3.1. Monitoring Diseases
Paul and Dredze proposed a new associative topic model for identifying tweets regarding ailments (
Table 1) [
13]. This model, called the Ailment Topic Aspect Model (ATAM), identifies relevant tweets by using a combination of keywords and associated topics. ATAM learns the symptoms and treatments that are associated with specific ailments, and organizes the health terms into ailment groups. It then separates the coherent ailment groups from the more general topics. ATAM identifies latent topic information from a large dataset and enables browsing frequently co-occurring words [
14]. In testing, both ATAM and latent Dirichlet allocation (LDA) methods were applied to the same dataset. Human intelligence was used to review the ATAM and LDA labels for ailment-related tweets. For the LDA method, 45% agreed with the labels; for the ATAM method, 70% agreed with the labels. The ATAM method produces more detailed ailment information through the inclusion of symptoms and treatments as well. The data from this method was compared to influenza-like illness (ILI) data from the Centers for Disease Control and Prevention (CDC). The Pearson’s correlation coefficient between the ATAM frequencies and the CDC data was 0.934 (Google Flu Trends yielded a correlation of 0.932 with the CDC). These results show that the ATAM method is capable of monitoring disease and providing detailed information on occurring ailments.
Gesualdo et al. designed and tested a minimally-trained algorithm for identifying ILI on Twitter [
15]. Using the definition of an ILI case from the European Centre for Disease Prevention and Control, the authors created a Boolean search query for Twitter data. This query identifies all of the tweets reporting a combination of symptoms that satisfies the query. The algorithm learns technical and naïve terms to identify all of the jargon expressions that are related to a specific technical term. It was trained based on pattern generalization using term pairs (one technical and one naïve; e.g., emesis–vomiting). After training, the algorithm was able to extract basic health-related term patterns from the web. The performance of this algorithm was manually evaluated by experts. One hundred tweets satisfying the query were selected along with 500 random symptom-containing tweets. These were evaluated by three of the authors independently, and the overall rate of precision was 0.97. When compared to influenza trends reported by the United States (U.S.) Outpatient ILI Surveillance Network (ILINet), the trends that the query found yielded a correlation coefficient of 0.981. The tweets were also selected for geolocation purposes by identifying those with GPS, time zone, place code, etc. The geolocated tweets were compared to the ILINet data to return a correlation coefficient of 0.980.
Coppersmith, Dredze, and Harman analyzed mental health phenomena on Twitter through simple natural language processing methods [
16]. The focus of their study was on four mental health conditions: (1) post-traumatic stress disorder (PTSD), (2) depression, (3) bipolar disorder, and (4) seasonal affective disorder (SAD). Self-expressions of mental illness diagnosis were used to identify the sample of users for this study. Diagnosis tweets were manually assessed and labeled as genuine or not. Three methods of analysis were conducted. The first was pattern-of-life. This method looked at social engagement and exercise as positive influences and insomnia as a sign of negative outcomes. Sentiment analysis was also used to determine positive or negative outlooks. Pattern-of-life analysis performs especially poorly in detecting depression, but surprisingly, it performs especially well in detecting SAD. Another analysis method utilized was linguistic inquiry word count (LIWC), which is a tool for the psychometric analysis of language data. LIWC is able to provide quantitative data regarding the state of a patient from the patient’s writing. LIWC generally performed on par with pattern-of-life analysis. A third means of analysis was language models (LMs). LMs were used to estimate the likelihood of a given sequence of words. The LMs had superior performance compared to the other analysis methods. The purpose of this study was to generate proof-of-concept results for the quantification of mental health signals through Twitter.
Denecke et al. presented a prototype implementation of a disease surveillance system called M-Eco that processes social media data for relevant disease outbreak information [
17]. The M-Eco system uses a pool of data from Twitter, blogs, forums, television, and radio programs. The data is continuously filtered for keywords. Texts containing keywords are further analyzed to determine their relevance to disease outbreaks, and signals are automatically generated by unexpected behaviors. Signals are only generated when the threshold for the number of texts with the same word or phrase has been exceeded. These signals, which are mainly generated from news agencies’ tweets, are again analyzed for false alarms and visualized through geolocation, tag clouds, and time series. The M-Eco system allows for searching and filtering the signals by various criteria.
