Inhibition of let-7c Regulates Cardiac Regeneration after Cryoinjury in Adult Zebrafish
Abstract
1. Introduction
2. Materials and Methods
2.1. Zebrafish
2.2. Cryoinjury
2.3. let-7c Inhibition
2.4. Echocardiography
2.5. RNA Isolation and Quantitative Real-Time PCR (qPCR)
2.6. 5-Bromo-2-Deoxyuridine (BRDU) Baths
2.7. Histology
2.8. Immunohistochemistry
2.9. Statistical Analysis
3. Results
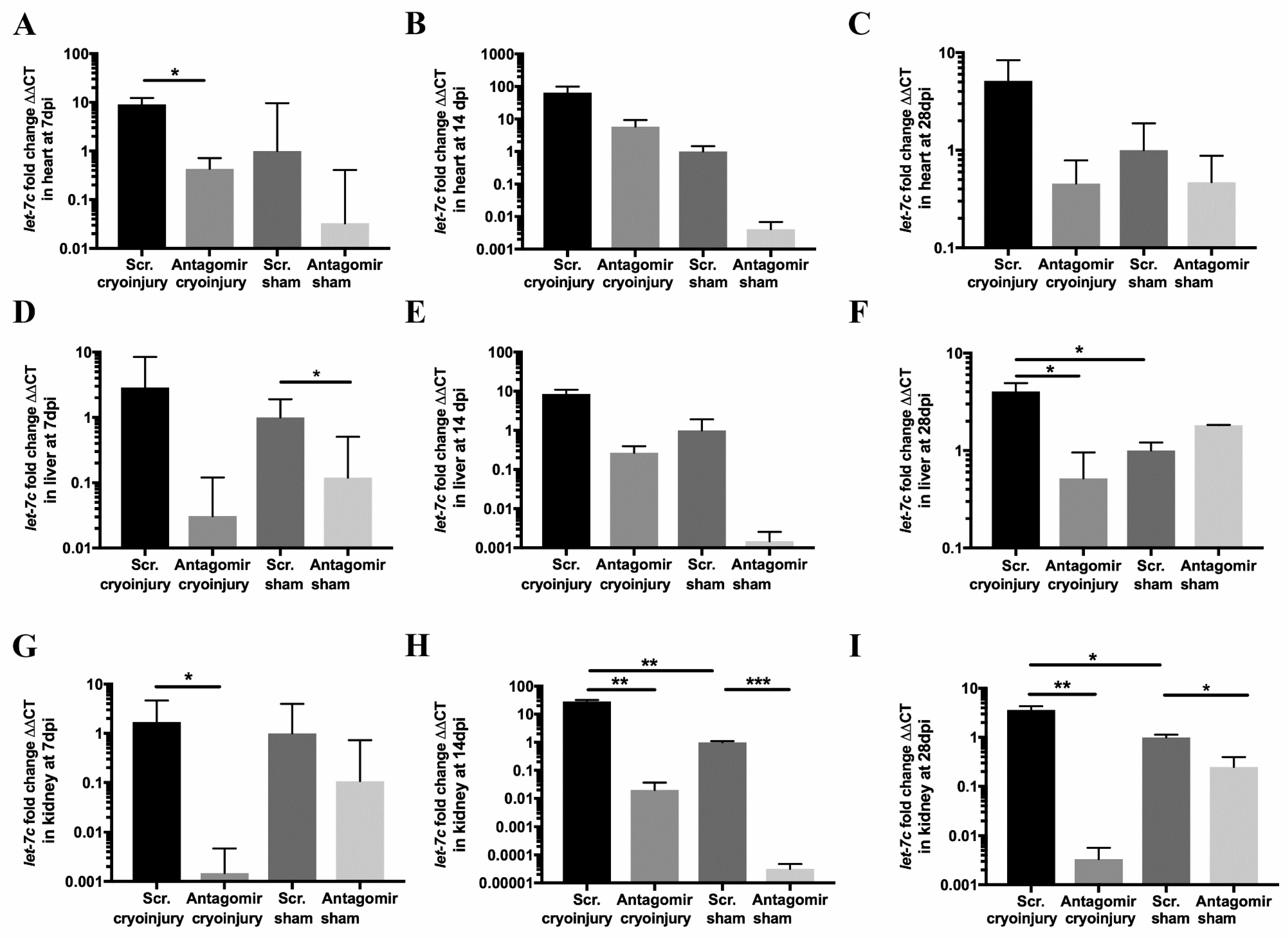
3.1. let-7c Expression after Cryoinjury
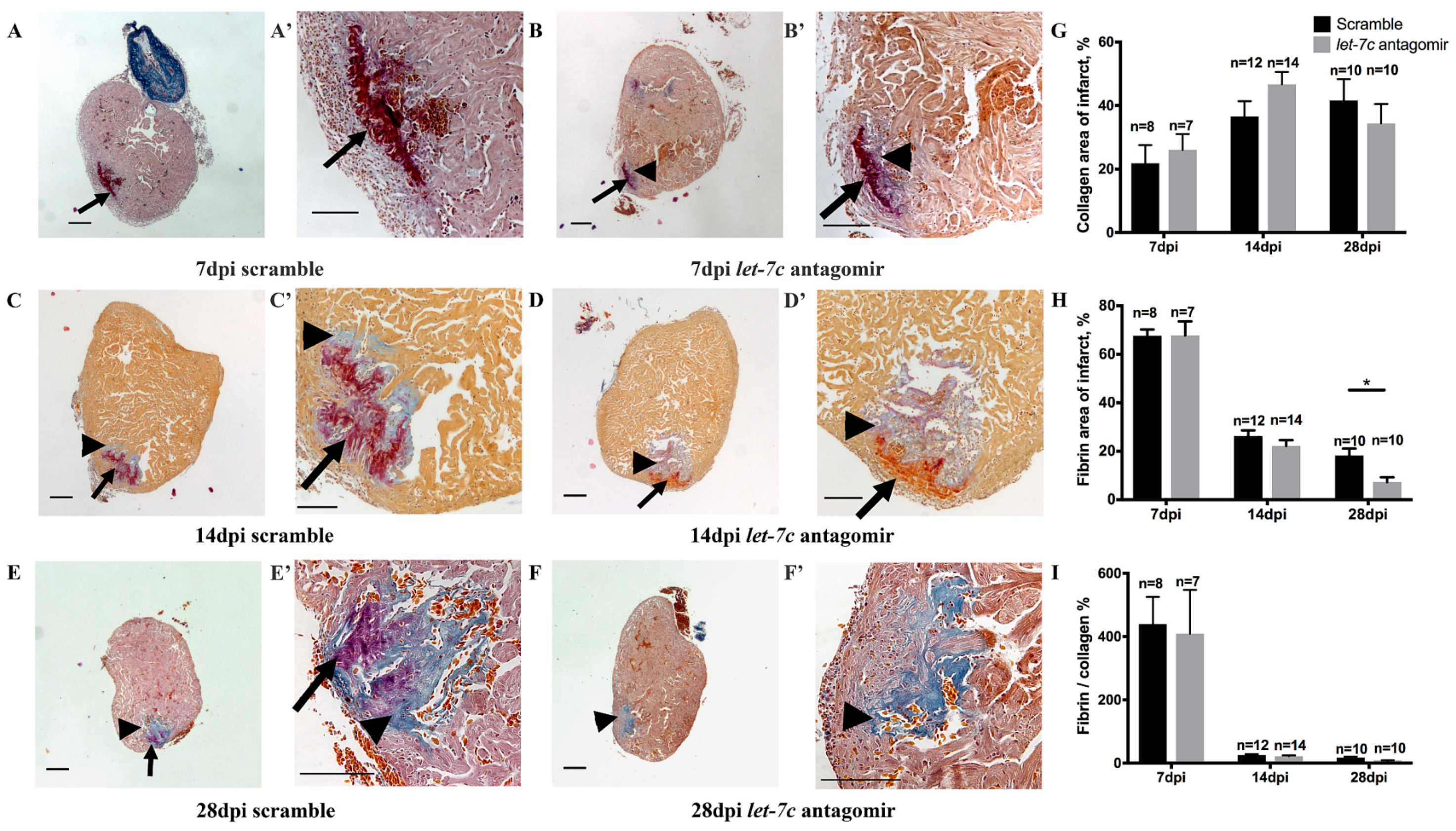
3.2. Fibrin and Collagen Deposition after Cryoinjury
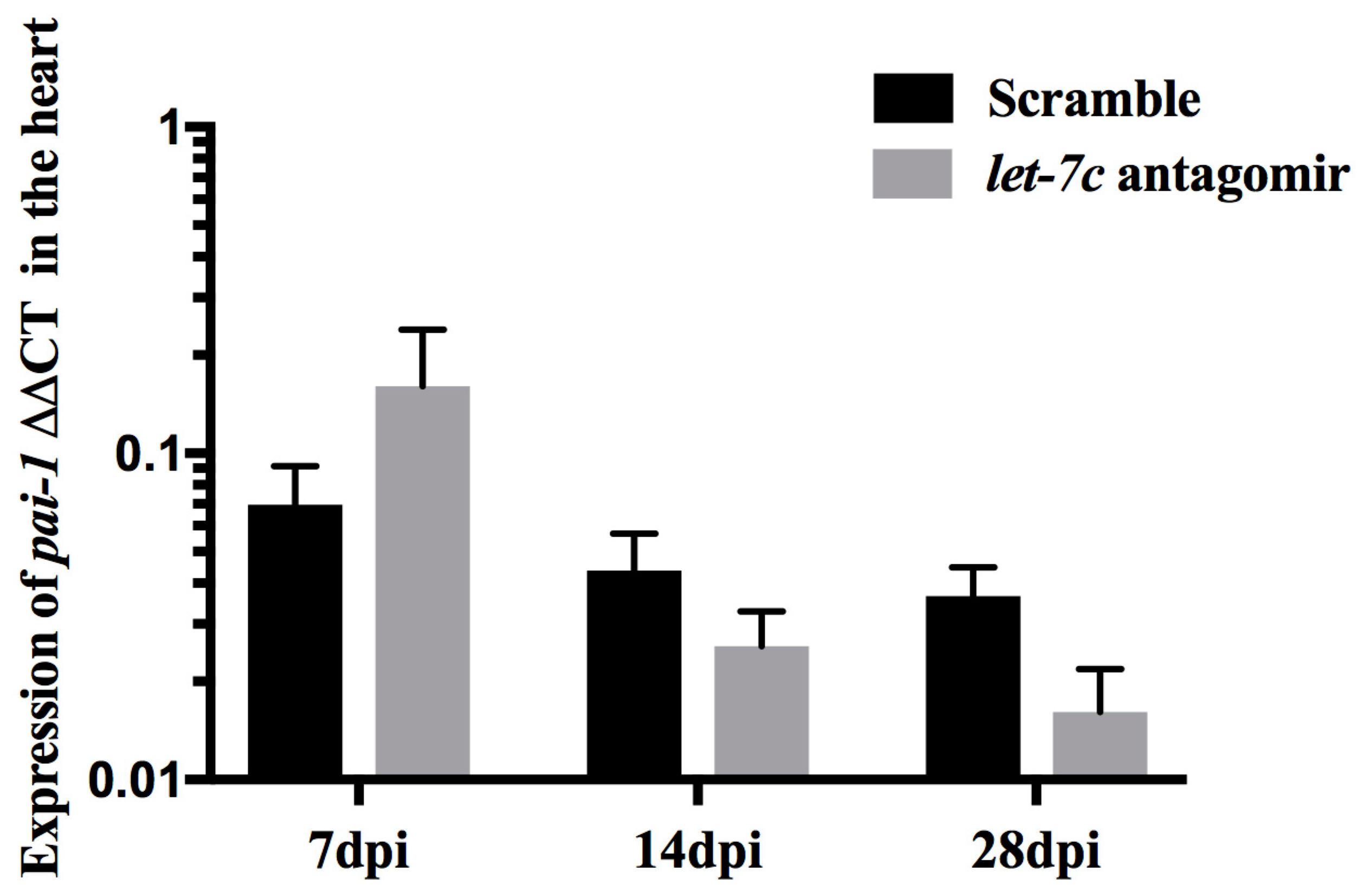
3.3. Expression of pai-1
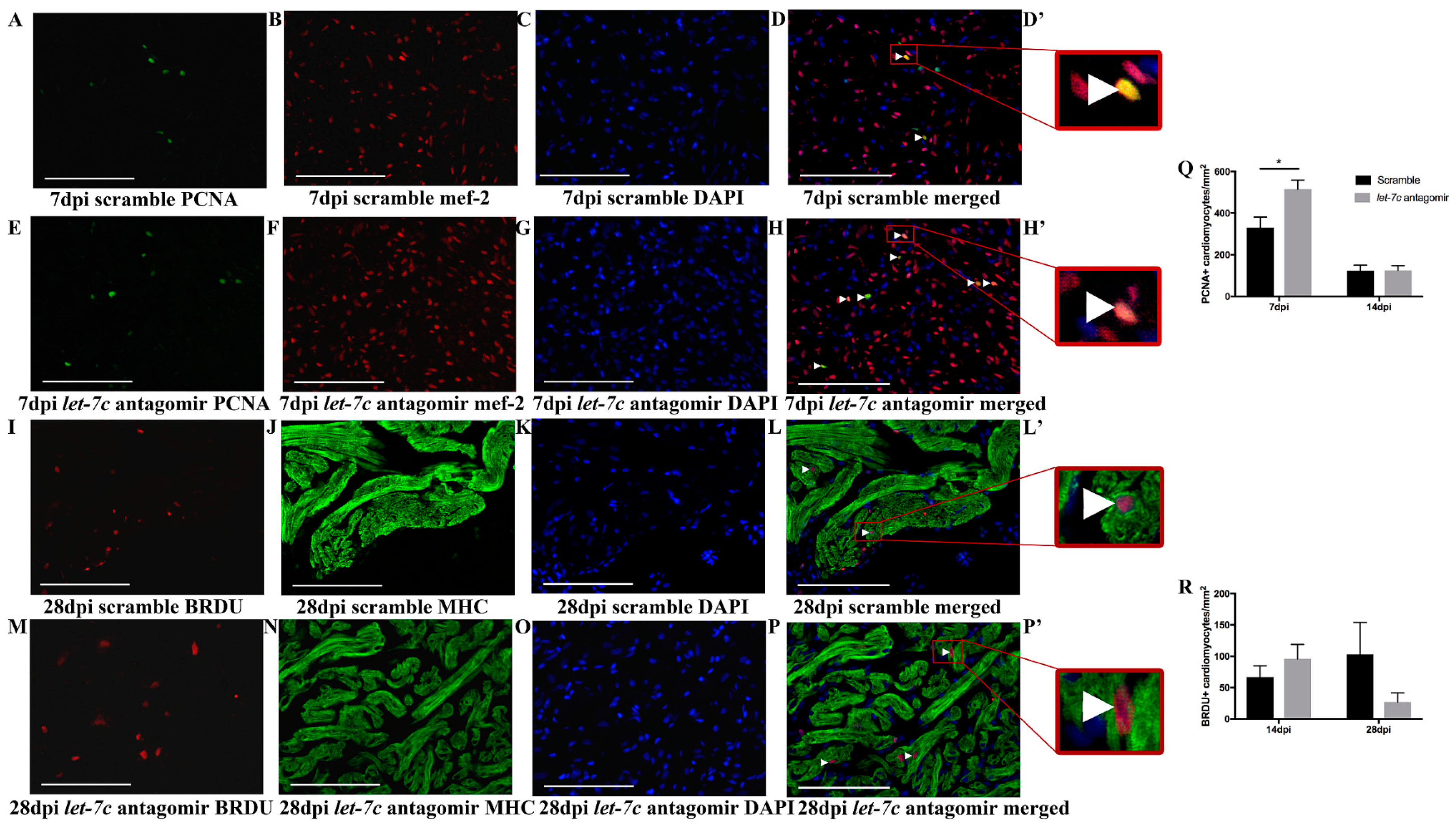
3.4. Proliferating Cardiomyocytes
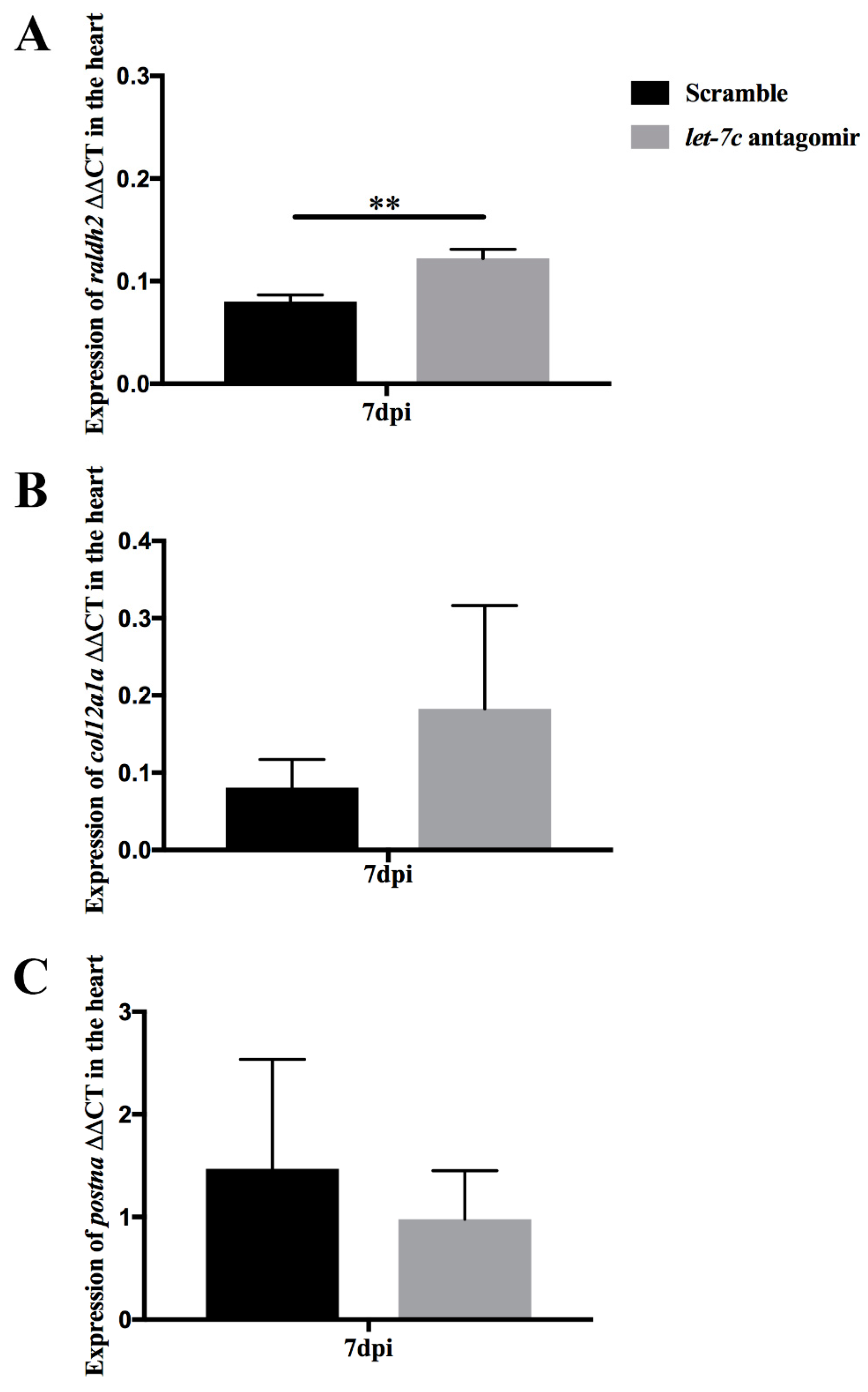
3.5. Epicardial Activation through raldh2 Expression
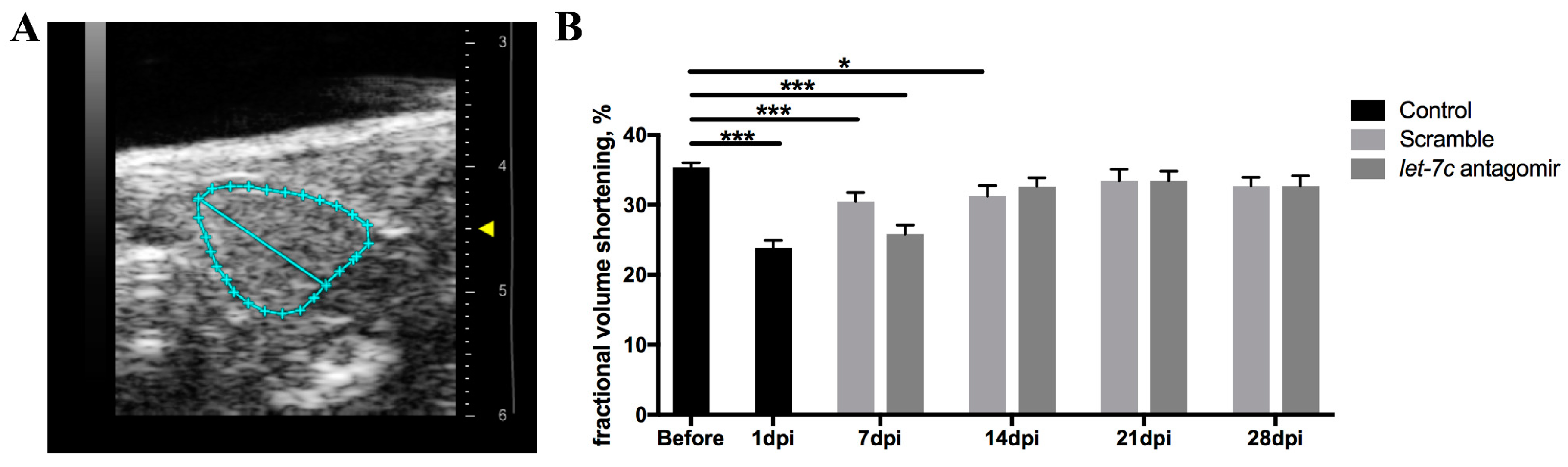
3.6. Echocardiography
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Benjamin, E.J.; Virani, S.S.; Callaway, C.W.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Chiuve, S.E.; Cushman, M.; Delling, F.N.; Deo, R.; et al. American Heart Association Council on Epidemiology and Prevention Statistics Committee and Stroke Statistics Subcommittee Heart Disease and Stroke Statistics-2018 Update: A Report from the American Heart Association. Circulation 2018, 137, e67–e492. [Google Scholar] [CrossRef]
- Laflamme, M.A.; Murry, C.E. Heart regeneration. Nature 2011, 473, 326–335. [Google Scholar] [CrossRef]
- Senyo, S.E.; Steinhauser, M.L.; Pizzimenti, C.L.; Yang, V.K.; Cai, L.; Wang, M.; Wu, T.D.; Guerquin-Kern, J.L.; Lechene, C.P.; Lee, R.T. Mammalian heart renewal by pre-existing cardiomyocytes. Nature 2013, 493, 433–436. [Google Scholar] [CrossRef]
