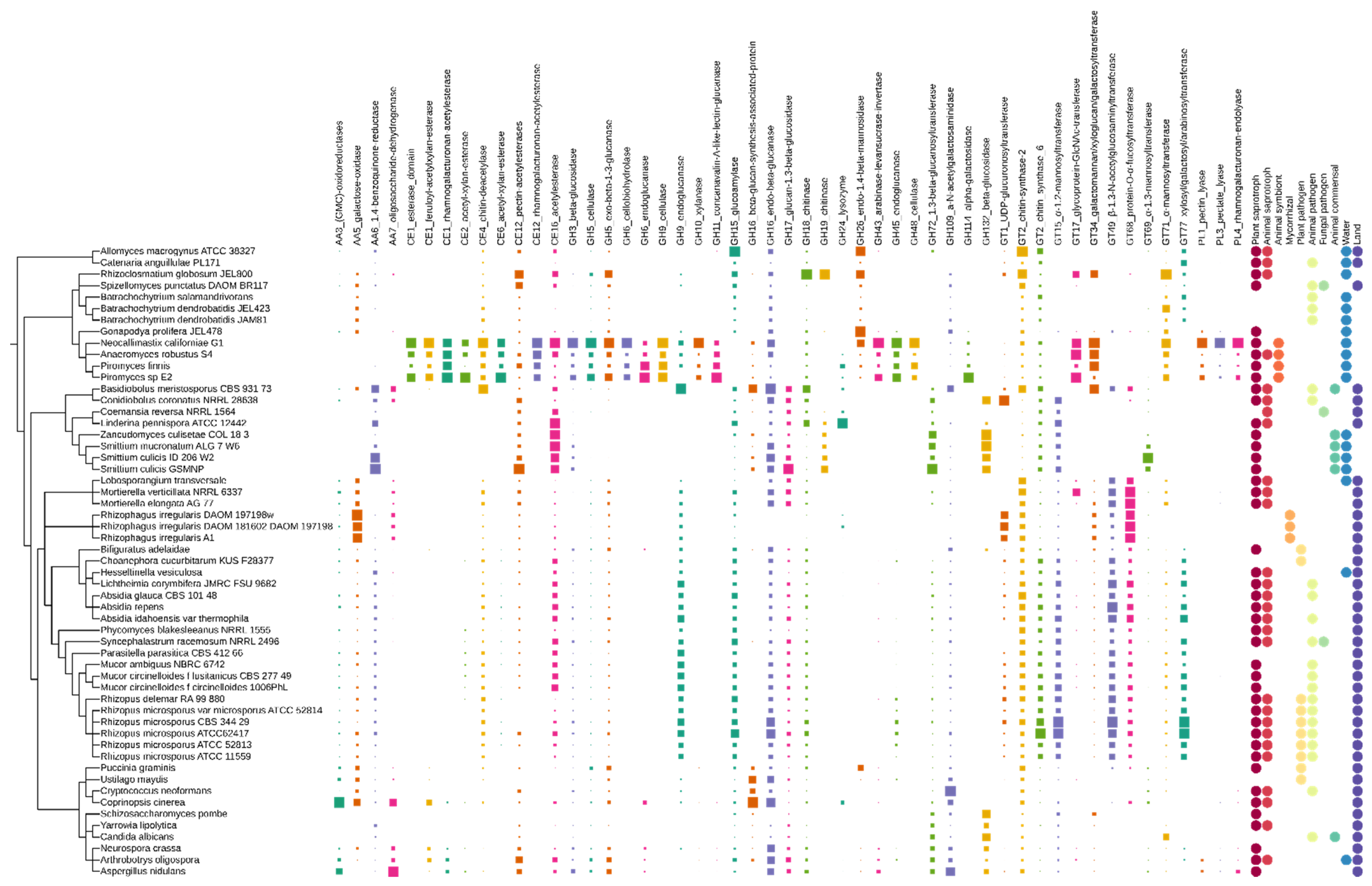
3.1. Carbohydrate-Active Enzymes Families (CAZymes)
3.1.1. CAZymes with Auxiliary Activity (AA)
3.1.2. Carbohydrate Esterases (CE)
3.1.3. Glycoside Hydrolases (GH)
3.1.4. Glycosyl Transferases (GT)
3.1.5. CAZyme Distribution and Abundance Can Be Linked to the Ecology of Particular EDF Lineages
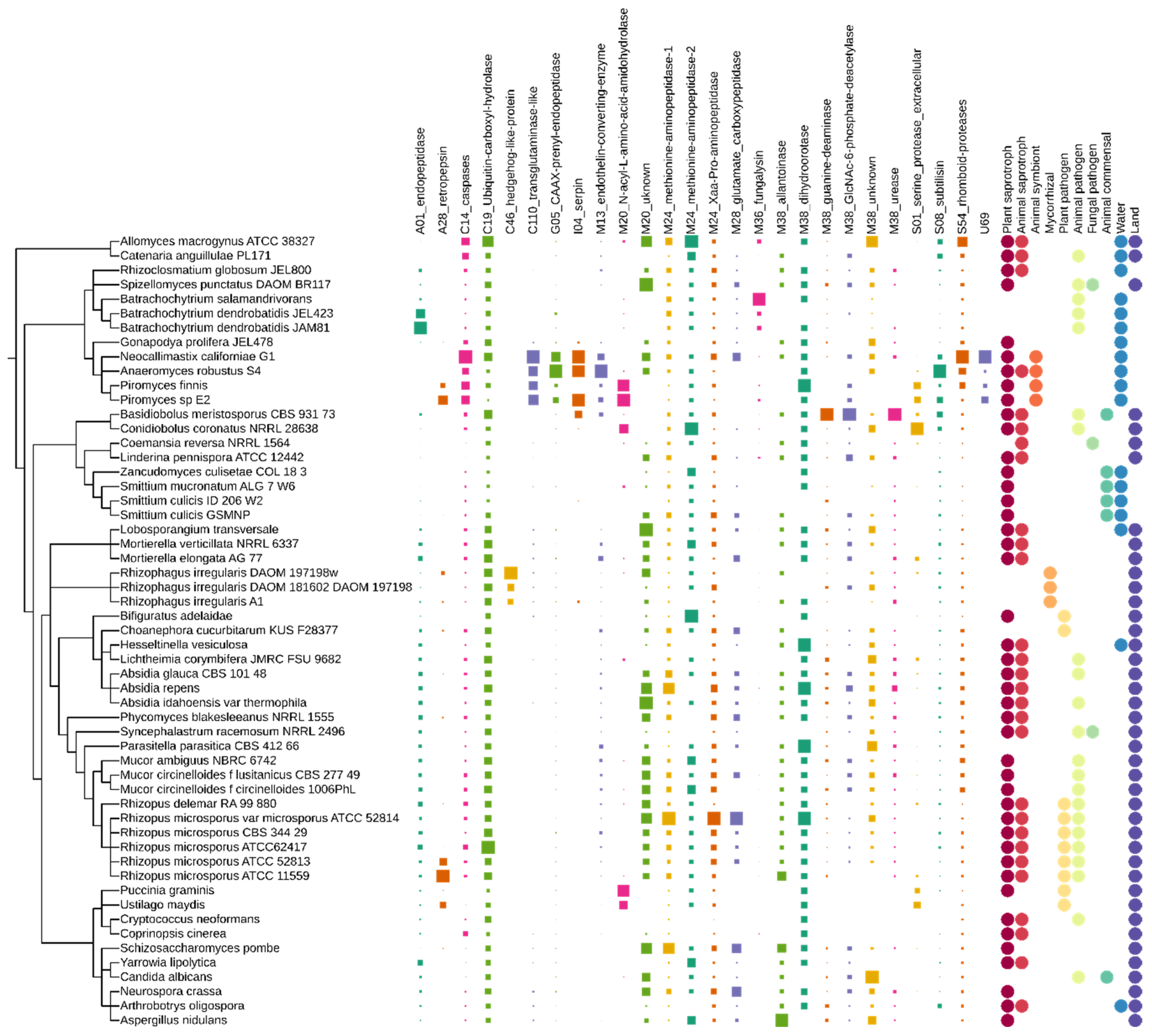
3.2. Peptidases
3.2.1. Aspartic Peptidases
3.2.2. Cysteine Peptidases
3.2.3. Glutamic Peptidases
3.2.4. Inhibitors
3.2.5. Metallo Peptidases
3.2.6. Serine Peptidases
3.2.7. Unknown Peptidases
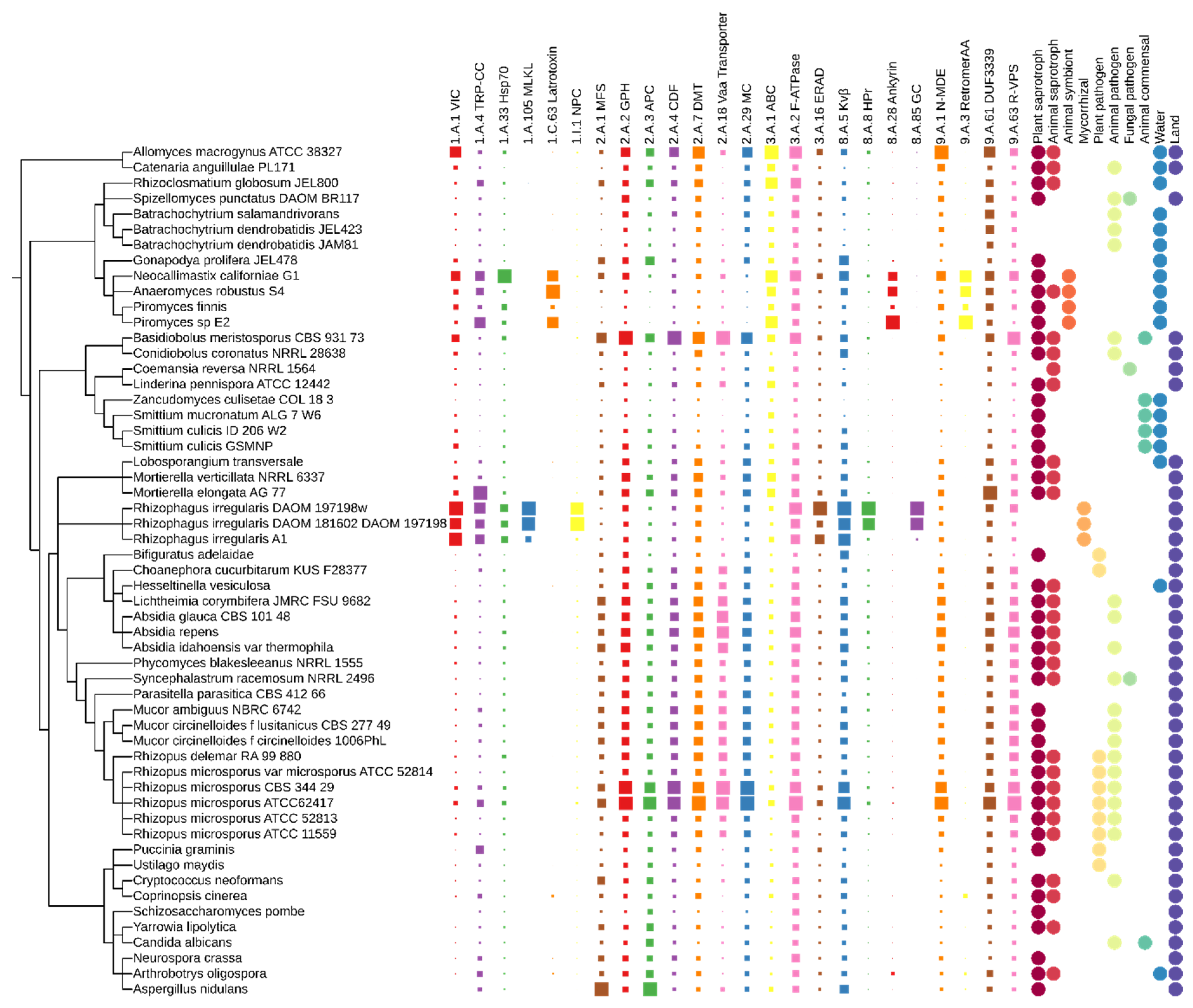
3.3. Transporters
3.3.1. Essential Transporters
3.3.2. Less Frequent Fungal Transporters
3.3.3. Uncharacterized Fungal Transporters
3.4. Protein Expansions and Ecology
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, Y.; Steenwyk, J.L.; Chang, Y.; Wang, Y.; James, T.Y.; Stajich, J.E.; Spatafora, J.W.; Groenewald, M.; Dunn, C.W.; Hittinger, C.T.; et al. A genome-scale phylogeny of the kingdom Fungi. Curr. Biol. 2021, 31, 1653–1665.e5. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Chávez, M.J.; Pérez-García, L.A.; Niño-Vega, G.A.; Mora-Montes, H.M. Fungal Strategies to Evade the Host Immune Recognition. J. Fungi 2017, 3, 51. [Google Scholar] [CrossRef]
- Kamel, L.; Tang, N.; Malbreil, M.; San Clemente, H.; Le Marquer, M.; Roux, C.; Frei dit Frey, N. The Comparison of Expressed Candidate Secreted Proteins from Two Arbuscular Mycorrhizal Fungi Unravels Common and Specific Molecular Tools to Invade Different Host Plants. Front. Plant Sci. 2017, 8, 124. [Google Scholar] [CrossRef]
- Hurdeal, V.G.; Gentekaki, E.; Hyde, K.D.; Jeewon, R. Where are the basal fungi? Current status on diversity, ecology, evolution, and taxonomy. Biologia 2021, 76, 421–440. [Google Scholar] [CrossRef]
- Kiss, E.; Hegedüs, B.; Virágh, M.; Varga, T.; Merényi, Z.; Kószó, T.; Bálint, B.; Prasanna, A.N.; Krizsán, K.; Kocsubé, S.; et al. Comparative genomics reveals the origin of fungal hyphae and multicellularity. Nat. Commun. 2019, 10, 4080. [Google Scholar] [CrossRef]
- Põlme, S.; Abarenkov, K.; Nilsson, R.H.; Lindahl, B.D.; Clemmensen, K.E.; Kauserud, H.; Nguyen, N.; Kjøller, R.; Bates, S.T.; Baldrian, P.; et al. FungalTraits: A user-friendly traits database of fungi and fungus-like stramenopiles. Fungal Divers. 2020, 105, 1–16. [Google Scholar] [CrossRef]
- Demuth, J.P.; Hahn, M.W. The life and death of gene families. Bioessays 2009, 31, 29–39. [Google Scholar] [CrossRef]
- Albertin, W.; Marullo, P. Polyploidy in fungi: Evolution after whole-genome duplication. Proc. Biol. Sci. 2012, 279, 2497–2509. [Google Scholar] [CrossRef]
- Spanu, P.D.; Abbott, J.C.; Amselem, J.; Burgis, T.A.; Soanes, D.M.; Stüber, K.; van Themaat, E.V.L.; Brown, J.K.; Butcher, S.A.; Gurr, S.J.; et al. Genome expansion and gene loss in powdery mildew fungi reveal tradeoffs in extreme parasitism. Science 2010, 330, 1543–1546. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Zhang, X.; Li, K.; Wang, C.; Cai, L.; Zhuang, W.; Xiang, M.; Liu, X. Introgression and gene family contraction drive the evolution of lifestyle and host shifts of hypocrealean fungi. Mycology 2018, 9, 176–188. [Google Scholar] [CrossRef] [PubMed]
- Tunlid, A.; Talbot, N.J. Genomics of parasitic and symbiotic fungi. Curr. Opin. Microbiol. 2002, 5, 513–519. [Google Scholar] [CrossRef]
- Hage, H.; Miyauchi, S.; Virágh, M.; Drula, E.; Min, B.; Chaduli, D.; Navarro, D.; Favel, A.; Norest, M.; Lesage-Meessen, L.; et al. Gene family expansions and transcriptome signatures uncover fungal adaptations to wood decay. Environ. Microbiol. 2021, 23, 5716–5732. [Google Scholar] [CrossRef]
- Andreis, F.C.; Schrank, A.; Thompson, C.E. Molecular evolution of Pr1 proteases depicts ongoing diversification in Metarhizium spp. Mol. Genet. Genom. 2019, 294, 901–917. [Google Scholar] [CrossRef] [PubMed]
- Sommer, B.; Overy, D.P.; Kerr, R.G. Identification and characterization of lipases from Malassezia restricta, a causative agent of dandruff. FEMS Yeast Res. 2015, 15, fov078. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.D.; Waller, M.; Barrett, A.J.; Bateman, A. MEROPS: The database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 2014, 42, D503–D509. [Google Scholar] [CrossRef]
- Lombard, V.; Ramulu, H.G.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef] [PubMed]
- Saier, M.H., Jr.; Reddy, V.S.; Moreno-Hagelsieb, G.; Hendargo, K.J.; Zhang, Y.; Iddamsetty, V.; Lam, K.J.K.; Tian, N.; Russum, S.; Wang, J.; et al. The Transporter Classification Database (TCDB): 2021 update. Nucleic Acids Res. 2021, 49, D461–D467. [Google Scholar] [CrossRef]
- Seppey, M.; Manni, M.; Zdobnov, E.M. BUSCO: Assessing Genome Assembly and Annotation Completeness. Methods Mol. Biol. 2019, 1962, 227–245. [Google Scholar]
- Ellinghaus, D.; Kurtz, S.; Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinform. 2008, 9, 18. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Potter, S.C.; Luciani, A.; Eddy, S.R.; Park, Y.; Lopez, R.; Finn, R.D. HMMER web server: 2018 update. Nucleic Acids Res. 2018, 46, W200–W204. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.D.; Tomii, K.; Katoh, K. Application of the MAFFT sequence alignment program to large data—Reexamination of the usefulness of chained guide trees. Bioinformatics 2016, 32, 3246–3251. [Google Scholar] [CrossRef]
