Molecular and Morphological Assessment of Septoria Species Associated with Ornamental Plants in Yunnan Province, China
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungus Collection and Isolation
2.2. Morphological Studies
2.3. DNA Extraction, Amplification (PCR), and Sequencing
2.4. Phylogenetic Analyses
2.5. Genealogical Concordance Phylogenetic Species Recognition Analysis
3. Results
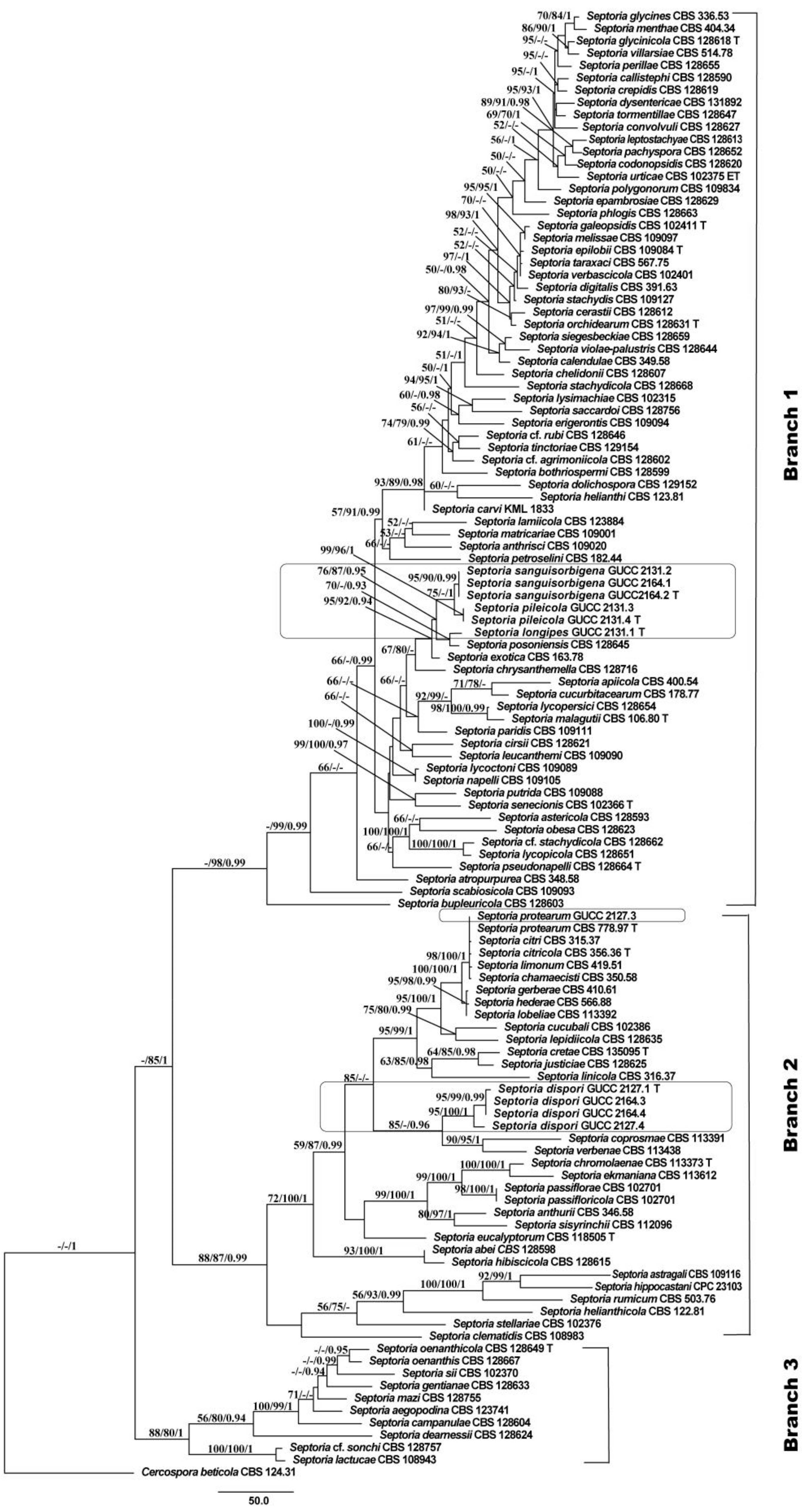
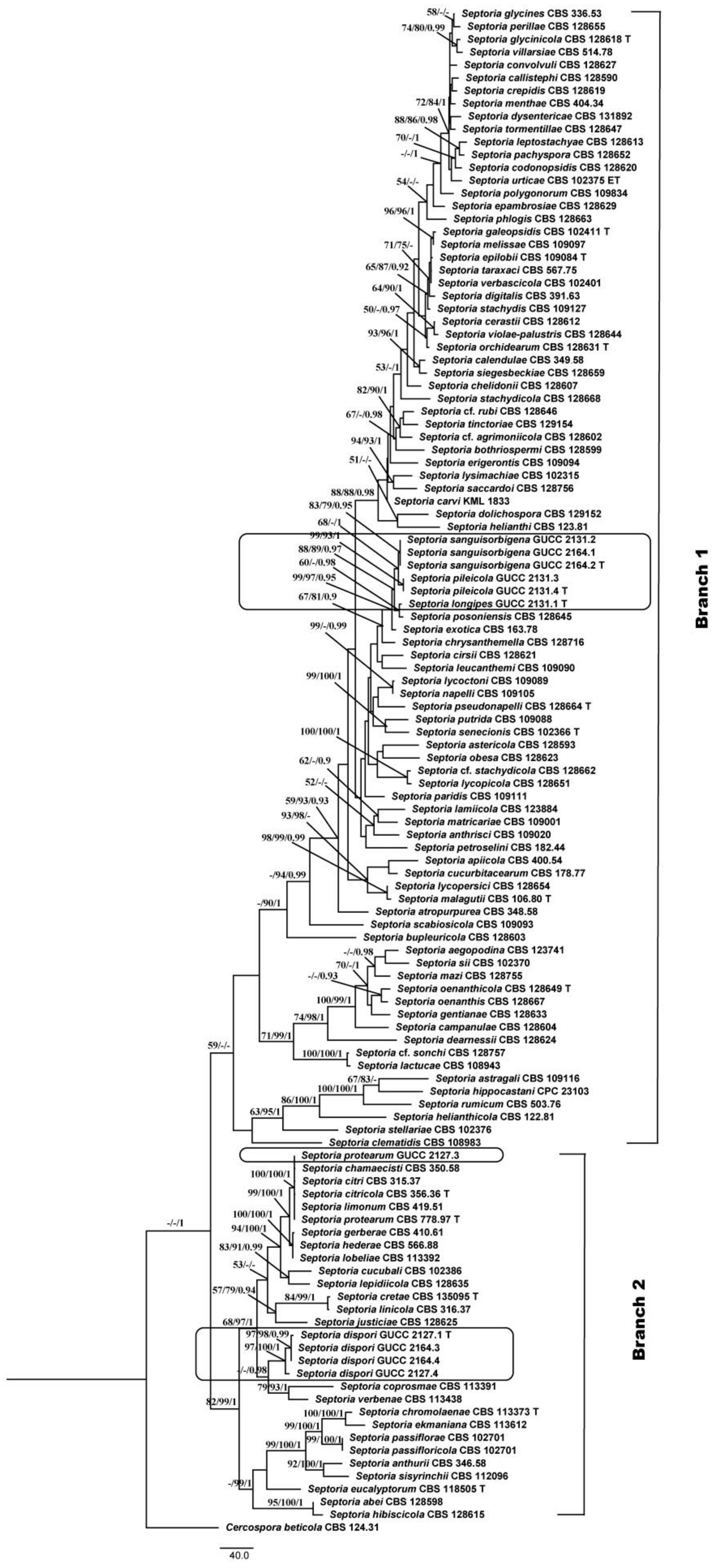
3.1. Phylogenetic Analyses
3.2. Genealogical Concordance Phylogenetic Species Recognition
3.3. Taxonomy
- (1)
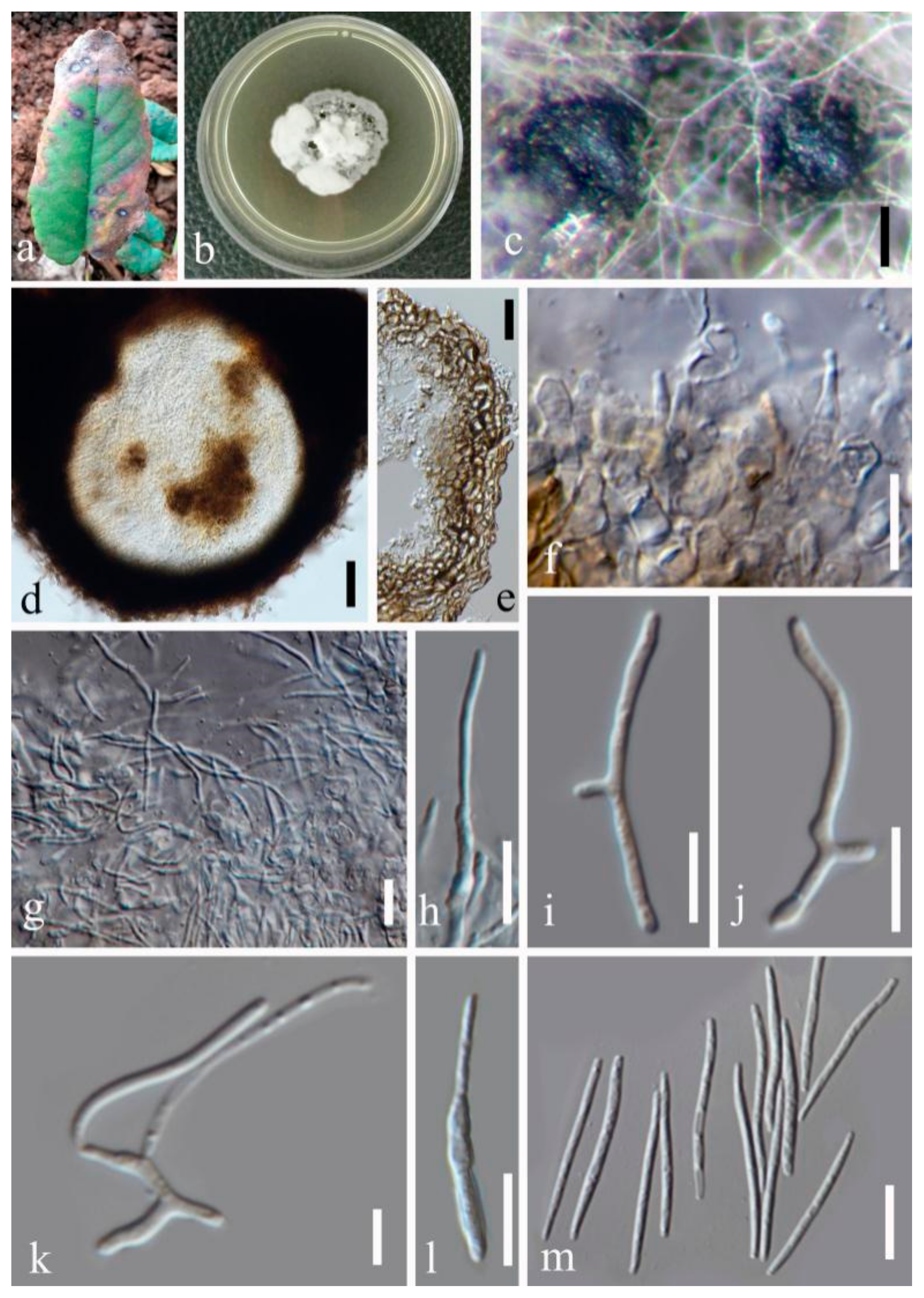
- Septoria sanguisorbigena Y.Y. An & Yong Wang bis, sp. nov. (Figure 3)
