Multilocus Genotyping and Intergenic Spacer Single Nucleotide Polymorphisms of Amylostereum areolatum (Russulales: Amylostereacea) Symbionts of Native and Non-Native Sirex Species
Abstract
1. Introduction
2. Materials and Methods
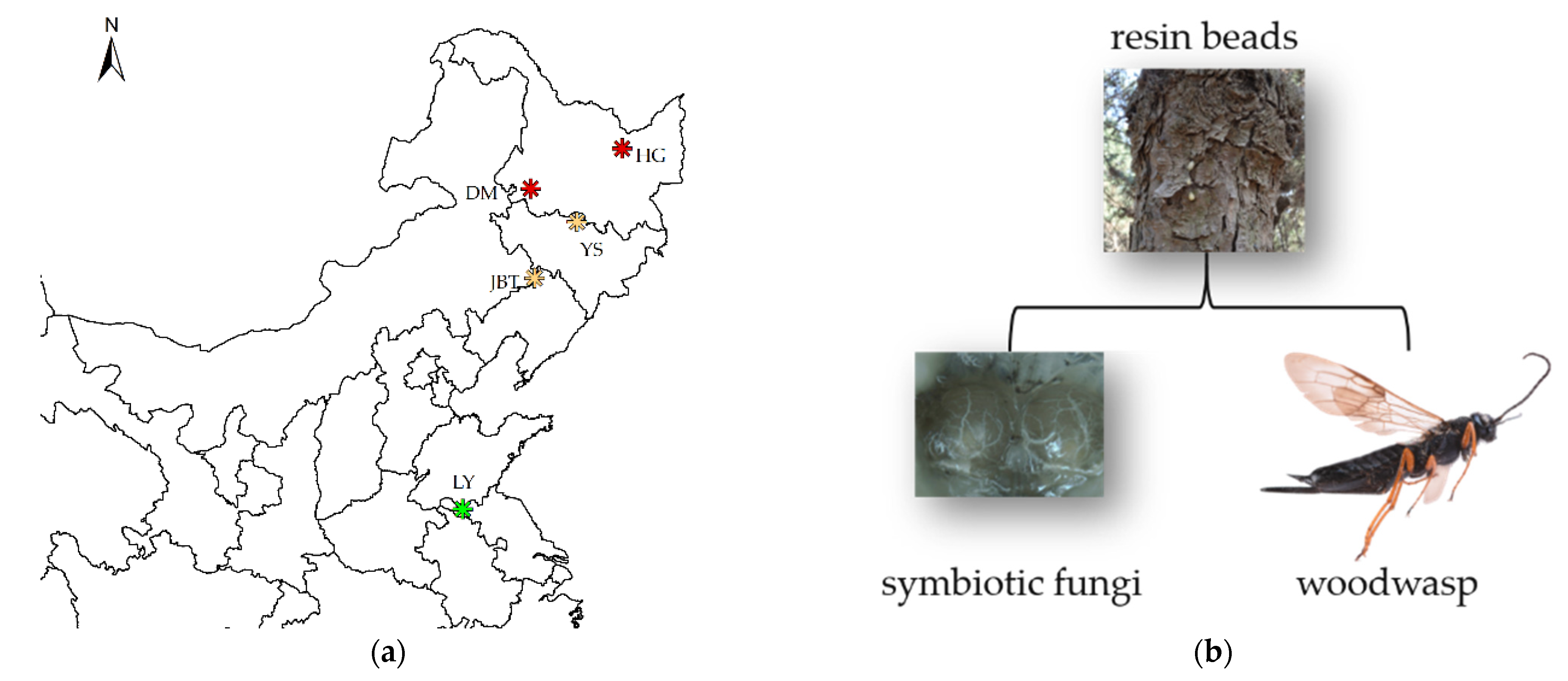
2.1. Sample Collection and Symbiont Isolation
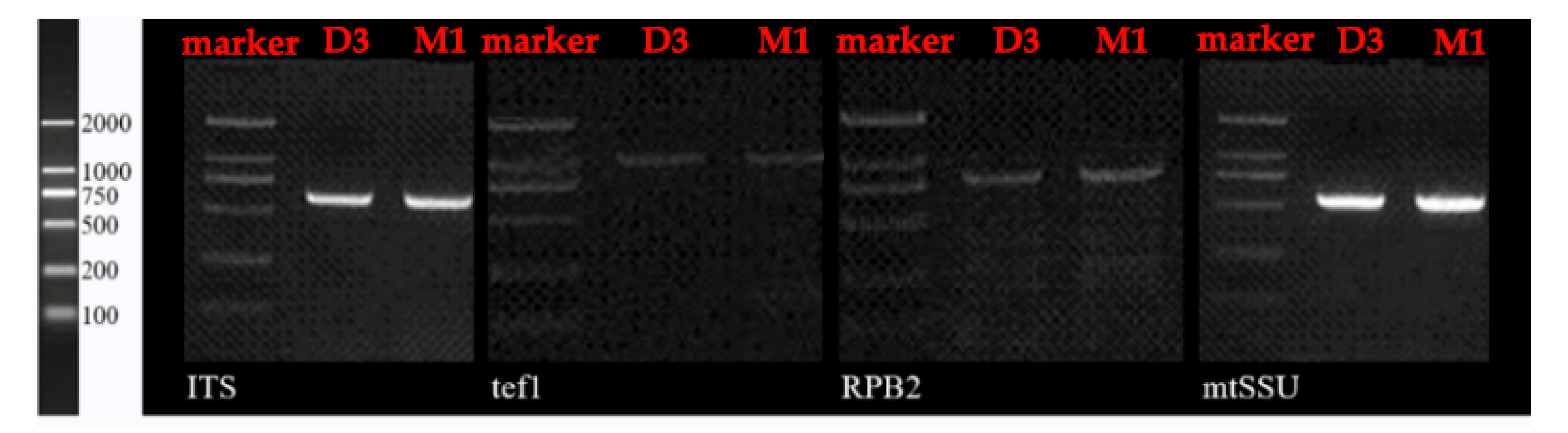
2.2. Colony Polymerase Chain Reaction (PCR) and Sequencing
2.3. Multilocus Genotyping
2.4. Fragment Analysis of Intergenic Spacer (IGS) Types
2.5. Data Analysis
3. Results
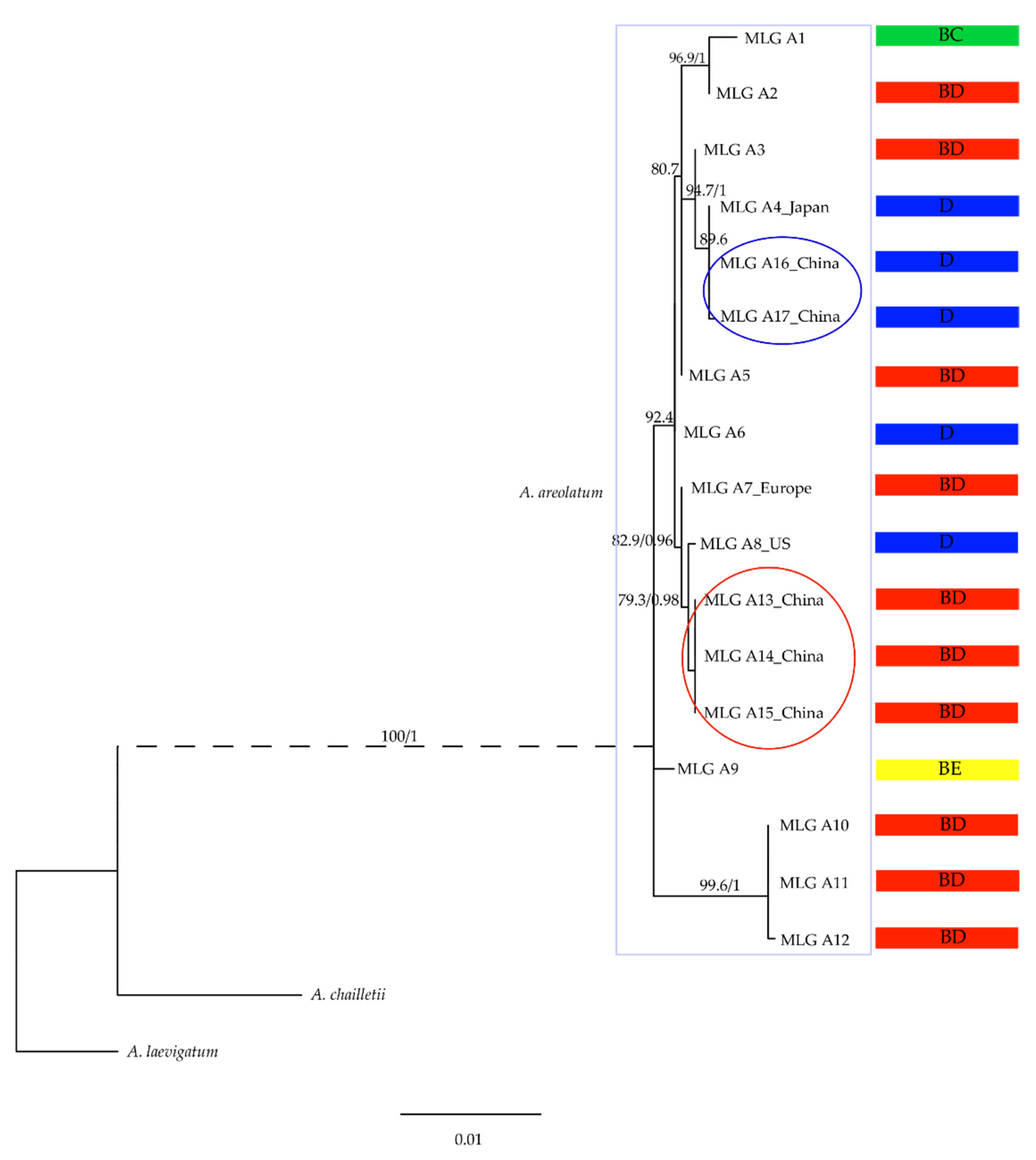
3.1. Phylogenetic Relationship in Amylostereum
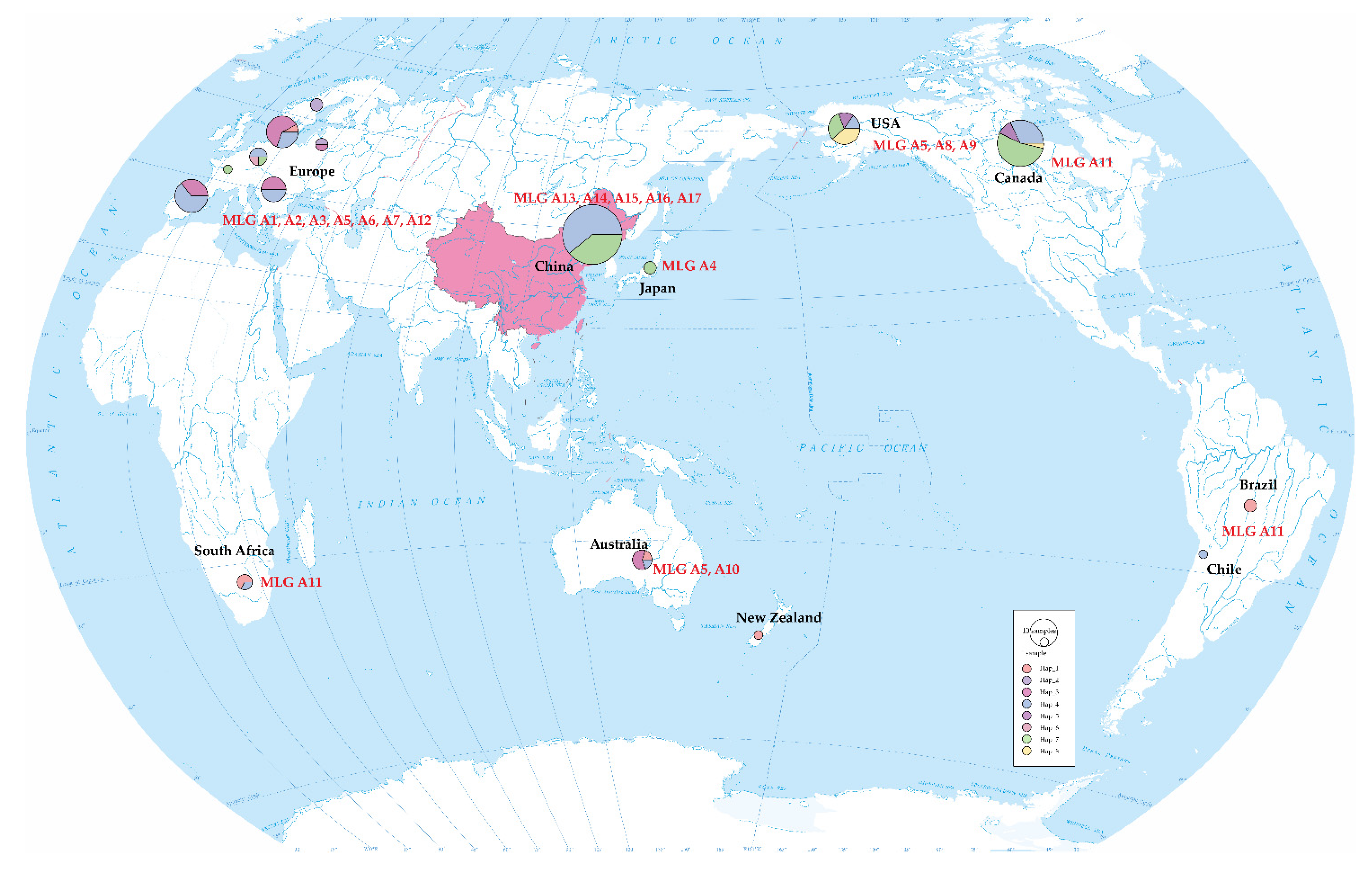
3.2. Haplotype Relationships Based on IGS Sequences and Fragment Analysis Data
3.3. Co-Infestation of S. noctilio and S. nitobei in Pinus
4. Discussion
4.1. Multilocus Genotyping and IGS Heterogeneity of A. areolatum
4.2. Sirex noctilio and S. nitobei-A. areolatum Association
4.3. Focus for Further Research
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Boidin, J. Hétérobasidiomycètes saprophytes et homobasidiomycètes résupinés. VII.—Essai sur le genre «Stereum sensu lato» (Troisième contribution). Bull. Mens. Soc. Linn. Lyon 1959, 28, 205–222. [Google Scholar] [CrossRef]
- Pechmann, H.V. Untersuchungen uber die Rotstreifigkeit des Fichtenholzes. Summ. Forstw. Cbl. Beih. 1967, 27, 112. [Google Scholar]
- Siepmann, R. Artdiagnose holzzerstörender Hymenomyceten an Hand vonReinkulturen. V. Nova Hedwig. 1979, 30, 69–78. [Google Scholar] [CrossRef]
- Ginns, J.; Stalpers, J.A. Identification of Wood-Inhabiting Aphyllophorales in Pure Culture. Mycologia 1979, 71, 224. [Google Scholar] [CrossRef]
- Thomsen, I. Fruitbody characters and cultural characteristics useful for recognizing Amylostereum areolatum and A. chailletii. Mycotaxon 1998, 69, 419–428. [Google Scholar]
- Morgan, F. Bionomics of Siricidae. Annu. Rev. Entomol. 1968, 13, 239–256. [Google Scholar] [CrossRef]
- Cartwright, K. Notes on a fungus associated with Sirex cyaneus. Ann. Appl. Biol. 1929, 16, 182–187. [Google Scholar]
- Coutts, M.P. The Mechanism of Pathogenicity of Sirex noctilio on Pinus radiata I. Effects of the Symbiotic Fungus Amylostereum sp. (Thelophoraceae). Aust. J. Biol. Sci. 1969, 22, 915–924. [Google Scholar] [CrossRef]
