Analysis of Transposable Elements in Coccidioides Species
Abstract
1. Introduction
2. Materials and Methods
2.1. Genomes
2.2. Analysis tools
3. Results
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Teixeira, M.M.; Barker, B.M. Use of Population Genetics to Assess the Ecology, Evolution, and Population Structure of Coccidioides. Emerg. Infect. Dis. 2016, 22, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Fisher, M.C.; Koenig, G.; White, T.J.; Taylor, J.W. A test for concordance between the multilocus genealogies of genes and microsatellites in the pathogenic fungus Coccidioides immitis. Mol. Biol. Evol. 2000, 17, 1164–1174. [Google Scholar] [CrossRef] [PubMed]
- Neafsey, D.E.; Barker, B.M.; Sharpton, T.J.; Park, D.J.; Whiston, E.; Hung, C.-Y.; McMahan, C.; White, J.; Sykes, S.; Heiman, D.; et al. Population genomic sequencing of Coccidioides Fungi reveals recent hybridization and transposon control. Genome Res. 2010, 20, 938–946. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.A.; Tarcha, E.J.; Cole, G.T.; Inglis, D.O.; Sil, A.; Heitman, J. Evolution of the mating type locus: Insights gained from the dimorphic primary fungal pathogens Histoplasma capsulatum, Coccidioides immitis, and Coccidioides posadasii. Eukaryot. Cell 2007, 6, 622–629. [Google Scholar] [CrossRef] [PubMed]
- Mandel, M.A.; Barker, B.M.; Kroken, S.; Rounsley, S.D.; Orbach, M.J. Genomic and population analyses of the mating type loci in Coccidioides species reveal evidence for sexual reproduction and gene acquisition. Eukaryot. Cell 2007, 6, 1189–1199. [Google Scholar] [CrossRef] [PubMed]
- Kirkland, T.N.; Fierer, J. Coccidioidomycosis: A reemerging infectious disease. Emerg. Infect. Dis. 1996, 2, 192–199. [Google Scholar] [CrossRef] [PubMed]
- Valdivia, L.; Nix, D.; Wright, M.; Lindberg, E.; Fagan, T.; Lieberman, D.; Stoffer, T.; Ampel, N.M.; Galgiani, J.N. Coccidioidomycosis as a common cause of community-acquired pneumonia. Emerg. Infect. Dis. 2006, 12, 958–962. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, C.; Barker, B.M.; Hoover, S.; Nix, D.E.; Ampel, N.M.; Frelinger, J.A.; Orbach, M.J.; Galgiani, J.N. Recent advances in our understanding of the environmental, epidemiological, immunological, and clinical dimensions of coccidioidomycosis. Clin. Microbiol. Rev. 2013, 26, 505–525. [Google Scholar] [CrossRef] [PubMed]
- Whiston, E.; Zhang Wise, H.; Sharpton, T.J.; Jui, G.; Cole, G.T.; Taylor, J.W. Comparative transcriptomics of the saprobic and parasitic growth phases in Coccidioides spp. PLoS ONE 2012, 7, e41034. [Google Scholar] [CrossRef] [PubMed]
- Converse, J.L. Effect of physico-chemical environment of spherulation of Coccidioides immitis in a chemically defined medium. J. Bacteriol. 1956, 72, 784–792. [Google Scholar] [PubMed]
- Viriyakosol, S.; Singhania, A.; Fierer, J.; Goldberg, J.; Kirkland, T.N.; Woelk, C.H. Gene expression in human fungal pathogen Coccidioides immitis changes as arthroconidia differentiate into spherules and mature. BMC Microbiol. 2013, 13, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kirkland, T.N. A few shared up-regulated genes may influence conidia to yeast transformation in dimorphic fungal pathogens: Table 1. Med. Mycol. 2016, 54, 648–653. [Google Scholar] [CrossRef] [PubMed]
- Boyce, K.J.; McLauchlan, A.; Schreider, L.; Andrianopoulos, A. Intracellular Growth Is Dependent onTyrosine Catabolism in the Dimorphic FungalPathogen Penicillium marneffei. PLoS Pathog. 2015, 11, e1004790. [Google Scholar] [CrossRef] [PubMed]