3.2. Public Reaction
Adrover et al. [
18] attempted to identify Twitter users who have HIV and determine if drug treatments and their associated sentiments could be detected through Twitter (
Table 2). Beginning with a dataset of approximately 40 million tweets, they used a combination of human and computational approaches, including keyword filtering, crowdsourcing, computational algorithms, and machine learning, to filter the noise from the original data. The narrowed sample consisted of only 5443 tweets. The small sample size and extensive manual hours dedicated to filtering, tagging, and processing the data limited this method. However, the analysis of this data led to the identification of 512 individual users who self-reported HIV and the effects of HIV treatment drugs, as well as a community of 2300 followers with strong, friendly ties. Around 93% of tweets provided information on adverse drug effects. It was found that 238 of the 357 tweets were associated with negative sentiment, with only 78 positive and 37 neutral tweets.
Ginn et al. presented a corpus of 10,822 tweets mentioning adverse drug reactions (ADRs) for training Twitter mining tools [
19]. These tweets were mined from the Twitter application programming interface (API) and manually annotated by experts with medical and biological science backgrounds. The annotation was a two-step process. First, the original corpus of tweets was processed through a binary annotation system to identify mentions of ADRs. ADRs, which are defined as “injuries resulting from medical drug use”, were carefully distinguished from the disease, symptom, or condition that caused the patient to use the drug initially. The Kappa value for binary classification was 0.69. Once the ADR-mentioning tweets were identified, the second step, full annotation, began. The tweets were annotated for identification of the span of expressions regarding ADRs and labeled with the Unified Medical Language System for IDs. The final annotated corpus of tweets was then used to train two different machine learning algorithms: Naïve Bayes and support vector machines (SVMs). Analysis was conducted by observing the frequency and distribution of ADR mentions, the agreement between the two annotators, and the performance of the text-mining classifiers. The performance was modest, setting a baseline for future development.
Sarker and Gonzalez proposed a method of classifying ADRs for public health data by using advanced natural language processing (NLP) techniques [
20]. Three datasets were developed for the task of identifying ADRs from user-posted internet data: one consisted of annotated sentences from medical reports, and the remaining two were built in-house on annotated posts from Twitter and the DailyStrength online health community, respectively. The data from each of the three corpora were combined into a single training set to utilize in machine learning algorithms. The ADR classification performance of the combined dataset was significantly better than the existing benchmarks with an F-score of 0.812 (compared to the previous 0.77). Semantic features such as topics, concepts, sentiments, and polarities were annotated in the dataset as well, providing a basis for the high performance levels of the classifiers.
Behera and Eluri proposed a method of sentiment analysis to monitor the spread of diseases according to location and time [
21]. The goal of their research was to measure the degree of concern in tweets regarding three diseases: malaria, swine flu, and cancer. The tweets were subjected to a two-step sentiment classification process to identify negative personal tweets. The first step of classification consisted of a subjectivity clue-based algorithm to determine which tweets were personal and which were non-personal (e.g., advertisements and news sources) The second step involved applying lexicon-based and Naïve Bayes classifiers to the dataset. These classifiers distinguished negative sentiment from non-negative (positive or neutral) sentiment. To improve the performance of these classifiers, negation handling and Laplacian Smoothing techniques were combined with the algorithms. The best performance came from the combination of Naïve Bayes and negation handling for a precision of 92.56% and an accuracy of 95.67%. After isolating the negative personal tweets, the degree of concern was measured.