- Coutts, M.P. The Mechanism of Pathogenicity of Sirex noctilio on Pinus radiata II. Effects of S. noctilio mucus. Aust. J. Biol. Sci. 1969, 22, 1153–1162. [Google Scholar] [CrossRef]
- Hanson, H.S. Ecological Notes on the Sirex Wood Wasps and their Parasites. Bull. Entomol. Res. 2009, 30, 27–65. [Google Scholar] [CrossRef]
- Talbot, P.H.B. The Sirex-Amylostereum-Pinus Association. Annu. Rev. Phytopathol. 1977, 15, 41–54. [Google Scholar] [CrossRef]
- Stoddart, J.A.; Taylor, J.F. Genotypic diversity: Estimation and prediction in samples. Genetics 1988, 118, 705–711. [Google Scholar] [CrossRef]
- Spradbery, J.P.; Kirk, A.A. Aspects of the ecology of siricid woodwasps (Hymenoptera: Siricidae) in Europe, North Africa and Turkey with special reference to the biological control of Sirex noctilio F. in Australia. Bull. Entomol. Res. 1978, 68, 341–359. [Google Scholar] [CrossRef]
- Wang, M.; Bao, M.; Ao, T.G.; Ren, L.L.; Luo, Y.Q. Population distribution patterns and ecological niches of two Sirex species damaging Pinus sylvestris var. mongolica. Chin. J. Appl. Entomol. 2017, 54, 924–932. [Google Scholar] [CrossRef]
- Hall, M.J. A Survey of Siricid Attack on Radiata Pine in Europe. Aust. For. 1968, 32, 155–162. [Google Scholar] [CrossRef]
- Spradbery, J.P. A comparative study of the phytotoxic effects of siricid woodwasps on conifers. Ann. Appl. Biol. 1973, 75, 309–320. [Google Scholar] [CrossRef]
- Lantschner, M.V.; Villacide, J.M.; Garnas, J.R.; Croft, P.; Carnegie, A.J.; Liebhold, A.M.; Corley, J.C. Temperature explains variable spread rates of the invasive woodwasp Sirex noctilio in the Southern Hemisphere. Biol. Invasions 2014, 16, 329–339. [Google Scholar] [CrossRef]
- Spradbery, J.P. The oviposition biology of siricid woodwasps in Europe. Ecol. Entomol. 1977, 2, 225–230. [Google Scholar] [CrossRef]
- Carnegie, A.J.; Bashford, R. Sirex Woodwasp in Australia: Current Management Strategies, Research and Emerging Issues. In The Sirex Woodwasp and Its Fungal Symbiont; Springer: Berlin/Heidelberg, Germany, 2012; pp. 175–201. [Google Scholar] [CrossRef]