- Chablais, F.; Veit, J.; Rainer, G.; Jazwinska, A. The zebrafish heart regenerates after cryoinjury-induced myocardial infarction. BMC Dev. Biol. 2011, 11, 21. [Google Scholar] [CrossRef]
- Gonzalez-Rosa, J.M.; Martin, V.; Peralta, M.; Torres, M.; Mercader, N. Extensive scar formation and regression during heart regeneration after cryoinjury in zebrafish. Development 2011, 138, 1663–1674. [Google Scholar] [CrossRef]
- Lieschke, G.J.; Currie, P.D. Animal models of human disease: Zebrafish swim into view. Nat. Rev. Genet. 2007, 8, 353–367. [Google Scholar] [CrossRef]
- Bakkers, J. Zebrafish as a model to study cardiac development and human cardiac disease. Cardiovasc. Res. 2011, 91, 279–288. [Google Scholar] [CrossRef]
- Chico, T.J.; Ingham, P.W.; Crossman, D.C. Modeling cardiovascular disease in the zebrafish. Trends Cardiovasc. Med. 2008, 18, 150–155. [Google Scholar] [CrossRef]
- Jopling, C.; Sleep, E.; Raya, M.; Marti, M.; Raya, A.; Izpisua Belmonte, J.C. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature 2010, 464, 606–609. [Google Scholar] [CrossRef]
- Kikuchi, K.; Holdway, J.E.; Major, R.J.; Blum, N.; Dahn, R.D.; Begemann, G.; Poss, K.D. Retinoic acid production by endocardium and epicardium is an injury response essential for zebrafish heart regeneration. Dev. Cell 2011, 20, 397–404. [Google Scholar] [CrossRef]
- Kikuchi, K.; Holdway, J.E.; Werdich, A.A.; Anderson, R.M.; Fang, Y.; Egnaczyk, G.F.; Evans, T.; Macrae, C.A.; Stainier, D.Y.; Poss, K.D. Primary contribution to zebrafish heart regeneration by gata4(+) cardiomyocytes. Nature 2010, 464, 601–605. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Panakova, D.; Kikuchi, K.; Holdway, J.E.; Gemberling, M.; Burris, J.S.; Singh, S.P.; Dickson, A.L.; Lin, Y.F.; Sabeh, M.K.; et al. The regenerative capacity of zebrafish reverses cardiac failure caused by genetic cardiomyocyte depletion. Development 2011, 138, 3421–3430. [Google Scholar] [CrossRef] [PubMed]
