Multiple Roles of Pitx2 in Cardiac Development and Disease
Abstract
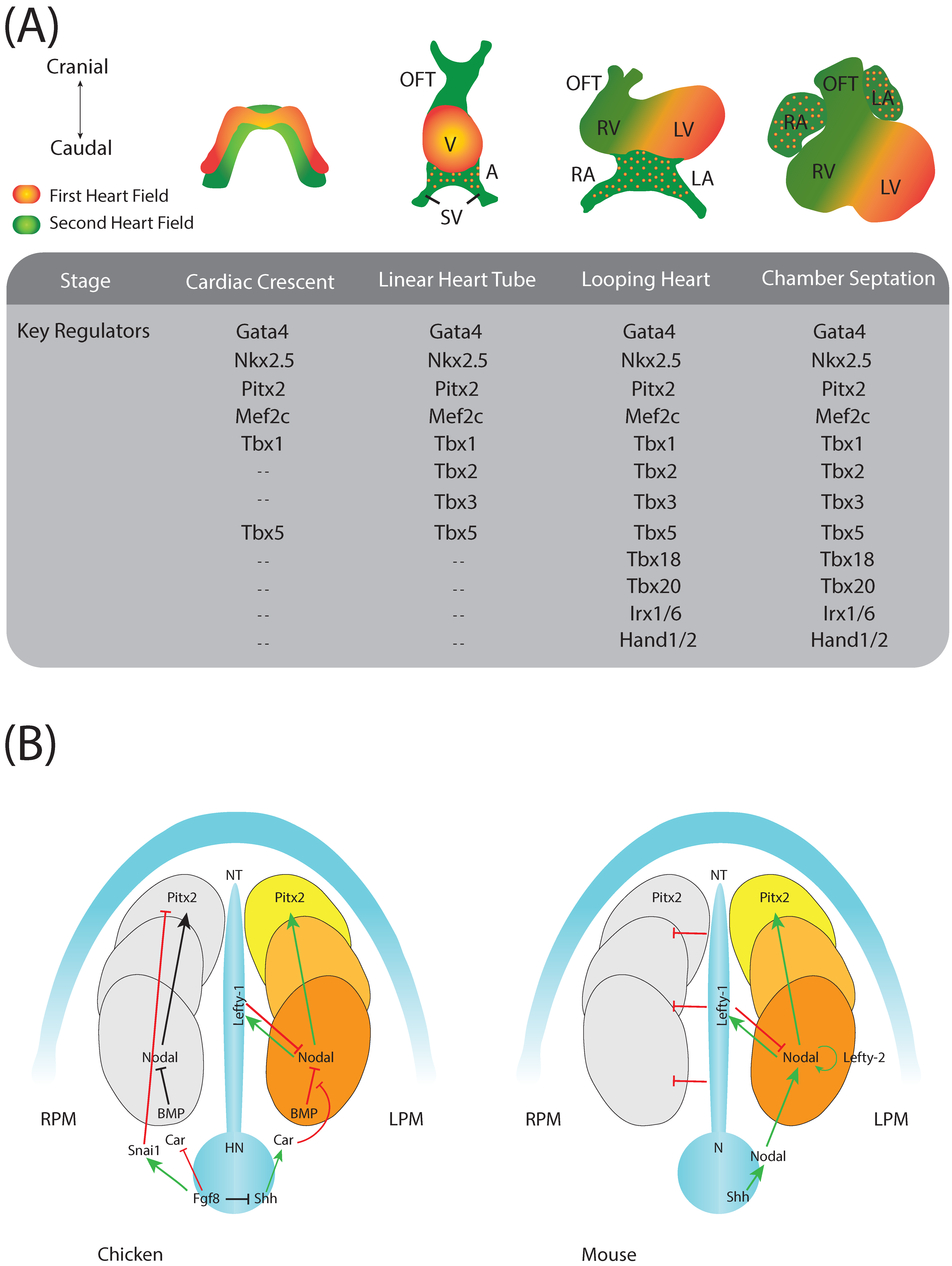
:1. Introduction to Cardiac Development
2. Cardiac Regulatory Mechanisms
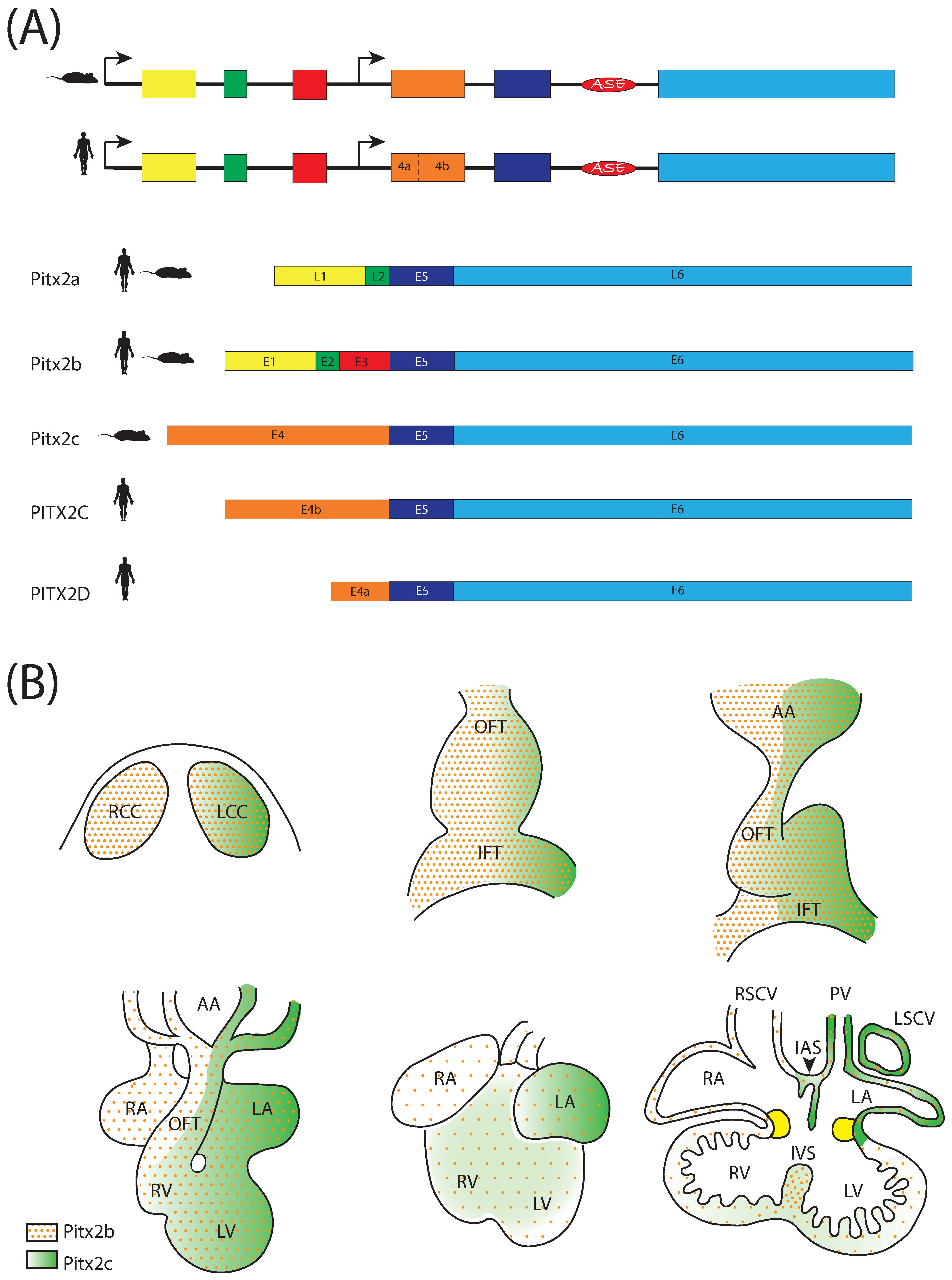
3. The Homeobox Transcription Factor Pitx2
4. Pitx2 and Left-Right Signalling
5. Genetics of Congenital Heart Diseases
6. Left-Right Signalling and Congenital Heart Diseases
7. Pitx2 and Congenital Heart Diseases
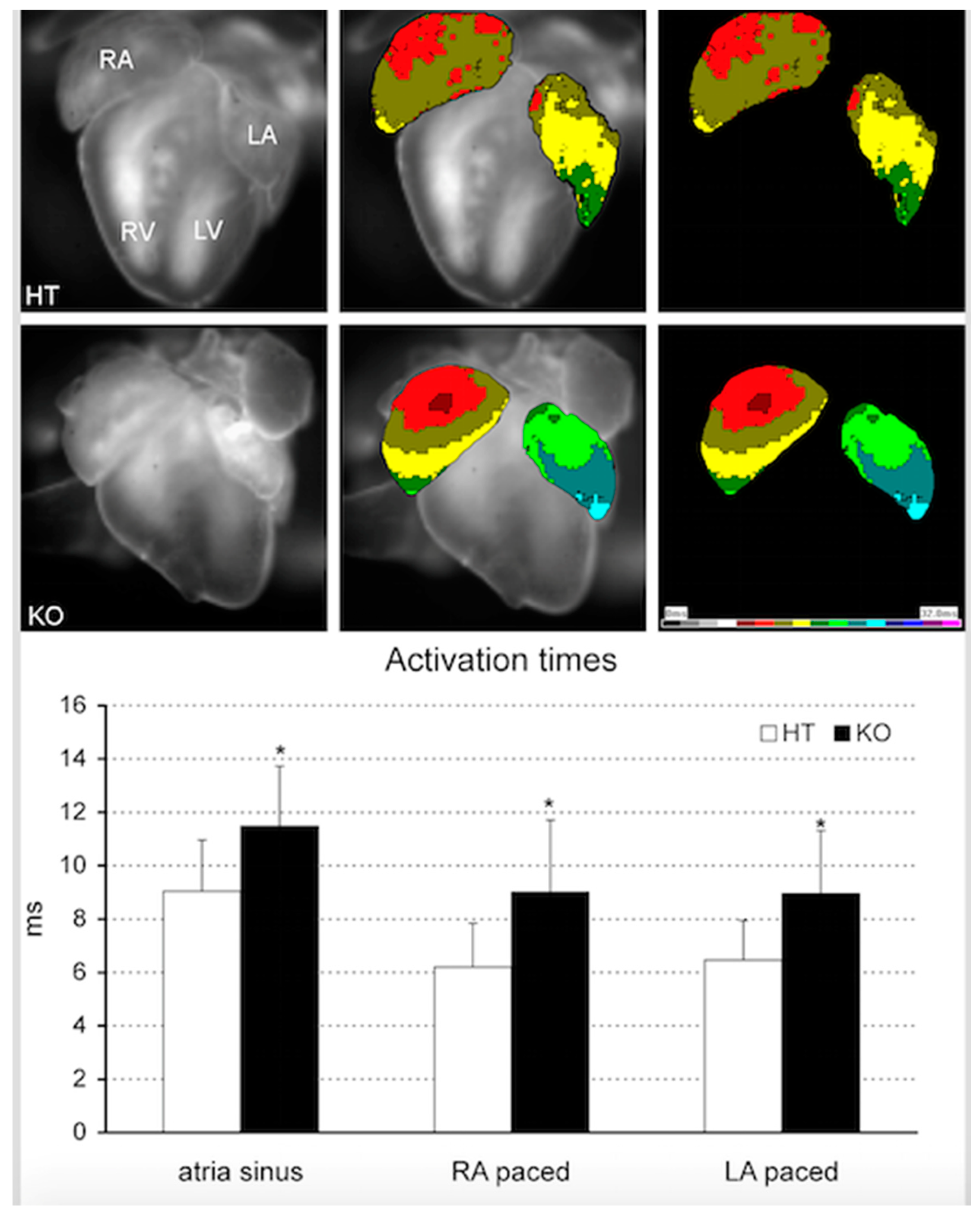
8. Beyond CHD: Pitx2 and Atrial Fibrillation
9. Conclusions and Future Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Moorman, A.F.; Christoffels, V.M. Cardiac chamber formation: Development, genes, and evolution. Physiol. Rev. 2003, 83, 1223–1267. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Martinez, V.; Schoenwolf, G.C. Positional control of mesoderm movement and fate during avian gastrulation and neurulation. Dev. Dyn. 1992, 193, 249–256. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Martinez, V.; Schoenwolf, G.C. Primitive-streak origin of the cardiovascular system in avian embryos. Dev. Biol. 1993, 159, 706–719. [Google Scholar] [CrossRef] [PubMed]
- Franco, D.; Campione, M.; Kelly, R.; Zammit, P.S.; Buckingham, M.; Lamers, W.H.; Moorman, A.F. Multiple transcriptional domains, with distinct left and right components, in the atrial chambers of the developing heart. Circ. Res. 2000, 87, 984–991. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.H.; Webb, S.; Brown, N.A.; Lamers, W.; Moorman, A. Development of the heart: (3) Formation of the ventricular outflow tracts, arterial valves, and intrapericardial arterial trunks. Heart 2003, 89, 1110–1118. [Google Scholar] [CrossRef] [PubMed]
- Anderson, R.H.; Webb, S.; Brown, N.A.; Lamers, W.; Moorman, A. Development of the heart: (2) Septation of the atriums and ventricles. Heart 2003, 89, 949–958. [Google Scholar] [CrossRef] [PubMed]
- Campione, M.; Ros, M.A.; Icardo, J.M.; Piedra, E.; Christoffels, V.M.; Schweickert, A.; Blum, M.; Franco, D.; Moorman, A.F. Pitx2 expression defines a left cardiac lineage of cells: Evidence for atrial and ventricular molecular isomerisms in the iv/iv mice. Dev. Biol. 2001, 231, 252–264. [Google Scholar] [CrossRef] [PubMed]
- Kathiriya, I.S.; Srivastava, D. Left-right asymmetry and cardiac looping: Implications for cardiac development and congenital heart disease. Am. J. Med. Genet. 2000, 97, 271–279. [Google Scholar] [CrossRef]
- Franco, D.; Campione, M. The role of Pitx2 during cardiac development: Linking left-right signaling and congenital heart diseases. Trends Cardiovasc. Med. 2003, 13, 157–163. [Google Scholar] [CrossRef]
- Campione, M.; Acosta, L.; Martínez, S.; Icardo, J.M.; Aránega, A.; Franco, D. Pitx2 and cardiac development: A molecular link between left/right signaling and congenital heart disease. Cold Spring Harb. Symp. Quant. Biol. 2002, 67, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Paige, S.L.; Plonowska, K.; Xu, A.; Wu, S.M. Molecular regulation of cardiomyocyte differentiation. Circ. Res. 2015, 116, 341–353. [Google Scholar] [CrossRef] [PubMed]
- Meyers, E.N.; Martin, G.R. Differences in left-right axis pathways in mouse and chick: Functions of FGF8 and SHH. Science 1999, 285, 403–406. [Google Scholar] [CrossRef] [PubMed]
- Schlueter, J.; Brand, T. Left/right assymetrical development of the proepicardiu. J. Dev. Biol. 2013, 1, 126–140. [Google Scholar] [CrossRef]
- Lopez-Sanchez, C.; Climent, V.; Schoenwolf, G.C.; Alvarez, I.S.; Garcia-Martinez, V. Induction of cardiogenesis by Hensen's node and fibroblast growth factors. Cell Tissue Res. 2002, 309, 237–249. [Google Scholar] [CrossRef] [PubMed]
- Andrée, B.; Duprez, D.; Vorbusch, B.; Arnold, H.H.; Brand, T. BMP-2 induces ectopic expression of cardiac lineage markers and interferes with somite formation in chicken embryos. Mech. Dev. 1998, 70, 119–131. [Google Scholar] [CrossRef]
- Ladd, A.N.; Yatskievych, T.A.; Antin, P.B. Regulation of avian cardiac myogenesis by activin/TGFbeta and bone morphogenetic proteins. Dev. Biol. 1998, 204, 407–419. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, D. HAND proteins: Molecular mediators of cardiac development and congenital heart disease. Trends Cardiovasc. Med. 1999, 9, 11–18. [Google Scholar] [CrossRef]
- Schlesinger, J.; Schueler, M.; Grunert, M.; Fischer, J.J.; Zhang, Q.; Krueger, T.; Lange, M.; Tönjes, M.; Dunkel, I.; Sperling, S.R. The cardiac transcription network modulated by Gata4, Mef2a, Nkx2.5, Srf, histone modifications, and microRNAs. PLoS Genet. 2011, 7, e1001313. [Google Scholar] [CrossRef] [PubMed]
- Ieda, M.; Fu, J.D.; Delgado-Olguin, P.; Vedantham, V.; Hayashi, Y.; Bruneau, B.G.; Srivastava, D. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 2010, 142, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.D.; Stone, N.R.; Liu, L.; Spencer, C.I.; Qian, L.; Hayashi, Y.; Delgado-Olguin, P.; Ding, S.; Bruneau, B.G.; Srivastava, D. Direct reprogramming of human fibroblasts toward a cardiomyocyte-like state. Stem Cell Rep. 2013, 1, 235–247. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, D. Making or breaking the heart: From lineage determination to morphogenesis. Cell 2006, 126, 1037–1048. [Google Scholar] [CrossRef] [PubMed]
- Baldini, A.; Fulcoli, F.G.; Illingworth, E. Tbx1: Transcriptional and Developmental Functions. Curr. Top. Dev. Biol. 2017, 122, 223–243. [Google Scholar] [PubMed]