- Xu, Q. Bio-Ecological Characteristics, Monitoring and Silvicultural Control Technology of Invasive Alien Species Sirex noctilio in China. Ph.D. Thesis, Beijing Forestry University, Beijing, China, 2020. [Google Scholar] [CrossRef]
- Gilbertson, R.L. Relationships between insects and wood-rotting Basidiomycetes. In Fungus-Insect Relationships, Perspectives in Ecology and Evolution; Columbia University Press: New York, NY, USA, 1984. [Google Scholar]
- Vasiliauskas, R.; Stenlid, J.A.N.; Thomsen, I.M. Clonality and genetic variation in Amylostereum areolatum and A. chailletii from northern Europe. New Phytol. 1998, 139, 751–758. [Google Scholar] [CrossRef]
- Thomsen, I.M. Amylostereum areolatum and Amylostereum chailletii: Symbiotic Fungi of Woodwasps (Sirex sp. and Urocerus sp.). Ph.D. Thesis, Danish Forest and Landscape Research Institute, Hoersholm, Denmark, 1996. [Google Scholar]
- Slippers, B.; Coutinho, T.A.; Wingfield, B.D.; Wingfield, M.J. A review of the genus Amylostereum and its association with woodwasps. S. Arf. J. Sci. 2003, 99, 70–74. [Google Scholar] [CrossRef]
- Margrete Thomsen, I.; Koch, J. Somatic compatibility in Amylostereum areolatum and A. chailletii as a consequence of symbiosis with siricid woodwasps. Mycol. Res. 1999, 103, 817–823. [Google Scholar] [CrossRef]
- Slippers, B.; Wingfield, M.J.; Coutinho, T.A.; Wingfield, B.D. Population structure and possible origin of Amylostereum areolatum in South Africa. Plant Pathol. 2010, 50, 206–210. [Google Scholar] [CrossRef]
- Wang, Y.; Lu, J.; Beattie, G.A.; Islam, M.R.; Om, N.; Dao, H.T.; Van Nguyen, L.; Zaka, S.M.; Guo, J.; Tian, M.Y. Phylogeography of Diaphorina citri (Hemiptera: Liviidae) and its primary endosymbiont, ‘Candidatus Carsonella ruddii’: An evolutionary approach to host-endosymbiont interaction. Pest Manag. Sci. 2018, 74, 2185–2194. [Google Scholar] [CrossRef]
- Wang, M.; Wang, L.X.; Li, D.P.; Fu, N.N.; Li, C.C.; Luo, Y.Q.; Ren, L.L. Advances in the Study of Mutualism Relationship Between Amylostereum areolatum and Sirex noctilio. J. Temp. For. Res. 2020, 3, 1–11. [Google Scholar]
- Fitza, K.N.E.; Tabata, M.; Kanzaki, N.; Kimura, K.; Garnas, J.; Slippers, B. Host specificity and diversity of Amylostereum associated with Japanese siricids. Fungal. Ecol. 2016, 24, 76–81. [Google Scholar] [CrossRef]