- (2)
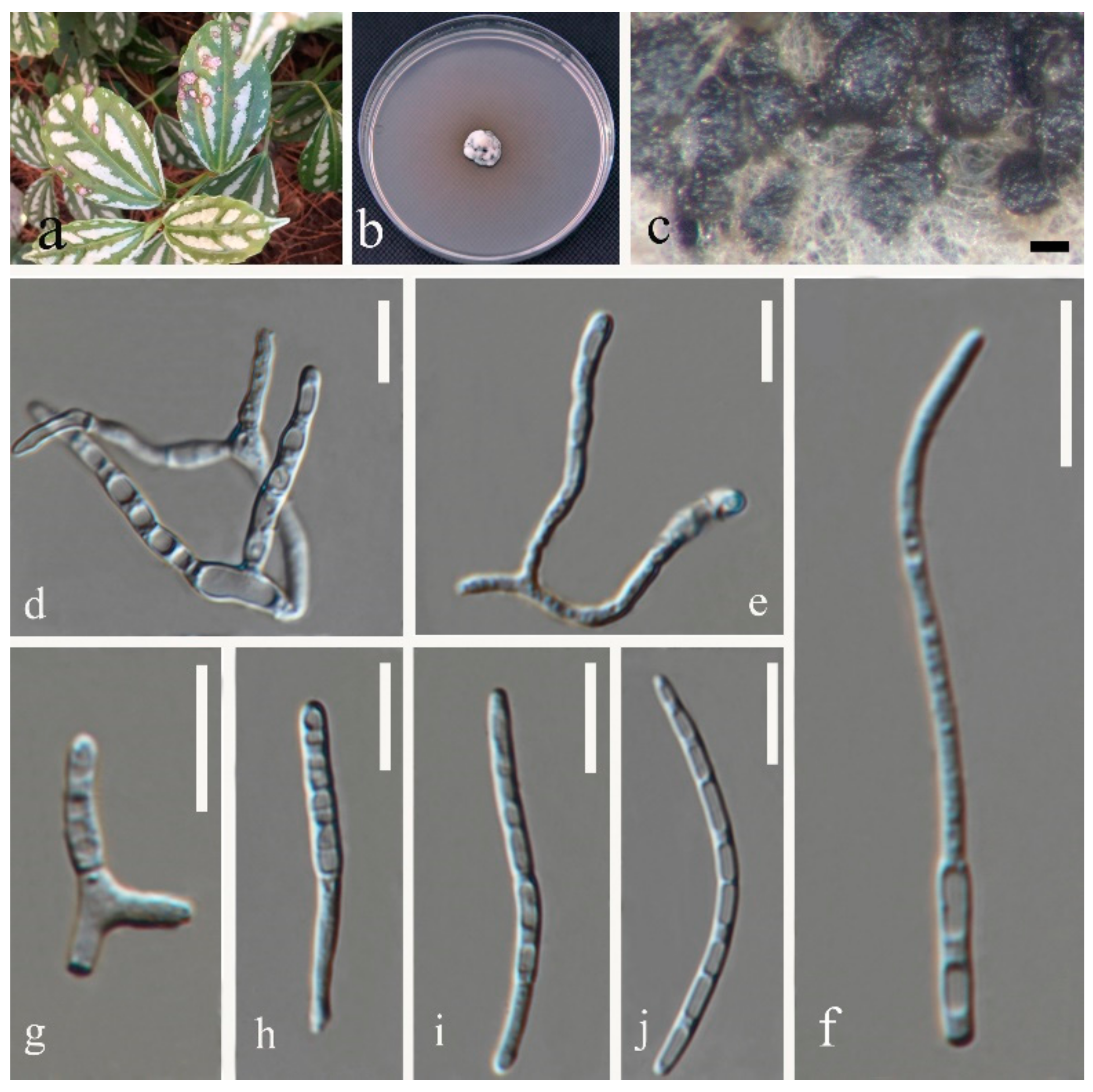
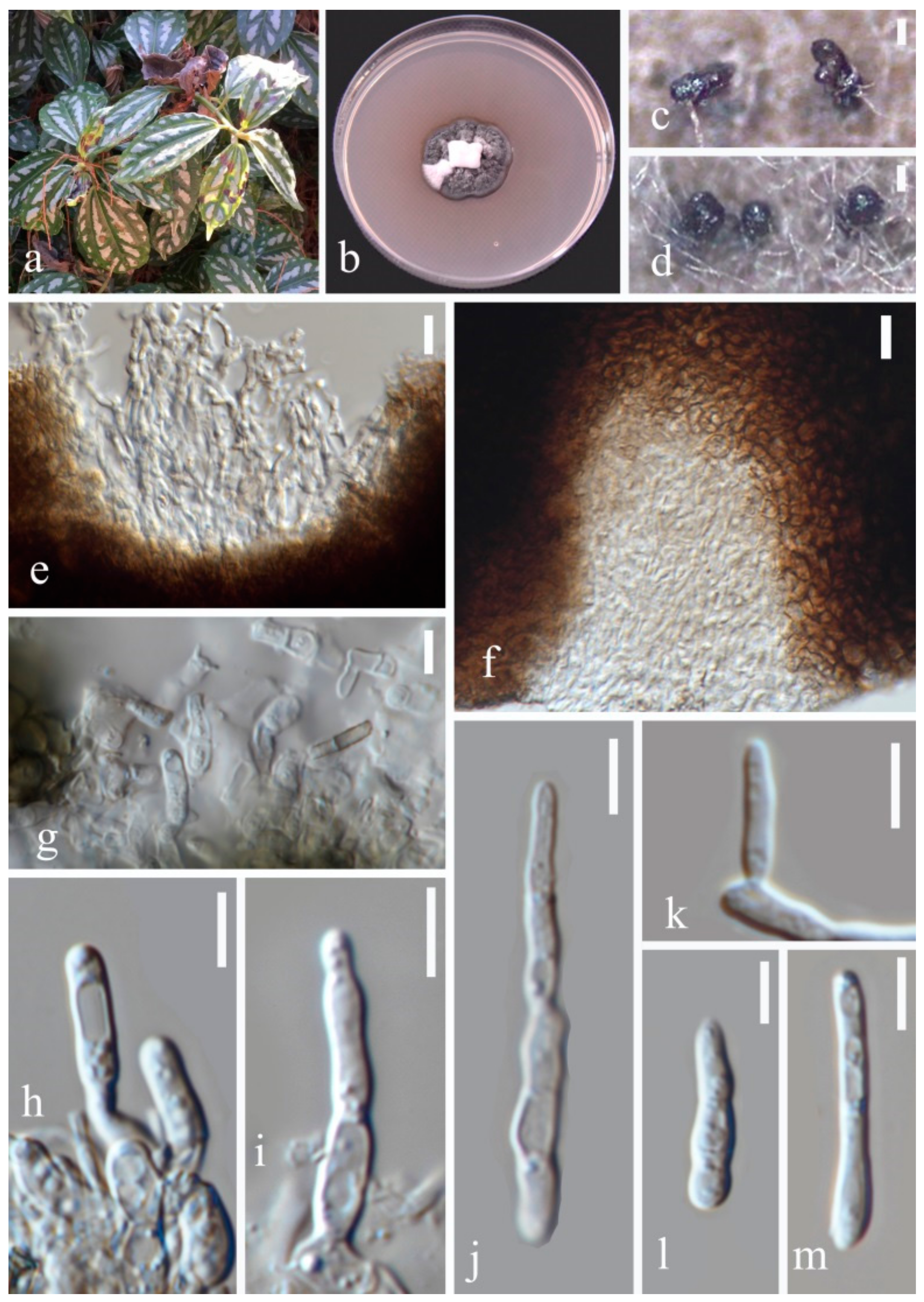
- Septoria pileicola Y.Y. An & Yong Wang bis sp. nov. (Figure 4)
- (3)
- Septoria longipes Y.Y. An & Yong Wang bis sp. nov. (Figure 5)
- (4)
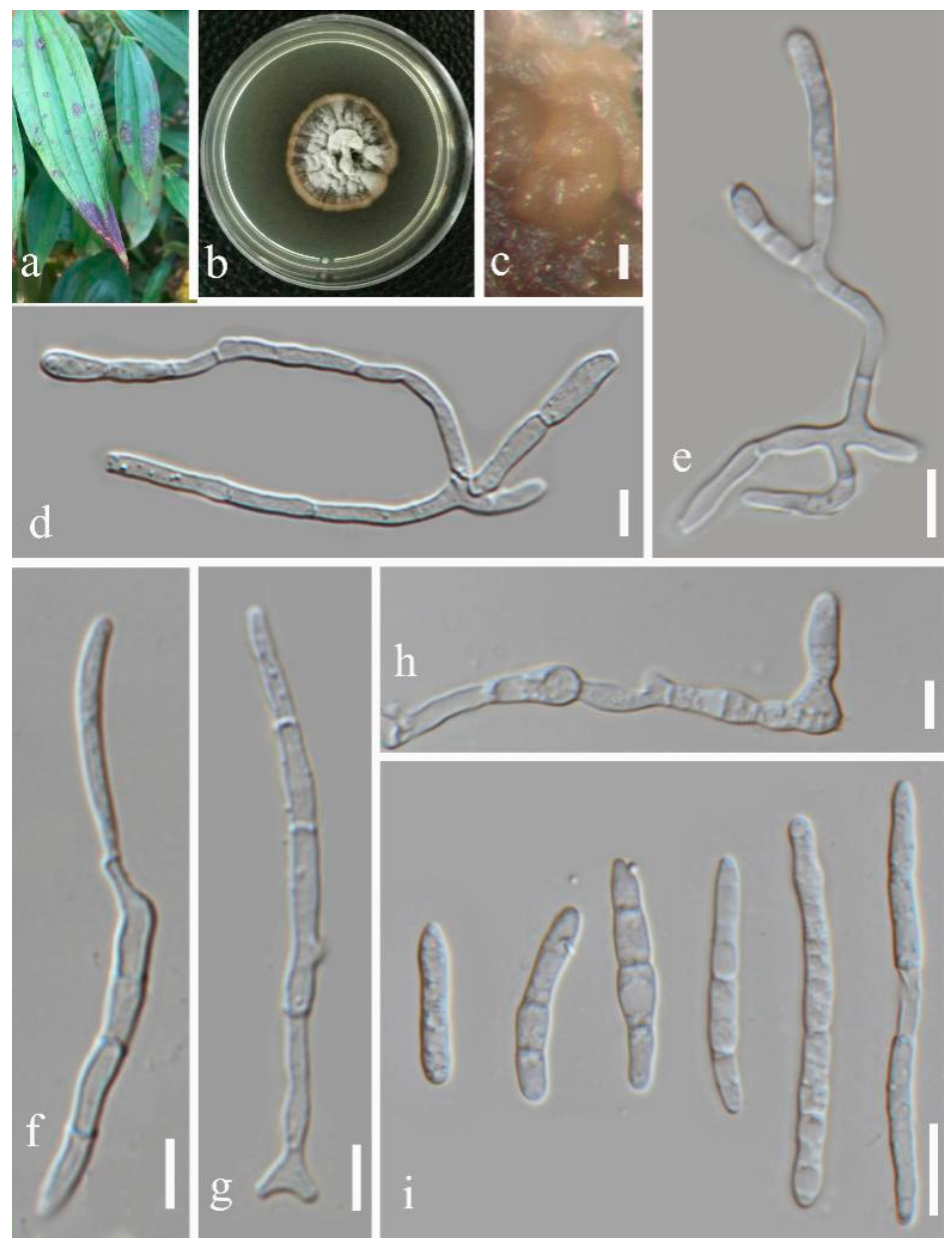
- Septoria dispori Y.Y. An & Yong Wang bis sp. nov. (Figure 6)