- Fedoroff, N.V. Presidential address. Transposable elements, epigenetics, and genome evolution. Science 2012, 338, 758–767. [Google Scholar] [CrossRef] [PubMed]
- Feschotte, C.; Pritham, E.J. DNA Transposons and the Evolution of Eukaryotic Genomes. Annu. Rev. Genet. 2007, 41, 331–368. [Google Scholar] [CrossRef] [PubMed]
- Rebollo, R.; Romanish, M.T.; Mager, D.L. Transposable Elements: An Abundant and Natural Source of Regulatory Sequences for Host Genes. Annu. Rev. Genet. 2012, 46, 21–42. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Leem, Y.-E.; Levin, H.L. Transposon integration enhances expression of stress response genes. Nucleic Acids Res. 2012, 41, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Casacuberta, E.; González, J. The impact of transposable elements in environmental adaptation. Mol. Ecol. 2013, 22, 1503–1517. [Google Scholar] [CrossRef] [PubMed]
- Biemont, C. A Brief History of the Status of Transposable Elements: From Junk DNA to Major Players in Evolution. Genetics 2010, 186, 1085–1093. [Google Scholar] [CrossRef] [PubMed]
- Kempken, F.; Kück, U. Transposons in filamentous Fungi—Facts and perspectives. Bioessays 1998, 20, 652–659. [Google Scholar] [CrossRef]
- Parlange, F.; Oberhaensli, S.; Breen, J.; Platzer, M.; Taudien, S.; Šimková, H.; Wicker, T.; Doležel, J.; Keller, B. A major invasion of transposable elements accounts for the large size of the Blumeria graminis f.sp. tritici genome. Funct. Integr. Genom. 2011, 11, 671–677. [Google Scholar] [CrossRef] [PubMed]
- Seberg, O.; Petersen, G. A unified classification system for eukaryotic transposable elements should reflect their phylogeny. Nat. Rev. Genet. 2009, 10, 276. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.-W.; Wessler, S.R. The catalytic domain of all eukaryotic cut-and-paste transposase superfamilies. Proc. Natl. Acad. Sci. USA 2011, 108, 7884–7889. [Google Scholar] [CrossRef] [PubMed]
- Santana, M.F.; Silva, J.C.; Mizubuti, E.S.; Araújo, E.F.; Condon, B.J.; Turgeon, B.; Queiroz, M.V. Characterization and potential evolutionary impact of transposable elements in the genome of Cochliobolus heterostrophus. BMC Genom. 2014, 15, 536. [Google Scholar] [CrossRef] [PubMed]
- Muszewska, A.; Hoffman-Sommer, M.; Grynberg, M. LTR Retrotransposons in Fungi. PLoS ONE 2011, 6, e29425. [Google Scholar] [CrossRef] [PubMed]
- Daboussi, M.-J.; Capy, P. Transposable Elements in Filamentous Fungi. Annu. Rev. Microbiol. 2003, 57, 275–299. [Google Scholar] [CrossRef] [PubMed]
- Grandaubert, J.; Balesdent, M.-H.; Rouxel, T. Evolutionary and Adaptive Role of Transposable Elements in Fungal Genomes, 1st ed.; Elsevier Ltd.: Amsterdam, The Netherlands, 2014; Volume 70, pp. 79–107. [Google Scholar]
- Atkinson, P.W. hAT Transposable Elements. Microbiol. Spectr. 2015, 3, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Sharpton, T.J.; Rounsley, S.D.; Gardner, M.J.; Wortman, J.R.; Jordar, V.S.; Maiti, R.; Kodira, C.D.; Neafsey, D.E.; Zeng, Q.; Hung, C.-Y.; et al. Comparative genomic analyses of the human fungal pathogens Coccidioides and their relatives. Genome Res. 2009, 19, 1722–1731. [Google Scholar] [CrossRef] [PubMed]
- Harris, T.; Brunk, B.P.; Brestelli, J.; Fischer, S.; Harb, O.S.; Kissinger, J.C.; Li, W.; Nayak, V.; Pinney, D.F.; Stoeckert, C.J.; et al. FungiDB: An integrated functional genomics database for Fungi. Nucleic Acids Res. 2012, 40, D675–D681. [Google Scholar]
- FingiDB. Available online: http://fungidb.org/fungidb/ (accessed on 10 January 2017).