- Greulich, F.; Rudat, C.; Kispert, A. Mechanisms of T-box gene function in the developing heart. Cardiovasc. Res. 2011, 91, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Mori, A.D.; Bruneau, B.G. TBX5 mutations and congenital heart disease: Holt-Oram syndrome revealed. Curr. Opin. Cardiol. 2004, 19, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Steimle, J.D.; Moskowitz, I.P. TBX5: A Key Regulator of Heart Development. Curr. Top. Dev. Biol. 2017, 122, 195–221. [Google Scholar] [PubMed]
- Christoffels, V.M.; Keijser, A.G.; Houweling, A.C.; Clout, D.E.; Moorman, A.F. Patterning the embryonic heart: Identification of five mouse Iroquois homeobox genes in the developing heart. Dev. Biol. 2000, 224, 263–274. [Google Scholar] [CrossRef] [PubMed]
- Houweling, A.C.; Dildrop, R.; Peters, T.; Mummenhoff, J.; Moorman, A.F.; Rüther, U.; Christoffels, V.M. Gene and cluster-specific expression of the Iroquois family members during mouse development. Mech. Dev. 2001, 107, 169–174. [Google Scholar] [CrossRef]
- Bruneau, B.G.; Bao, Z.Z.; Fatkin, D.; Xavier-Neto, J.; Georgakopoulos, D.; Maguire, C.T.; Berul, C.I.; Kass, D.A.; Kuroski-de Bold, M.L.; de Bold, A.J.; et al. Cardiomyopathy in Irx4-deficient mice is preceded by abnormal ventricular gene expression. Mol. Cell. Biol. 2001, 21, 1730–1736. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.H.; Rosen, A.; Bruneau, B.G.; Hui, C.C.; Backx, P.H. Iroquois homeodomain transcription factors in heart development and function. Circ. Res. 2012, 110, 1513–1524. [Google Scholar] [CrossRef] [PubMed]
- Ryan, A.K.; Blumberg, B.; Rodríguez-Esteban, C.; Yonei-Tamura, S.; Tamura, K.; Tsukui, T.; de la Peña, J.; Sabbagh, W.; Greenwald, J.; Choe, S.; et al. Pitx2 determines left-right asymmetry of internal organs in vertebrates. Nature 1998, 394, 545–551. [Google Scholar] [PubMed]
- Logan, M.; Pagán-Westphal, S.M.; Smith, D.M.; Paganessi, L.; Tabin, C.T. The transcription factor Pitx2 mediates situs-specific morphogenesis in response to left-right asymmetric signals. Cell 1998, 94, 307–317. [Google Scholar] [CrossRef]
- Piedra, M.E.; Icardo, J.M.; Albajar, M.; Rodriguez-Rey, J.C.; Ros, M.A. Pitx2 participates in the late phase of the pathway controlling left-right asymmetry. Cell 1998, 94, 319–324. [Google Scholar] [CrossRef]
- Campione, M.; Steinbeisser, H.; Schweickert, A.; Deissler, K.; van Bebber, F.; Lowe, L.A.; Nowotschin, S.; Viebahn, C.; Haffter, P.; Kuehn, M.R.; et al. The homeobox gene Pitx2: Mediator of asymmetric left-right signaling in vertebrate heart and gut looping. Development 1999, 126, 1225–1234. [Google Scholar] [PubMed]
- Scott, M.P.; Tamkun, J.W.; Hartzell, I.G.W. The structure and function of the homeodomain. Biochem. Biophys. Acta 1989, 989, 25–48. [Google Scholar] [CrossRef]
- Gage, P.J.; Suh, H.; Camper, S.A. The bicoid-related Pitx gene family in development. Mamm. Genome 1999, 10, 197–200. [Google Scholar] [CrossRef] [PubMed]
- Gehring, W.J.; Qian, Y.Q.; Billeter, M.; Furukubo-Tokunaga, K.; Schier, A.F.; Resendez-Perez, D.; Affolter, M.; Otting, G.; Wüthrich, K. Homeodo- main-DNA recognition. Cell 1994, 78, 211–223. [Google Scholar] [CrossRef]
- Gage, P.J.; Suh, H.; Camper, S.A. Dosage requirement of Pitx2 for development of multiple organs. Development 1999, 126, 4643–4651. [Google Scholar] [PubMed]
- Lanctot, C.; Lamolet, B.; Drouin, J. The bicoid-related homeoprotein Ptx1 defines the most anterior domain of the embryo and differentiates posterior from anterior lateral mesoderm. Development 1997, 124, 2807–2817. [Google Scholar] [PubMed]
- Logan, M.; Tabin, C.J. Role of Pitx1 upstream of Tbx4 in specification of hindlimb identity. Science 1999, 283, 1736–1739. [Google Scholar] [CrossRef] [PubMed]
- Szeto, D.P.; Rodriguez-Esteban, C.; Ryan, A.K.; O’Connell, S.M.; Liu, F.; Kioussi, C.; Gleiberman, A.S.; Izpisua-Belmonte, J.C.; Rosenfeld, M.G. Role of the Bicoid-related homeodomain factor Pitx1 in specifying hindlimb morphogenesis and pituitary development. Genes Dev. 1999, 13, 484–494. [Google Scholar] [CrossRef] [PubMed]
- Kioussi, C.; Briata, P.; Baek, S.H.; Rose, D.W.; Hamblet, N.S.; Herman, T.; Ohgi, K.A.; Lin, C.; Gleiberman, A.; Wang, J.; et al. Identification of a Wnt/Dvl/beta-Catenin → Pitx2 pathway mediating cell-type-specific proliferation during development. Cell 2002, 111, 673–985. [Google Scholar] [CrossRef]
- Kioussi, C.; Briata, P.; Baek, S.H.; Wynshaw-Boris, A.; Rose, D.W.; Rosenfeld, M.G. Pitx genes during cardiovascular development. Cold Spring Harb. Symp. Quant. Biol. 2002, 67, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, K.; Miura, H.; Miyagawa-Tomita, S.; Yanazawa, M.; Katoh-Fukui, Y.; Suzuki, R.; Ohuchi, H.; Suehiro, A.; Motegi, Y.; Nakahara, Y.; et al. Mouse Pitx2 deficiency leads to anomalies of the ventral body wall, heart, extra- and periocular mesoderm and right pulmonary isomerism. Development 1999, 126, 5749–5758. [Google Scholar] [PubMed]
- Kitamura, K.; Miura, H.; Yanazawa, M.; Miyashita, T.; Kato, K. Expression patterns of Brx1 (Rieg gene), Sonic hedgehog, Nkx2.2, Dlx1 and Arx during zona limitans intrathalamica and embryonic ventral lateral geniculate nuclear formation. Mech. Dev. 1997, 67, 83–96. [Google Scholar] [CrossRef]
- Lu, M.F.; Pressman, C.; Dyer, R.; Johnson, R.L.; Martin, J.F. Function of Rieger syndrome gene in left-right asymmetry and craniofacial development. Nature 1999, 401, 276–278. [Google Scholar] [PubMed]
- Muccielli, M.L.; Martinez, S.; Pattyn, A.; Goridis, C.; Brunet, J.F. Otlx2, an Otx-related homeobox gene expressed in the pituitary gland and in a restricted pattern in the forebrain. Mol. Cell. Neurosci. 1996, 8, 258–271. [Google Scholar] [CrossRef] [PubMed]
- Qiu, H.Y.; Guo, C.; Cheng, X.W.; Huang, Y.; Xiong, Z.Q.; Ding, Y.Q. Pitx3-CreER mice showing restricted Cre expression in developing ocular lens and skeletal muscles. Genesis 2008, 46, 324–328. [Google Scholar] [CrossRef] [PubMed]