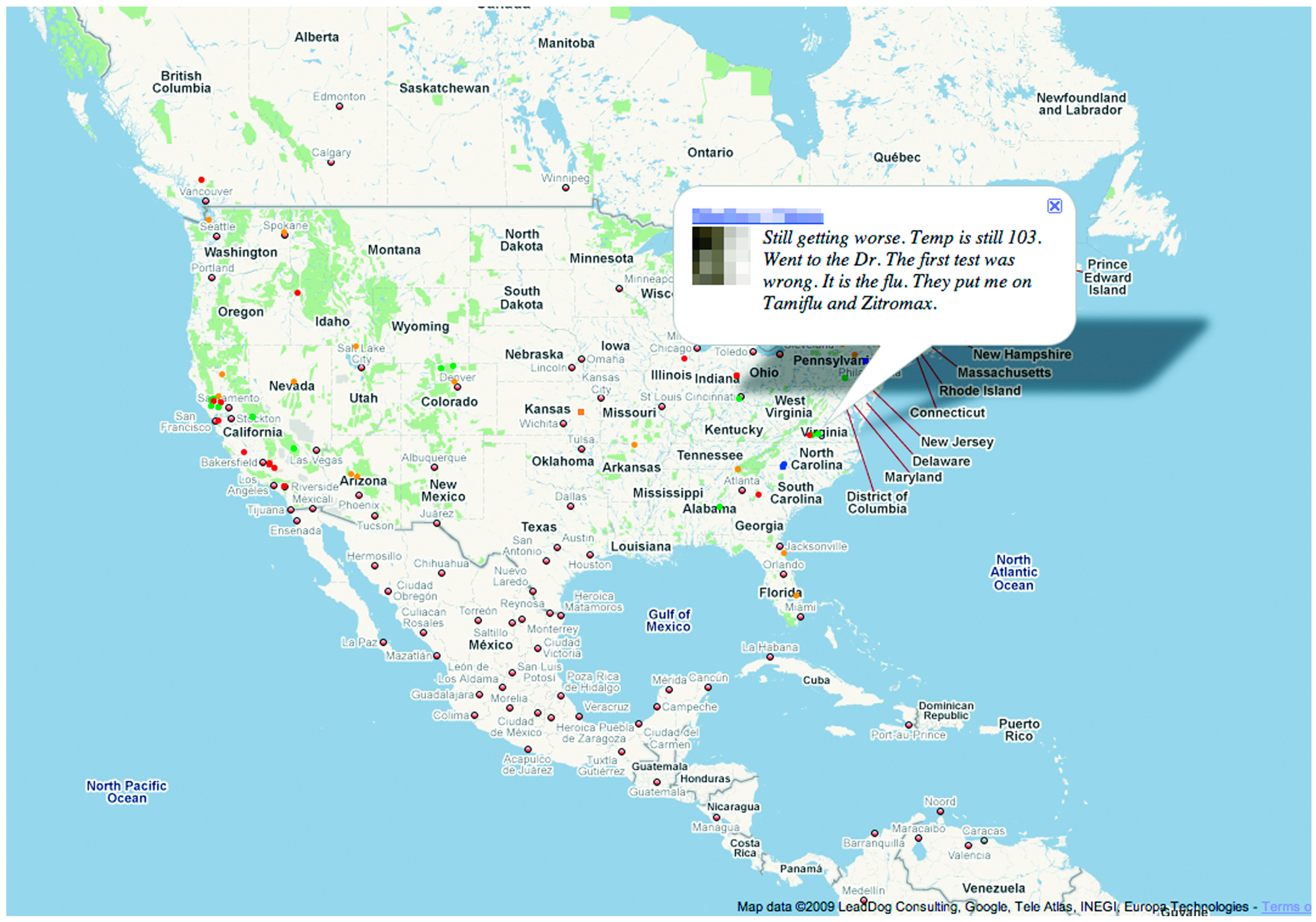
Signorini, Segre, and Polgreen studied the usefulness of Twitter data in tracking the rapidly evolving public sentiment regarding H1N1 influenza and the actual disease activity [
22]. Using keywords to filter the Twitter API and obtain a dataset of over 950,000 tweets, they time-stamped and geolocated each tweet using the author’s self-declared home location. A JavaScript application was developed to display a continuously updating Google map of influenza and H1N1-related tweets according to their geographical context. The tweets and sentiments are depicted as color-coded dots on the map, as shown in
Figure 2. Users can scroll over the dots to read the related tweets (see
Figure 2). Estimates of ILI occurrence rates performed with an average error of 0.28%. When the geolocations of the tweets were factored in, the dataset was reduced due to the rarity of geotagged tweets. The average error for regional ILI estimates was slightly higher at 0.37%. This study demonstrated the concept that Twitter traffic can be used to track public sentiment and concern, and potentially estimate the real-time disease activity of H1N1 and ILIs.