- RepeatMasker. Available online: http://www.repeatmasker.org/ (accessed on 6 January 2017).
- Castanera, R.; López-Varas, L.; Borgognone, A.; LaButti, K.; Lapidus, A.; Schmutz, J.; Grimwood, J.; Pérez, G.; Pisabarro, A.G.; Grigoriev, I.V.; et al. Transposable Elements versus the Fungal Genome: Impact on Whole-Genome Architecture and Transcriptional Profiles. PLoS Genet. 2016, 12, e1006108. [Google Scholar] [CrossRef] [PubMed]
- GitHub. Available online: https://github.com/hyphaltip/cocci_repeats/ (accessed on 10 January 2017).
- Galaxy. Available online: https://usegalaxy.org/ (accessed on 9 January 2017).
- iTOL. Available online: https://itol.embl.de/ (accessed on 7 January 2017).
- Integrative Genomics Viewer. Available online: http://software.broadinstitute.org/software/igv/userguide (accessed on 12 October 2017).
- Clutterbuck, A.J. Genomic evidence of repeat-induced point mutation (RIP) in filamentous ascomycetes. Fungal Genet. Biol. 2011, 48, 306–326. [Google Scholar] [CrossRef] [PubMed]
- Baucom, R.S.; Estill, J.C.; Chaparro, C.; Upshaw, N.; Jogi, A.; Deragon, J.-M.; Westerman, R.P.; SanMiguel, P.J.; Bennetzen, J.L. Exceptional Diversity, Non-Random Distribution, and Rapid Evolution of Retroelements in the B73 Maize Genome. PLoS Genet. 2009, 5, e1000732. [Google Scholar] [CrossRef] [PubMed]
- Dhillon, B.; Gill, N.; Hamelin, R.C.; Goodwin, S.B. The landscape of transposable elements in the finished genome of the fungal wheat pathogen Mycosphaerella graminicola. BMC Genom. 2014, 15, 1132–1149. [Google Scholar] [CrossRef] [PubMed]
- Labbé, J.; Murat, C.; Morin, E.; Tuskan, G.A.; Le Tacon, F.; Martin, F. Characterization of Transposable Elements in the Ectomycorrhizal Fungus Laccaria bicolor. PLoS ONE 2012, 7, e40197. [Google Scholar] [CrossRef] [PubMed]