- (5)
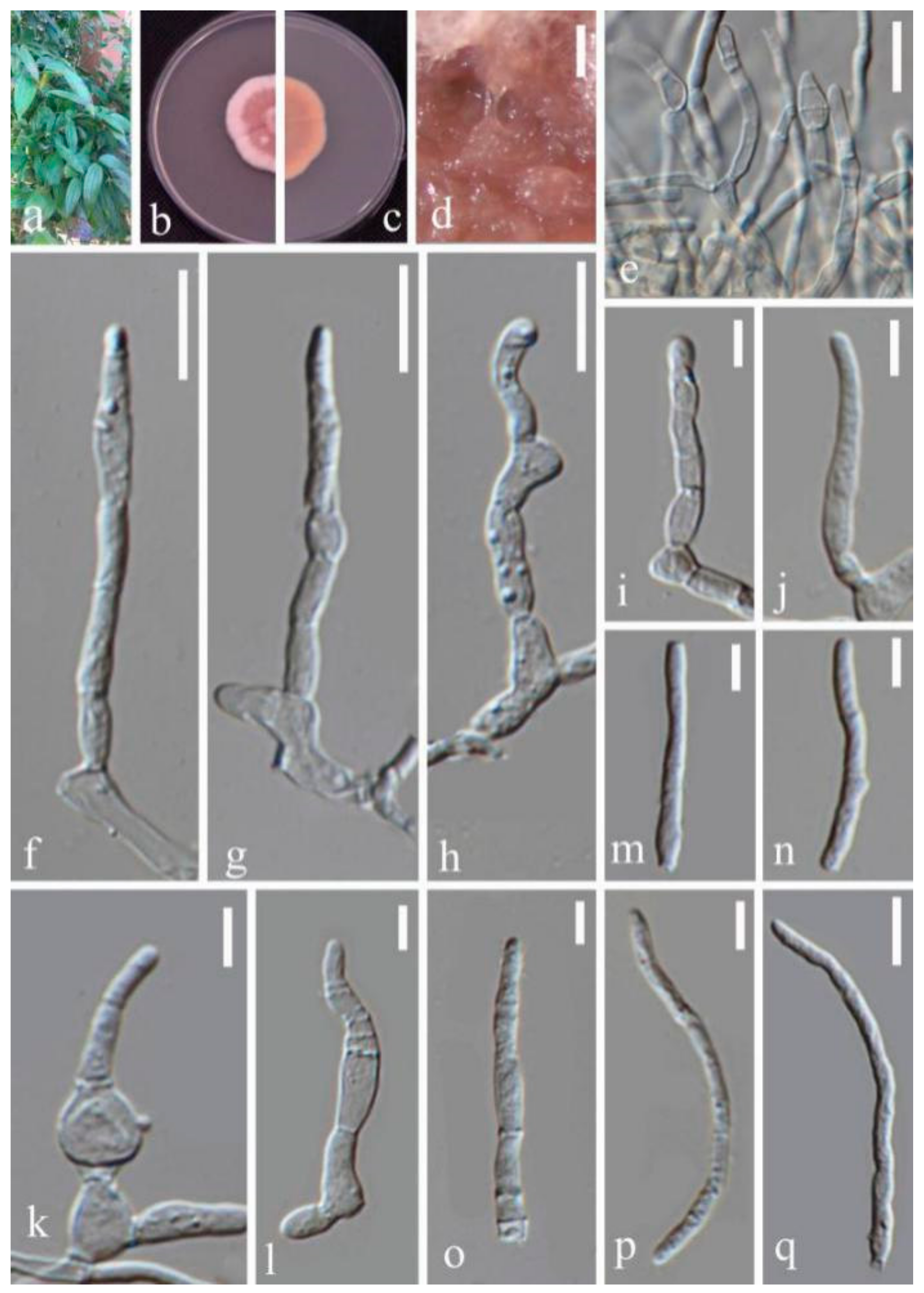
- Septoria protearum Viljoen & Crous, in Swart, Crous, Denman & Palm, S. Afr. J. Bot. 64(2): 144 (1998) (Figure 7)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Index Fungorum 2020. Available online: http://www.indexfungorum.org/names/Names.asp (accessed on 8 October 2020).