- Bergeron, M.J.; Leal, I.; Foord, B.; Ross, G.; Davis, C.; Slippers, B.; de Groot, P.; Hamelin, R.C. Putative origin of clonal lineages of Amylostereum areolatum, the fungal symbiont associated with Sirex noctilio, retrieved from Pinus sylvestris, in eastern Canada. Fungal. Biol. 2011, 115, 750–758. [Google Scholar] [CrossRef] [PubMed]
- Schiff, N.M.; Goulet, H.; Smith, D.R.; Boudreault, C.; Scheffler, B.E. Siricidae (Hymenoptera: Symphyta: Siricoidea) of the Western Hemisphere. Can. J. Arthropod Identif. 2012, 21, 1–305. [Google Scholar] [CrossRef]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specificity for basidiomycetes—Application to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef]
- Morehouse, E.A.; James, T.Y.; Ganley, A.R.; Vilgalys, R.; Berger, L.; Murphy, P.J.; Longcore, J.E. Multilocus sequence typing suggests the chytrid pathogen of amphibians is a recently emerged clone. Mol. Ecol. 2003, 12, 395–403. [Google Scholar] [CrossRef]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- White, T.; Bruns, T.; Lee, S.; Taylor, F.; White, T.; Lee, S.H.; Taylor, L.; Shawetaylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nuclc Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar] [CrossRef]
- Xia, X. DAMBE6: New Tools for Microbial Genomics, Phylogenetics, and Molecular Evolution. J. Hered. 2017, 108, 431–437. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Lemey, P. The Phylogenetic Handbook: Assessing Substitution Saturation with DAMBE; Cambridge University Press: Cambridge, UK, 2009. [Google Scholar]
- Xia, X.; Xie, Z.; Salemi, M.; Chen, L.; Wang, Y. An index of substitution saturation and its application. Mol. Phylogenet. Evol. 2003, 26, s1055–s7903. [Google Scholar] [CrossRef]
- Hsiau, T.W. The Taxonomy and Phylogeny of the Mycangial Fungi from Dendroctonus brevicomis and D. frontalis (Coleoptera: Scolytidae). Ph.D. Thesis, Iowa State University, Ames, IA, USA, 1996. [Google Scholar]
- Slippers, B.; Wingfield, B.D.; Coutinho, T.A.; Wingfield, M.J. DNA sequence and RFLP data reflect geographical spread and relationships of Amylostereum areolatum and its insect vectors. Mol. Ecol. 2002, 11, 1845–1854. [Google Scholar] [CrossRef] [PubMed]