Myslín et al. studied the public sentiment toward tobacco and tobacco-related products through Twitter data [
23]. Tweets were manually classified by two annotators to identify genre, theme, and sentiment. From a cohort of 7362 tweets mined through the Twitter API, 57.3% (4215) were classified as tobacco-related. The tweets were then used to train machine learning classifiers to distinguish between tobacco-related and irrelevant tweets as well as positive, negative, or neutral sentiment in tweets. Three machine learning algorithms were tested in this study: SVM, Naïve Bayes, and K-Nearest Neighbors (KNN). The F-score for discriminating between tobacco-related and irrelevant tweets was 0.85. The SVMs yielded the highest performance. Overall, sentiment toward tobacco was found to be more positive (1939/4215, 46%) than negative (1349/4215, 32%) or neutral (see
Figure 3). These values were found even after the advertising tweets (9%) were excluded. Words relating to hookah or e-cigarettes were highly predictive of positive sentiment, while more general terms related to tobacco were predictive of negative sentiment. This suggests gaps in public knowledge regarding newer tobacco products. This study was limited by the number of keywords that were used to find tobacco-related tweets. While the novelty effects of hookah and e-cigarettes were not considered in the analysis, this work demonstrated the capabilities of machine learning classifiers trained on Twitter data to determine public sentiment and identify areas to direct public health information dissemination.
Ji et al. used Twitter to track the spread of public concern regarding epidemics [
24]. Their methods included separating tweets into personal and news (non-personal) categories to focus on public concern. The personal tweets were further classified into personal negative and personal non-negative, depending on the sentiment detected. Training data auto-generated from an emotion-oriented, clue-based method was used to train and test three different machine learning models. The tweets that were classified as personal negative were used to generate a Measure of Concern (MOC) and format the MOC to a timeline. The MOC timeline was compared to a news timeline. The peaks were compared to find a Jaccard correlation coefficient with a range of 0.2–0.3. These results are insufficient for predictions. However, some MOC peaks aligned with news peaks on the timeline, suggesting that the general public expresses negative emotions when news activity increases.
Colleta et al. studied the public sentiment classification of tweets using a combination of SVM and cluster ensemble techniques [
25]. This algorithm, named the C
3E-SL, is capable of combining classifiers with cluster ensembles to refine tweet classifications from additional information provided by the clusters. Four different categories of tweets were used to train and test the C
3E-SL algorithm. The first set consisted of 621 training tweets (215 positive and 406 negative) related to the topic of health care reform. The second set, the Obama–McCain debate, was made up of 3238 tweets. Neutral tweets were removed, leaving only 1906 (710 positive and 1196 negative) to be used for training. The third set contained 1224 tweets (570 positive and 654 negative) related to Apple, Google, Microsoft, and Twitter. The final set consisted of 359 manually annotated tweets (182 positive and 177 negative) from a study completed at Stanford [
26]. The results demonstrated that the C
3E-SL algorithm performed better than the SVM classifier alone and was competitive with the highest performances found in the literature.
3.3. Outbreak and Emergency
France and Christopher Cheong used Twitter to conduct a social network analysis case study for the floods of Queensland, New South Wales, and Victoria, Australia, from March 2010 to February 2011 (
Table 3) [
27]. The research goal was to identify the main active users during these events, and determine their effectiveness in disseminating critical information regarding the crisis. Two types of networks were generated for each of the three flood-affected sites: a “user” network based on the responses of users to certain tweets, and a “user-resources” network connecting user tweets to the included resource links. The most active users were found to be local authorities, political personalities, social media volunteers, traditional media reporters, and nonprofit, humanitarian, and community organizations.
Odlum and Yoon collected over 42,000 tweets related to Ebola during the outbreak in summer 2014 [
28]. This Twitter data was analyzed to monitor the trends of information spread, examine early epidemic detection, and determine public knowledge and attitudes regarding Ebola. Throughout the summer, a gradual increase was detected in the rate of information dissemination. An increase in Ebola-related Twitter activity occurred in the days prior to the official news alert. This increase is indicative of Twitter’s potential in supporting early warning systems in the outbreak surveillance effort. The four main topics found in Ebola-related tweets during the epidemic were risk factors, prevention education, disease trends, and compassion toward affected countries and citizens. The public concern regarding Ebola nearly doubled on the day after the CDC’s health advisory.
