A Cardiac Cell Outgrowth Assay for Evaluating Drug Compounds Using a Cardiac Spheroid-on-a-Chip Device
Abstract
1. Introduction
2. Materials and Methods
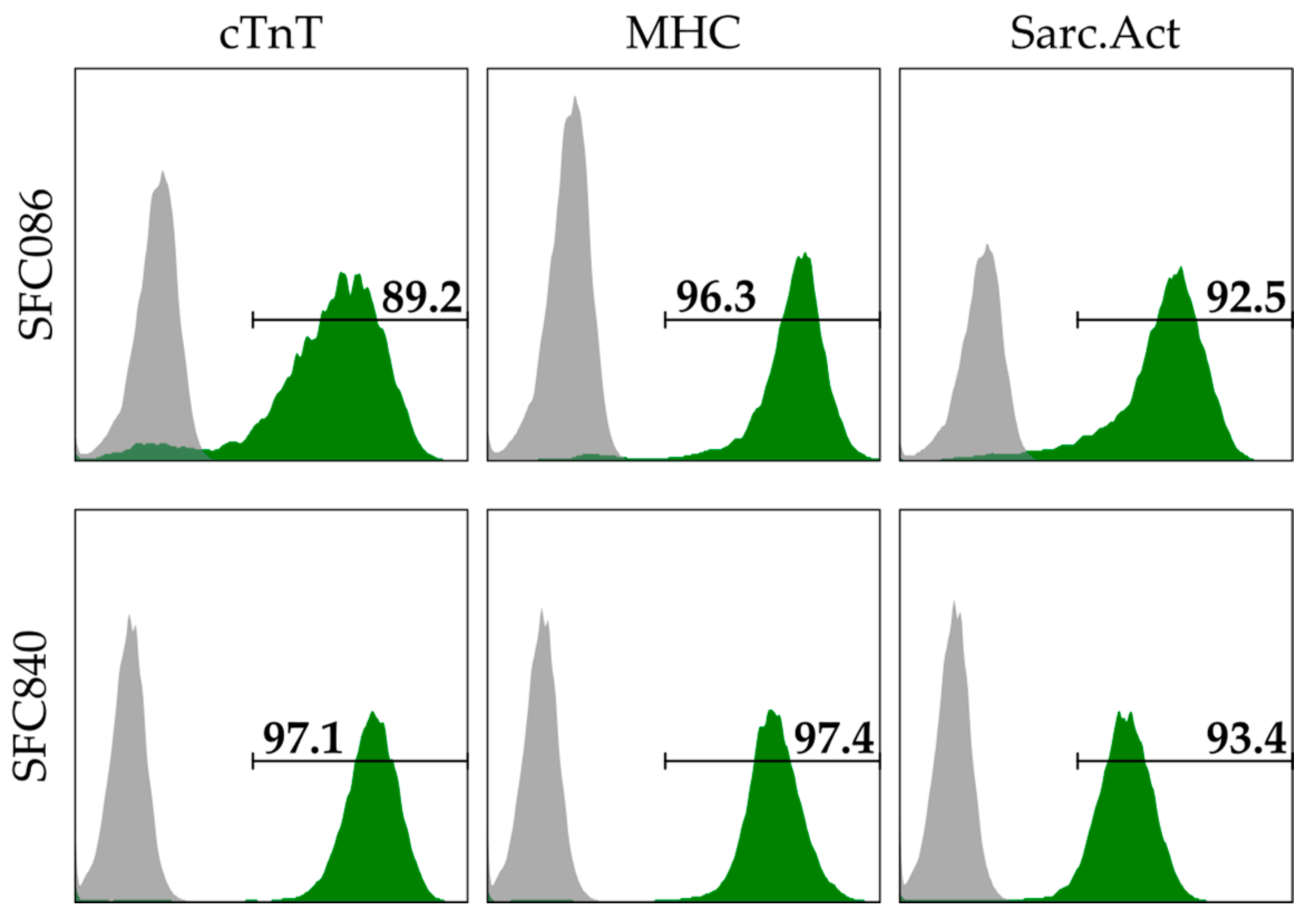
2.1. Cell Lines and Preparation of Cardiac Spheroids