- Lepilina, A.; Coon, A.N.; Kikuchi, K.; Holdway, J.E.; Roberts, R.W.; Burns, C.G.; Poss, K.D. A dynamic epicardial injury response supports progenitor cell activity during zebrafish heart regeneration. Cell 2006, 127, 607–619. [Google Scholar] [CrossRef] [PubMed]
- Karra, R.; Poss, K.D. Redirecting cardiac growth mechanisms for therapeutic regeneration. J. Clin. Investig. 2017, 127, 427–436. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, C.E.; Bakovic, M.; Karra, R. Endothelial Contributions to Zebrafish Heart Regeneration. J. Cardiovasc. Dev. Dis. 2018, 5, 56. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Rosa, J.M.; Peralta, M.; Mercader, N. Pan-epicardial lineage tracing reveals that epicardium derived cells give rise to myofibroblasts and perivascular cells during zebrafish heart regeneration. Dev. Biol. 2012, 370, 173–186. [Google Scholar] [CrossRef] [PubMed]
- Chablais, F.; Jazwinska, A. Induction of myocardial infarction in adult zebrafish using cryoinjury. J. Vis. Exp. 2012, 62, 3666. [Google Scholar] [CrossRef] [PubMed]
- Chablais, F.; Jazwinska, A. The regenerative capacity of the zebrafish heart is dependent on TGFβ signalling. Development 2012, 139, 1921–1930. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Iranzo, H.; Galardi-Castilla, M.; Sanz-Morejon, A.; Gonzalez-Rosa, J.M.; Costa, R.; Ernst, A.A.; Sainz de Aja, J.; Langa, X.; Mercader, N. Transient fibrosis resolves via fibroblast inactivation in the regenerating zebrafish heart. Proc. Natl. Acad. Sci. USA 2018, 115, 4188–4193. [Google Scholar] [CrossRef]