- Frickey, T.; Lupas, A. CLANS: A Java application for visualizing protein families based on pairwise similarity. Bioinformatics 2004, 20, 3702–3704. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; Von Haeseler, A.; Lanfear, R. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.-J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Armenteros, J.J.A.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; Von Heijne, G.; Nielsen, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 2019, 37, 420–423. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Close, D.W.; D’Angelo, S.; Bradbury, A.R.M. A new family of β-helix proteins with similarities to the polysaccharide lyases. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 2583–2592. [Google Scholar] [CrossRef]
- Gryganskyi, A.P.; Golan, J.; Dolatabadi, S.; Mondo, S.; Robb, S.; Idnurm, A.; Muszewska, A.; Steczkiewicz, K.; Masonjones, S.; Liao, H.-L.; et al. Phylogenetic and Phylogenomic Definition of Species. G3 2018, 8, 2007–2018. [Google Scholar] [CrossRef]
- Kuuskeri, J.; Häkkinen, M.; Laine, P.; Smolander, O.P.; Tamene, F.; Miettinen, S.; Nousiainen, P.; Kemell, M.; Auvinen, P.; Lundell, T. Time-scale dynamics of proteome and transcriptome of the white-rot fungus Phlebia radiata: Growth on spruce wood and decay effect on lignocellulose. Biotechnol. Biofuels 2016, 9, 192. [Google Scholar] [CrossRef]
- Jensen, K.A., Jr.; Ryan, Z.C.; Vanden Wymelenberg, A.; Cullen, D.; Hammel, K.E. An NADH: Quinone oxidoreductase active during biodegradation by the brown-rot basidiomycete Gloeophyllum trabeum. Appl. Environ. Microbiol. 2002, 68, 2699–2703. [Google Scholar] [CrossRef]
- Pedrini, N.; Ortiz-Urquiza, A.; Huarte-Bonnet, C.; Fan, Y.; Juárez, M.P.; Keyhani, N.O. Tenebrionid secretions and a fungal benzoquinone oxidoreductase form competing components of an arms race between a host and pathogen. Proc. Natl. Acad. Sci. USA 2015, 112, E3651–E3660. [Google Scholar] [CrossRef] [PubMed]
- Rühl, M.; Majcherczyk, A.; Kües, U. Lcc1 and Lcc5 are the main laccases secreted in liquid cultures of Coprinopsis cinerea strains. Antonie Van Leeuwenhoek 2013, 103, 1029–1039. [Google Scholar] [CrossRef]
- Kameshwar, A.K.S.; Qin, W. Systematic review of publicly available non-Dikarya fungal proteomes for understanding their plant biomass-degrading and bioremediation potentials. Bioresour. Bioprocess. 2019, 6, 30. [Google Scholar] [CrossRef]
- Prakash, O.; Mahabare, K.; Yadav, K.K.; Sharma, R. Fungi from extreme environments: A potential source of laccases group of extremozymes. In Fungi in Extreme Environments: Ecological Role and Biotechnological Significance; Springer International Publishing: Cham, Switzerland, 2019; pp. 441–462. [Google Scholar]
- Momeni, M.H.; Fredslund, F.; Bissaro, B.; Raji, O.; Vuong, T.V.; Meier, S.; Nielsen, T.S.; Lombard, V.; Guigliarelli, B.; Biaso, F.; et al. Discovery of fungal oligosaccharide-oxidising flavo-enzymes with previously unknown substrates, redox-activity profiles and interplay with LPMOs. Nat Commun. 2021, 12, 2132. [Google Scholar] [CrossRef] [PubMed]
- Andlar, M.; Rezić, T.; Marđetko, N.; Kracher, D.; Ludwig, R.; Šantek, B. Lignocellulose degradation: An overview of fungi and fungal enzymes involved in lignocellulose degradation. Eng. Life Sci. 2018, 18, 768–778. [Google Scholar] [CrossRef] [PubMed]
- Nagy, L.G.; Riley, R.; Tritt, A.; Adam, C.; Daum, C.; Floudas, D.; Sun, H.; Yadav, J.S.; Pangilinan, J.; Larsson, K.H.; et al. Comparative Genomics of Early-Diverging Mushroom-Forming Fungi Provides Insights into the Origins of Lignocellulose Decay Capabilities. Mol. Biol. Evol. 2016, 33, 959–970. [Google Scholar] [CrossRef]
- Hess, M.; Paul, S.S.; Puniya, A.K.; Van Der Giezen, M.; Shaw, C.; Edwards, J.E.; Fliegerová, K. Anaerobic Fungi: Past, Present, and Future. Front. Microbiol. 2020, 11, 584893. [Google Scholar] [CrossRef]
- Murphy, C.L.; Youssef, N.H.; Hanafy, R.; Couger, M.B.; Stajich, J.E.; Wang, Y.; Baker, K.; Dagar, S.S.; Griffith, G.W.; Farag, I.F.; et al. Horizontal Gene Transfer as an Indispensable Driver for Evolution of Neocallimastigomycota into a Distinct Gut-Dwelling Fungal Lineage. Appl. Environ. Microbiol. 2019, 85, e00988-19. [Google Scholar] [CrossRef]
- Muszewska, A.; Piłsyk, S.; Perlińska-Lenart, U.; Kruszewska, J.S. Diversity of Cell Wall Related Proteins in Human Pathogenic Fungi. J. Fungi 2017, 4, 6. [Google Scholar] [CrossRef] [PubMed]
- Bellincampi, D.; Cervone, F.; Lionetti, V. Plant cell wall dynamics and wall-related susceptibility in plant-pathogen interactions. Front. Plant Sci. 2014, 5, 228. [Google Scholar] [CrossRef]
- Andrés, E.; Albesa-Jové, D.; Biarnés, X.; Moerschbacher, B.M.; Guerin, M.E.; Planas, A. Structural basis of chitin oligosaccharide deacetylation. Angew. Chem. Weinheim. Bergstr. Ger. 2014, 126, 7002–7007. [Google Scholar] [CrossRef]
- Lastovetsky, O.A.; Krasnovsky, L.D.; Qin, X.; Gaspar, M.L.; Gryganskyi, A.P.; Huntemann, M.; Clum, A.; Pillay, M.; Palaniappan, K.; Varghese, N.; et al. Molecular Dialogues between Early Divergent Fungi and Bacteria in an Antagonism versus a Mutualism. mBio 2020, 11, e02088-20. [Google Scholar] [CrossRef]