Missier et al. studied the performance of two different approaches to detecting Twitter data relevant to dengue and other
Aedes-borne disease outbreaks in Brazil [
29]; both supervised classification and unsupervised clustering using topic modeling performed well. The supervised classifier identified four different classes of topics: (1) mosquito focus was the most directly actionable class; (2) sickness was the most informative class; (3) news consisted of indirectly actionable information; and (4) jokes made up approximately 20% of the tweets studied, and were regarded as noise. It was difficult to distinguish jokes from relevant tweets due to the prevalence of common words and topics. A training set of 1000 tweets was manually annotated and used to train the classifier. Another set of 1600 tweets was used to test the classifier, and resulted in an accuracy range of 74–86% depending on the class. Over 100,000 tweets were harvested for the LDA-based clustering. A range of two to eight clusters were formed, and interclustering and intraclustering were calculated to determine the level of distinction between clusters. The intraclustering was found to be over double that of interclustering, indicating that the clusters were well separated. Overall, clustering using topic modeling was found to offer less control over the content of the topics than a traditional classifier. However, the classifier required a lot of manual annotations, and was thus costlier than the clustering method.
Schulz et al. presented an analysis of a multi-label learning method for classification of incident-related Twitter data [
30]. Tweets were processed using three different methods (binary relevance, classifier chains, and label powerset) to identify four labels: (S) Shooting, (F) Fire, (C) Crash, and (I) Injury. Each approach was analyzed for precision, recall, exact match, and h-loss. Keyword-based filtering yielded poor results in each evaluation category, indicating that it is inadequate for multi-label classification. It was found that the correlation between labels needs to be taken into account for classification. The classifier chains method is able to outperform the other methods if a cross-validation is performed on the training data. Overall, it was found that multiple labels were able to be detected with an exact match of 84.35%.
Gomide et al. proposed a four-dimensional active surveillance methodology for tracking dengue epidemics in Brazil using Twitter [
31]. The four dimensions were volume (the number of tweets mentioning “dengue”), time (when these tweets were posted), location (the geographic information of the tweets), and public perception (overall sentiment toward dengue epidemics). The number of dengue-related tweets was compared to official statistics from the same time period obtained from the Brazilian Health Ministry, and an R2 value of 0.9578 was obtained. The time and location information were combined to predict areas of outbreak. A clustering approach was used to find cities in close proximity to each other with similar dengue incidence rates at the same time. The Rand index value was found to be 0.8914.
3.4. Prediction
Santos and Matos investigated the use of tweets and search engine queries to estimate the incidence rate of influenza (
Table 4) [
32]. In this study, tweets regarding ILI were manually classified as positive or negative according to whether the message indicated that the author had the flu. These tweets were then used to train machine learning models to make the positive or negative classification for the entire set of 14 million tweets. After classification, the Twitter-generated influenza incidence rate was compared to epidemiological results from Influenzanet, which is a European-wide network for flu surveillance. In addition to the Twitter data, 15 million search queries from the SAPO ((Online Portuguese Links Server)) search platform were included in the analysis. A linear regression model was applied to the predicted influenza trend and the Influenzanet data to result in a correlation value of approximately 0.85.
To test the accuracy of the models in predicting influenza incidence from one flu season to the next, more linear regression models were implemented. The data generated was then compared to the weekly incidence rate reported by the European Influenza Surveillance Network (EISN). The predicted trend appeared to be a week ahead of the EISN report. Interestingly, in this comparison, the flu trend was overestimated by the model in week nine. The EINS did not show the exaggerated rate of influenza; however, media reports and the National Institute of Health demonstrate a high incidence rate in Portugal at the time. This study demonstrated the ability of the models to correlate as well as 0.89 to Influenzanet and across seasons, with a Pearson correlation coefficient (r) value of 0.72.
