Repository Approaches to Improving the Quality of Shared Data and Code
Abstract
1. Introduction
2. Approaches for Advancing Dataset Quality
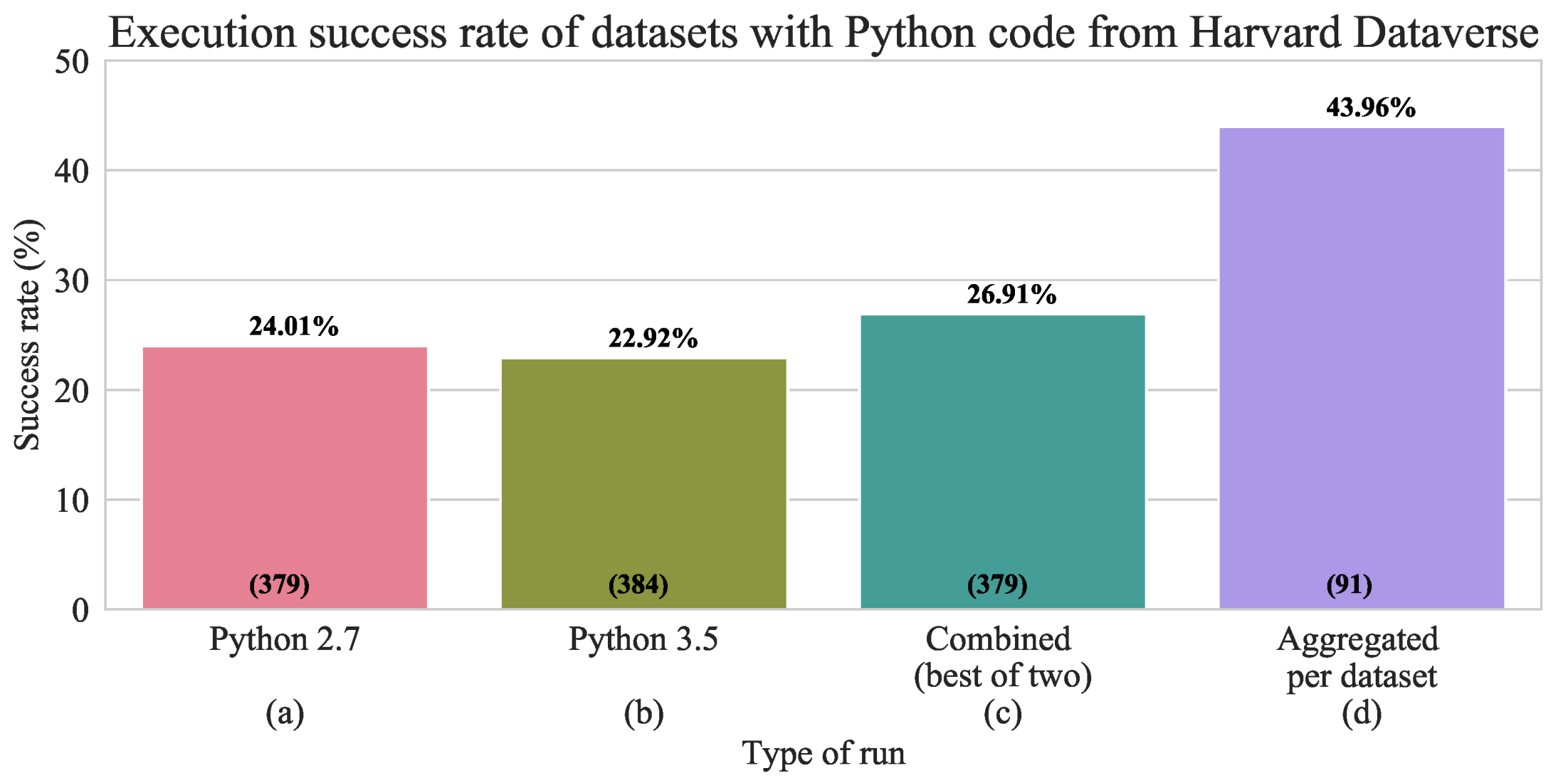
2.1. Ensure Research Code Completeness
- We look for files such as “requirements.txt” or “environment.yml” inside the dataset because these filenames are common conventions for documenting needed code dependencies for Python. If such files were not found, we scan the Python code looking for the used libraries, and create a new requirements file. We attempt to install all libraries from the requirements file.