- Van Rooij, E.; Kauppinen, S. Development of microRNA therapeutics is coming of age. EMBO Mol. Med. 2014, 6, 851–864. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Van Rooij, E.; Olson, E.N. MicroRNA therapeutics for cardiovascular disease: Opportunities and obstacles. Nat. Rev. Drug Discov. 2012, 11, 860–872. [Google Scholar] [CrossRef]
- Seeger, T.; Xu, Q.F.; Muhly-Reinholz, M.; Fischer, A.; Kremp, E.M.; Zeiher, A.M.; Dimmeler, S. Inhibition of let-7 augments the recruitment of epicardial cells and improves cardiac function after myocardial infarction. J. Mol. Cell Cardiol. 2016, 94, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ridzon, D.; Lee, C.T.; Blake, J.; Sun, Y.; Strauss, W.M. Defining embryonic stem cell identity using differentiation-related microRNAs and their potential targets. Mamm. Genome 2007, 18, 316–327. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.W.; Zhou, L.F.; Liu, Y.L.; Wan, S.M.; Gao, Z.X. Evolution of Fish Let-7 MicroRNAs and Their Expression Correlated to Growth Development in Blunt Snout Bream. Int. J. Mol. Sci. 2017, 18, 646. [Google Scholar] [CrossRef] [PubMed]
- Kuppusamy, K.T.; Jones, D.C.; Sperber, H.; Madan, A.; Fischer, K.A.; Rodriguez, M.L.; Pabon, L.; Zhu, W.Z.; Tulloch, N.L.; Yang, X.; et al. Let-7 family of microRNA is required for maturation and adult-like metabolism in stem cell-derived cardiomyocytes. Proc. Natl. Acad. Sci. USA 2015, 112, E2785–E2794. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, A.; Montserrat, N.; Zacchigna, S.; Nivet, E.; Hishida, T.; Krause, M.N.; Kurian, L.; Ocampo, A.; Vazquez-Ferrer, E.; Rodriguez-Esteban, C.; et al. In vivo activation of a conserved microRNA program induces mammalian heart regeneration. Cell Stem Cell 2014, 15, 589–604. [Google Scholar] [CrossRef]