- Wijayawardene, N.N.; Hyde, K.D.; Al-Ani, L.K.T.; Tedersoo, L.; Haelewaters, D.; Rajeshkumar, K.C.; Zhao, R.L.; Aptroot, A.; Leontyev, D.V.; Saxena, R.K.; et al. Outline of Fungi and fungus-like taxa. Mycosphere 2020, 11, 1060–1456. [Google Scholar] [CrossRef]
- Verkley, G.J.M.; Quaedvlieg, W.; Shin, H.D.; Crous, P.W. A new approach to species delimitation in Septoria. Stud. Mycol. 2013, 75, 213–305. [Google Scholar] [CrossRef]
- Sutton, B.C. The Coelomycetes. Fungi Imperfecti with Pycnidia, Acervuli and Stromata; Commonwealth Mycological Institute: Kew, UK, 1980. [Google Scholar]
- Constantinescu, O. Taxonomic revision of Septoria-like fungi parasitic on Betulaceae. Trans. Br. Mycol. Soc. 1984, 83, 383–398. [Google Scholar] [CrossRef]
- Sutton, B.C.; Pascoe, I.G. Septoria species on Acacia. Trans. Br. Mycol. Soc. 1987, 89, 521–532. [Google Scholar] [CrossRef]
- Sutton, B.C.; Pascoe, I.G. Some Septoria species on native Australian plants. Stud. Mycol. 1989, 31, 177–186. [Google Scholar]
- Farr, D.F. Septoria species on Cornus. Mycologia 1991, 83, 611–623. [Google Scholar] [CrossRef]
- Farr, D.F. Species of Septoria on the Fabaceae, subfamily Faboidae, tribe Genistae. Sydowia 1992, 44, 13–31. [Google Scholar]
- Quaedvlieg, W.; Verkley, G.J.M.; Shin, H.D.; Barretto, R.W.; Alfenas, A.C.; Swart, W.J.; Groenewald, J.Z.; Crous, P.W. Sizing up Septoria. Stud. Mycol. 2013, 75, 307–390. [Google Scholar] [CrossRef]
- Crous, P.W.; Shivas, R.G.; Quaedvlieg, W.; van der Bank, M.; Zhang, Y.; Summerell, B.A.; Guarro, J.; Wingfield, M.J.; Wood, A.R.; Alfenas, A.C.; et al. Fungal Planet Description Sheets: 214–280. Persoonia 2014, 32, 184–306. [Google Scholar] [CrossRef]
- Crous, P.W.; Schumacher, R.K.; Wingfield, M.J.; Lombard, L.; Giraldo, A.; Christensen, M.; Gardiennet, A.; Nakashima, C.; Pereira, O.; Smith, A.J.; et al. Fungal Systematics and Evolution: FUSE 1. Sydowia 2015, 67, 81–118. [Google Scholar]
- Senanayake, I.C.; Rathnayaka, A.R.; Marasinghe, D.S.; Calabon, M.; Vedprakash, H.G. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. Mycosphere 2020, 11, 2678–2754. [Google Scholar] [CrossRef]
- Braun, U.; Nakashima, C.; Bakhshi, M.; Zare, R.; Shin, H.D.; Alves, R.F.; Sposito, M.B. Taxonomy and phylogeny of cercosporoid ascomycetes on Diospyros spp. with special emphasis on Pseudocercospora spp. Fungal Syst. Evol. 2020, 6, 95–127. [Google Scholar] [CrossRef]
- Dissanayake, A.J.; Bhunjun, C.S.; Maharachchikumbura, S.S.N.; Liu, J.K. Applied aspects of methods to infer phylogenetic relationships amongst fungi. Mycosphere 2020, 11, 2652–2676. [Google Scholar] [CrossRef]
- Kuraku, S.; Zmasek, C.M.; Nishimura, O.; Katoh, K. Leaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res. 2013, 41, W22–W28. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017, 20, 1160–1166. [Google Scholar] [CrossRef]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in fifilamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- O’Donnell, K.; Kistler, H.C.; Cigelnik, E.; Ploetz, R.C. Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proc. Natl. Acad. Sci. USA 1998, 95, 2044–2049. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol. Phylogenetics Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Stukenbrock, E.H.; Quaedvlieg, W.; Javan-Nikhah, M.; Zala, M.; Crous, P.W.; McDonald, B.A. Zymoseptoria ardabilia and Z. pseudotritici, two progenitor species of the Septoria tritici leaf blotch fungus Z. tritici (synonym: Mycosphaerella graminicola). Mycologia 2012, 104, 1397–1407. [Google Scholar] [CrossRef]
- Liu, Y.; Whelen, S.; Hall, B. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Quaedvlieg, W.; Kema, G.H.J.; Groenewald, J.Z.; Verkley, G.J.M.; Seifbarghi, S.; Razavi, M.; Gohari, A.M.; Mehrabi, R.; Crous, P.W. Zymoseptoria gen. nov.: A new genus to accommodate Septoria-like species occurring on graminicolous hosts. Persoonia 2011, 26, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Crous, P.W.; Braun, U.; Wingfield, M.J.; Wood, A.R.; Shin, H.D.; Summerell, B.A.; Alfenas, A.C.; Cumagun, C.J.; Groenewald, J.Z. Phylogeny and taxonomy of obscure genera of microfungi. Persoonia 2009, 22, 139–161. [Google Scholar] [CrossRef]
- Vilgalys, R.; Hester, M. Rapid genetic identifification and mapping of enzymatically amplifified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplifification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfland, D.H., Sininsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP—Phylogenetic Analysis Using Parsimony and Other Methods; Version 4; Sinauer Associates: Sunderland, MA, USA, 2003. [Google Scholar]
- Hillis, D.M.; Bull, J.J. An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst. Biol. 1993, 42, 182–192. [Google Scholar] [CrossRef]
- CIPRES Science Gateway. Available online: https://www.phylo.org/portal2/login.action (accessed on 22 June 2020).