- We automatically (naively) re-execute the Python files first with Python 2.7 and then with Python 3.5 with a time limit of 10 minutes per each Python file. If the file executes successfully in the allocated time, we record a success; if it crashes with error, we record the error; and if it exceeds the allocated time, we record ’time limit exceeded’ (TLE) or null result (which are ignored in the success analysis as we cannot be certain whether the file would successfully execute or not).
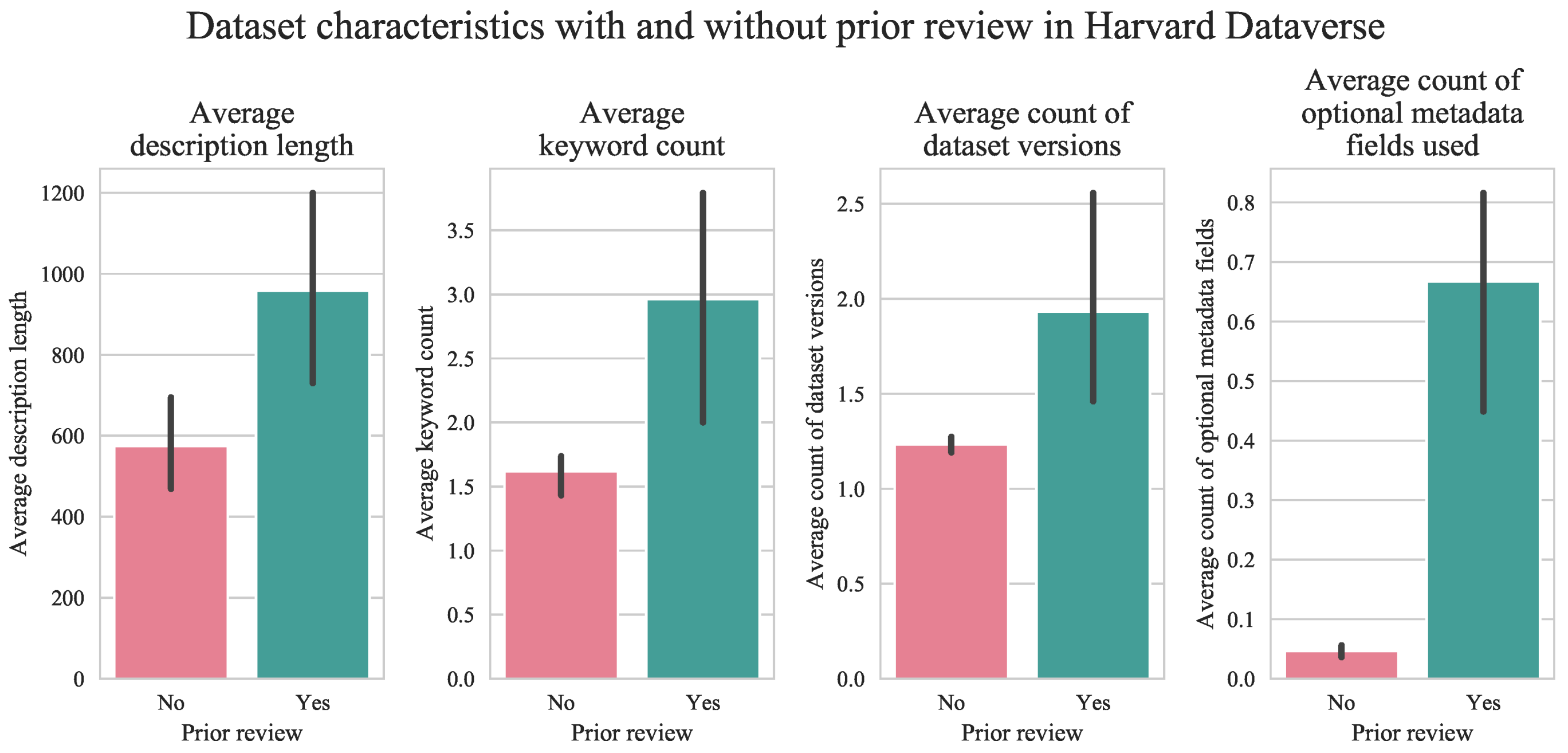
2.2. Encourage Use of Curation Features and Pre-Submission Dataset Review
- Optional metadata blocks. A well-curated dataset should have at least one optional metadata block to support its discoverability and reuse.
- Keywords. A well-curated dataset should also have at least one keyword.