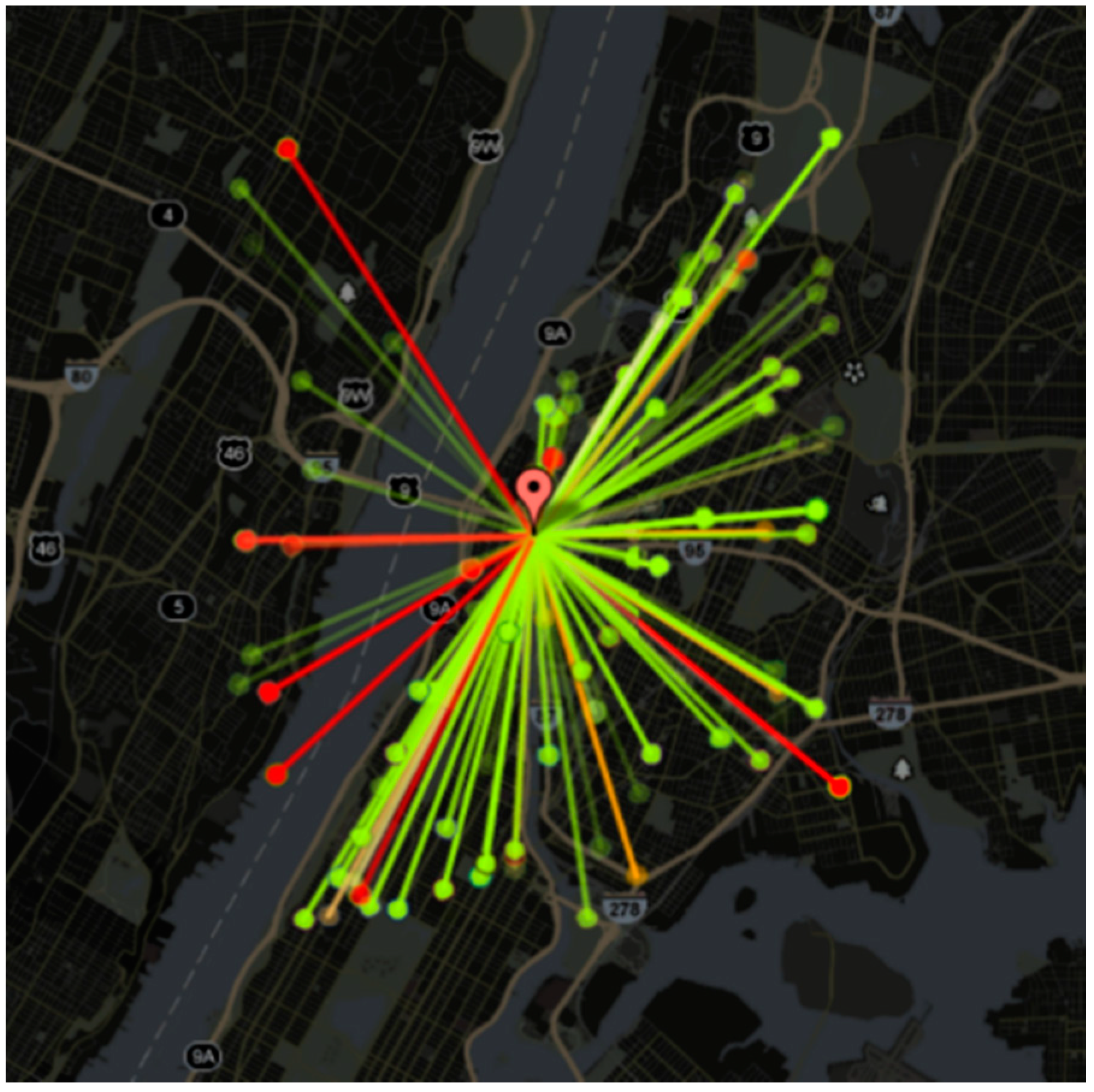
Kautz and Sadilek proposed a model to predict the future health status (“sick” or “healthy”) of an individual with accuracy up to 91% [
33]. This study was conducted using 16 million tweets from one month of collection in New York City. Users who posted more than 100 GPS-tagged tweets in the collection month (totaling 6237 individual users) were investigated by data mining regarding their online communication, open accounts, and geolocated activities to describe the individual’s behavior. Specifically, the locations, environment, and social interactions of the users were identified. Locations were determined through GPS monitoring, and used to count visits to different ‘venues’ (bars, gyms, public transportation, etc.), physical encounters with sick individuals (defined as co-located within 100 m), and the ZIP code of the individual (found by analyzing the mean location of a user between the hours of 01:00–06:00). The environment of the user was also determined through GPS, as well as the relative distance of the user to pollution sources (factories, power plants, transportation hubs, etc.). The social interactions of a user were determined through their online communication (
Figure 4). Social status was analyzed using the number of reciprocated ‘follows’ on Twitter, mentions of the individual’s name, number of ‘likes’ and retweets, and through the PageRank calculation. Applying machine learning techniques to mined data, researchers were able to find the feature that was most strongly correlated with poor health: the proximity to pollution sources. Higher social status was strongly correlated with better health, while visits to public parks was also positively correlated with improved health. Overall, the model explained more than 54% of the variance in people’s health.