- Smidt, M.P.; Smits, S.M.; Bouwmeester, H.; Hamers, F.P.; van der Linden, A.J.; Hellemons, A.J.; Graw, J.; Burbach, J.P. Early developmental failure of substantia nigra dopamine neurons in mice lacking the homeodomain gene Pitx3. Development 2004, 131, 1145–1155. [Google Scholar] [CrossRef] [PubMed]
- Smidt, M.P.; van Schaick, H.S.; Lanctot, C.; Tremblay, J.J.; Cox, J.J.; van der Kleij, A.A.; Wolterink, G.; Drouin, J.; Burbach, J.P. A homeodomain gene Ptx3 has highly restricted brain expression in mesencephalic dopaminergic neurons. Proc. Natl. Acad. Sci. USA 1997, 94, 13305–13310. [Google Scholar] [CrossRef] [PubMed]
- Gage, P.J.; Camper, S.A. Pituitary homeobox 2, a novel member of the bicoid-related family of homeobox genes, is a potential regulator of anterior structure formation. Hum. Mol. Genet. 1997, 6, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Arakawa, H.; Nakamura, T.; Zhadanov, A.B.; Fidanza, V.; Yano, T.; Bullrich, F.; Shimizu, M.; Blechman, J.; Mazo, A.; Canaani, E.; et al. Identification and characterization of the ARP1 gene, a target for the human acute leukemia ALL1 gene. Proc. Natl. Acad. Sci. USA 1998, 95, 4573–4578. [Google Scholar] [CrossRef] [PubMed]
- Cox, C.J.; Espinoza, H.M.; McWilliams, B.; Chappell, K.; Morton, L.; Hjalt, T.A.; Semina, E.V.; Amendt, B.A. Differential regulation of gene expression by PITX2 isoforms. J. Biol. Chem. 2002, 277, 25001–25010. [Google Scholar] [CrossRef] [PubMed]
- Semina, E.V.; Reiter, R.; Leysens, N.J.; Alward, W.L.; Small, K.W.; Datson, N.A.; Siegel-Bartelt, J.; Bierke-Nelson, D.; Bitoun, P.; Zabel, B.U.; et al. Cloning and characterization of a novel bicoid-related homeobox transcription factor gene, RIEG, involved in Rieger syndrome. Nat. Genet. 1996, 14, 392–399. [Google Scholar] [CrossRef] [PubMed]
- Shiratori, H.; Sakuma, R.; Watanabe, M.; Hashiguchi, H.; Mochida, K.; Sakai, Y.; Nishino, J.; Saijoh, Y.; Whitman, M.; Hamada, H. Two-step regulation of left-right asymmetric expression of Pitx2: Initiation by nodal signaling and maintenance by Nkx2. Mol. Cell 2001, 7, 137–149. [Google Scholar] [CrossRef]
- Yoshioka, H.; Meno, C.; Koshiba, K.; Sugihara, M.; Itoh, H.; Ishimaru, Y.; Inoue, T.; Ohuchi, H.; Semina, E.V. Pitx2, a bicoid-type homeobox gene, is involved in a lefty-signaling pathway in determination of left-right asymmetry. Cell 1998, 94, 299–305. [Google Scholar] [CrossRef]
- Semina, E.V.; Reiter, R.S.; Murray, J.C. Isolation of a new homeobox gene belonging to the Pitx/Rieg family: Expression during lens development and mapping to the aphakia region on mouse chromosome 19. Hum. Mol. Genet. 1997, 6, 2109–2116. [Google Scholar] [CrossRef] [PubMed]
- Amand, T.R.S.; Ra, J.; Zhang, Y.; Hu, Y.; Baber, S.; Qiu, M.; Chen, Y. Cloning and expression pattern of chicken Pitx2: A new component in the SHH signalling pathway controlling embryonic heart looping. Biochem. Biophys. Res. Commun. 1998, 247, 100–105. [Google Scholar] [CrossRef]
- Lin, C.R.; Kioussi, C.; O’Connell, S.; Briata, P.; Szeto, D.; Liu, F.; Izpisúa-Belmonte, J.C.; Resenfield, M.G. Pitx2 regulates lung asymmetry, cardiac positioning and pituitary and tooth morphogenesis. Nature 1999, 401, 279–282. [Google Scholar] [PubMed]
- Cunningham, E.T., Jr.; Eliott, D.; Miller, N.R.; Maumenee, I.H.; Green, W.R. Familial Axenfeld-Rieger anormaly, atrial septal defect and sensorineural hearing loss: A possible new genetic syndrome. Arch. Ophthalmol. 1998, 1666, 78–82. [Google Scholar]
- Mammi, I.; De Giorgio, P.; Clementi, M.; Tenconi, R. Cardiovascular anomaly in Rieger syndrome: Heterogeneity or contiguity? Acta Ophthalmol. Scand. 1998, 76, 509–512. [Google Scholar] [CrossRef] [PubMed]
- Tsai, J.C.; Grajewski, A.L. Cardiac valvular disease and Axenfeld-Rieger syndrome. Am. J. Ophthalmol. 1994, 118, 255–256. [Google Scholar] [CrossRef]
- Spéder, P.; Petzoldt, A.; Suzanne, M.; Noselli, S. Strategies to establish left/right asymmetry in vertebrates and invertebrates. Curr. Opin. Genet. Dev. 2007, 17, 351–358. [Google Scholar] [CrossRef] [PubMed]
- Bakkers, J.; Verhoeven, M.C.; Abdelilah-Seyfried, S. Shaping the zebrafish heart: From left-right axis specification to epithelial tissue morphogenesis. Dev. Biol. 2009, 330, 213–220. [Google Scholar] [CrossRef] [PubMed]
- Namigai, E.K.; Kenny, N.J.; Shimeld, S.M. Right across the tree of life: The evolution of left-right asymmetry in the Bilateria. Genesis 2014, 52, 458–470. [Google Scholar] [CrossRef] [PubMed]
- Meno, C.; Ito, Y.; Saijoh, Y.; Matsuda, Y.; Tashiro, K.; Kuhara, S.; Hamada, H. Two closely-related left-right asymmetrically expressed genes, lefty-1 and lefty-2: Their distinct expression domains, chromosomal linkage and direct neutralizing activity in Xenopus embryos. Genes Cells 1997, 2, 513–524. [Google Scholar] [CrossRef] [PubMed]
- Meno, C.; Shimono, A.; Saijoh, Y.; Yashiro, K.; Mochida, K.; Ohishi, S.; Noji, S.; Kondoh, H.; Hamada, H. lefty-1 is required for left-right determination as a regulator of lefty-2 and nodal. Cell 1998, 94, 287–297. [Google Scholar] [CrossRef]
- Oki, S.; Kitajima, K.; Marques, S.; Belo, J.A.; Yokoyama, T.; Hamada, H.; Meno, C. Reversal of left-right asymmetry induced by aberrant Nodal signaling in the node of mouse embryos. Development 2009, 136, 3917–3925. [Google Scholar] [CrossRef] [PubMed]
- Adachi, H.; Saijoh, Y.; Mochida, K.; Ohishi, S.; Hashiguchi, H.; Hirao, A.; Hamada, H. Determination of left/right asymmetric expression of nodal by a left side-specific enhancer with sequence similarity to a lefty-2 enhancer. Genes Dev. 1999, 13, 1589–1600. [Google Scholar] [CrossRef] [PubMed]