- Stalpers, J. Identification of Wood-Inhabiting Aphyllophorales in Pure Culture Studies in Mycology 16; CBS: Baarn, The Netherlands; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 1978; p. 224. [Google Scholar]
- Siepmann, R. Artdiagnose einiger holzzerstörender Hymenomyceten an Hand von Reinkulturen. IV. Nova Hedwig. Allem. Da. 1971, 21, 843–875. [Google Scholar]
- Hajek, A.E.; Nielsen, C.; Kepler, R.M.; Long, S.J.; Castrillo, L. Fidelity among Sirex woodwasps and their fungal symbionts. Microb. Ecol. 2013, 65, 735–762. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kimura, M. Estimation of evolutionary distances between homologous nucleotide sequences. Proc. Natl. Acad. Sci. USA 1981, 78, 454–458. [Google Scholar] [CrossRef]
- Felsenstein, J. Evolutionary trees from DNA sequences: A maximum likelihood approach. J. Mol. Evol. 1981, 17, 368–376. [Google Scholar] [CrossRef]
- Zhang, D.; Gao, F.; Jakovlic, I.; Zou, H.; Zhang, J.; Li, W.X.; Wang, G.T. PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol. Ecol. Resour. 2020, 20, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Minh, B.Q.; Nguyen, M.A.; von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 2013, 30, 1188–1195. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Leigh, J.W.; Bryant, D. Popart: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Olatinwo, R.O.; Schowalter, T.D.; Doucet, D.; Bowman, S.; Johnson, W.C.; Allison, J.D.; Morisette, J. Intergenic Spacer Single Nucleotide Polymorphisms for Genotyping Amylostereum areolatum (Russulales: Amylostereacea) Symbionts of Native and Non-native Sirex Species. Ann. Entomol. Soc. Am. 2020, 113, 280–287. [Google Scholar] [CrossRef]
- Nielsen, C.; Williams, D.W.; Hajek, A.E. Putative source of the invasive Sirex noctilio fungal symbiont, Amylostereum areolatum, in the eastern United States and its association with native siricid woodwasps. Mycol. Res. 2009, 113, 1242–1253. [Google Scholar] [CrossRef] [PubMed]