- Yadav, C.B.; Bonthala, V.S.; Muthamilarasan, M.; Pandey, G.; Khan, Y.; Prasad, M. Genome-wide development of transposable elements-based markers in foxtail millet and construction of an integrated database. DNA Res. 2015, 22, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Loftus, B.J.; Fung, E.; Roncaglia, P.; Rowley, D.; Amedeo, P.; Bruno, D.; Vamathevan, J.; Miranda, M.; Anderson, I.J.; Fraser, J.A.; et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 2005, 307, 1321–1324. [Google Scholar] [CrossRef] [PubMed]
- Devine, S.E.; Boeke, J.D. Integration of the yeast retrotransposon Ty1 is targeted to regions upstream of genes transcribed by RNA polymerase III. Genes Dev. 1996, 10, 620–633. [Google Scholar] [CrossRef] [PubMed]
- Elbarbary, R.A.; Lucas, B.A.; Maquat, L.E. Retrotransposons as regulators of gene expression. Science 2016, 351, aac7247. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Weigel, D.; Smith, L.M. Transposon Variants and Their Effects on Gene Expression in Arabidopsis. PLoS Genet. 2013, 9, e1003255. [Google Scholar] [CrossRef] [PubMed]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory activities of transposable elements: From conflicts to benefits. Nat. Rev. Genet. 2017, 18, 71–86. [Google Scholar] [CrossRef] [PubMed]
- Makarevitch, I.; Waters, A.J.; West, P.T.; Stitzer, M.; Hirsch, C.N.; Ross-Ibarra, J.; Springer, N.M. Transposable Elements Contribute to Activation of Maize Genes in Response to Abiotic Stress. PLoS Genet. 2015, 11, e1004915. [Google Scholar] [CrossRef] [PubMed]