- Mélida, H.; Sain, D.; Stajich, J.E.; Bulone, V. Deciphering the uniqueness of Mucoromycotina cell walls by combining biochemical and phylogenomic approaches. Environ. Microbiol. 2015, 17, 1649–1662. [Google Scholar] [CrossRef] [PubMed]
- van den Brink, J.; de Vries, R.P. Fungal enzyme sets for plant polysaccharide degradation. Appl. Microbiol. Biotechnol. 2011, 91, 1477–1492. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Liu, H.; Wang, C.; Xu, J.-R. Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genom. 2013, 14, 274. [Google Scholar] [CrossRef]
- Pawłowska, J.; Okrasińska, A.; Kisło, K.; Aleksandrzak-Piekarczyk, T.; Szatraj, K.; Dolatabadi, S.; Muszewska, A. Carbon assimilation profiles of mucoralean fungi show their metabolic versatility. Sci. Rep. 2019, 9, 11864. [Google Scholar] [CrossRef]
- Mewis, K.; Lenfant, N.; Lombard, V.; Henrissat, B. Dividing the Large Glycoside Hydrolase Family 43 into Subfamilies: A Motivation for Detailed Enzyme Characterization. Appl. Environ. Microbiol. 2016, 82, 1686–1692. [Google Scholar] [CrossRef]
- Moussa, S.H.; Kuznetsov, V.; Tran, T.A.T.; Sacchettini, J.C.; Young, R. Protein determinants of phage T4 lysis inhibition. Protein Sci. 2012, 21, 571–582. [Google Scholar] [CrossRef]
- Briard, B.; Muszkieta, L.; Latgé, J.-P.; Fontaine, T. Galactosaminogalactan of Aspergillus fumigatus, a bioactive fungal polymer. Mycologia 2016, 108, 572–580. [Google Scholar] [CrossRef]
- Balestrini, R.; Bonfante, P. Cell wall remodeling in mycorrhizal symbiosis: A way towards biotrophism. Front. Plant Sci. 2014, 5, 237. [Google Scholar] [CrossRef]
- Hopke, A.; Brown, A.J.P.; Hall, R.A.; Wheeler, R.T. Dynamic Fungal Cell Wall Architecture in Stress Adaptation and Immune Evasion. Trends Microbiol. 2018, 26, 284–295. [Google Scholar] [CrossRef] [PubMed]
- Bamford, N.C.; Le Mauff, F.; Subramanian, A.S.; Yip, P.; Millán, C.; Zhang, Y.; Zacharias, C.; Forman, A.; Nitz, M.; Codée, J.D.; et al. Ega3 from the fungal pathogen is an endo-α-1,4-galactosaminidase that disrupts microbial biofilms. J. Biol. Chem. 2019, 294, 13833–13849. [Google Scholar] [CrossRef]
- Spatafora, J.W.; Chang, Y.; Benny, G.L.; Lazarus, K.; Smith, M.E.; Berbee, M.L.; Bonito, G.; Corradi, N.; Grigoriev, I.; Gryganskyi, A.; et al. A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 2016, 108, 1028–1046. [Google Scholar] [CrossRef]
- Han, B.; Zhou, K.; Li, Z.; Sun, B.; Ni, Q.; Meng, X.; Pan, G.; Li, C.; Long, M.; Li, T.; et al. Characterization of the First Fungal Glycosyl Hydrolase Family 19 Chitinase (NbchiA) from Nosema bombycis (Nb). J. Eukaryot. Microbiol. 2016, 63, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Yano, S.; Kanno, H.; Tsuhako, H.; Ogasawara, S.; Suyotha, W.; Konno, H.; Makabe, K.; Uechi, K.; Taira, T. Cloning, expression, and characterization of a GH 19-type chitinase with antifungal activity from Lysobacter sp. MK9-1. J. Biosci. Bioeng. 2021, 131, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Norice, C.T.; Smith, F.J., Jr.; Solis, N.; Filler, S.G.; Mitchell, A.P. Requirement for Candida albicans Sun41 in biofilm formation and virulence. Eukaryot. Cell 2007, 6, 2046–2055. [Google Scholar] [CrossRef]
- Schwerdt, J.; Qiu, H.; Shirley, N.; Little, A.; Bulone, V. Phylogenomic Analyses of Nucleotide-Sugar Biosynthetic and Interconverting Enzymes Illuminate Cell Wall Composition in Fungi. mBio 2021, 12, e03540-20. [Google Scholar] [CrossRef]
- Corfield, A.P.; Berry, M. Glycan variation and evolution in the eukaryotes. Trends Biochem. Sci. 2015, 40, 351–359. [Google Scholar] [CrossRef]
- Brockington, M.; Torelli, S.; Prandini, P.; Boito, C.; Dolatshad, N.F.; Longman, C.; Brown, S.; Muntoni, F. Localization and functional analysis of the LARGE family of glycosyltransferases: Significance for muscular dystrophy. Hum. Mol. Genet. 2005, 14, 657–665. [Google Scholar] [CrossRef] [PubMed]
- Valdivieso, M.H.; Henar Valdivieso, M.; Durán, A.; Roncero, C. Chitin synthases in yeast and fungi. Chitin Chitinases 1999, 87, 55–69. [Google Scholar] [CrossRef]
- Ikeda, Y.; Ohashi, T.; Tanaka, N.; Takegawa, K. Identification and characterization of a gene required for alpha1,2-mannose extension in the O-linked glycan synthesis pathway in Schizosaccharomyces pombe. FEMS Yeast Res. 2009, 9, 115–125. [Google Scholar] [CrossRef][Green Version]
- Sommer, U.; Liu, H.; Doering, T.L. An alpha-1,3-mannosyltransferase of Cryptococcus neoformans. J. Biol. Chem. 2003, 278, 47724–47730. [Google Scholar] [CrossRef]
- Yip, C.L.; Welch, S.K.; Klebl, F.; Gilbert, T.; Seidel, P.; Grant, F.J.; O’Hara, P.J.; MacKay, V.L. Cloning and analysis of the Saccharomyces cerevisiae MNN9 and MNN1 genes required for complex glycosylation of secreted proteins. Proc. Natl. Acad. Sci. USA 1994, 91, 2723–2727. [Google Scholar] [CrossRef]
- Holdener, B.C.; Haltiwanger, R.S. Protein O-fucosylation: Structure and function. Curr. Opin. Struct. Biol. 2019, 56, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Muszewska, A.; Okrasińska, A.; Steczkiewicz, K.; Drgas, O.; Orłowska, M.; Perlińska-Lenart, U.; Aleksandrzak-Piekarczyk, T.; Szatraj, K.; Zielenkiewicz, U.; Piłsyk, S.; et al. Metabolic Potential, Ecology and Presence of Associated Bacteria Is Reflected in Genomic Diversity of Mucoromycotina. Front. Microbiol. 2021, 12, 636986. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, P.M.; Andersen, M.R.; Kolenova, K.; Vankuyk, P.A.; Benoit, I.; Gruben, B.S.; Trejo-Aguilar, B.; Visser, H.; Van Solingen, P.; Pakula, T. Post-genomic insights into the plant polysaccharide degradation potential of Aspergillus nidulans and comparison to Aspergillus niger and Aspergillus oryzae. Fungal Genet. Biol. 2009, 46 (Suppl. 1), S161–S169. [Google Scholar] [CrossRef] [PubMed]