- Bao, M.H.; Feng, X.; Zhang, Y.W.; Lou, X.Y.; Cheng, Y.; Zhou, H.H. Let-7 in cardiovascular diseases, heart development and cardiovascular differentiation from stem cells. Int. J. Mol. Sci. 2013, 14, 23086–23102. [Google Scholar] [CrossRef]
- Topkara, V.K.; Mann, D.L. Role of MicroRNAs in cardiac remodelling and heart failure. Cardiovasc. Drugs Ther. 2011, 25, 171. [Google Scholar] [CrossRef]
- Tolonen, A.M.; Magga, J.; Szabo, Z.; Viitala, P.; Gao, E.; Moilanen, A.M.; Ohukainen, P.; Vainio, L.; Koch, W.J.; Kerkela, R.; et al. Inhibition of Let-7 microRNA attenuates myocardial remodeling and improves cardiac function postinfarction in mice. Pharmacol. Res. Perspect. 2014, 2, e00056. [Google Scholar] [CrossRef]
- Li, R.; Xiao, J.; Qing, X.; Xing, J.; Xia, Y.; Qi, J.; Liu, X.; Zhang, S.; Sheng, X.; Zhang, X.; et al. Sp1 Mediates a Therapeutic Role of MiR-7a/b in Angiotensin II-Induced Cardiac Fibrosis via Mechanism Involving the TGF-beta and MAPKs Pathways in Cardiac Fibroblasts. PLoS ONE 2015, 10, e0125513. [Google Scholar] [CrossRef]
- Sallinen, V.; Sundvik, M.; Reenilä, I.; Peitsaro, N.; Khrustalyov, D.; Anichtchik, O.; Toleikyte, G.; Kaslin, J.; Panula, P. Hyperserotonergic phenotype after monoamine oxidase inhibition in larval zebrafish. J. Neurochem. 2009, 109, 403–415. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Rosa, J.M.; Guzman-Martinez, G.; Marques, I.J.; Sanchez-Iranzo, H.; Jimenez-Borreguero, L.J.; Mercader, N. Use of echocardiography reveals reestablishment of ventricular pumping efficiency and partial ventricular wall motion recovery upon ventricular cryoinjury in the zebrafish. PLoS ONE 2014, 9, e115604. [Google Scholar] [CrossRef] [PubMed]