- Wooding, A.L.; Wingfield, M.J.; Hurley, B.P.; Garnas, J.R.; de Groot, P.; Slippers, B. Lack of fidelity revealed in an insect-fungal mutualism after invasion. Biol. Lett. 2013, 9, 20130342. [Google Scholar] [CrossRef] [PubMed]
- Olatinwo, R.; Allison, J.; Meeker, J.; Johnson, W.; Streett, D.; Aime, M.C.; Carlton, C. Detection and identification of Amylostereum areolatum (Russulales: Amylostereaceae) in the mycangia of Sirex nigricornis (Hymenoptera: Siricidae) in central Louisiana. Environ. Entomol. 2013, 42, 1246–1256. [Google Scholar] [CrossRef] [PubMed]
- Li, D.P.; Shi, J.; Luo, Y.Q. Mutualism between the Eurasian woodwasp, Sirex noctilio (Hymenoptera: Siricidae) and its fungal symbiont Amylostereum areolatum (Russulales: Amylostereaceae). Acta Entomol. Sin. 2015, 58, 1019–1029. [Google Scholar] [CrossRef]
- Sun, X.; Tao, J.; Roques, A.; Luo, Y. Invasion History of Sirex noctilio Based on COI Sequence: The First Six Years in China. Insects 2020, 11, 111. [Google Scholar] [CrossRef]
- Castrillo, L.A.; Hajek, A.E.; Pajares, J.A.; Thomsen, I.M.; Csoka, G.; Kenaley, S.C.; Kepler, R.M.; Zamora, P.; Angeli, S. Multilocus genotyping of Amylostereum spp. associated with Sirex noctilio and other woodwasps from Europe reveal clonal lineage introduced to the US. Fungal. Biol. 2015, 119, 595–604. [Google Scholar] [CrossRef] [PubMed]
- Aanen, D.K.; Eggleton, P.; Rouland-Lefevre, C.; Guldberg-Frøslev, T.; Rosendahl, S.; Boomsma, J.J. The evolution of fungus-growing termites and their mutualistic fungal symbionts. Proc. Natl. Acad. Sci. USA 2002, 99, 14887–14892. [Google Scholar] [CrossRef] [PubMed]
- Harrington, T.C. 11. Ecology and evolution of mycophagous bark beetles and their fungal partners. Insect Fungal Associations. Available online: https://www.semanticscholar.org/paper/Ecology-and-evolution-of-mycophagous-bark-beetles-Harrington-Vega/f3ce0b8255adbe467a6e1cbe4ac5c66130d1cd21 (accessed on 9 December 2021).
- Harrington, T.C.; Aghayeva, D.N.; Fraedrich, S.W. New combinations in Raffaelea, Ambrosiella, and Hyalorhinocladiella, and four new species from the redbay ambrosia beetle, Xyleborus glabratus. Mycotaxon 2010, 111, 337–361. [Google Scholar] [CrossRef]
- Hajek, A.E.; Harris, D.C.; Bittner, T.D. Symbiont Spillover from Invasive to Native Woodwasps. Microb. Ecol. 2018, 75, 7–9. [Google Scholar] [CrossRef] [PubMed]