- Kohler, A.; Kuo, A.; Nagy, L.G.; Morin, E.; Barry, K.W.; Buscot, F.; Canbäck, B.; Choi, C.; Cichocki, N.; Clum, A.; et al. Convergent losses of decay mechanisms and rapid turnover of symbiosis genes in mycorrhizal mutualists. Nat. Genet. 2015, 47, 410–415. [Google Scholar] [CrossRef]
- McDonald, C.A.; Ellison, A.R.; Toledo, L.F.; James, T.Y.; Zamudio, K.R. Gene expression varies within and between enzootic and epizootic lineages of Batrachochytrium dendrobatidis (Bd) in the Americas. Fungal Biol. 2020, 124, 34–43. [Google Scholar] [CrossRef]
- Fisher, M.C.; Pasmans, F.; Martel, A. Virulence and Pathogenicity of Chytrid Fungi Causing Amphibian Extinctions. Annu. Rev. Microbiol. 2021, 75, 673–693. [Google Scholar] [CrossRef] [PubMed]
- Pellegrin, C.; Morin, E.; Martin, F.M.; Veneault-Fourrey, C. Comparative Analysis of Secretomes from Ectomycorrhizal Fungi with an Emphasis on Small-Secreted Proteins. Front. Microbiol. 2015, 6, 1278. [Google Scholar] [CrossRef] [PubMed]
- Llorens, C.; Futami, R.; Renaud, G.; Moya, A. Bioinformatic flowchart and database to investigate the origins and diversity of clan AA peptidases. Biol. Direct 2009, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Muszewska, A.; Hoffman-Sommer, M.; Grynberg, M. LTR Retrotransposons in Fungi. PLoS ONE 2011, 6, e29425. [Google Scholar] [CrossRef] [PubMed]
- Muszewska, A.; Steczkiewicz, K.; Stepniewska-Dziubinska, M.; Ginalski, K. Transposable elements contribute to fungal genes and impact fungal lifestyle. Sci. Rep. 2019, 9, 4307. [Google Scholar] [CrossRef]
- Gonçalves, A.P.; Heller, J.; Daskalov, A.; Videira, A.; Glass, N.L. Regulated Forms of Cell Death in Fungi. Front. Microbiol. 2017, 8, 1837. [Google Scholar] [CrossRef]
- Kamlangdee, N. Identifying Target Proteins of the CreB Deubiquitination Enzyme in the Fungus Aspergillus Nidulans. Ph.D. Thesis, University of Adelaide, Adelaide, Australia, 2007. [Google Scholar]
- Katz, M.E.; Bernardo, S.M.; Cheetham, B.F. The interaction of induction, repression and starvation in the regulation of extracellular proteases in Aspergillus nidulans: Evidence for a role for CreA in the response to carbon starvation. Curr. Genet. 2008, 54, 47–55. [Google Scholar] [CrossRef]
- Denton, J.A.; Kelly, J.M. Disruption of Trichoderma reesei cre2, encoding an ubiquitin C-terminal hydrolase, results in increased cellulase activity. BMC Biotechnol. 2011, 11, 103. [Google Scholar] [CrossRef]
- Mateus, I.D.; Rojas, E.C.; Savary, R.; Dupuis, C.; Masclaux, F.G.; Aletti, C.; Sanders, I.R. Coexistence of genetically different Rhizophagus irregularis isolates induces genes involved in a putative fungal mating response. ISME J. 2020, 14, 2381–2394. [Google Scholar] [CrossRef]
- Green, C.M.; Novikova, O.; Belfort, M. The dynamic intein landscape of eukaryotes. Mob. DNA 2018, 9, 4. [Google Scholar] [CrossRef] [PubMed]
- Xin, F.; Wang, S.; Song, L.; Liang, Q.; Qi, Q. Molecular identification and characterization of peptide: N-glycanase from Schizosaccharomyces pombe. Biochem. Biophys. Res. Commun. 2008, 368, 907–912. [Google Scholar] [CrossRef]
- Seppälä, S.; Lankiewicz, T.S.; Saxena, M.; Henske, J.K.; Salamov, A.A.; Grigoriev, I.V.; O’Malley, M.A. Genomic and proteomic biases inform metabolic engineering strategies for anaerobic fungi. Metab. Eng. Commun. 2020, 10, e00107. [Google Scholar]
- Pei, J.; Grishin, N.V. Type II CAAX prenyl endopeptidases belong to a novel superfamily of putative membrane-bound metalloproteases. Trends Biochem. Sci. 2001, 26, 275–277. [Google Scholar] [CrossRef]
- Steenbakkers, P.J.; Irving, J.A.; Harhangi, H.R.; Swinkels, W.J.; Akhmanova, A.; Dijkerman, R.; Jetten, M.S.; van der Drift, C.; Whisstock, J.C.; Camp, H.J.O.D. A serpin in the cellulosome of the anaerobic fungus Piromyces sp. strain E2. Mycol. Res. 2008, 112, 999–1006. [Google Scholar] [CrossRef]
- Janer, C.; Arigoni, F.; Lee, B.H.; Peláez, C.; Requena, T. Enzymatic ability of Bifidobacterium animalis subsp. lactis to hydrolyze milk proteins: Identification and characterization of endopeptidase O. Appl. Environ. Microbiol. 2005, 71, 8460–8465. [Google Scholar] [CrossRef]
- Lindner, H.A.; Lunin, V.V.; Alary, A.; Hecker, R.; Cygler, M.; Ménard, R. Essential roles of zinc ligation and enzyme dimerization for catalysis in the aminoacylase-1/M20 family. J. Biol. Chem. 2003, 278, 44496–44504. [Google Scholar] [CrossRef]
- Zuo, S.; Guo, Q.; Ling, C.; Chang, Y.H. Evidence that two zinc fingers in the methionine aminopeptidase from Saccharomyces cerevisiae are important for normal growth. Mol. Gen. Genet. 1995, 246, 247–253. [Google Scholar] [CrossRef]
- Cottrell, G.S.; Hooper, N.M.; Turner, A.J. Cloning, expression, and characterization of human cytosolic aminopeptidase P: A single manganese(II)-dependent enzyme. Biochemistry 2000, 39, 15121–15128. [Google Scholar] [CrossRef]