- Soukup, V.; Yong, L.W.; Lu, T.M.; Huang, S.W.; Kozmik, Z.; Yu, J.K. The Nodal signaling pathway controls left-right asymmetric development in amphioxus. EvoDevo 2015, 6, 5. [Google Scholar] [CrossRef] [PubMed]
- Fujinaga, M.; Lowe, L.A.; Kuehn, M.R. Alpha (1)-Adrenergic stimulation perturbs the left-right asymmetric expression pattern of nodal during rat embryogenesis. Teratology 2000, 62, 317–324. [Google Scholar] [CrossRef]
- Long, S.; Ahmad, N.; Rebagliati, M. The zebrafish nodal-related gene southpaw is required for visceral and diencephalic left-right asymmetry. Development 2003, 130, 2303–2316. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Lualdi, M.; Lewandoski, M.; Kuehn, M.R. Broad mesodermal and endodermal deletion of Nodal at postgastrulation stages results solely in left/right axial defects. Dev. Dyn. 2008, 237, 3591–3601. [Google Scholar] [CrossRef] [PubMed]
- Heymer, J.; Kuehn, M.; Rüther, U. The expression pattern of nodal and lefty in the mouse mutant Ft suggests a function in the establishment of handedness. Mech. Dev. 1997, 66, 5–11. [Google Scholar] [CrossRef]
- Kawasumi, A.; Nakamura, T.; Iwai, N.; Yashiro, K.; Saijoh, Y.; Belo, J.A.; Shiratori, H.; Hamada, H. Left-right asymmetry in the level of active Nodal protein produced in the node is translated into left-right asymmetry in the lateral plate of mouse embryos. Dev. Biol. 2011, 353, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Tadjuidje, E.; Kofron, M.; Mir, A.; Wylie, C.; Heasman, J.; Cha, S.W. Nodal signalling in Xenopus: The role of Xnr5 in left/right asymmetry and heart development. Open Biol. 2016, 6, 150187. [Google Scholar] [CrossRef] [PubMed]
- Vincent, S.D.; Norris, D.P.; Le Good, J.A.; Constam, D.B.; Robertson, E.J. Asymmetric Nodal expression in the mouse is governed by the combinatorial activities of two distinct regulatory elements. Mech. Dev. 2004, 121, 1403–1415. [Google Scholar] [CrossRef] [PubMed]
- Gaio, U.; Schweickert, A.; Fischer, A.; Garratt, A.N.; Müller, T.; Ozcelik, C.; Lankes, W.; Strehle, M.; Britsch, S.; Blum, M.; et al. A role of the cryptic gene in the correct establishment of the left-right axis. Curr. Biol. 1999, 9, 1339–1342. [Google Scholar] [CrossRef]
- Hashimoto, H.; Rebagliati, M.; Ahmad, N.; Muraoka, O.; Kurokawa, T.; Hibi, M.; Suzuki, T. The Cerberus/Dan-family protein Charon is a negative regulator of Nodal signaling during left-right patterning in zebrafish. Development 2004, 131, 1741–1753. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez Esteban, C.; Capdevila, J.; Economides, A.N.; Pascual, J.; Ortiz, A.; Izpisúa Belmonte, J.C. The novel Cer-like protein Caronte mediates the establishment of embryonic left-right asymmetry. Nature 1999, 401, 243–251. [Google Scholar]
- Tsiairis, C.D.; McMahon, A.P. An Hh-dependent pathway in lateral plate mesoderm enables the generation of left/right asymmetry. Curr. Biol. 2009, 19, 1912–1917. [Google Scholar] [CrossRef] [PubMed]
- Inácio, J.M.; Marques, S.; Nakamura, T.; Shinohara, K.; Meno, C.; Hamada, H.; Belo, J.A. The dynamic right-to-left translocation of Cerl2 is involved in the regulation and termination of Nodal activity in the mouse node. PLoS ONE 2013, 8, e60406. [Google Scholar] [CrossRef] [PubMed]
- Larkins, C.E.; Long, A.B.; Caspary, T. Defective Nodal and Cerl2 expression in the Arl13b(hnn) mutant node underlie its heterotaxia. Dev. Biol. 2012, 367, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Marques, S.; Borges, A.C.; Silva, A.C.; Freitas, S.; Cordenonsi, M.; Belo, J.A. The activity of the Nodal antagonist Cerl-2 in the mouse node is required for correct L/R body axis. Genes Dev. 2004, 18, 2342–2347. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, T.; Saito, D.; Kawasumi, A.; Shinohara, K.; Asai, Y.; Takaoka, K.; Dong, F.; Takamatsu, A.; Belo, J.A.; Mochizuki, A.; et al. Fluid flow and interlinked feedback loops establish left-right asymmetric decay of Cerl2 mRNA. Nat. Commun. 2012, 3, 1322. [Google Scholar] [CrossRef] [PubMed]
- Patel, K.; Isaac, A.; Cooke, J. Nodal signalling and the roles of the transcription factors SnR and Pitx2 in vertebrate left-right asymmetry. Curr. Biol. 1999, 9, 609–612. [Google Scholar] [CrossRef]
- Kitajima, K.; Oki, S.; Ohkawa, Y.; Sumi, T.; Meno, C. Wnt signaling regulates left-right axis formation in the node of mouse embryos. Dev. Biol. 2013, 380, 222–232. [Google Scholar] [CrossRef] [PubMed]
- Supp, D.M.; Witte, D.P.; Potter, S.S.; Brueckner, M. Mutation of an axonemal dynein affects left-right asymmetry in inversus viscerum mice. Nature 1997, 389, 963–966. [Google Scholar] [CrossRef] [PubMed]
- Buceta, J.; Ibañes, M.; Rasskin-Gutman, D.; Okada, Y.; Hirokawa, N.; Izpisúa-Belmonte, J.C. Nodal cilia dynamics and the specification of the left/right axis in early vertebrate embryo development. Biophys. J. 2005, 89, 2199–2209. [Google Scholar] [CrossRef] [PubMed]
- Hjeij, R.; Lindstrand, A.; Francis, R.; Zariwala, M.A.; Liu, X.; Li, Y.; Damerla, R.; Dougherty, G.W.; Abouhamed, M.; Olbrich, H.; et al. ARMC4 mutations cause primary ciliary dyskinesia with randomization of left/right body asymmetry. Am. J. Hum. Genet. 2013, 93, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Burnicka-Turek, O.; Steimle, J.D.; Huang, W.; Felker, L.; Kamp, A.; Kweon, J.; Peterson, M.; Reeves, R.H.; Maslen, C.L.; Gruber, P.J.; et al. Cilia gene mutations cause atrioventricular septal defects by multiple mechanisms. Hum. Mol. Genet. 2016, 25, 3011–3028. [Google Scholar] [CrossRef] [PubMed]