- Slippers, B.; Hurley, B.P.; Wingfield, M.J. Sirex woodwasp: A model for evolving management paradigms of invasive forest pests. Annu. Rev. Entomol. 2015, 60, 601–619. [Google Scholar] [CrossRef]


| Composition | Dosage (μL) |
|---|---|
| The supernatant | 1 |
| Premix Taq (Ex Taq Version 2.0) 1.25 U/25 μL | 12.5 |
| Primer (F) 10 pmol/μL | 1 |
| Primer (R) 10 pmol/μL | 1 |
| Sterile distilled water | 9.5 |
| Target Gene | Primer Sequence (5′–3′) | Reference | PCR Amplification Conditions |
|---|---|---|---|
| ITS | 95 °C 4 min; 95 °C 1 min; 55 °C 1 min; 72 °C 1 min; 72 °C 10 min; 35 cycles | ||
| ITS1-F | 5′-CTTGGTCATTTAGAGGAAGTAA | Gardes and Bruns (1993) [32] | |
| ITS4-B | 5′-CAGGAGACTTGTACACGGTCCAG | ||
| tef1 | 95 °C 4 min; 95 °C 1 min; 55 °C 1 min; 72 °C 1 min; 72 °C 10 min; 35 cycles | ||
| tef1F | 5′-TACAAGTGCGGTGGTATTGACA | Morehouse et al. (2003) [33] | |
| tef1R | 5′-ACGGACTTGACTTCAGTGGT | ||
| RPB2 | 95 °C 4 min; 95 °C 1 min; 50 °C 2 min; 72 °C 2 min; 72 °C 10 min; 35 cycles | ||
| RPB2-6F | 5′-TGGGGTATGGTCTGTCCTGC | Liu et al. (1999) [34] | |
| fRPB2-7.c R | 5′-TGGGGTATGGTCTGTCCTGC | ||
| mtSSU | 95 °C 4 min; 95 °C 1 min; 55 °C 1 min; 72 °C 1 min; 72 °C 10 min; 35 cycles | ||
| MS1 | 5′-CAGCAGTCAAGAATATTAGTCAATG | White et al. (1990) [35] | |
| MS2 | 5′-GCGGATTATCGAATTAAATAAC |
| MLG 1 | Isolate Code | ITS-tef1-RPB2-mtSSU Types | Woodwasp/Tree Host | Collection Site | No. Samples | IGS |
|---|---|---|---|---|---|---|
| A. areolatum | ||||||
| A1 | RMK11-001 | A-A-A-A | S. juvencus, S. noctilio, U. albicornis, U. gigas | Denmark, Hungary, and Spain | 16 | BC |
| A2 | RMK11-011 | B-A-A-B | S. juvencus | Hungary | 1 | BD |
| A3 | RMK11-022 | B-A-B-B | S. noctilio | Hungary | 1 | BD |
| A4 | B1395 | C-A-B-B | S. nitobei | Japan | 2 | D2 |
| A5 | GR94 | B-A-C-B | S. juvencus, S. noctilio | Australia, Denmark, Hungary, Spain, and US | 14 | BD |
| A6 | B1352 | D-A-C-B | tree source (Picea abies) | Germany | 1 | D |
| A7 | B1385 | D-A-C-D | S. juvencus | Germany | 1 | BD |
| A8 | AH1-17 | B-A-C-E | S. noctilio | US | 1 | D |
| A9 | ScyME9/10 | E-B-C-C | S. nitidus | US | 1 | BE |