- Hein, S.J.; Lehmann, L.H.; Kossack, M.; Juergensen, L.; Fuchs, D.; Katus, H.A.; Hassel, D. Advanced echocardiography in adult zebrafish reveals delayed recovery of heart function after myocardial cryoinjury. PLoS ONE 2015, 10, e0122665. [Google Scholar] [CrossRef] [PubMed]
- Dash, S.N.; Lehtonen, E.; Wasik, A.A.; Schepis, A.; Paavola, J.; Panula, P.; Nelson, W.J.; Lehtonen, S. Sept7b is essential for pronephric function and development of left-right asymmetry in zebrafish embryogenesis. J. Cell Sci. 2014, 127, 1476–1486. [Google Scholar] [CrossRef]
- Li, M.; Naqvi, N.; Yahiro, E.; Liu, K.; Powell, P.C.; Bradley, W.E.; Martin, D.I.; Graham, R.M.; Dell’Italia, L.J.; Husain, A. C-Kit is Required for Cardiomyocyte Terminal Differentiation. Circ. Res. 2008, 102, 677–685. [Google Scholar] [CrossRef]
- Frost, R.J.; van Rooij, E. miRNAs as therapeutic targets in ischemic heart disease. J. Cardiovasc. Transl. Res. 2010, 3, 280–289. [Google Scholar] [CrossRef]
- Frost, R.J.; Olson, E.N. Control of glucose homeostasis and insulin sensitivity by the Let-7 family of microRNAs. Proc. Natl. Acad. Sci. USA 2011, 108, 21075–21080. [Google Scholar] [CrossRef]
- Munch, J.; Grivas, D.; Gonzalez-Rajal, A.; Torregrosa-Carrion, R.; de la Pompa, J.L. Notch signalling restricts inflammation and serpine1 expression in the dynamic endocardium of the regenerating zebrafish heart. Development 2017, 144, 1425–1440. [Google Scholar] [CrossRef] [PubMed]
- Dash, S.N.; Narumanchi, S.; Paavola, J.; Perttunen, S.; Wang, H.; Lakkisto, P.; Tikkanen, I.; Lehtonen, S. Sept7b is required for the subcellular organization of cardiomyocytes and cardiac function in zebrafish. Am. J. Physiol. Heart Circ. Physiol. 2017, 312, H1085–H1095. [Google Scholar] [CrossRef]