- Hadjantonakis, A.K.; Pisano, E.; Papaioannou, V.E. Tbx6 regulates left/right patterning in mouse embryos through effects on nodal cilia and perinodal signaling. PLoS ONE 2008, 3, e2511. [Google Scholar] [CrossRef] [PubMed]
- Gros, J.; Feistel, K.; Viebahn, C.; Blum, M.; Tabin, C.J. Cell movements at Hensen’s node establish left/right asymmetric gene expression in the chick. Science 2009, 324, 941–944. [Google Scholar] [CrossRef] [PubMed]
- Bisgrove, B.W.; Essner, J.J.; Yost, H.J. Multiple pathways in the midline regulate concordant brain, heart and gut left-right asymmetry. Development 2000, 127, 3567–3579. [Google Scholar] [PubMed]
- Bisgrove, B.W.; Yost, H.J. Classification of left-right patterning defects in zebrafish, mice, and humans. Am. J. Med. Genet. 2001, 101, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Branford, W.W.; Essner, J.J.; Yost, H.J. Regulation of gut and heart left-right asymmetry by context-dependent interactions between xenopus lefty and BMP4 signaling. Dev. Biol. 2000, 223, 291–306. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Liu, W.; Lu, M.F.; Brown, N.A.; Martin, J.F. Regulation of left-right asymmetry by thresholds of Pitx2c activity. Development 2001, 128, 2039–2048. [Google Scholar] [PubMed]
- Dagle, J.M.; Sabel, J.L.; Littig, J.L.; Sutherland, L.B.; Kolker, S.J.; Weeks, D.L. Pitx2c attenuation results in cardiac defects and abnormalities of intestinal orientation in developing Xenopus laevis. Dev. Biol. 2003, 262, 268–281. [Google Scholar] [CrossRef]
- Davis, N.M.; Kurpios, N.A.; Sun, X.; Gros, J.; Martin, J.F.; Tabin, C.J. The chirality of gut rotation derives from left-right asymmetric changes in the architecture of the dorsal mesentery. Dev. Cell 2008, 15, 134–145. [Google Scholar] [CrossRef] [PubMed]
- Essner, J.J.; Branford, W.W.; Zhang, J.; Yost, H.J. Mesendoderm and left-right brain, heart and gut development are differentially regulated by pitx2 isoforms. Development 2000, 127, 1081–1093. [Google Scholar] [PubMed]
- Guioli, S.; Lovell-Badge, R. PITX2 controls asymmetric gonadal development in both sexes of the chick and can rescue the degeneration of the right ovary. Development 2007, 134, 4199–4208. [Google Scholar] [CrossRef] [PubMed]
- Nowotschin, S.; Liao, J.; Gage, P.J.; Epstein, J.A.; Campione, M.; Morrow, B.E. Tbx1 affects asymmetric cardiac morphogenesis by regulating Pitx2 in the secondary heart field. Development 2006, 133, 1565–1573. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Selever, J.; Lu, M.F.; Martin, J.F. Genetic dissection of Pitx2 in craniofacial development uncovers new functions in branchial arch morphogenesis, late aspects of tooth morphogenesis and cell migration. Development 2003, 130, 6375–6385. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Amand, T.R.S.; Wang, S.; Li, G.; Zhang, Y.; Hu, Y.; Nguyen, L.; Qiu, M.; Chen, Y. Differential expression and functional analysis of Pitx2 isoforms in regulation of heart looping in the chick. Development 2001, 128, 1005–1101. [Google Scholar] [PubMed]
- Chinchilla, A.; Daimi, H.; Lozano-Velasco, E.; Domínguez, J.N.; Caballero, R.; Delpón, E.; Tamargo, J.; Cinca, J.; Hove-Madsen, L.; Aránega, A.E.; et al. Pitx2 insufficiency leads to electrical and structural remodelling linked to arrhythmogenesis. Circ. Cardiovasc. Genet. 2011, 4, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Klysik, E.; Sood, S.; Johnson, R.L.; Wehrens, X.H.; Martin, J.F. Pitx2 prevents susceptibility to atrial arrhythmias by inhibiting left-sided pacemarker specification. Proc. Natl. Acad. Sci. USA 2010, 107, 9753–9758. [Google Scholar] [CrossRef] [PubMed]
- Kirchhof, P.; Kahr, P.C.; Kaese, S.; Piccini, I.; Vokshi, I.; Scheld, H.H.; Rotering, H.; Fortmueller, L.; Laakmann, S.; Verheule, S.; et al. PITX2c is expressed in the adult left atrium, and reducing Pitx2c expression promotes atrial fibrillation inducibility and complex changes in gene expression. Circ. Cardiovasc. Genet. 2011, 4, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Boorman, C.J.; Shimeld, S.M. The evolution of left-right asymmetry in chordates. Bioessays 2002, 24, 1004–1011. [Google Scholar] [CrossRef] [PubMed]
- Schweickert, A.; Campione, M.; Steinbeisser, H.; Blum, M. Pitx2 isoforms: Involvement of Pitx2c but not Pitx2a or Pitx2b in vertebrate left-right asymmetry. Mech. Dev. 2000, 90, 41–51. [Google Scholar] [CrossRef]
- Linask, K.K.; Yu, X.; Chen, Y.; Han, M.D. Directionality of heart looping: Effects of Pitx2c misexpression on flectin asymmetry and midline structures. Dev. Biol. 2002, 246, 407–417. [Google Scholar] [CrossRef] [PubMed]
- Blue, G.M.; Kirk, E.P.; Giannoulatou, E.; Sholler, G.F.; Dunwoodie, S.L.; Harvey, R.P.; Winlaw, D.S. Advances in the Genetics of Congenital Heart Disease: A Clinician’s Guide. J. Am. Coll. Cardiol. 2017, 69, 859–870. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, S.; Brueckner, M. Genetics and Genomics of Congenital Heart Disease. Circ. Res. 2017, 120, 923–940. [Google Scholar] [CrossRef] [PubMed]
- Moorman, A.; Webb, S.; Brown, N.A.; Lamers, W.; Anderson, R.H. Development of the heart: (1) Formation of the cardiac chambers and arterial trunks. Heart 2003, 89, 806–814. [Google Scholar] [CrossRef] [PubMed]
- Furtado, M.B.; Biben, C.; Shiratori, H.; Hamada, H.; Harvey, R.P. Characterization of Pitx2c expression in the mouse heart using a reporter transgene. Dev. Dyn. 2011, 240, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Mommersteeg, M.T.; Brown, N.A.; Prall, O.W.; de Gier-de Vries, C.; Harvey, R.P.; Moorman, A.F.; Christoffels, V.M. Pitx2c and Nkx2–5 are required for the formation and identity of the pulmonary myocardium. Circ. Res. 2007, 101, 902–909. [Google Scholar] [CrossRef] [PubMed]