| A10 | Ecogrow | F-C-D-B | Ecogrow nematode culture | Australia | 1 | BD |
| A11 | DAOM:239281 | D-C-D-B | S. noctilio | Canada, South Africa, Chile | 8 | BD |
| A12 | DAOM:Francke | D-C-D-D | Germany | 1 | BD | |
| A13 | D18 | D-D-D-E | S. noctilio, S. nitobei | DM, JBT (this study) | 4 | BD |
| A14 | D3 | D-D-E-E | S. noctilio | DM (this study) | 1 | BD |
| A15 | D10 | D-E-D-E | S. noctilio | DM, YS, HG, and JBT (this study) | 11 | BD |
| A16 | M1 | G-E-B-B | S. nitobei | JBT, LY, YS (this study) | 9 | D |
| A17 | L29 | G-D-B-B | S. nitobei | LY (this study) | 1 | D |
| A. chailletii | U. gigas | Australia, Denmark, Hungary, Spain, and US | 1 | |||
| A. laevigatum | - | U. antennatus | Japan | 1 |
| Sirex Host | Samples, n | A. areolatum | A. chailletii | |
|---|---|---|---|---|
| IGS-D2 | IGS-B1D2 | |||
| Sirex noctilio | 26 | - | 26 | - |
| Sirex nitobei | 44 | 18 | 2 | 24 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, M.; Fu, N.; Gao, C.; Wang, L.; Ren, L.; Luo, Y. Multilocus Genotyping and Intergenic Spacer Single Nucleotide Polymorphisms of Amylostereum areolatum (Russulales: Amylostereacea) Symbionts of Native and Non-Native Sirex Species. J. Fungi 2021, 7, 1065. https://doi.org/10.3390/jof7121065
Wang M, Fu N, Gao C, Wang L, Ren L, Luo Y. Multilocus Genotyping and Intergenic Spacer Single Nucleotide Polymorphisms of Amylostereum areolatum (Russulales: Amylostereacea) Symbionts of Native and Non-Native Sirex Species. Journal of Fungi. 2021; 7(12):1065. https://doi.org/10.3390/jof7121065
Chicago/Turabian StyleWang, Ming, Ningning Fu, Chenglong Gao, Lixia Wang, Lili Ren, and Youqing Luo. 2021. "Multilocus Genotyping and Intergenic Spacer Single Nucleotide Polymorphisms of Amylostereum areolatum (Russulales: Amylostereacea) Symbionts of Native and Non-Native Sirex Species" Journal of Fungi 7, no. 12: 1065. https://doi.org/10.3390/jof7121065
APA StyleWang, M., Fu, N., Gao, C., Wang, L., Ren, L., & Luo, Y. (2021). Multilocus Genotyping and Intergenic Spacer Single Nucleotide Polymorphisms of Amylostereum areolatum (Russulales: Amylostereacea) Symbionts of Native and Non-Native Sirex Species. Journal of Fungi, 7(12), 1065. https://doi.org/10.3390/jof7121065