- Knedlík, T.; Vorlová, B.; Navrátil, V.; Tykvart, J.; Sedlák, F.; Vaculín, Š.; Franěk, M.; Šácha, P.; Konvalinka, J. Mouse glutamate carboxypeptidase II (GCPII) has a similar enzyme activity and inhibition profile but a different tissue distribution to human GCPII. FEBS Open Bio 2017, 7, 1362–1378. [Google Scholar] [CrossRef]
- Cheng, Y.; Zak, O.; Aisen, P.; Harrison, S.C.; Walz, T. Structure of the human transferrin receptor-transferrin complex. Cell 2004, 116, 565–576. [Google Scholar] [CrossRef]
- Rutherford, J.C. The emerging role of urease as a general microbial virulence factor. PLoS Pathog. 2014, 10, e1004062. [Google Scholar] [CrossRef]
- Hu, P.; Ding, H.; Shen, L.; He, G.-J.; Liu, H.; Tian, X.; Tao, C.; Bai, X.; Liang, J.; Jin, C.; et al. A unique cell wall synthetic response evoked by glucosamine determines pathogenicity-associated fungal cellular differentiation. PLoS Genet. 2021, 17, e1009817. [Google Scholar] [CrossRef] [PubMed]
- Dubovenko, A.G.; Dunaevsky, Y.E.; Belozersky, M.A.; Oppert, B.; Lord, J.C.; Elpidina, E.N. Trypsin-like proteins of the fungi as possible markers of pathogenicity. Fungal Biol. 2010, 114, 151–159. [Google Scholar] [CrossRef]
- Leger, R.J.S.; Joshi, L.; Roberts, D.W. Adaptation of proteases and carbohydrates of saprophytic, phytopathogenic and entomopathogenic fungi to the requirements of their ecological niches. Microbiology 1997, 143 Pt 6, 1983–1992. [Google Scholar] [CrossRef]
- Muszewska, A.; Stepniewska-Dziubinska, M.M.; Steczkiewicz, K.; Pawlowska, J.; Dziedzic, A.; Ginalski, K. Fungal lifestyle reflected in serine protease repertoire. Sci. Rep. 2017, 7, 9147. [Google Scholar] [CrossRef]
- Li, Q.; Zhang, N.; Zhang, L.; Ma, H. Differential evolution of members of the rhomboid gene family with conservative and divergent patterns. New Phytol. 2015, 206, 368–380. [Google Scholar] [CrossRef] [PubMed]
- Marger, M.D.; Saier, M.H. A major superfamily of transmembrane facilitators that catalyse uniport, symport and antiport. Trends Biochem. Sci. 1993, 18, 13–20. [Google Scholar] [CrossRef]
- Celenza, J.L.; Marshall-Carlson, L.; Carlson, M. The yeast SNF3 gene encodes a glucose transporter homologous to the mammalian protein. Proc. Natl. Acad. Sci. USA 1988, 85, 2130–2134. [Google Scholar] [CrossRef]
- Heiland, S.; Radovanovic, N.; Höfer, M.; Winderickx, J.; Lichtenberg, H. Multiple hexose transporters of Schizosaccharomyces pombe. J. Bacteriol. 2000, 182, 2153–2162. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.D.; Dickson, R.C. Primary structure of the lactose permease gene from the yeast Kluyveromyces lactis. Presence of an unusual transcript structure. J. Biol. Chem. 1988, 263, 16696–16703. [Google Scholar] [CrossRef]
- Yao, B.; Sollitti, P.; Marmur, J. Primary structure of the maltose-permease-encoding gene of Saccharomyces carlsbergensis. Gene 1989, 79, 189–197. [Google Scholar] [CrossRef]
- Mannhaupt, G. What’s in the genome of a filamentous fungus? Analysis of the Neurospora genome sequence. Nucleic Acids Res. 2003, 31, 1944–1954. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.M.; George, A.M. The ABC transporter structure and mechanism: Perspectives on recent research. Cell. Mol. Life Sci. 2004, 61, 682–699. [Google Scholar] [CrossRef] [PubMed]
- Fedorova, N.D.; Khaldi, N.; Joardar, V.S.; Maiti, R.; Amedeo, P.; Anderson, M.J.; Crabtree, J.; Silva, J.C.; Badger, J.H.; Albarraq, A.; et al. Genomic islands in the pathogenic filamentous fungus Aspergillus fumigatus. PLoS Genet. 2008, 4, e1000046. [Google Scholar] [CrossRef]
- Ghaemmaghami, S.; Huh, W.-K.; Bower, K.; Howson, R.W.; Belle, A.; Dephoure, N.; O’Shea, E.K.; Weissman, J.S. Global analysis of protein expression in yeast. Nature 2003, 425, 737–741. [Google Scholar] [CrossRef]
- Bowman, S.; Churcher, C.; Badcock, K.; Brown, D.; Chillingworth, T.; Connor, R.; Dedman, K.; Devlin, K.; Gentles, S.; Hamlin, N.; et al. The nucleotide sequence of Saccharomyces cerevisiae chromosome XIII. Nature 1997, 387, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.K.; Chiba, Y.; Takeuchi, M.; Jigami, Y. Characterization of Yeast Yea4p, a uridine diphosphate-N-acetylglucosamine transporter localized in the endoplasmic reticulum and required for chitin synthesis. J. Biol. Chem. 2000, 275, 13580–13587. [Google Scholar] [CrossRef]
- Nakanishi, H.; Nakayama, K.-I.; Yokota, A.; Tachikawa, H.; Takahashi, N.; Jigami, Y. Hut1 proteins identified inSaccharomyces cerevisiae andSchizosaccharomyces pombe are functional homologues involved in the protein-folding process at the endoplasmic reticulum. Yeast 2001, 18, 543–554. [Google Scholar] [CrossRef]
- Nishikawa, A.; Poster, J.B.; Jigami, Y.; Dean, N. Molecular and phenotypic analysis of CaVRG4, encoding an essential Golgi apparatus GDP-mannose transporter. J. Bacteriol. 2002, 184, 29–42. [Google Scholar] [CrossRef]
- Wood, V.; Gwilliam, R.; Rajandream, M.-A.; Lyne, M.; Lyne, R.; Stewart, A.; Sgouros, J.; Peat, N.; Hayles, J.; Baker, S.; et al. The genome sequence of Schizosaccharomyces pombe. Nature 2002, 415, 871–880. [Google Scholar] [CrossRef]
- Jacq, C.; Alt-Mörbe, J.; Andre, B.; Arnold, W.; Bahr, A.; Ballesta, J.P.G.; Bargues, M.; Baron, L.; Becker, A.; Biteau, N.; et al. The nucleotide sequence of Saccharomyces cerevisiae chromosome IV. Nature 1997, 387, 75–78. [Google Scholar] [CrossRef] [PubMed]