- Cowley, M.; Oakey, R.J. Transposable elements re-wire and fine-tune the transcriptome. PLoS Genet. 2013, 9, e1003234. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yazaki, J.; Sundaresan, A.; Cokus, S.; Chan, S.W.L.; Chen, H.; Henderson, I.R.; Shinn, P.; Pellegrini, M.; Jacobsen, S.E.; et al. Genome-wide High-Resolution Mapping and Functional Analysis of DNA Methylation in Arabidopsis. Cell 2006, 126, 1189–1201. [Google Scholar] [CrossRef] [PubMed]
- Aramayo, R.; Selker, E.U. Neurospora crassa, a Model System for Epigenetics Research. Cold Spring Harb. Perspect. Biol. 2013, 5, a017921. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, X.; He, K.; Ma, Y.; Su, N.; He, H.; Stolc, V.; Tongprasit, W.; Jin, W.; Jiang, J.; et al. High-Resolution Mapping of Epigenetic Modifications of the Rice Genome Uncovers Interplay between DNA Methylation, Histone Methylation, and Gene Expression. Plant Cell Online 2008, 20, 259–276. [Google Scholar] [CrossRef] [PubMed]
- Lippman, Z.; Gendrel, A.-V.; Black, M.; Vaughn, M.W.; Dedhia, N.; McCombie, W.R.; Lavine, K.; Mittal, V.; May, B.; Kasschau, K.D.; et al. Role of transposable elements in heterochromatin and epigenetic control. Nature 2004, 430, 471–476. [Google Scholar] [CrossRef] [PubMed]
- Nicolás, F.E.; Garre, V. RNA Interference in Fungi: Retention and Loss. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Zhu, Y.; Dai, J.; Fuerst, P.G.; Voytas, D.F. Controlling integration specificity of a yeast retrotransposon. Proc. Natl. Acad. Sci. USA 2003, 100, 5891–5895. [Google Scholar] [CrossRef] [PubMed]
- Hua-Van, A.; Le Rouzic, A.; Maisonhaute, C.; Capy, P. Abundance, distribution and dynamics of retrotransposable elements and transposons: Similarities and differences. Cytogenet. Genome Res. 2005, 110, 426–440. [Google Scholar] [CrossRef] [PubMed]