- FigTree. Available online: http://tree.bio.ed.ac.uk/software/figtree (accessed on 22 June 2020).
- Quaedvlieg, W.; Binder, M.; Groenewald, J.Z.; Summerell, B.A.; Carnegie, A.J.; Burgess, T.I.; Crous, P.W. Introducing the consolidated species concept to resolve species in the Teratosphaeriaceae. Persoonia 2014, 33, 1–40. [Google Scholar] [CrossRef] [PubMed]
- Huson, M.G. Physical properties of wool fibers in electrolyte solutions. Text. Res. J. 1998, 68, 595–605. [Google Scholar] [CrossRef]
- Huson, D.H.; Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 2006, 23, 254–267. [Google Scholar] [CrossRef]
- Saccardo, P.A. Supplementum Universale, Pars. III. Sylloge Fungorum 1895, 11, 1–753. [Google Scholar]
- Bäumler, J.A. Mycologisches aus Pressburg. Hedwigia 1885, 24, 75. [Google Scholar]
- Spegazzini, C. Fungi argentini. Pugillus secundus (Continuacion). An. Soc. Cient. Argent. 1880, 10, 5–33. [Google Scholar]
- Jeewon, R.; Hyde, K.D. Establishing species boundaries and new taxa among fungi: Recommendations to resolve taxonomic ambiguities. Mycosphere 2016, 7, 1669–1677. [Google Scholar] [CrossRef]
- Chethana, K.W.T.; Zhou, Y.; Zhang, W.; Liu, M.; Xing, Q.K.; Hyde, K.D.; Yan, J.Y.; Li, X.H. Coniella vitis sp. nov. is the common pathogen of white rot in Chinese vineyards. Plant Dis. 2017, 101, 2123–2136. [Google Scholar] [CrossRef]
- Cooke, M.C. Some exotic fungi. Grevillea 1886, 14, 129–130. [Google Scholar]
- Saccardo, P.A. Sylloge Fungorum: Sylloge Sphaeropsidearum et Melanconiearum; Sumptibus auctoris: Patavii, Italy, 1884; Volume 3, pp. 1–860. [Google Scholar]
- Swart, L.; Crous, P.W.; Denman, S.; Palm, M.E. Fungi occurring on Proteaceae. I. S. Afr. J. Bot. 1998, 64, 137–146. [Google Scholar] [CrossRef]

| Species | Isolate No. | GenBank Accession No. | ||||
|---|---|---|---|---|---|---|
| tef1 | tub2 | rpb2 | LSU | ITS | ||
| Cercospora beticola | CBS 124.31 | KF253246 | KF252780 | KF252304 | KF251802 | KF251298 |
| Septoria aegopodina | CBS 123741 | KF253282 | KF252807 | – | KF251838 | KF251334 |
| S. anthrisci | CBS 109020 | KF253286 | KF252811 | KF252340 | KF251843 | KF251339 |
| S. anthurii | CBS 346.58 | KF253288 | KF252813 | KF252342 | KF251845 | KF251341 |
| S. apiicola | CBS 400.54 | KF253292 | KF252817 | KF252346 | KF251849 | KF251345 |
| S. astericola | CBS 128593 | KF253294 | KF252819 | KF252348 | KF251851 | KF251347 |
| S. astragali | CBS 109116 | KF253298 | KF252823 | KF252352 | KF251855 | KF251351 |
| S. atropurpurea | CBS 348.58 | KF253299 | KF252824 | KF252353 | KF251856 | KF251352 |
| S. bothriospermi | CBS 128599 | KF253301 | KF252826 | KF252355 | KF251858 | KF251354 |
| S. bupleuricola | CBS 128603 | KF253303 | KF252828 | KF252357 | KF251860 | KF251356 |
| S. calendulae | CBS 349.58 | KF253304 | KF252829 | KF252358 | KF251861 | KF251357 |
| S. callistephi | CBS 128590 | KF253305 | KF252830 | KF252359 | KF251862 | KF251358 |
| S. campanulae | CBS 128604 | KF253308 | KF252833 | KF252362 | KF251865 | KF251361 |
| S. carvi | KML 1833 | – | – | – | – | KX453687 |
| S. cerastii | CBS 128612 | KF253311 | KF252836 | KF252365 | KF251868 | KF251364 |
| S. cf. agrimoniicola | CBS 128602 | KF253284 | KF252809 | KF252338 | KF251841 | KF251337 |
| S. cf. rubi | CBS 128646 | KF253314 | KF252839 | KF252368 | KF251871 | KF251367 |
| S. cf. sonchi | CBS 128757 | KF253500 | KF253020 | KF252546 | KF252057 | KF251552 |
| S. cf. stachydicola | CBS 128662 | KF253513 | KF253034 | KF252559 | KF252071 | KF251566 |
| S. chamaecisti | CBS 350.58 | KF253318 | KF252843 | KF252372 | KF251875 | KF251371 |
| S. chelidonii | CBS 128607 | KF253319 | KF252844 | KF252373 | KF251876 | KF251372 |
| S. chromolaenae | CBS 113373 T | KF253321 | KF252846 | KF252375 | KF251878 | KF251374 |
| S. chrysanthemella | CBS 128716 | KF253325 | KF252850 | KF252379 | KF251882 | KF251378 |
| S. cirsii | CBS 128621 | KF253328 | KF252853 | KF252382 | KF251885 | KF251381 |
| S. citri | CBS 315.37 | KF253465 | – | KF252511 | KF252021 | KF251516 |
| S. citricola | CBS 356.36 T | KF253329 | KF252854 | KF252383 | KF251886 | KF251382 |
| S. clematidis | CBS 108983 | KF253330 | KF252855 | KF252384 | KF251887 | KF251383 |
| S. codonopsidis | CBS 128620 | KF253333 | KF252858 | KF252387 | KF251890 | KF251386 |
| S. convolvuli | CBS 128627 | KF253336 | KF252861 | KF252390 | KF251893 | KF251389 |
| S. coprosmae | CBS 113391 | KF253255 | KF252787 | KF252313 | KF251812 | KF251308 |
| S. crepidis | CBS 128619 | KF253338 | KF252863 | KF252392 | KF251895 | KF251391 |
| S. cretae | CBS 135095 T | – | KF252720 | – | KF251736 | KF251233 |
| S. cruciatae | CBS 123747 | KF253340 | KF252865 | KF252394 | KF251897 | KF251393 |
| S. cucubali | CBS 102386 | KF253344 | KF252869 | KF252398 | KF251901 | KF251397 |
| S. cucurbitacearum | CBS 178.77 | KF253346 | – | KF252400 | KF251903 | KF251399 |