- Mommersteeg, M.T.; Hoogaars, W.M.; Prall, O.W.; de Gier-de Vries, C.; Wiese, C.; Clout, D.E.; Papaioannou, V.E.; Brown, N.A.; Harvey, R.P.; Moorman, A.F.; et al. Molecular pathway for the localized formation of the sinoatrial node. Circ. Res. 2007, 100, 354–362. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Liu, W.; Palie, J.; Lu, M.F.; Brown, N.A.; Martin, J.F. Pitx2c patterns anterior myocardium and aortic arch vessels and is required for local cell movement into atrioventricular cushions. Development 2002, 129, 5081–5091. [Google Scholar] [PubMed]
- Ai, D.; Liu, W.; Ma, L.; Dong, F.; Lu, M.F.; Wang, D.; Verzi, M.P.; Cai, C.; Gage, P.J.; Evans, S.; et al. Pitx2 regulates cardiac left-right asymmetry by patterning second cardiac lineage-derived myocardium. Dev. Biol. 2006, 296, 437–449. [Google Scholar] [CrossRef] [PubMed]
- Tessari, A.; Pietrobon, M.; Notte, M.; Cifelli, G.; Gage, P.J.; Schneider, M.D.; Lembi, G.; Campione, M. Myocardial Pitx2 Differentially Regulates the Left Atrial Identity and Ventricular Asymmetric Remodeling Programs. Circ. Res. 2008, 102, 813–822. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Zhang, M.; Li, L.; Bai, Y.; Zhou, Y.; Moon, A.M.; Kaminski, H.J.; Martin, J.F. Pitx2, an atrial fibrillation predisposition gene, directly regulates ion transport and intercalated disc genes. Circ. Cardiovasc. Genet. 2014, 7, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Ammirabile, G.; Tessari, A.; Pignataro, V.; Szumska, D.; Sardo, F.S.; Benes, J., Jr.; Balistreri, M.; Bhattacharya, S.; Sedmera, D.; Campione, M. Pitx2 confers left morphological, molecular, and functional identity to the sinus venosus myocardium. Cardiovasc. Res. 2012, 93, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.M.; Probst, F.J.; Fox, S.E.; Schimmenti, L.A.; Semina, E.V.; Hefner, M.A.; Belmont, J.W.; Camper, S.A. Exclusion of PITX2 mutations as a major cause of CHARGE association. Am. J. Med. Genet. 2002, 111, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Muncke, N.; Niesler, B.; Roeth, R.; Schön, K.; Rüdiger, H.J.; Goldmuntz, E.; Goodship, J.; Rappold, G. Mutational analysis of the PITX2 coding region revealed no common cause for transposition of the great arteries (dTGA). BMC Med. Genet. 2005, 6, 20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zaidi, S.; Choi, M.; Wakimoto, H.; Ma, L.; Jiang, J.; Overton, J.D.; Romano-Adesman, A.; Bjornson, R.D.; Breitbart, R.E.; Brown, K.K.; et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 2013, 498, 220–223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, F.; Zhao, L.; Wang, J.; Zhang, W.; Li, X.; Qiu, X.B.; Li, R.G.; Xu, Y.J.; Xu, L.; Qu, X.K.; et al. PITX2c loss-of-function mutations responsible for congenital atrial septal defects. Int. J. Med. Sci. 2013, 10, 1422–1429. [Google Scholar] [CrossRef] [PubMed]
- Wei, D.; Gong, X.H.; Qiu, G.; Wang, J.; Yang, Y.Q. Novel PITX2c loss-of-function mutations associated with complex congenital heart disease. Int. J. Mol. Med. 2014, 33, 1201–1208. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xin, Y.F.; Xu, W.J.; Liu, Z.M.; Qiu, X.B.; Qu, X.K.; Xu, L.; Li, X.; Yang, Y.Q. Prevalence and spectrum of PITX2c mutations associated with congenital heart disease. DNA Cell Biol. 2013, 32, 708–716. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.M.; Wang, J.; Qiu, X.B.; Yuan, F.; Xu, Y.J.; Li, R.G.; Qu, X.K.; Huang, R.T.; Xue, S.; Yang, Y.Q. PITX2 loss-of-function mutation contributes to tetralogy of Fallot. Gene 2016, 577, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Pan, H.; Guan, L.; Su, D.; Ma, X. CITED2 mutation links congenital heart defects to dysregulation of the cardiac gene VEGF and PITX2C expression. Biochem. Biophys. Res. Commun. 2012, 423, 895–899. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.M.; Peng, L.Y.; Li, L.; Liu, X.Y.; Wang, J.; Zhang, X.L.; Yuan, F.; Li, R.G.; Qiu, X.B.; Yang, Y.Q. PITX2 Loss-of-Function Mutation Contributes to Congenital Endocardial Cushion Defect and Axenfeld-Rieger Syndrome. PLoS ONE 2015, 10, e0124409. [Google Scholar] [CrossRef] [PubMed]
- Hirayama-Yamada, K.; Kamisago, M.; Akimoto, K.; Aotsuka, H.; Nakamura, Y.; Tomita, H.; Furutani, M.; Imamura, S.; Takao, A.; Nakazawa, M.; et al. Phenotypes with GATA4 or NKX2.5 mutations in familial atrial septal defect. Am. J. Med. Genet. Part A 2005, 135, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Kirk, E.P.; Hyun, C.; Thomson, P.C.; Lai, D.; Castro, M.L.; Biben, C.; Buckley, M.F.; Martin, I.C.; Moran, C.; Harvey, R.P. Quantitative trait loci modifying cardiac atrial septal morphology and risk of patent foramen ovale in the mouse. Circ. Res. 2006, 98, 651–658. [Google Scholar] [CrossRef] [PubMed]
- Posch, M.G.; Perrot, A.; Berger, F.; Ozcelik, C. Molecular genetics of congenital atrial septal defects. Clin. Res. Cardiol. 2010, 99, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Gebbia, M.; Ferrero, G.B.; Pilia, G.; Bassi, M.T.; Aylsworth, A.; Penman-Splitt, M.; Bird, L.M.; Bamforth, J.S.; Burn, J.; Schlessinger, D.; et al. X-linked situs abnormalities result from mutations in ZIC3. Nat. Genet. 1997, 17, 305–308. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, E.J.; Levy, D.; Vaziri, S.M.; D’Agostino, R.B.; Belanger, A.J.; Wolf, P.A. Independent risk factors for atrial fibrillation in a population-based cohort: The Framingham Heart Study. JAMA 1994, 271, 840–844. [Google Scholar] [CrossRef] [PubMed]
- Go, A.S.; Hylek, E.M.; Philips, K.A.; Chang, Y.; Henault, L.E.; Selby, J.V. Prevalence of diagnosed atrial fibrillation in adults: National implications for rhythm management and stroke prevention: The AnTicoagulation and Risk Factors in Atrial Fibrillation (ATRIA) Study. JAMA 2001, 285, 2370–2375. [Google Scholar] [CrossRef] [PubMed]