- Description. A well-curated dataset should have a description. Like keywords, descriptions help to facilitate its discovery and reuse.
- Open file formats. A well-curated dataset should use open file formats, where possible.
- Discipline standard file formats. Not all disciplines use open standards, but at minimum, datasets should adhere to best practices for discipline file formats.
- Supplemental Files. A well-curated dataset should have either a codebook or a readme file that provides insight into the datasets’ internals, such as descriptions of its variables.
- Submission review. A well-curated dataset may undergo an additional review by the collection owner prior to publication. In contrast to the previous six, this characteristic might be considered a direct indicator of dataset quality [24].
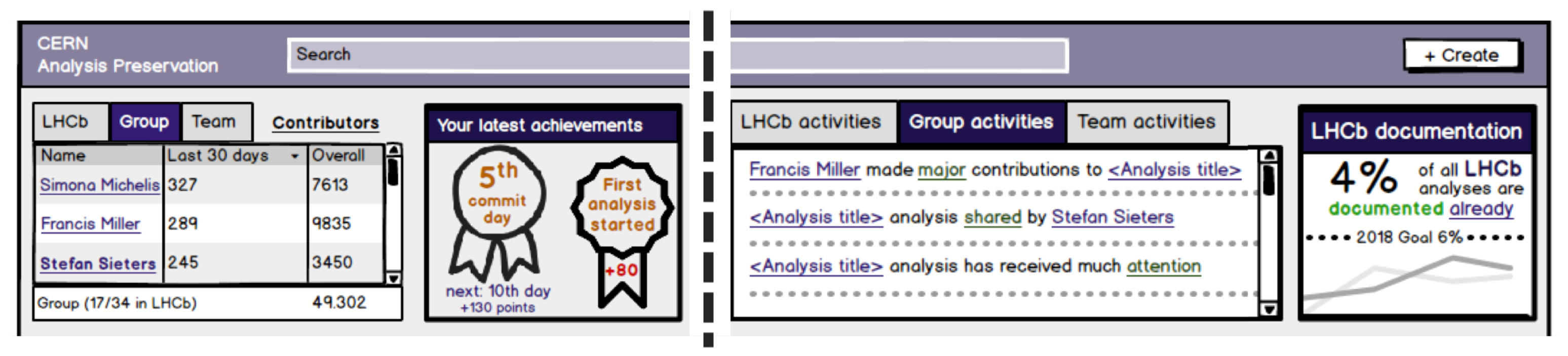
2.3. Incorporate Gamified Design Elements
3. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Borgman, C.L. Big Data, Little Data, No Data: Scholarship in the Networked World; MIT Press: Cambridge, MA, USA, 2015. [Google Scholar]
- Baker, M. 1500 scientists lift the lid on reproducibility. Nature 2016, 533, 452–454. [Google Scholar] [CrossRef] [PubMed]
- Stodden, V.; Seiler, J.; Ma, Z. An empirical analysis of journal policy effectiveness for computational reproducibility. Proc. Natl. Acad. Sci. USA 2018, 115, 2584–2589. [Google Scholar] [CrossRef] [PubMed]
- Pimentel, J.F.; Murta, L.; Braganholo, V.; Freire, J. A large-scale study about quality and reproducibility of jupyter notebooks. In Proceedings of the 2019 IEEE/ACM 16th International Conference on Mining Software Repositories (MSR), Montreal, QC, Canada, 25–31 May 2019; pp. 507–517. [Google Scholar]
- Assante, M.; Candela, L.; Castelli, D.; Tani, A. Are Scientific Data Repositories Coping with Research Data Publishing? Data Sci. J. 2016, 15, 6. [Google Scholar] [CrossRef]
- Crosas, M. The Dataverse Network®: An Open-Source Application for Sharing, Discovering and Preserving Data. D-Lib Mag. 2011, 17. [Google Scholar] [CrossRef]
- King, G. An Introduction to the Dataverse Network as an Infrastructure for Data Sharing. Sociol. Methods Res. 2007. [Google Scholar] [CrossRef]
- Marchionini, G.; Lee, C.A.; Bowden, H.; Lesk, M. Curating for Quality: Ensuring Data Quality to Enable New Science; Final Report: Invitational Workshop Sponsored by the National Science Foundation; National Science Foundation: Arlington, VA, USA, 2012. [Google Scholar]
- Cai, L.; Zhu, Y. The challenges of data quality and data quality assessment in the big data era. Data Sci. J. 2015, 14. [Google Scholar] [CrossRef]
- Martin, E.G.; Law, J.; Ran, W.; Helbig, N.; Birkhead, G.S. Evaluating the quality and usability of open data for public health research: A systematic review of data offerings on 3 open data platforms. J. Public Health Manag. Pract. 2017, 23, e5–e13. [Google Scholar] [CrossRef]
- Ferguson, A.R.; Nielson, J.L.; Cragin, M.H.; Bandrowski, A.E.; Martone, M.E. Big data from small data: Data-sharing in the ’long tail’ of neuroscience. Nat. Neurosci. 2014, 17, 1442–1447. [Google Scholar] [CrossRef]
- Heidorn, P.B. Shedding Light on the Dark Data in the Long Tail of Science. Libr. Trends 2008, 57, 280–299. [Google Scholar] [CrossRef]
- Palmer, C.L.; Cragin, M.H.; Heidorn, P.B.; Smith, L.C. Data Curation for the Long Tail of Science: The Case of Environmental Sciences. In Proceedings of the Third International Digital Curation Conference, Washington, DC, USA, 11–13 December 2007; pp. 11–13. [Google Scholar]