- Miyao, A. Target Site Specificity of the Tos17 Retrotransposon Shows a Preference for Insertion within Genes and against Insertion in Retrotransposon-Rich Regions of the Genome. Plant Cell Online 2003, 15, 1771–1780. [Google Scholar] [CrossRef]
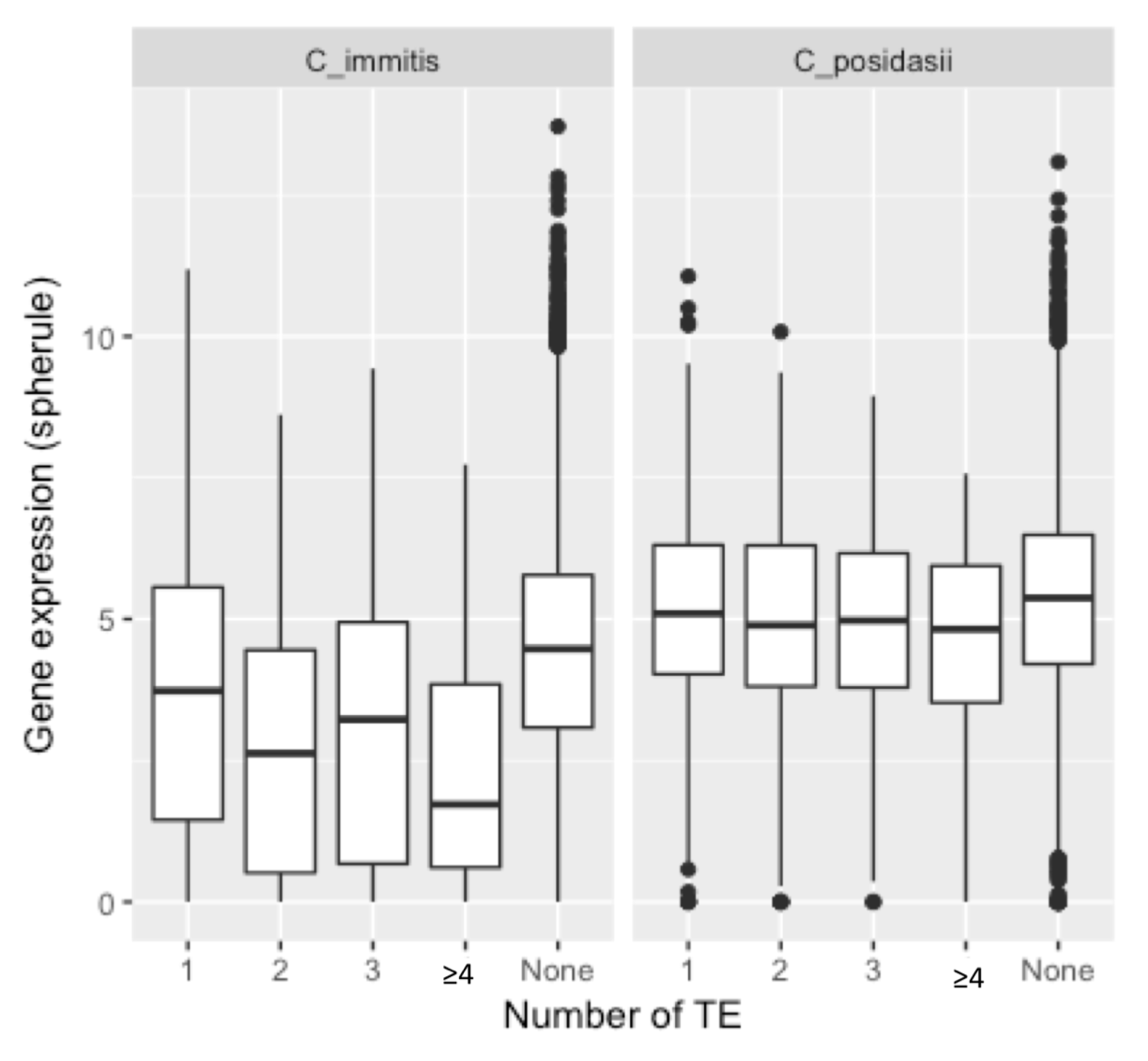
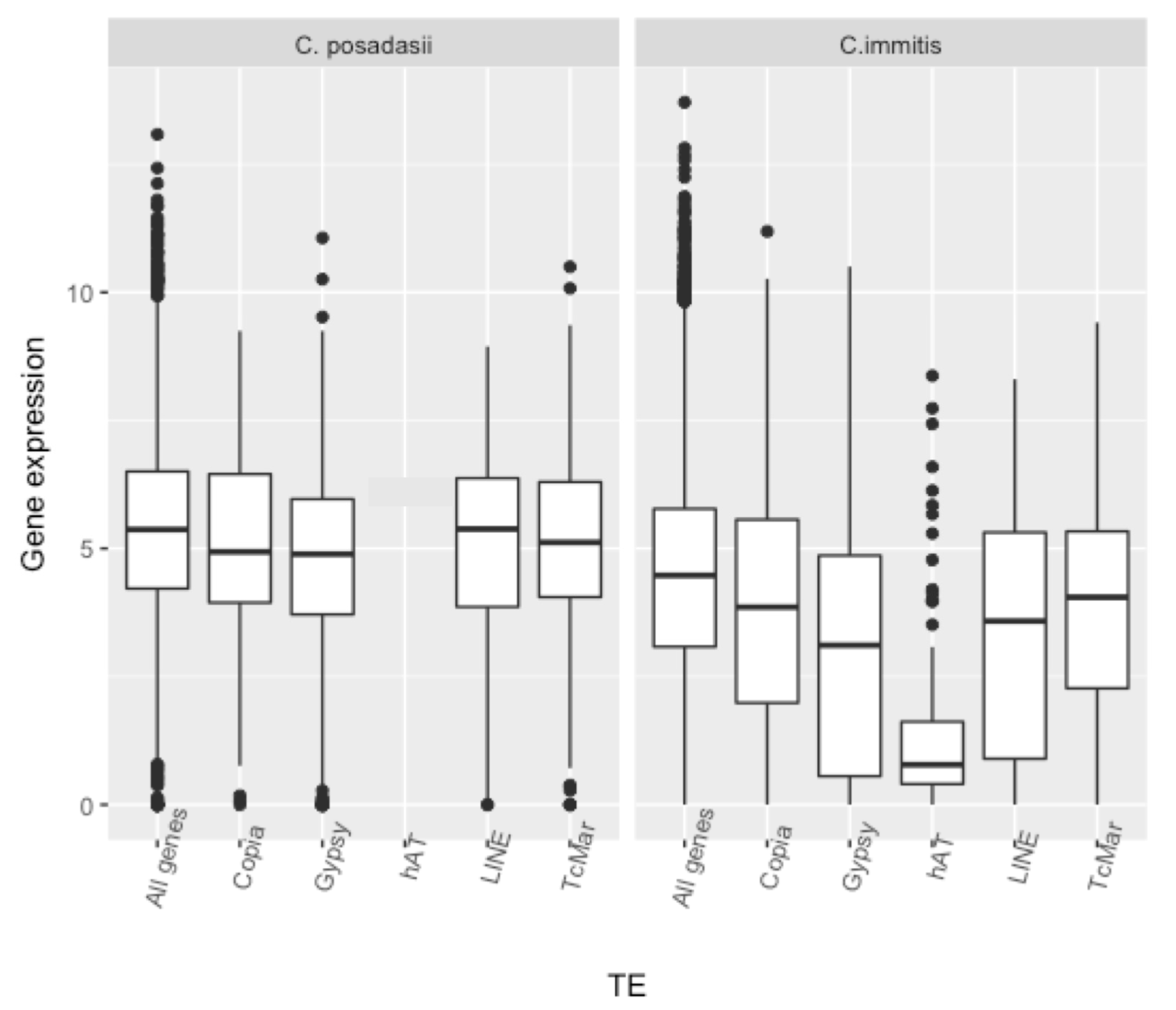
| C. immitis | C. posadasii | |||||||
|---|---|---|---|---|---|---|---|---|
| Type | Number | Mean Length | ORFs > 300 nt (%) | GC Content (%) | Number | Mean length | ORF > 300 nt (%) | GC Content (%) |
| DNA/TcMar | 286 | 884 | 113 (40) | 37 | 575 | 991 | 456 (79) | 36 |
| -Ant1 | 63 | 31 | ||||||
| -Fot1 | 134 | 430 | ||||||
| -Pogo | 27 | 29 | ||||||
| -Tc1 | 66 | 83 | ||||||
| DNA/hAT | 100 | 2416 | 80 (80) | 36 | 37 | 1232 | 15 (40) | 31 |
| LTR/Gypsy | 1204 | 2089 | 1557 (130) | 33 | 1199 | 2046 | 1387 (116) | 34 |
| LTR/Copia | 287 | 1351 | 151 (52) | 38 | 190 | 937 | 131 (69) | 39 |
| LINE | 364 | 1331 | 239 (66) | 28 | 225 | 1237 | 144 (64) | 31 |
| C. immitis | C. posadasii | |||||
|---|---|---|---|---|---|---|
| Total | Pol Domain | None | Total | Pol Domain | None | |
| Number of TEs | 1204 | 338 (28%) | 866 (72%) | 1199 | 260 (22%) | 938 (78%) |
| Associated loci | 571 | 60 (11%) | 511 (89%) | 341 | 24 (7%) | 317 (91%) |
| Number of TEs | |||||
|---|---|---|---|---|---|
| C. posadasii orthologs | ≥4 | 3 | 2 | 1 | None |
| 27 | 17 | 2 | 5 | 4 | 1 |
| Control | Upstream a | Overlap b | Downstream c | ||||
|---|---|---|---|---|---|---|---|
| Mean | Number | Mean | Number | Mean | Number | Mean | |
