Detection of Methicillin-Resistant Staphylococci Isolated from Food Producing Animals: A Public Health Implication
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Sampling and Isolation Method
2.3. Bacterial Identification of Staphylococci
2.4. Molecular Identification by PCR
2.4.1. Genomic DNA Extraction
2.4.2. PCR for Identification of Staphylococcal Isolates
2.4.3. Sequencing of the 16S rRNA Genes
2.4.4. Specific PCR for the Identification of Methicillin-Resistant Staphylococcus aureus
2.4.5. Detection Luk S/F gene Sequences that Encode Virulence Determinants in Staphylococci
2.5. Determination of the Antibiotic Resistance Profiles of Staphylococcal Isolates
2.5.1. Antibiotic Susceptibility Testing
2.5.2. Statistical Analysis
3. Results
3.1. Specific PCR for the Identification of Staphylococcal Isolates
3.2. Prevalence and Species Diversity of Staphylococcal Isolates
3.3. Antibiotic Resistance of Staphylococcal Isolates from Nasal and Raw Milk Samples
3.4. Multiple Antibiotic Resistance Phenotypes of Staphylococcal Isolates
3.5. Detection of Resistance and Virulence Genes in Staphylococcal Isolates
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- El-Jakee, J.K.; Atta, N.S.; Sammy, A.A.; Elgabry, E.A.; Kandil, M.M.; El-Said, W.A. Antimicrobial resistance in clinical isolates of Staphylococcus aureus from bovine and human sources in Egypt. Glob. Vet. 2011, 7, 581–586. [Google Scholar]
- Hulya, T.; Senay, E.; Dilek, O. Antibiotic resistance of Staphylococcus aureus and coagulase-negative staphylococci isolated from bovine mastitis. Bull. Vet. Inst. Pulawy 2006, 50, 41–45. [Google Scholar]
- Feßler, A.; Scott, C.; Kadlec, K.; Ehricht, R.; Monecke, S.; Schwarz, S. Characterization of methicillin-resistant Staphylococcus aureus ST398 from cases of bovine mastitis. J. Antimicrob. Chemother. 2010, 65, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Vishnupriya, S.; Anthony, P.X.; Mukhopadhyay, H.K.; Pillai, R.M.; Thanislass, J.; Vivek, S.V.M.; Sumanth, K.R. Methicillin-resistant staphylococci associated with bovine mastitis and their zoonotic importance. Vet. World 2014, 7, 422–427. [Google Scholar] [CrossRef]