- Cragin, M.H.; Palmer, C.L.; Carlson, J.R.; Witt, M. Data sharing, small science and institutional repositories. Philos. Trans. Math. Phys. Eng. Sci. 2010, 368, 4023–4038. [Google Scholar] [CrossRef]
- Chen, X.; Dallmeier-Tiessen, S.; Dasler, R.; Feger, S.; Fokianos, P.; Gonzalez, J.B.; Hirvonsalo, H.; Kousidis, D.; Lavasa, A.; Mele, S.; et al. Open is not enough. Nat. Phys. 2019, 15, 113–119. [Google Scholar] [CrossRef]
- Gregory, K.; Groth, P.; Scharnhorst, A.; Wyatt, S. Lost or Found? Discovering Data Needed for Research. Harv. Data Sci. Rev. 2020. [Google Scholar] [CrossRef]
- Pasquetto, I.V.; Borgman, C.L.; Wofford, M.F. Uses and reuses of scientific data: The data creators’ advantage. Harv. Data Sci. Rev. 2019, 2019, 1. [Google Scholar]
- Borgman, C.L.; Wallis, J.C.; Enyedy, N. Little Science Confronts the Data Deluge: Habitat Ecology, Embedded Sensor Networks, and Digital Libraries. Cent. Embed. Netw. Sens. 2006, 7, 17–30. [Google Scholar] [CrossRef]
- Borgman, C.L. The conundrum of sharing research data. J. Am. Soc. Inf. Sci. Technol. 2012, 63, 1059–1078. [Google Scholar] [CrossRef]
- Wallis, J.C.; Rolando, E.; Borgman, C.L. If We Share Data, Will Anyone Use Them? Data Sharing and Reuse in the Long Tail of Science and Technology. PLoS ONE 2013, 8, e67332. [Google Scholar] [CrossRef]
- National Academies of Sciences, Engineering, and Medicine. Reproducibility and Replicability in Science; National Academies Press: Washington, DC, USA, 2019. [CrossRef]
- Trisovic, A. Replication Data for: Repository approaches to improving quality of shared data and code. Harvard Dataverse, 13 October 2020. [Google Scholar] [CrossRef]
- Trisovic, A.; Durbin, P.; Schlatter, T.; Durand, G.; Barbosa, S.; Brooke, D.; Crosas, M. Advancing Computational Reproducibility in the Dataverse Data Repository Platform. In Proceedings of the 3rd International Workshop on Practical Reproducible Evaluation of Computer Systems, P-RECS ’20, Stockholm, Sweden, 23 June 2020; Association for Computing Machinery: New York, NY, USA, 2020; pp. 15–20. [Google Scholar] [CrossRef]
- Hense, A.; Quadt, F. Acquiring high quality research data. D-Lib Mag. 2011, 17. [Google Scholar] [CrossRef]
- Wilkinson, M.D.; Dumontier, M.; Aalbersberg, I.J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.W.; da Silva Santos, L.B.; Bourne, P.E.; et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 2016, 3, 1–9. [Google Scholar] [CrossRef]
- Boyd, C. Harvard Dataverse Optional Feature Use Data. Harvard Dataverse, 2 October 2020. [Google Scholar] [CrossRef]
- Koshoffer, A.; Neeser, A.E.; Newman, L.; Johnston, L.R. Giving datasets context: A comparison study of institutional repositories that apply varying degrees of curation. Int. J. Digit. Curation 2018, 13, 15–34. [Google Scholar] [CrossRef]
- Bishop, B.W.; Hank, C.; Webster, J.; Howard, R. Scientists’ data discovery and reuse behavior: (Meta)data fitness for use and the FAIR data principles. Proc. Assoc. Inf. Sci. Technol. 2019, 56, 21–31. [Google Scholar] [CrossRef]
- Smit, E. Abelard and Héloise: Why Data and Publications Belong Together. D-Lib Mag. 2011, 17. [Google Scholar] [CrossRef]
- Faniel, I.M.; Jacobsen, T.E. Reusing Scientific Data: How Earthquake Engineering Researchers Assess the Reusability of Colleagues’ Data. Comput. Support. Coop. Work. (CSCW) 2010, 19, 355–375. [Google Scholar] [CrossRef]
- Deterding, S.; Khaled, R.; Nacke, L.E.; Dixon, D. Gamification: Toward a definition. In Proceedings of the CHI 2011 Gamification Workshop Proceedings, Vancouver, BC, Canada, 7–12 May 2011; Volume 12. [Google Scholar]