| S. dearnessii | CBS 128624 | KF253347 | KF252871 | KF252401 | KF251904 | KF251400 |
| S. digitalis | CBS 391.63 | KF253349 | KF252873 | KF252403 | KF251906 | KF251402 |
| S. dispori | GUCC 2127.1 T | MT996515 | MT984348 | MT993632 | MT985366 | MT974584 |
| S. dispori | GUCC 2164.3 | MT996523 | MT984357 | MT993641 | MT985375 | MT974593 |
| S. dispori | GUCC 2164.4 | MT996524 | MT984358 | MT993642 | MT985376 | MT974594 |
| S. dispori | GUCC 2127.4 | MT996517 | MT984350 | MT993634 | MT985368 | MT974586 |
| S. dolichospora | CBS 129152 | KF253350 | KF252874 | – | KF251907 | KF251403 |
| S. dysentericae | CBS 131892 | KF253353 | KF252877 | KF252406 | KF251910 | KF251406 |
| S. ekmaniana | CBS 113612 | KF253355 | KF252879 | – | KF251912 | KF251408 |
| S. epambrosiae | CBS 128629 | KF253356 | KF252880 | KF252407 | KF251913 | KF251409 |
| S. epilobii | CBS 109084 T | KF253358 | KF252882 | KF252409 | KF251915 | KF251411 |
| S. erigerontis | CBS 109094 | KF253360 | KF252884 | KF252411 | KF251917 | KF251413 |
| S. eucalyptorum | CBS 118505 T | KF253365 | KF252889 | KF252415 | KF251921 | KF251417 |
| S. exotica | CBS 163.78 | KF253366 | KF252890 | KF252416 | KF251922 | KF251418 |
| S. galeopsidis | CBS 102411 T | KF253372 | KF252896 | KF252422 | KF251928 | KF251424 |
| S. gentianae | CBS 128633 | KF253374 | KF252898 | KF252424 | KF251930 | KF251426 |
| S. gerberae | CBS 410.61 | KF253468 | KF252988 | KF252514 | KF252024 | KF251519 |
| S. glycines | CBS 336.53 | KF253377 | KF252901 | – | KF251933 | KF251429 |
| S. glycinicola | CBS 128618 T | KF253378 | KF252902 | KF252427 | KF251934 | KF251430 |
| S. hederae | CBS 566.88 | KF253470 | KF252990 | KF252515 | KF252026 | KF251521 |
| S. helianthi | CBS 123.81 | KF253379 | KF252903 | KF252428 | KF251935 | KF251431 |
| S. helianthicola | CBS 122.81 | KF253380 | KF252904 | KF252429 | KF251936 | KF251432 |
| S. hibiscicola | CBS 128615 | KF253382 | KF252906 | KF252431 | KF251938 | KF251434 |
| S. hippocastani | CPC 23103 | KF253510 | KF253031 | KF252556 | KF252068 | KF251563 |
| S. justiciae | CBS 128625 | KF253385 | KF252909 | KF252434 | KF251941 | KF251437 |
| S. lactucae | CBS 108943 | KF253387 | KF252911 | KF252436 | KF251943 | KF251439 |
| S. lamiicola | CBS 123884 | KF253397 | KF252921 | KF252446 | KF251953 | KF251449 |
| S. lepidiicola | CBS 128635 | KF253398 | KF252922 | KF252447 | KF251954 | KF251450 |
| S. leptostachyae | CBS 128613 | KF253399 | KF252923 | KF252448 | KF251955 | KF251451 |
| S. leucanthemi | CBS 109090 | KF253403 | KF252927 | KF252452 | KF251959 | KF251455 |
| S. limonum | CBS 419.51 | KF253407 | KF252931 | KF252456 | KF251963 | KF251459 |
| S. linicola | CBS 316.37 | KF253408 | KF252932 | KF252457 | KF251964 | KF251460 |
| S. lobeliae | CBS 113392 | KF253460 | KF252981 | KF252507 | KF252016 | KF251511 |
| S. longipes | GUCC 2131.1 T | – | MT984351 | MT993635 | MT985369 | MT974587 |
| S. lycoctoni | CBS 109089 | KF253409 | KF252933 | KF252458 | KF251965 | KF251461 |
| S. lycopersici | CBS 128654 | KF253410 | KF252934 | KF252459 | KF251966 | KF251462 |
| S. lycopicola | CBS 128651 | KF253412 | KF252936 | KF252461 | KF251968 | KF251464 |
| S. lysimachiae | CBS 102315 | KF253413 | KF252937 | KF252462 | KF251969 | KF251465 |
| S. malagutii | CBS 106.80 T | KF253418 | – | KF252467 | KF251974 | KF251470 |
| S. matricariae | CBS 109001 | KF253420 | KF252943 | KF252469 | KF251976 | KF251472 |
| S. mazi | CBS 128755 | KF253422 | KF252945 | KF252471 | KF251978 | KF251474 |
| S. melissae | CBS 109097 | KF253423 | KF252946 | KF252472 | KF251979 | KF251475 |
| S. menthae | CBS 404.34 | KF253424 | KF252947 | – | KF251980 | KF251476 |
| S. napelli | CBS 109105 | KF253426 | KF252949 | KF252474 | KF251982 | KF251478 |
| S. obesa | CBS 128623 | KF253429 | KF252952 | KF252477 | KF251985 | KF251481 |
| S. oenanthicola | CBS 128649 T | KF253433 | KF252954 | KF252239 | KF251737 | KF251234 |
| S. oenanthis | CBS 128667 | KF253432 | KF252953 | - | KF251989 | KF251485 |
| S. orchidearum | CBS 128631 T | KF253434 | KF252955 | KF252482 | KF251990 | KF251486 |
| S. pachyspora | CBS 128652 | KF253437 | KF252958 | KF252485 | KF251993 | KF251488 |
| S. paridis | CBS 109111 | KF253438 | KF252959 | KF252486 | KF251994 | KF251489 |
| S. passifloricola | CBS 102701 | KF253442 | KF252963 | KF252490 | KF251998 | KF251493 |
| S. perillae | CBS 128655 | KF253444 | KF252965 | KF252491 | KF252000 | KF251495 |
| S. petroselini | CBS 182.44 | KF253446 | KF252967 | KF252493 | KF252002 | KF251497 |
| S. phlogis | CBS 128663 | KF253448 | KF252969 | KF252495 | KF252004 | KF251499 |
| S. pileicola | GUCC 2131.3 | MT996519 | MT984353 | MT993637 | MT985371 | MT974589 |
| S. pileicola | GUCC 2131.4 T | MT996520 | MT984354 | MT993638 | MT985372 | MT974590 |