- Goytain, A.; Hines, R.M.; El-Husseini, A.; Quamme, G.A. NIPA1(SPG6), the basis for autosomal dominant form of hereditary spastic paraplegia, encodes a functional Mg2+ transporter. J. Biol. Chem. 2007, 282, 8060–8068. [Google Scholar] [CrossRef] [PubMed]
- Uemura, S.; Kihara, A.; Inokuchi, J.-I.; Igarashi, Y. Csg1p and newly identified Csh1p function in mannosylinositol phosphorylceramide synthesis by interacting with Csg2p. J. Biol. Chem. 2003, 278, 45049–45055. [Google Scholar] [CrossRef]
- Greenwald, J.; Buhtz, C.; Ritter, C.; Kwiatkowski, W.; Choe, S.; Maddelein, M.-L.; Ness, F.; Cescau, S.; Soragni, A.; Leitz, D.; et al. The mechanism of prion inhibition by HET-S. Mol. Cell. 2010, 38, 889–899. [Google Scholar] [CrossRef]
- Bankaitis, V.A.; Johnson, L.M.; Emr, S.D. Isolation of yeast mutants defective in protein targeting to the vacuole. Proc. Natl. Acad. Sci. USA 1986, 83, 9075–9079. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, D.L.; Rizzo, J.; Joffe, L.S.; Godinho, R.M.C.; Rodrigues, M.L. Where do they come from and where do they go: Candidates for regulating extracellular vesicle formation in fungi. Int. J. Mol. Sci. 2013, 14, 9581–9603. [Google Scholar] [CrossRef]
- Zhu, X.-M.; Li, L.; Wu, M.; Liang, S.; Shi, H.-B.; Liu, X.-H.; Lin, F.-C. Current opinions on autophagy in pathogenicity of fungi. Virulence 2019, 10, 481–489. [Google Scholar] [CrossRef]
- Trip, H.; Evers, M.E.; Driessen, A.J.M. PcMtr, an aromatic and neutral aliphatic amino acid permease of Penicillium chrysogenum. Biochim. Biophys. Acta 2004, 1667, 167–173. [Google Scholar] [CrossRef]
- Lewis, A.; McCrossan, Z.A.; Manville, R.W.; Popa, M.O.; Cuello, L.G.; Goldstein, S.A.N. TOK channels use the two gates in classical K channels to achieve outward rectification. FASEB J. 2020, 34, 8902–8919. [Google Scholar] [CrossRef]
- Paidhungat, M.; Garrett, S. A homolog of mammalian, voltage-gated calcium channels mediates yeast pheromone-stimulated Ca2+ uptake and exacerbates the cdc1(Ts) growth defect. Mol. Cell. Biol. 1997, 17, 6339–6347. [Google Scholar] [CrossRef]
- Denis, V.; Cyert, M.S. Internal Ca2+ release in yeast is triggered by hypertonic shock and mediated by a TRP channel homologue. J. Cell Biol. 2002, 156, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Protchenko, O.; Rodriguez-Suarez, R.; Androphy, R.; Bussey, H.; Philpott, C.C. A screen for genes of heme uptake identifies the FLC family required for import of FAD into the endoplasmic reticulum. J. Biol. Chem. 2006, 281, 21445–21457. [Google Scholar] [CrossRef]
- Palmer, C.P.; Aydar, E.; Djamgoz, M.B.A. A microbial TRP-like polycystic-kidney-disease-related ion channel gene. Biochem. J. 2005, 387, 211–219. [Google Scholar] [CrossRef]
- Pawlowska, T.E.; Taylor, J.W. Organization of genetic variation in individuals of arbuscular mycorrhizal fungi. Nature 2004, 427, 733–737. [Google Scholar] [CrossRef] [PubMed]
- Reinders, A.; Ward, J.M. Functional characterization of the alpha-glucoside transporter Sut1p from Schizosaccharomyces pombe, the first fungal homologue of plant sucrose transporters. Mol. Microbiol. 2001, 39, 445–454. [Google Scholar] [CrossRef]
- Gawaz, M.; Douglas, M.G.; Klingenberg, M. Structure-function studies of adenine nucleotide transport in mitochondria. II. Biochemical analysis of distinct AAC1 and AAC2 proteins in yeast. J. Biol. Chem. 1990, 265, 14202–14208. [Google Scholar] [CrossRef]
- Kakhniashvili, D.; Mayor, J.A.; Gremse, D.A.; Xu, Y.; Kaplan, R.S. Identification of a novel gene encoding the yeast mitochondrial dicarboxylate transport protein via overexpression, purification, and characterization of its protein product. J. Biol. Chem. 1997, 272, 4516–4521. [Google Scholar] [CrossRef]
- Tzagoloff, A.; Jang, J.; Glerum, D.M.; Wu, M. FLX1 codes for a carrier protein involved in maintaining a proper balance of flavin nucleotides in yeast mitochondria. J. Biol. Chem. 1996, 271, 7392–7397. [Google Scholar] [CrossRef]
- Kim, H.; Melén, K.; Osterberg, M.; von Heijne, G. A global topology map of the Saccharomyces cerevisiae membrane proteome. Proc. Natl. Acad. Sci. USA 2006, 103, 11142–11147. [Google Scholar] [CrossRef]
- Kolaj-Robin, O.; Russell, D.; Hayes, K.A.; Pembroke, J.T.; Soulimane, T. Cation Diffusion Facilitator family: Structure and function. FEBS Lett. 2015, 589, 1283–1295. [Google Scholar] [CrossRef]
- Swanson, R.; Locher, M.; Hochstrasser, M. A conserved ubiquitin ligase of the nuclear envelope/endoplasmic reticulum that functions in both ER-associated and Matalpha2 repressor degradation. Genes Dev. 2001, 15, 2660–2674. [Google Scholar] [CrossRef]
- Dibrova, D.V.; Galperin, M.Y.; Mulkidjanian, A.Y. Characterization of the N-ATPase, a distinct, laterally transferred Na -translocating form of the bacterial F-type membrane ATPase. Bioinformatics 2010, 26, 1473–1476. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sun, L.; Wang, H.; Wang, Z.; He, S.; Chen, S.; Liao, D.; Wang, L.; Yan, J.; Liu, W.; Lei, X.; et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell 2012, 148, 213–227. [Google Scholar] [CrossRef]
- Radons, J. The human HSP70 family of chaperones: Where do we stand? Cell Stress Chaperones 2016, 21, 379–404. [Google Scholar] [CrossRef]
- Bussink, H.-J.; Clark, A.; Oliver, R. The Cladosporium Fulvum Bap1 Gene: Evidence for a Novel Class of Yap-related Transcription Factors with Ankyrin Repeats in Phytopathogenic Fungi. Eur. J. Plant Pathol. 2001, 107, 655–659. [Google Scholar] [CrossRef]