- Krahn, A.D.; Manfreda, J.; Tate, R.B.; Mathewson, F.A.; Cuddy, T.E. The natural history of atrial fibrillation: Incidence, risk factors and prognosis in the Manitoba Follow-Up Study. Am. J. Med. 1995, 98, 476–484. [Google Scholar] [CrossRef]
- Gudbjartsson, D.F.; Arnar, D.O.; Helgadottir, A.; Gretarsdottir, S.; Holm, H.; Sigurdsson, A.; Jonasdottir, A.; Baker, A.; Thorleifsson, G.; Kristjansson, K.; et al. Var-iants conferring risk of atrial fibrillation on chromosome 4q25. Nature 2007, 448, 353–357. [Google Scholar] [CrossRef] [PubMed]
- Gudbjartsson, D.F.; Holm, H.; Gretarsdottir, S.; Thorleifsson, G.; Walters, G.B.; Thorgeirsson, G.; Gulcher, J.; Mathiesen, E.B.; Njølstad, I.; Nyrnes, A.; et al. A sequence variant in ZFHX3 on 16q22 associates with atrial fibrillation and ischemic stroke. Nat. Genet. 2009, 41, 876–878. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, E.J.; Rice, K.M.; Arking, D.E.; Pfeufer, A.; van Noord, C.; Smith, A.V.; Schnabel, R.B.; Bis, J.C.; Boerwinkle, E.; Sinner, M.F.; et al. Variants in ZFHX3 are associated with atrial fibrilla- tion in individuals of European ancestry. Nat. Genet. 2009, 41, 879–881. [Google Scholar] [CrossRef] [PubMed]
- Ellinor, P.T.; Lunetta, K.L.; Glazer, N.L.; Pfeufer, A.; Alonso, A.; Chung, M.K.; Sinner, M.F.; de Bakker, P.I.; Mueller, M.; Lubitz, S.A.; et al. Common variants in KCNN3 are associated with lone atrial fibrillation. Nat. Genet. 2010, 42, 240–244. [Google Scholar] [CrossRef] [PubMed]
- Schnabel, R.B.; Kerr, K.F.; Lubitz, S.A.; Alkylbekova, E.L.; Marcus, G.M.; Sinner, M.F.; Magnani, J.W.; Wolf, P.A.; Deo, R.; Lloyd-Jones, D.M.; et al. Large-scale candidate gene analysis in whites and African Americans identifies IL6R polymorphism in relation to atrial fibrillation: The National Heart, Lung, and Blood Institute’s Candidate Gene Association Resource (CARe) project. Circ. Cardiovasc. Genet. 2011, 4, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Kääb, S.; Darbar, D.; van Noord, C.; Dupuis, J.; Pfeufer, A.; Newton-Cheh, C.; Schnabel, R.; Makino, S.; Sinner, M.F.; Kannankeril, P.J.; et al. Large scale replication and meta-analysis of variants on chromosome 4q25 associated with atrial fibrillation. Eur. Heart J. 2009, 30, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Lubitz, S.A.; Lunetta, K.L.; Lin, H.; Arking, D.E.; Trompet, S.; Li, G.; Krijthe, B.P.; Chasman, D.I.; Barnard, J.; Kleber, M.E.; et al. Novel genetic markers associate with atrial fibrillation risk in Europeans and Japanese. J. Am. Coll. Cardiol. 2014, 63, 1200–1210. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Sun, L.; Zhang, Y.; Liang, H.; Li, X.; Cai, R.; Wang, L.; Du, W.; Zhang, R.; Li, J.; et al. Overexpression of microRNA-1 causes atrioventricular block in rodents. Int. J. Biol. Sci. 2013, 9, 455–462. [Google Scholar] [CrossRef] [PubMed]
- Mahida, S.; Ellinor, P.T. New advances in the genetic basis of atrial fibrillation. J. Cardiovasc. Electrophysiol. 2012, 23, 1400–1406. [Google Scholar] [CrossRef] [PubMed]
- Mahida, S.; Mills, R.W.; Tucker, N.R.; Simonson, B.; Macri, V.; Lemoine, M.D.; Das, S.; Milan, D.J.; Ellinor, P.T. Overexpression of KCNN3 results in sudden cardiac death. Cardiovasc. Res. 2014, 101, 326–334. [Google Scholar] [CrossRef] [PubMed]
- Lozano-Velasco, E.; Hernández-Torres, F.; Daimi, H.; Serra, S.A.; Herraiz, A.; Hove-Madsen, L.; Aránega, A.; Franco, D. Pitx2 impairs calcium handling in a dose-dependent manner by modulating Wnt signalling. Cardiovasc. Res. 2016, 109, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wang, C.; Yao, Y.; Zuo, X.; Chen, S.; Xu, C.; Zhang, H.; Lu, Q.; Chang, L.; Wang, F.; et al. Molecular Basis of Gene-Gene Interaction: Cyclic Cross-Regulation of Gene Expression and Post-GWAS Gene-Gene Interaction Involved in Atrial Fibrillation. PLoS Genet. 2015, 11, e1005393. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.M.; Yang, F.; Mahida, S.; Zhao, L.; Chen, X.; Zhang, M.L.; Sun, Z.; Yao, Y.; Zhang, Y.X.; Zheng, G.Y.; et al. TBX5 mutations contribute to early-onset atrial fibrillation in Chinese and Caucasians. Cardiovasc. Res. 2016, 109, 442–450. [Google Scholar] [CrossRef] [PubMed]
- Nadadur, R.D.; Broman, M.T.; Boukens, B.; Mazurek, S.R.; Yang, X.; van den Boogaard, M.; Bekeny, J.; Gadek, M.; Ward, T.; Zhang, M.; et al. Pitx2 modulates a Tbx5-dependent gene regulatory network to maintain atrial rhythm. Sci. Transl. Med. 2016, 8, 354ra115. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Bai, Y.; Li, N.; Ye, W.; Zhang, M.; Greene, S.B.; Tao, Y.; Chen, Y.; Wehrens, X.H.; Martin, J.F. Pitx2-microRNA pathway that delimits sinoatrial node development and inhibits predisposition to atrial fibrillation. Proc. Natl. Acad. Sci. USA 2014, 111, 9181–9186. [Google Scholar] [CrossRef] [PubMed]
- Xia, M.; Jin, Q.; Bendahhou, S.; He, Y.; Larroque, M.M.; Chen, Y.; Zhou, Q.; Yang, Y.; Liu, Y.; Liu, B.; et al. A Kir2.1 gain-of-function mutation underlies familial atrial fibrillation. Biochem. Biophys. Res. Commun. 2005, 332, 1012–1019. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.H.; Xu, S.J.; Bendahhou, S.; Wang, X.L.; Wang, Y.; Xu, W.Y.; Jin, H.W.; Sun, H.; Su, X.Y.; Zhuang, Q.N.; et al. KCNQ1 gain-of-function mutation in familial atrial fibrillation. Science 2003, 299, 251–254. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, L.A.; Alonso, M.E.; Badía-Careaga, C.; Rollán, I.; Arias, C.; Fernández-Miñán, A.; López-Jiménez, E.; Aránega, A.; Gómez-Skarmeta, J.L.; Franco, D.; et al. Long-range regulatory interactions at the 4q25 atrial fibrillation risk locus involve PITX2c and ENPEP. BMC Biol. 2015, 13, 26. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Tucker, N.R.; Weng, L.C.; Clauss, S.; Lubitz, S.A.; Ellinor, P.T. A Functional Variant Associated with Atrial Fibrillation Regulates PITX2c Expression through TFAP2a. Am. J. Hum. Genet. 2016, 99, 1281–1291. [Google Scholar] [CrossRef] [PubMed]