The methods used in this study infer “sick” versus “healthy” from brief messages, leaving room for misinterpretation. The visits to certain venues and interactions with sick individuals may be false positives. In addition, some illness may be overreported or underreported via social media. Thus, controlling for misrepresentations of the occurrence of illnesses must be improved through cross-referencing social media reports with other sources of data.
3.5. Public Lifestyle
Pennacchiotti and Popescu proposed a system for user classification in social media (
Table 5) [
34]. This team focused on classifying users according to three criteria: political affiliation (Democrat or Republican), race (African American or other, in this case), and potential as a customer for a particular business (Starbucks). Their machine learning framework relied on data from user profile accounts, user tweeting behavior (i.e., number of tweets per day, number of replies, etc.), linguistic content (main topics and lexical usage), and the social network of the user. The combination of all the features is more successful in classifying users than any individual feature. This framework was most successful in identifying the political affiliation of users. The features that were most accurate for this task were the social network and followers of the user, followed by the linguistic and profile features. The most difficult category was race, with values near 0.6–0.7. Linguistic features were most accurate for this task.
Prier et al. proposed the use of LDA for topic modeling Twitter data [
14]. LDA was used to analyze terms and topics from a dataset of over two million tweets. The topic model identified a series of conversational topics related to public health, including physical activity, obesity, substance abuse, and healthcare. Unfortunately, the LDA method of analysis was unable to detect less common topics, such as the targeted topic of tobacco use. Instead, the researchers built their own query list by which to find tweets. The query list included terms such as “tobacco”, “smoking”, “cigarette”, “cigar”, and “hookah”. By topic modeling this tobacco data subset, they were able to gain understanding of how Twitter users are discussing tobacco usage.
3.6. Geolocation
Dredze et al. introduced a system to determine the geographic location of tweets through the analysis of “Place” tags, GPS positions, and user profile data (
Table 6) [
35]. The purpose of the proposed system, called Carmen, was to assign a location to each tweet from a database of structured location information. “Place” tags on tweets associate a location with the message. These tags may include information such as the country, city, geographical coordinates, business name, or street address. Other tweets are GPS-tagged, and include the latitude and longitude coordinates of the location. The user profile contains a field where the user can announce their primary location. However, the profiles are subject to false information or nonsensical entries (i.e., “Candy Land”). User profiles are insufficient in accounting for travel as well. Carmen uses a combination of factors to infer the origin of the tweet. This system analyzes the language of the tweet, the “Place” and GPS tags, and the profile of the user. This information can provide the country, state, county, and city from which the tweet originated. Health officials may utilize Carmen’s geolocation to track the occurrence of disease rates and prevent and manage outbreaks. Traditional systems rely on patient clinical visits, which take up to two weeks to publish. However, with this system, officials can use Twitter to find the possible areas of outbreaks in real time, improving reaction time.