- Schnabel, K.; Wu, C.C.; Kurth, T.; Weidinger, G. Regeneration of cryoinjury induced necrotic heart lesions in zebrafish is associated with epicardial activation and cardiomyocyte proliferation. PLoS ONE 2011, 6, e18503. [Google Scholar] [CrossRef] [PubMed]
- Koth, J.; Maguire, M.L.; McClymont, D.; Diffley, L.; Thornton, V.L.; Beech, J.; Patient, R.K.; Riley, P.R.; Schneider, J.E. High-Resolution Magnetic Resonance Imaging of the Regenerating Adult Zebrafish Heart. Sci. Rep. 2017, 7, 2917. [Google Scholar] [CrossRef] [PubMed]
- Hu, N.; Yost, H.J.; Clark, E.B. Cardiac morphology and blood pressure in the adult zebrafish. Anat. Rec. 2001, 264, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Klett, H.; Jurgensen, L.; Most, P.; Busch, M.; Gunther, F.; Dobreva, G.; Leuschner, F.; Hassel, D.; Busch, H.; Boerries, M. Delineating the Dynamic Transcriptome Response of mRNA and microRNA during Zebrafish Heart Regeneration. Biomolecules 2018, 9, 11. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Rosa, J.M.; Burns, C.E.; Burns, C.G. Zebrafish heart regeneration: 15 years of discoveries. Regeneration 2017, 4, 105–123. [Google Scholar] [CrossRef] [PubMed]
| Target | Primer Sequences | Source | Reference |
|---|---|---|---|
| Plasminogen activator inhibitor–1 (pai-1) | Forward 5′-GAGCGTCCCACACCAGATAG-3′ Reverse 5′-GCACTCCAGATGGGAGGAAC-3′ | Sigma-Aldrich | This study |
| Collagen 12ala (col12a1a) | Forward 5′-GGTGAAAGAGGAGACACTGCGT-3′ Reverse 5′-AGTTGCTGGGGATCTGGTT-3′ | Sigma-Aldrich | This study |
| Periostin (postna) | Forward 5′-CAAGGATCAAGACGAAGAGCAAG-3′ Reverse 5′-ATCTCAGGGTCTCCATTCATCT-3′ | Sigma-Aldrich | This study |