| All genes | 4.032 | 992 | 773 | ||||
| TcMar | 115 | 3.734 | 131 | 3.759 | 115 | 3.734 | |
| hAT | 75 | 1.134 | 73 | 1.135 | 77 | 1.292 | |
| Copia | 119 | 3.242 | 69 | 3.343 | 137 | 4.229 | |
| Gypsy | 272 | 3.027 | 217 | 2.001 | 312 | 2.156 | |
| C. immitisb | C. posadasii | ||||
|---|---|---|---|---|---|
| ID | Name | Odds Ratio | p (Bonferroni) | Odds Ratio | p (Bonferroni) |
| GO:0016310 | Phosphorylation | 3.35 | 1.43 × 10−7 | 3.34 | 7.35 × 10−7 |
| GO:0006468 | protein phosphorylation | 3.47 | 1.56 × 10−7 | 3.60 | 2.10 × 10−6 |
| GO:0036211 | protein modification process | 2.72 | 3.87 × 10−6 | 2.48 | 2.90 × 10−4 |
| GO:0006464 | cellular protein modification process | 2.72 | 3.87 × 10−6 | 2.48 | 2.90 × 10−4 |
| GO:0006796 | phosphate-containing compound metabolic process | 2.45 | 1.17 × 10−5 | 2.07 | 4.41 × 10−3 |
| GO:0006793 | phosphorus metabolic process | 2.43 | 1.36 × 10−5 | 2.05 | 5.00 × 10−3 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kirkland, T.N.; Muszewska, A.; Stajich, J.E. Analysis of Transposable Elements in Coccidioides Species. J. Fungi 2018, 4, 13. https://doi.org/10.3390/jof4010013
Kirkland TN, Muszewska A, Stajich JE. Analysis of Transposable Elements in Coccidioides Species. Journal of Fungi. 2018; 4(1):13. https://doi.org/10.3390/jof4010013
Chicago/Turabian StyleKirkland, Theo N., Anna Muszewska, and Jason E. Stajich. 2018. "Analysis of Transposable Elements in Coccidioides Species" Journal of Fungi 4, no. 1: 13. https://doi.org/10.3390/jof4010013
APA StyleKirkland, T. N., Muszewska, A., & Stajich, J. E. (2018). Analysis of Transposable Elements in Coccidioides Species. Journal of Fungi, 4(1), 13. https://doi.org/10.3390/jof4010013