- Torrado, M.; Franco, D.; Lozano-Velasco, E.; Hernández-Torres, F.; Calviño, R.; Aldama, G.; Centeno, A.; Castro-Beiras, A.; Mikhailov, A. A MicroRNA-Transcription Factor Blueprint for Early Atrial Arrhythmogenic Remodeling. BioMed Res. Int. 2015, 2015, 263151. [Google Scholar] [CrossRef] [PubMed]
- Syeda, F.; Holmes, A.P.; Yu, T.Y.; Tull, S.; Kuhlmann, S.M.; Pavlovic, D.; Betney, D.; Riley, G.; Kucera, J.; Jousset, F.; et al. Pitx2 modulates atrial membrane potential and reduced PITX2 potentiates the anti-arrhythmic effects of socium-channel blockers. J. Am. Coll. Cardiol. 2016, 68, 1881–1894. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Yang, Y.-Q.; Gong, X.Q.; Wang, X.-H.; Li, R.-G.; Tan, H.-W.; Liu, X.; Fang, W.-Y.; Bai, D. Novel germline GJA5/connexin40 mutations associated with the lone atrial fibrillation impair gap junctional intercellular communication. Hum. Mutat. 2013, 34, 603–609. [Google Scholar] [PubMed]
- Pérez-Hernández, M.; Matamoros, M.; Barana, A.; Amorós, I.; Gómez, R.; Núñez, M.; Sacristán, S.; Pinto, A.; Fernández-Avilés, F.; Tamargo, J.; et al. Pitx2c increases in atrial myocites from chronic atrial fibrillation patients enhancing IKs and devreasin ICa, L. Cardiovasc. Res. 2016, 109, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Girmatsion, Z.; Biliczki, P.; Bonauer, A.; Wimmer-Greinecker, G.; Scherer, M.; Moritz, A.; Bukowska, A.; Goette, A.; Nattel, S.; Hohnloser, S.H.; et al. Changes in microRNA-1 expression and IK1 up-regulation in human atrial fibrillation. Heart Rhythm 2009, 6, 1802–1809. [Google Scholar] [CrossRef] [PubMed]
- Cooley, N.; Cowley, M.J.; Lin, R.C.; Marasco, S.; Wong, C.; Kaye, D.M.; Dart, A.M.; Woodcock, E.A. Influence of atrial fibrillation on microRNA expression profiles in left and right atria from patients with valvular heart disease. Physiol. Genom. 2012, 44, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Scridon, A.; Fouilloux-Meugnier, E.; Loizon, E.; Rome, S.; Julien, C.; Barrès, C.; Chevalier, P. Long-standing arterial hypertension is associated with Pitx2 down-regulation in a rat model of spontaneous atrial tachyarrhythmias. Europace 2015, 17, 160–165. [Google Scholar] [CrossRef] [PubMed]
- Fakhro, K.A.; Choi, M.; Ware, S.M.; Belmont, J.W.; Towbin, J.A.; Lifton, R.P.; Khokha, M.K.; Brueckner, M. Rare copy number variations in congenital heart disease patients identify unique genes in left-right patterning. Proc. Natl. Acad. Sci. USA 2011, 108, 2915–2920. [Google Scholar] [CrossRef] [PubMed]
- Franco, D.; Chinchilla, A.; Daimi, H.; Dominguez, J.N.; Aránega, A. Modulation of conductive elements by Pitx2 and their impact on atrial arrhythmogenesis. Cardiovasc. Res. 2011, 91, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Franco, D.; Chinchilla, A.; Aránega, A.E. Transgenic insights linking pitx2 and atrial arrhythmias. Front. Physiol. 2012, 3, 206. [Google Scholar] [CrossRef] [PubMed]
- Franco, D.; Christoffels, V.M.; Campione, M. Homeobox transcription factor Pitx2: The rise of an asymmetry gene in cardiogenesis and arrhythmogenesis. Trends Cardiovasc. Med. 2014, 24, 23–31. [Google Scholar] [CrossRef] [PubMed]
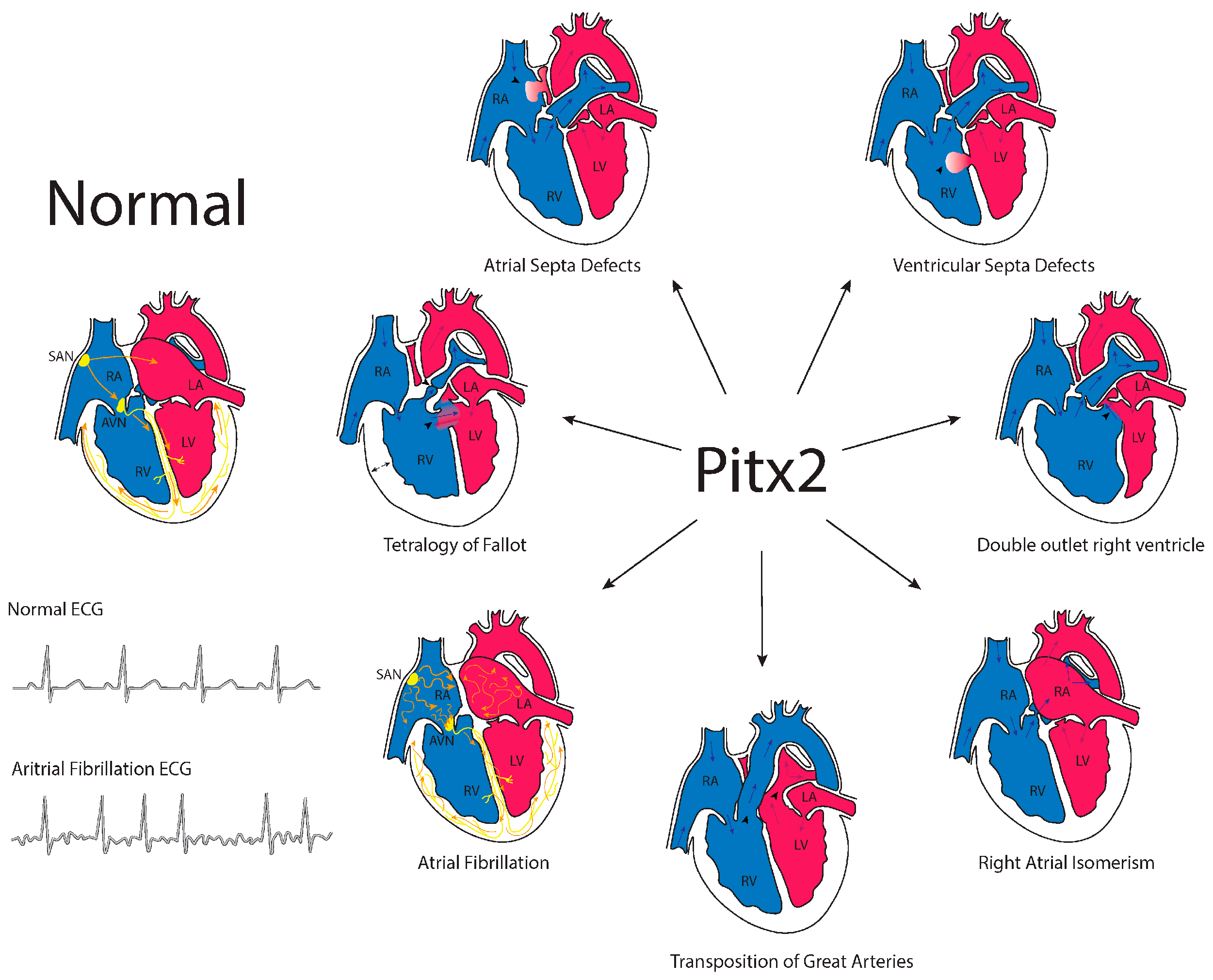
| Congenital Heart Disease | Type | References |
|---|---|---|
| Atrial Septa Defect (ASD) | Ostium primum ASD Ostium secundum ASD Patent foramen ovale Sinus venosus ASD Common or single atrium | [124] [130,131,132] |
| Ventricular Septa Defect (VSD) | Perimembranous/membranous VSD (type 2) Inlet and AV canal VSD (type 3) Left ventricular to right atrial communication (Gerbode type) | [125] [126] |
| Double Outlet Right Ventricle (DORV) | DORV with subaortic VSD DORV with subpulmonic VSD | [7,10] [38,39] |
| Right Atrial Isomerism (RAI) | [27,28] [133] | |
| Transposition of Great Arteries (TGA) | [125] | |
| Tetralogy of Fallot (TOF) | [131] |
| Cardiac Disease | Type | Related Pathways | References |
|---|---|---|---|
| Atrial Fibrillation (AF) | Paroxysmal Persistent Permanent | Pitx2 regulates AF through modulation of multiple genes and miRNAs implicated: → KCNN3, TBX5, HCN4, KCNJ2, CAV-1, KCNQ1, ZFHX3, SYNE2, IL6R, ENPEP, SCN5A, GJA1, RYR2, DSP, ATP2A2, SHOX2, TBX3, WNT8A → miR17-92, miR106b-25, miR-1, miR-26b, miR-29a, miR-106a/b, miR-13 | [104,105,106] [137,138,139,140,141,142,143] [145,146,147] [154] [147] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Franco, D.; Sedmera, D.; Lozano-Velasco, E. Multiple Roles of Pitx2 in Cardiac Development and Disease. J. Cardiovasc. Dev. Dis. 2017, 4, 16. https://doi.org/10.3390/jcdd4040016
Franco D, Sedmera D, Lozano-Velasco E. Multiple Roles of Pitx2 in Cardiac Development and Disease. Journal of Cardiovascular Development and Disease. 2017; 4(4):16. https://doi.org/10.3390/jcdd4040016
Chicago/Turabian StyleFranco, Diego, David Sedmera, and Estefanía Lozano-Velasco. 2017. "Multiple Roles of Pitx2 in Cardiac Development and Disease" Journal of Cardiovascular Development and Disease 4, no. 4: 16. https://doi.org/10.3390/jcdd4040016
APA StyleFranco, D., Sedmera, D., & Lozano-Velasco, E. (2017). Multiple Roles of Pitx2 in Cardiac Development and Disease. Journal of Cardiovascular Development and Disease, 4(4), 16. https://doi.org/10.3390/jcdd4040016