| Retinaldehyde dehydrogenase 2 (raldh2) | Forward 5′-ACAGTGCTTACCTTGCTACCC-3′ Reverse 5′-CTTATCTGCCCATCCAGCGT-3′ | Oligomer Oy | [35] |
| Ribosomal protein S3 (rps3) | Forward 5′-CGTGTCACACCAACAAGA-3′ Reverse 5′-CAGCTTGTAGCGCAGAGA-3′ | Oligomer Oy | [35] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Narumanchi, S.; Kalervo, K.; Perttunen, S.; Wang, H.; Immonen, K.; Kosonen, R.; Laine, M.; Ruskoaho, H.; Tikkanen, I.; Lakkisto, P.; et al. Inhibition of let-7c Regulates Cardiac Regeneration after Cryoinjury in Adult Zebrafish. J. Cardiovasc. Dev. Dis. 2019, 6, 16. https://doi.org/10.3390/jcdd6020016
Narumanchi S, Kalervo K, Perttunen S, Wang H, Immonen K, Kosonen R, Laine M, Ruskoaho H, Tikkanen I, Lakkisto P, et al. Inhibition of let-7c Regulates Cardiac Regeneration after Cryoinjury in Adult Zebrafish. Journal of Cardiovascular Development and Disease. 2019; 6(2):16. https://doi.org/10.3390/jcdd6020016
Chicago/Turabian StyleNarumanchi, Suneeta, Karri Kalervo, Sanni Perttunen, Hong Wang, Katariina Immonen, Riikka Kosonen, Mika Laine, Heikki Ruskoaho, Ilkka Tikkanen, Päivi Lakkisto, and et al. 2019. "Inhibition of let-7c Regulates Cardiac Regeneration after Cryoinjury in Adult Zebrafish" Journal of Cardiovascular Development and Disease 6, no. 2: 16. https://doi.org/10.3390/jcdd6020016
APA StyleNarumanchi, S., Kalervo, K., Perttunen, S., Wang, H., Immonen, K., Kosonen, R., Laine, M., Ruskoaho, H., Tikkanen, I., Lakkisto, P., & Paavola, J. (2019). Inhibition of let-7c Regulates Cardiac Regeneration after Cryoinjury in Adult Zebrafish. Journal of Cardiovascular Development and Disease, 6(2), 16. https://doi.org/10.3390/jcdd6020016