- Carfora, V.; Caprioli, A.; Marri, N.; Sagrafoli, D.; Boselli, C.; Giacinti, G.; Giangolini, G.; Sorbara, L.; Dottarelli, S.; Battisti, A.; et al. Enterotoxin genes, enterotoxin production, and methicillin resistance in Staphylococcus aureus isolated from milk and dairy products in central Italy. Int. Dairy J. 2015, 42, 12–15. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA). Assessment of the public health significance of methicillin-resistant Staphylococcus aureus (MRSA) in animals and foods-scientific opinion of the panel on biological hazards. Eur. Food Saf. Auth. J. 2009, 993, 1–73. [Google Scholar]
- Rinsky, J.L.; Nadimpalli, M.; Wing, S.; Hall, D.; Baron, D.; Price, L.B.; Larsen, J.; Stegger, M.; Stewart, J.; Heaney, C.D. Livestock-associated methicillin and multidrug resistant Staphylococcus aureus is present among industrial, not antibiotic-free livestock operation workers in North Carolina. PLoS ONE 2013, 8, e67641. [Google Scholar] [CrossRef] [PubMed]
- Cruickshank, R.; Duguid, J.P.; Marmion, B.R.; Swain, R.H. Medical Microbiology, 12th ed.; Churchill-Living Stone: London, UK, 1975. [Google Scholar]
- Shuiep, E.S.; Kanbar, T.; Eissa, N.; Alber, J.; Lammler, C.; Zschock, M.; El zubeir, I.E.M.; Weiss, R. Phenotypic and genotypic characterization of Staphylococcus aureus isolated from raw camel milk sample. Res. Vet. Sci. 2009, 86, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Ma, A.; Lv, D.; Zhuang, X.; Zhuang, G. Quorum quenching in culturable phyllosphere bacteria from tobacco. Int. J. Mol. Sci. 2013, 14, 14607–14619. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nel, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSIBLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, B.O.; Elmekki, A.M.; Omer, E.E.; Elhassan, M. Molecular detection of methicillin-resistant Staphylococcus aureus in patients with urinary tract infections in Khartoum State. J. Sci. Technol. 2014, 15, 1–8. [Google Scholar]
- McDonald, R.R.; Antonishyn, A.N.; Hansen, T.; Snook, A.L.; Nagle, E.; Mulvey, R.M.; Levett, N.P.; Horsman, B.G. Development of a triplex real-time PCR assay for detection of panton-valentine leucocidin toxin genes in clinical isolates of methicillin-resistant Staphylococcus aureus. J. Clin. Microbiol. 2005, 43, 6147–6149. [Google Scholar] [CrossRef] [PubMed]