- Hamari, J.; Koivisto, J.; Sarsa, H. Does gamification work?—A literature review of empirical studies on gamification. In Proceedings of the 2014 47th Hawaii International Conference on System Sciences (HICSS), Waikoloa, HI, USA, 6–9 January 2014; pp. 3025–3034. [Google Scholar]
- Knaving, K.; Woźniak, P.W.; Niess, J.; Poguntke, R.; Fjeld, M.; Björk, S. Understanding grassroots sports gamification in the wild. In Proceedings of the 10th Nordic Conference on Human-Computer Interaction, Oslo, Norway, 1–3 September 2018; pp. 102–113. [Google Scholar]
- Oprescu, F.; Jones, C.; Katsikitis, M. I PLAY AT WORK—Ten principles for transforming work processes through gamification. Front. Psychol. 2014, 5, 14. [Google Scholar] [CrossRef]
- Ibanez, M.B.; Di-Serio, A.; Delgado-Kloos, C. Gamification for Engaging Computer Science Students in Learning Activities: A Case Study. IEEE Trans. Learn. Technol. 2014, 7, 291–301. [Google Scholar] [CrossRef]
- Eveleigh, A.; Jennett, C.; Lynn, S.; Cox, A.L. “I want to be a captain! I want to be a captain!”: Gamification in the old weather citizen science project. In Proceedings of the First International Conference on Gameful Design, Research, and Applications— Gamification ’13, Toronto, ON, Canada, 2–4 October 2013; pp. 79–82. [Google Scholar] [CrossRef]
- Bowser, A.; Hansen, D.; Preece, J.; He, Y.; Boston, C.; Hammock, J. Gamifying citizen science: A study of two user groups. In Proceedings of the 17th ACM Conference on Computer Supported Cooperative Work and Social Computing, CSCW 2014, Baltimore, MD, USA, 15–19 February 2014; pp. 137–140. [Google Scholar] [CrossRef]
- Nicholson, S. A recipe for meaningful gamification. In Gamification in Education and Business; Springer: New York, NY, USA, 2015; pp. 1–20. [Google Scholar] [CrossRef]
- Feger, S.; Dallmeier-Tiessen, S.; Woźniak, P.; Schmidt, A. Just Not The Usual Workplace: Meaningful Gamification in Science. In Proceedings of the Mensch und Computer 2018-Workshopband, Dresden, Germany, 2–5 September 2018. [Google Scholar]
- Feger, S.S.; Dallmeier-Tiessen, S.; Woźniak, P.W.; Schmidt, A. Gamification in Science: A Study of Requirements in the Context of Reproducible Research. In Proceedings of the 2019 CHI Conference on Human Factors in Computing Systems, Glasgow, UK, 4–9 May 2019; pp. 1–14. [Google Scholar] [CrossRef]
- Kidwell, M.C.; Lazarević, L.B.; Baranski, E.; Hardwicke, T.E.; Piechowski, S.; Falkenberg, L.S.; Kennett, C.; Slowik, A.; Sonnleitner, C.; Hess-Holden, C.; et al. Badges to acknowledge open practices: A simple, low-cost, effective method for increasing transparency. PLoS Biol. 2016, 14, e1002456. [Google Scholar] [CrossRef]
- Rowhani-Farid, A.; Allen, M.; Barnett, A.G. What incentives increase data sharing in health and medical research? A systematic review. Res. Integr. Peer Rev. 2017, 2, 4. [Google Scholar] [CrossRef]
- Borges, H.; Valente, M.T. What’s in a GitHub star? understanding repository starring practices in a social coding platform. J. Syst. Softw. 2018, 146, 112–129. [Google Scholar] [CrossRef]
| 1. | There were additional 15 datasets that contained Python files and were visible through the API but could not be retrieved due to restricted authorization or connection error. |
| 2. | AJPS Data Policy: https://ajps.org/ajps-verification-policy/. |
| 3. | In practice, a publication reference is often placed in the dataset description field. |
| 4. | |
| 5. |
| Cai & Zhu | Martin et al. | Examples of Common Data Repository Features |
|---|---|---|