- Peleg, Y.; Aramayo, R.; Kang, S.; Hall, J.G.; Metzenberg, R.L. NUC-2, a component of the phosphate-regulated signal transduction pathway inNeurospora crassa, is an ankyrin repeat protein. Mol. Gen. Genet. 1996, 252, 709–716. [Google Scholar] [CrossRef]
- Ichtchenko, K.; Khvotchev, M.; Kiyatkin, N.; Simpson, L.; Sugita, S.; Südhof, T.C. alpha-latrotoxin action probed with recombinant toxin: Receptors recruit alpha-latrotoxin but do not transduce an exocytotic signal. EMBO J. 1998, 17, 6188–6199. [Google Scholar] [CrossRef] [PubMed]
- Mosavi, L.K.; Cammett, T.J.; Desrosiers, D.C.; Peng, Z.-Y. The ankyrin repeat as molecular architecture for protein recognition. Protein Sci. 2004, 13, 1435–1448. [Google Scholar] [CrossRef] [PubMed]
- Rettig, J.; Heinemann, S.H.; Wunder, F.; Lorra, C.; Parcej, D.N.; Dolly, J.O.; Pongs, O. Inactivation properties of voltage-gated K channels altered by presence of β-subunit. Nature 1994, 369, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Postma, P.W.; Lengeler, J.W.; Jacobson, G.R. Phosphoenolpyruvate: Carbohydrate phosphotransferase systems of bacteria. Microbiol. Rev. 1993, 57, 543–594. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, M. Molecular Physiology of Membrane Guanylyl Cyclase Receptors. Physiol. Rev. 2016, 96, 751–804. [Google Scholar] [CrossRef] [PubMed]
- Rincón, E.; Santos, T.; Ávila-Flores, A.; Albar, J.P.; Lalioti, V.; Lei, C.; Hong, W.; Mérida, I. Proteomics Identification of Sorting Nexin 27 as a Diacylglycerol Kinase ζ-associated Protein. Mol. Cell. Proteom. 2007, 6, 1073–1087. [Google Scholar] [CrossRef] [PubMed]
- Hage, H.; Rosso, M.-N. Evolution of Fungal Carbohydrate-Active Enzyme Portfolios and Adaptation to Plant Cell-Wall Polymers. J. Fungi 2021, 7, 185. [Google Scholar] [CrossRef]
- Banasiak, J.; Jamruszka, T.; Murray, J.D.; Jasiński, M. A roadmap of plant membrane transporters in arbuscular mycorrhizal and legume-rhizobium symbioses. Plant Physiol. 2021, 187, 2071–2091. [Google Scholar] [CrossRef]
- Ene, I.V.; Walker, L.A.; Schiavone, M.; Lee, K.K.; Martin-Yken, H.; Dague, E.; Gow, N.A.; Munro, C.A.; Brown, A.J. Cell Wall Remodeling Enzymes Modulate Fungal Cell Wall Elasticity and Osmotic Stress Resistance. mBio 2015, 6, e00986. [Google Scholar] [CrossRef]
- Rosenblum, E.B.; Poorten, T.J.; Joneson, S.; Settles, M. Substrate-specific gene expression in Batrachochytrium dendrobatidis, the chytrid pathogen of amphibians. PLoS ONE 2012, 7, e49924. [Google Scholar] [CrossRef]
- Monod, M. Secreted proteases from dermatophytes. Mycopathologia 2008, 166, 285–294. [Google Scholar] [CrossRef]
- St Leger, R.J.; Cooper, R.M.; Charnley, A.K. Cuticle-degrading enzymes of entomopathogenic fungi: Regulation of production of chitinolytic enzymes. Microbiology 1986, 132, 1509–1517. [Google Scholar] [CrossRef][Green Version]
- Xiong, J.; Feng, J.; Yuan, D.; Zhou, J.; Miao, W. Tracing the structural evolution of eukaryotic ATP binding cassette transporter superfamily. Sci. Rep. 2015, 5, 16724. [Google Scholar] [CrossRef]
- Vishwakarma, P.; Banerjee, A.; Pasrija, R.; Prasad, R.; Lynn, A.M. Phylogenetic and conservation analyses of MFS transporters. 3 Biotech 2018, 8, 462. [Google Scholar] [CrossRef] [PubMed]
- Solomon, K.V.; Haitjema, C.H.; Henske, J.K.; Gilmore, S.P.; Borges-Rivera, D.; Lipzen, A.; Brewer, H.M.; Purvine, S.O.; Wright, A.T.; Theodorou, M.K.; et al. Early-branching gut fungi possess a large, comprehensive array of biomass-degrading enzymes. Science 2016, 351, 1192–1195. [Google Scholar] [CrossRef] [PubMed]
| MEROPS | ||
|---|---|---|
| MEROPS ID | Predicted Name | Putative Role in Fungi |
| A01 | endopeptidase | Nutrition, pathogenicity |
| A28 | retropepsin | Unknown |
| C110 | peptide-N(4)-(N-acetyl-β-glucosaminyl) asparagine amidase | Protein degradation |
| C14 | caspases | Controlled cell death |
| C19 | ubiquitin-carboxyl-hydrolase | Nutrition |
| C46 | hint domain containing | Putative mating response |
| G05 | type II CAAX-prenyl-endopeptidase | Protein modification |
| I04 | serpin | Peptidase inhibitor |
| M13 | unknown peptidase | Nutrition |
| M20 | N-acyl-l-amino-acid-amidohydrolase | Nutrition |
| unknown peptidase | Unknown | |
| M24 | MAP1 | Maturation of the nascent polypeptide during translation |
| MAP2 | ||
| Xaa-Pro aminopeptidase | Nutrition | |
| M28 | glutamate carboxypeptidase | Unknown |
| M36 | fungalysin | Nutrition, pathogenicity |
| M38 | allantoinase | Uric acid degradation pathway, pathogenicity |
| unknown peptidase | ||
| dihydroorotase | ||
| guanine deaminase | ||
| urease | ||
| N-acetylglucosamine-6-phosphate-deacetylase | Chitin degradation | |
| S01 | serine protease (extracellular) | Nutrition |
| S08 | subtilisin | Nutrition |
| S54 | rhomboid proteases | Mitochondrial endopeptidase |
| U69 | polysaccharide lyases | Nutrition |
| CAZY | ||
| CAZY ID | Predicted Name | Putative Role in Fungi |
| AA3 | glucose-methanol-choline (GMC) oxidoreductases | Nutrition |
| AA5 | galactose oxidase | Nutrition |
| AA6 | 1,4-benzoquinone reductase | Nutrition, interaction with insects |
| AA7 | glucooligosaccharide-oxidase/chitooligosaccharide-oxidase/cellooligosaccharide-dehydrogenase | Cell wall remodelling |