- Bauer, A.W.; Kirby, M.M.W.; Sherris, J.C.; Turck, M. Antibiotics susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 1996, 36, 493–496. [Google Scholar]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing; Sixteen International Supplements; CLSI document M100 S16. 26(3); M7-A7, 26(2); M2-A9, 26(1); Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2006; pp. 118–123. [Google Scholar]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Vanderhaeghen, W.; Hermans, K.; Haesebrouck, F.; Butaye, P. Methicillin-resistant Staphylococcus aureus (MRSA) in food production animals. Epidemiol. Infect. 2010, 138, 606–625. [Google Scholar] [CrossRef] [PubMed]
- Weese, S.J.; van-Duijkeren, E. Methicillin-resistant Staphylococcus aureus (MRSA) and Staphylococcus pseudintermedius in veterinary medicine. Vet. Microbiol. 2010, 140, 418–429. [Google Scholar] [CrossRef] [PubMed]
- Pehlivanoglu, F.; Yardimci, H. Detection of methicillin and vancomycin resistance in Staphylococcus strains isolated from Bovine milk samples with mastitis. Kafkas Univ. Vet. Fak. Derg. 2012, 18, 849–855. [Google Scholar] [CrossRef]
- Garipcin, M.; Seker, E. Nasal carriage of methicillin-resistant Staphylococcus aureus in Cattle and farm workers in Turkey. Vet. Arh. 2015, 85, 117–129. [Google Scholar]
- Abd-Al-Azeem, M.W.; Shaheen, H.M.; Abdel-Hameed, K.G.; Helmy, M.M. Penicillin resistance against Staphylococcal isolates recovered from subclinical mastitis in Sohag City, Egypt. Am. J. Res. Commun. 2013, 1, 116–130. [Google Scholar]
- Jahan, M.; Rahman, M.; Parvej, S.; Chowdhury, Z.H.; Haque, E.; Talukder, A.K.; Ahmed, S. Isolation and characterization of Staphylococcus aureus from raw cow milk in Bangladesh. J. Adv. Vet. Anim. Res. 2015, 2, 49–55. [Google Scholar] [CrossRef]
- Nakamura, K.R.; Tompkins, E. Methicillin-resistant Staphylococcus aureus infections. In Compendium: Continuing Education for Veterinarians; Vetlearn.com: Lawndale, CA, USA, 2012; pp. E1–E11. [Google Scholar]
- Mehndiratta, P.L.; Bhalla, P. Use of antibodies in animal agriculture and emergence of methicillin-resistant Staphylococcus aureus (MRSA) clones: Need to assess the impact on public health. Indian J. Med. Res. 2014, 140, 339–344. [Google Scholar] [PubMed]
- Abdulgader, M.S.; Shittu, O.A.; Kaba, M. Molecular epidemiology of methicillin-resistant Staphylococcus aureus in Africa: A systematic review. Front. Microbiol. 2015, 6, 1–21. [Google Scholar] [CrossRef] [PubMed]
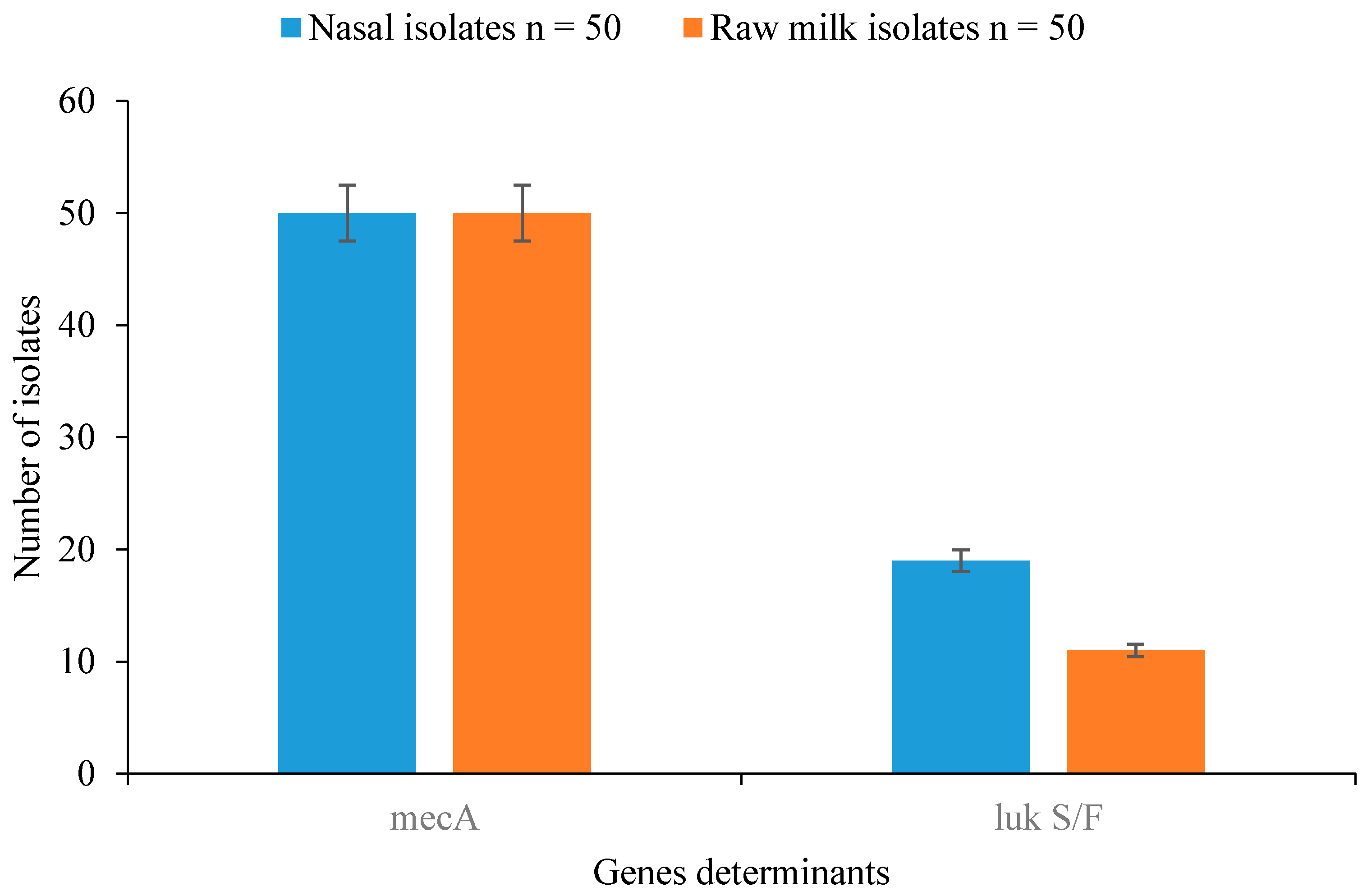
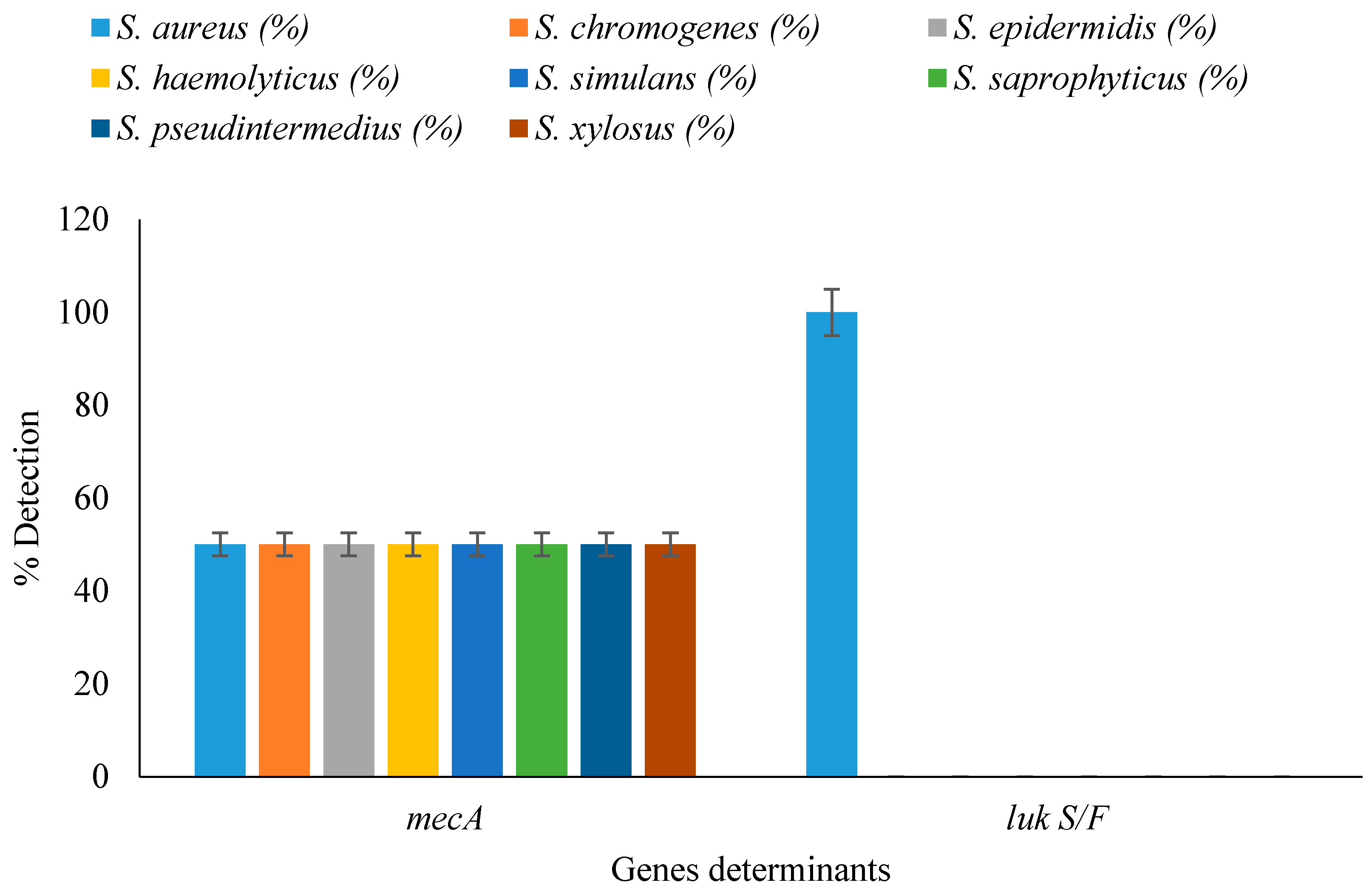
| Staphylococcal Isolates | Nasal Isolates n = 50 | Raw Milk Isolates n = 50 | Total n = 100 | p-Value |