Yepes et al. proposed a method for analyzing Twitter data for health-related surveillance [
36]. To conduct their analysis, this group obtained 12 billion raw tweets from 2014. These tweets were filtered to include tweets only in the English language and excluded all retweets. Prior to filtering, heuristics were applied to the dataset. An in-domain medical named entity recognizer, called Micromed, was used to identify all of the relevant tweets. Micromed uses supervised learning, having been trained on 1300 manually annotated tweets. This system was able to recognize three medical entities: diseases, symptoms, and pharmacological substances. After filtering the tweets, MALLET (machine learning for language toolkit) was used to group the tweets by topic. An adapted geotagging system (LIW-meta) was used to determine geographic information from the posts. LIW-meta uses a combination of explicit location terms, implicit location-indicative words (LIW), and user profile data to infer geolocations from the tweets that lack GPS labels. The results of their work yielded geotagging with 0.938 precision. Yepes also observed that tweets mentioning terms such as “heart attack” are frequently used in the figurative sense more than in the medical sense when posting on social media. Other figurative usage of terms includes the use of “tired” to mean bored or impatient rather than drowsiness as a symptom. However, the usage of some pharmacological substance words, such as “marijuana” and “caffeine” are more likely to be indicative of the frequency of people using these substances.
Prieto et al. proposed an automated method for measuring the incidence of certain health conditions by obtaining Twitter data that was relevant to the presence of the conditions [
37]. A two-step process was used to obtain the tweets. First, the data was defined and filtered according to specially crafted regular expressions. Secondly, the tweets were manually labeled as positive or negative for training classifiers to recognize the four health states. The health conditions that were studied were influenza, depression, pregnancy, and eating disorders. To begin the filtering, tweets originating in Portugal and Spain were selected using Twitter search API and geocoding information from the Twitter metadata. A language detection library was used to filter tweets that were not in Portuguese or Spanish. Once the tweets of the correct origin and language were identified, machine learning was applied to the data in order to filter out tweets that were not indicative of the person having the health condition. Finally, feature selection was applied to the data. Classification results of 0.7–0.9 in the area under the receiver operating characteristic (ROC) curve (AUC) and F-measure were obtained. The number of features was reduced by 90% by feature selection algorithms such as correlation-based feature selection (CFS), Pearson correlation, Gain Ration, and Relief. Classification results were improved with the feature selection algorithms by 18% in AUC and 7% in F-measure.
3.7. General
Tuarob et al. proposed a combination of five heterogeneous base classifiers to address the limitations of the traditional bag-of-words approach to discover health-related information in social media (
Table 7) [
38]. The five classifiers that were used were random forest, SVM, repeated incremental pruning to produce error reduction, Bernoulli Naïve Bayes, and multinomial Naïve Bayes. Over 5000 hand-labeled tweets were used to train the classifiers and cross-validate the models. A small-scale and a large-scale evaluation were performed to investigate the proposed model’s abilities. The small-scale evaluation used a 10-fold cross-validation to tune the parameters of the proposed model and compare it with the state-of-the-art method. The proposed model outperformed the traditional method by 18.61%. The large-scale evaluation tested the trained classifiers on real-world data to verify the ability of the proposed model. This evaluation demonstrated a performance improvement of 46.62%.
Sriram developed a new method of classifying Twitter messages using a small set of authorship features that were included to improve the accuracy [
39]. Tweets were classified into one of five categories focused on user intentions: news, events, opinions, deals, and private messages. The features extracted from the author’s profile and the text were used to classify the tweets through three different classifiers. The fourth classifier, bag-of-words (BOW), was used to process tweets without the authorship features. It was considered a baseline because of its popularity in text classification. Compared to the BOW approach, each classifier that used the authorship features had significantly improved accuracy and processing time. The greatest number of misclassified tweets was found between News and Opinions categories.
Lee et al. proposed a method of classification of tweets based on Twitter Trending Topics [
40]. Tweets were analyzed using text-based classification and network-based classification to fit into one of 18 categories such as sports, politics, technology, etc. For text-based classification, the BOW approach was implemented. In network-based classification, the top five similar topics for a given topic were identified through the number of common influential users. Each tweet could only be designated as falling into one category, which led to increased errors.
Parker et al. proposed a framework for tracking public health conditions and concerns via Twitter [
41]. This framework uses frequent term sets from health-related tweets, which were filtered according to over 20,000 keywords or phrases, as search queries for open-source resources such as Wikipedia, Mahout, and Lucene. The retrieval of medical-related articles was considered an indicator of a health-related condition. The fluctuating frequent term sets were monitored over time to detect shifts in public health conditions and concern. This method was found to identify seasonal afflictions. However, no quantitative data was reported.