| CE1 | esterase domain containing | Nutrition |
| feruloyl/acetylxylan esterase | Nutrition | |
| rhamnogalacturonan-acetylesterase | Nutrition | |
| CE12 | rhamnogalacturonan-acetylesterase | Nutrition |
| CE16 | acetylesterase | Nutrition |
| CE2 | acetyl-xylan esterase | Nutrition |
| CE4 | chitin deacetylase | Cell wall remodelling |
| CE6 | acetyl-xylan esterase | Nutrition |
| GH10 | xylanase | Nutrition |
| GH109 | α-N-acetylgalactosaminidase | Glycolipids modification |
| GH11 | endo-1-4-β-xylanase | Nutrition |
| GH114 | α-galactosidase | Nutrition |
| GH132 | β-glucosidase | Pathogenicity |
| GH15 | glucoamylase | Nutrition |
| GH16 | endo-β-glucanase | Nutrition |
| GH17 | glucan-1,3-β-glucosidase | Cell wall remodelling |
| GH18 | chitinase | Cell wall remodelling |
| GH19 | chitinase | Interaction with insects |
| GH24 | lysozyme | Cell wall remodelling |
| GH26 | endo-1,4-β-mannosidase | Cell wall remodelling, nutrition |
| GH3 | β-glucosidase | Nutrition |
| GH43 | arabinase/levansucrase/invertase | Nutrition |
| GH45 | endoglucanase | Nutrition |
| GH48 | cellulase | Nutrition |
| GH5 | cellulase | Nutrition |
| exo-β-1,3-glucanase | Cell wall remodelling | |
| GH6 | cellobiohydrolase | Nutrition |
| endoglucanase | Nutrition | |
| GH72 | 1,3-β-glucanosyltransferase | Cell wall remodelling |
| GH9 | cellulase | Nutrition |
| GT1 | UDP-glucuronosyltransferase | Cell wall remodelling |
| GT15 | α-1,2-mannosyltransferase | Cell wall remodelling, pathogenicity |
| GT17 | β-1,4-mannosyl-glycoprotein-β-1,4-N-acetylglucosaminyltransferase | Cell wall remodelling |
| GT2 | chitin synthase 2 | Cell wall remodelling |
| chitin synthase 6 | Cell wall remodelling | |
| GT34 | galactomannan-α-1,6-galactosyltransferase/xyloglucan-α-1,6-xylosyltransferase/α-1,2-galactosyltransferase | Cell wall remodelling |
| GT49 | β-1,3-N-acetylglucosaminyltransferase | Glycosylation |
| GT68 | GT68_protein-O-α-fucosyltransferase | Protein modification |
| GT69 | α-1,3-mannosyltransferase | Cell wall remodelling, pathogenicity |
| GT71 | α-mannosyltransferase | Cell wall remodelling, protein modification |
| GT77 | α-xylosyltransferase/α-1,3-galactosyltransferase/arabinosyltransferase | Protein modification |
| PL1 | pectin lyase | Nutrition |
| PL3 | pectate lyase | Nutrition |
| PL4 | rhamnogalacturonan endolyase | Nutrition |
| TCDB | ||
| TCDB ID | Family Name | Putative Role in Fungi |
| 1.A.1 | The Voltage-gated Ion Channel (VIC) Superfamily | Response to stress |
| 1.A.105 | The Mixed Lineage Kinase Domain-like (MLKL) Family | Programmed cell death |
| 1.A.33 | The Cation Channel-forming Heat Shock Protein-70 (Hsp70) Family | Response to stress |
| 1.A.4 | The Transient Receptor Potential Ca2+ Channel (TRP-CC) Family | Response to stress |
| 1.C.63 | The α-Latrotoxin (Latrotoxin) Family | Unknown |
| 1.I.1 | The Nuclear Pore Complex (NPC) Family [formerly 1.A.75] | Unknown |
| 2.A.1 | The Major Facilitator Superfamily (MFS) | Nutrition, drug and metabolites transport |
| 2.A.18 | Vacuolar amino acid transporter | Nutrition |
| 2.A.2 | The Glycoside-Pentoside-Hexuronide (GPH): Cation Symporter Family | Nutrition |
| 2.A.29 | The Mitochondrial Carrier (MC) Family | Molecules transfer to mitochondria |
| 2.A.3 | The Amino Acid-Polyamine-Organocation (APC) Superfamily | Nutrition |
| 2.A.4 | The Cation Diffusion Facilitator (CDF) Family | Heavy metal transport |
| 2.A.7 | The Drug/Metabolite Transporter (DMT) Superfamily | Drug/ion resistance, cell wall remodelling |
| 3.A.1 | The ATP-binding Cassette (ABC) Superfamily | Essential for many processes in the cell |
| 3.A.16 | The Endoplasmic Reticular Retrotranslocon (ER-RT or ERAD) Family | Protein degradation |
| 3.A.2 | The H+- or Na+-translocating F-type, V-type and A-type ATPase (F-ATPase) Superfamily | Decomposition of ATP to ADP |
| 8.A.28 | The Ankyrin (Ankyrin) Family | Unknown |
| 8.A.5 | The Voltage-gated K+ Channel β-subunit (Kvβ) Family | Unknown |
| 8.A.8 | The Phosphotransferase System HPr (HPr) Family | Unknown |
| 8.A.85 | The Guanylate Cyclase (GC) Family | Unknown |
| 9.A.1 | The Non ABC Multidrug Exporter (N-MDE) Family | Drug resistance |
| 9.A.3 | The Retromer Assembly Apparatus (RetromerAA) Family | Protein recycling |
| 9.A.63 | The Retromer-dependent Vacuolar Protein Sorting (R-VPS) Family | Intracellular sorting |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Orłowska, M.; Muszewska, A. In Silico Predictions of Ecological Plasticity Mediated by Protein Family Expansions in Early-Diverging Fungi. J. Fungi 2022, 8, 67. https://doi.org/10.3390/jof8010067
Orłowska M, Muszewska A. In Silico Predictions of Ecological Plasticity Mediated by Protein Family Expansions in Early-Diverging Fungi. Journal of Fungi. 2022; 8(1):67. https://doi.org/10.3390/jof8010067
Chicago/Turabian StyleOrłowska, Małgorzata, and Anna Muszewska. 2022. "In Silico Predictions of Ecological Plasticity Mediated by Protein Family Expansions in Early-Diverging Fungi" Journal of Fungi 8, no. 1: 67. https://doi.org/10.3390/jof8010067
APA StyleOrłowska, M., & Muszewska, A. (2022). In Silico Predictions of Ecological Plasticity Mediated by Protein Family Expansions in Early-Diverging Fungi. Journal of Fungi, 8(1), 67. https://doi.org/10.3390/jof8010067