|---|---|---|---|---|
| S. aureus | 19 (38) | 11 (22) | 30 (30) | 0.188 |
| S. chromogenes | 0 (0) | 8 (16) | 8 (8) | 0.015 |
| S. epidermidis | 13 (26) | 4 (8) | 17 (17) | 0.188 |
| S. haemolyticus | 0 (0) | 15 (30) | 15 (15) | 0.225 |
| S. simulans | 0 (0) | 7 (14) | 7 (7) | 0.250 |
| S. saprophyticus | 8 (16) | 5 (10) | 13 (13) | 0.580 |
| S. pseudintermedius | 6 (12) | 0 (0) | 6 (6) | 0.184 |
| S. xylosus | 4 (8) | 0 (0) | 4 (4) | 0.184 |
| Antibiotics | Resistant Profile (%) | p-Value | |||||||
|---|---|---|---|---|---|---|---|---|---|
| S. aureus n = 30 | S. chromogenes n = 8 | S. epidermidis n = 17 | S. haemolyticus n = 15 | S. simulans n = 7 | S. saprophyticus n = 13 | S. pseudintermedius n = 6 | S. xylosus n = 4 | ||
| MET | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 1.000 |
| PEN | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 1.000 |
| CLX | 100 | 100 | 100 | 100 | 100 | 92.30 | 100 | 75 | 0.043 |
| AMX | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 1.000 |
| ERY | 90 | 87.50 | 88.20 | 100 | 100 | 100 | 83.33 | 100 | 0.645 |
| GEN | 0 | 0 | 11.76 | 13.33 | 0 | 0 | 0 | 0 | 0.015 |
| KAN | 90 | 100 | 94.12 | 100 | 100 | 92.31 | 100 | 100 | 0.794 |
| CLN | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 1.000 |
| CHL | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 1.000 |
| SXT | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 100 | 1.000 |
| VAN | 13.30 | 25 | 5.88 | 26.67 | 57.1 | 7.69 | 0 | 0 | 0.000 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Igbinosa, E.O.; Beshiru, A.; Akporehe, L.U.; Ogofure, A.G. Detection of Methicillin-Resistant Staphylococci Isolated from Food Producing Animals: A Public Health Implication. Vet. Sci. 2016, 3, 14. https://doi.org/10.3390/vetsci3030014
Igbinosa EO, Beshiru A, Akporehe LU, Ogofure AG. Detection of Methicillin-Resistant Staphylococci Isolated from Food Producing Animals: A Public Health Implication. Veterinary Sciences. 2016; 3(3):14. https://doi.org/10.3390/vetsci3030014
Chicago/Turabian StyleIgbinosa, Etinosa O., Abeni Beshiru, Lucy U. Akporehe, and Abraham G. Ogofure. 2016. "Detection of Methicillin-Resistant Staphylococci Isolated from Food Producing Animals: A Public Health Implication" Veterinary Sciences 3, no. 3: 14. https://doi.org/10.3390/vetsci3030014