| S. polygonorum | CBS 109834 | KF253453 | KF252974 | KF252500 | KF252009 | KF251504 |
| S. posoniensis | CBS 128645 | KF253456 | KF252977 | KF252503 | KF252012 | KF251507 |
| S. protearum | CBS 778.97 T | KF253472 | KF252992 | KF252517 | KF252028 | KF251523 |
| S. protearum | GUCC 2127.3 | MT996516 | MT984349 | MT993633 | MT985367 | MT974585 |
| S. pseudonapelli | CBS 128664 T | KF253475 | KF252995 | KF252520 | KF252031 | KF251526 |
| S. putrida | CBS 109088 | KF253477 | KF252997 | KF252522 | KF252033 | KF251528 |
| S. rumicum | CBS 503.76 | KF253478 | KF252998 | KF252523 | KF252034 | KF251529 |
| S. saccardoi | CBS 128756 | KF253479 | KF252999 | KF252524 | KF252035 | KF251530 |
| S. sanguisorbigena | GUCC 2131.2 | MT996518 | MT984352 | MT993636 | MT985370 | MT974588 |
| S. sanguisorbigena | GUCC 2164.1 | MT996521 | MT984355 | MT993639 | MT985373 | MT974591 |
| S. sanguisorbigena | GUCC 2164.2 T | MT996522 | MT984356 | MT993640 | MT985374 | MT974592 |
| S. scabiosicola | CBS 109093 | KF253487 | KF253007 | KF252532 | KF252043 | KF251538 |
| S. senecionis | CBS 102366 T | KF253492 | KF253012 | KF252538 | KF252049 | KF251544 |
| S. siegesbeckiae | CBS 128659 | KF253494 | KF253014 | KF252540 | KF252051 | KF251546 |
| S. sii | CBS 102370 | KF253497 | KF253017 | KF252543 | KF252054 | KF251549 |
| S. sisyrinchii | CBS 112096 | KF253499 | KF253019 | KF252545 | KF252056 | KF251551 |
| S. stachydicola | CBS 128668 | KF253512 | KF253033 | KF252558 | KF252070 | KF251565 |
| S. stachydis | CBS 109127 | KF253517 | KF253038 | KF252563 | KF252075 | KF251570 |
| S. stellariae | CBS 102376 | KF253521 | KF253042 | KF252567 | KF252079 | KF251574 |
| S. taraxaci | CBS 567.75 | KF253524 | KF253045 | KF252570 | KF252082 | KF251577 |
| S. tinctoriae | CBS 129154 | KF253525 | KF253046 | KF252571 | KF252083 | KF251578 |
| S. tormentillae | CBS 128647 | KF253527 | KF253048 | KF252573 | KF252085 | KF251580 |
| S. urticae | CBS 102375 T | KF253530 | KF253051 | KF252576 | KF252088 | KF251583 |
| S. verbascicola | CBS 102401 | KF253531 | KF253052 | KF252577 | KF252089 | KF251584 |
| S. verbenae | CBS 113438 | KF253532 | KF253053 | KF252578 | KF252090 | KF251585 |
| S. villarsiae | CBS 514.78 | KF253534 | KF253055 | KF252580 | KF252092 | KF251587 |
| S. violae-palustris | CBS 128644 | KF253537 | KF253058 | KF252583 | KF252095 | KF251590 |
| Locus | Primer | Primer Sequence 5′ to 3′ | Annealing Temperature (°C) | Direction | Reference |
|---|---|---|---|---|---|
| tef1 | EF1-728F | CATCGAGAAGTTCGAGAAGG | 52 | Forward | [18] |
| EF-2 | GGARGTACCAGTSATCATGTT | Reverse | [19] | ||
| tub2 | T1 | AACATGCGTGAGATTGTAAGT | 52 | Forward | [20] |
| β-Sandy-R | GCRCGNGGVACRTACTTGTT | Reverse | [21] | ||
| rpb2 | fRPB2-5F | GAYGAYMGWGATCAYTTYGG | 49 | Forward | [22] |
| fRPB2-414R | ACMANNCCCCARTGNGWRTTRTG | Reverse | [23] | ||
| LSU | LSU1Fd | GRATCAGGTAGGRATACCCG | 52 | Forward | [24] |
| LR5 | TCCTGAGGGAAACTTCG | Reverse | [25] | ||
| ITS | ITS5 | GGAAGTAAAAGTCGTAACAAGG | 52 | Forward | [26] |
| ITS4 | TCCTCCGCTTATTGATATGC | Reverse | [26] |
| Total Characters | Number of Parsimony-Informative Characters | TL | CI | RI | HI | RC | |
|---|---|---|---|---|---|---|---|
| ITS | 486 | 43 | 176 | 0.642 | 0.76 | 0.358 | 0.488 |
| LSU | 799 | 31 | 112 | 0.625 | 0.863 | 0.375 | 0.539 |
| rpb2 | 345 | 18 | 780 | 0.273 | 0.746 | 0.727 | 0.204 |
| tef1 | 469 | 231 | 1498 | 0.379 | 0.707 | 0.621 | 0.268 |
| tub2 | 325 | 165 | 1221 | 0.326 | 0.774 | 0.674 | 0.252 |
| tef1 + rpb2 + tub2 + ITS | 1625 | 548 | 3927 | 0.328 | 0.716 | 0.672 | 0.235 |
| tef1 + rpb2 + tub2 + ITS + LSU | 2434 | 567 | 4075 | 0.330 | 0.720 | 0.670 | 0.238 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
An, Y.-Y.; Dayarathne, M.C.; Zeng, X.-Y.; Phillips, A.J.L.; Hyde, K.D.; Wang, Y. Molecular and Morphological Assessment of Septoria Species Associated with Ornamental Plants in Yunnan Province, China. J. Fungi 2021, 7, 483. https://doi.org/10.3390/jof7060483
An Y-Y, Dayarathne MC, Zeng X-Y, Phillips AJL, Hyde KD, Wang Y. Molecular and Morphological Assessment of Septoria Species Associated with Ornamental Plants in Yunnan Province, China. Journal of Fungi. 2021; 7(6):483. https://doi.org/10.3390/jof7060483
Chicago/Turabian StyleAn, Yuan-Yan, Monika C. Dayarathne, Xiang-Yu Zeng, Alan J. L. Phillips, Kevin D. Hyde, and Yong Wang. 2021. "Molecular and Morphological Assessment of Septoria Species Associated with Ornamental Plants in Yunnan Province, China" Journal of Fungi 7, no. 6: 483. https://doi.org/10.3390/jof7060483
APA StyleAn, Y.-Y., Dayarathne, M. C., Zeng, X.-Y., Phillips, A. J. L., Hyde, K. D., & Wang, Y. (2021). Molecular and Morphological Assessment of Septoria Species Associated with Ornamental Plants in Yunnan Province, China. Journal of Fungi, 7(6), 483. https://doi.org/10.3390/jof7060483