| Availability: accessibility, timeliness, authorization | Accessibility, timeliness, representational consistency, visibility, platform functionality | Capturing data citation information, minting DOIs |
| Usability: documentation, credibility, metadata | Intended use, subject matter expertise, technical skills, metadata quality (standards & consistency); learnability, believability & reputation, confidentiality, etc. | Supporting documentation, reuse licensing, terms of access/restrictions |
| Reliability: accuracy, integrity, consistency, completeness, auditability | Data accuracy, validity, reliability, completeness, missing data, timing and frequency, collection methods, format & layout, sample size & method, representation, study design, unit of analysis, etc. | Metadata standards, variable level metadata support |
| Relevance: fitness | Relevancy, value added | Reuse metrics, granular description |
| Presentation quality: readability & structure | Concise representation, ease of understanding, ease of manipulation, user-friendliness | Preview options, UI/UX reviews |
| Platform promotion and user training: availability of information, capacity to respond to feedback, financial resources, legal protections and interpretations, platform training and promotion, policies and regulation, political support for developing and releasing data | Support services, preservation policies, governance, and legal policies |
| File | Count (Out of 92) |
|---|---|
| environment.yml | 0 |
| requirement.txt | 6 |
| Dockerfile | 0 |
| README, instructions or codebook | 57 |
| Category I: Basic | Category II: Enhanced | Category III: Comprehensive |
|---|---|---|
| Default curation support supplied or enforced by the Dataverse software | Optional curation support provided by Dataverse software | Research-dependent data quality characteristics |
| E.g., Default rights statements, Required metadata fields, Reliable storage and access, Persistent identifiers (DOIs), Data citations | E.g., File versioning, Optional keywords, Optional description, Optional metadata blocks, Optional rights statements, Optional supporting documentation | E.g., Comprehensible variable names, Confirmed valid data values, Up-to-date codebook, Well-documented code |
| Characteristic | Value (n) | % of Total |
|---|---|---|
| Total published datasets (N) | 29,295 | 100% |
| Total file count in datasets | 383,685 | 100% |
| Contain optional metadata blocks | 8380 | 28.6% |
| Contain keywords | 14,593 | 49.8% |
| Contain description | 24,661 | 84.2% |
| Dataset linked to a publication | 6742 | 23% |
| Required review before before publishing | 25,938 | 89% |
| Affiliation | ||
| - Associated with groups | 15,368 | 52.5% |
| - Associated with individuals | 6125 | 20.9% |
| - Uncategorized | 7802 | 26.6% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trisovic, A.; Mika, K.; Boyd, C.; Feger, S.; Crosas, M. Repository Approaches to Improving the Quality of Shared Data and Code. Data 2021, 6, 15. https://doi.org/10.3390/data6020015
Trisovic A, Mika K, Boyd C, Feger S, Crosas M. Repository Approaches to Improving the Quality of Shared Data and Code. Data. 2021; 6(2):15. https://doi.org/10.3390/data6020015
Chicago/Turabian StyleTrisovic, Ana, Katherine Mika, Ceilyn Boyd, Sebastian Feger, and Mercè Crosas. 2021. "Repository Approaches to Improving the Quality of Shared Data and Code" Data 6, no. 2: 15. https://doi.org/10.3390/data6020015
APA StyleTrisovic, A., Mika, K., Boyd, C., Feger, S., & Crosas, M. (2021). Repository Approaches to Improving the Quality of Shared Data and Code. Data, 6(2), 15. https://doi.org/10.3390/data6020015