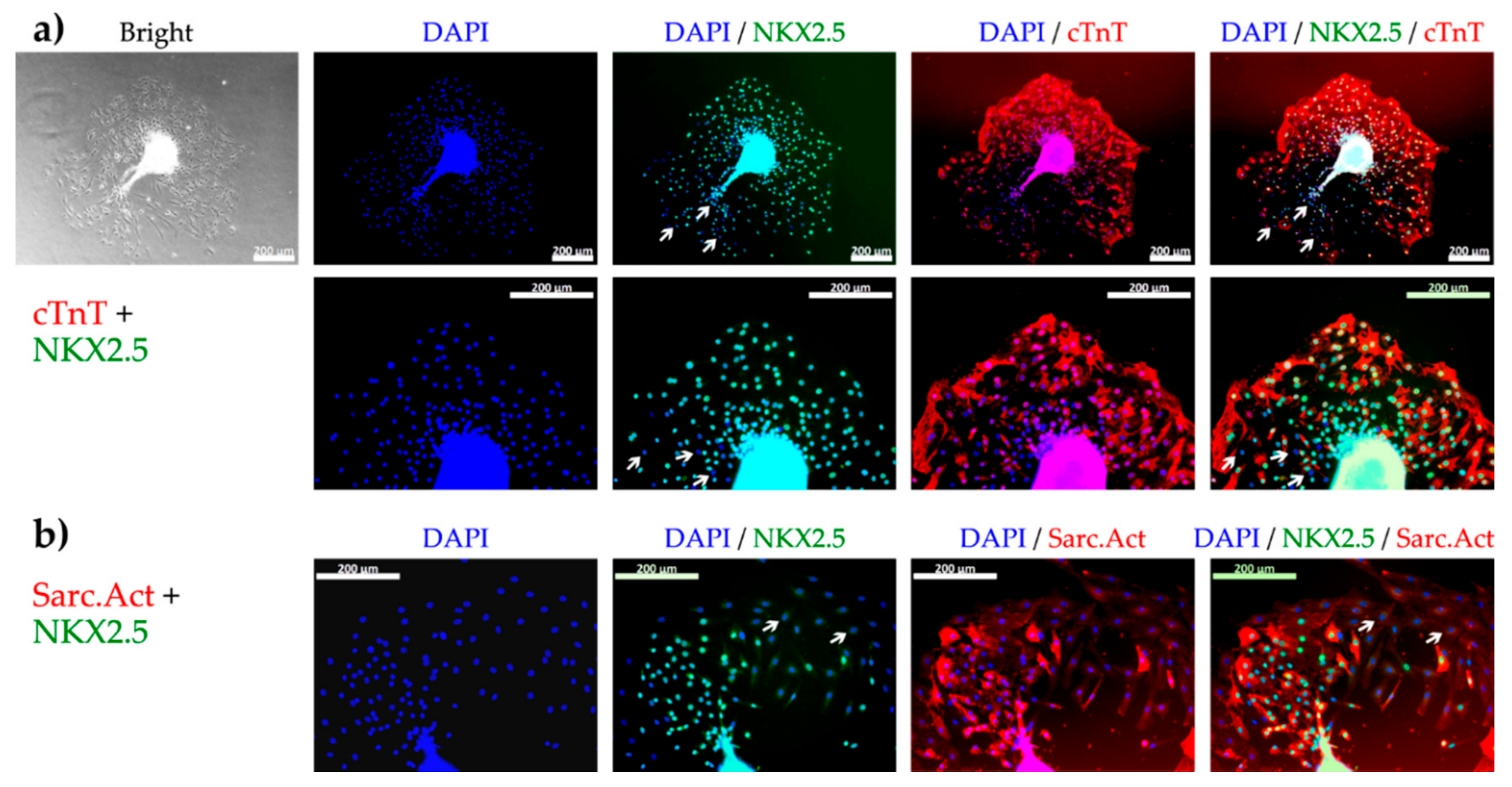
2.2. Flow Cytometry and Immunofluorescent Staining
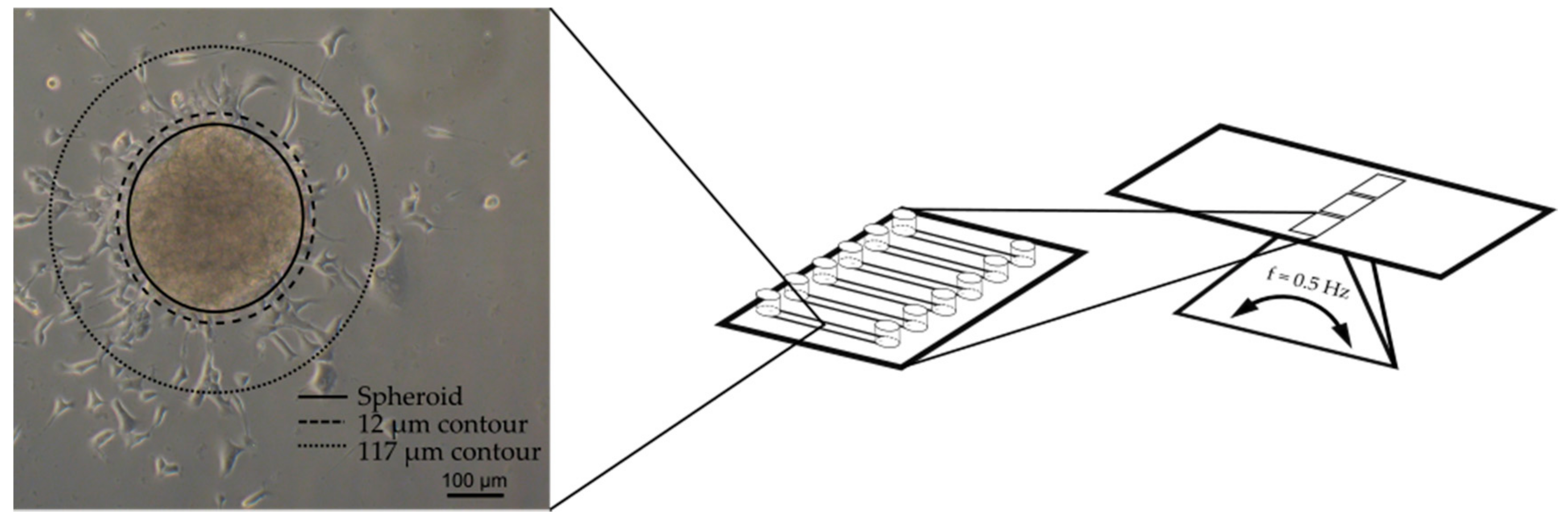
2.3. Microfluidic System and Experimental Procedure for Compound Testing
2.4. Image Processing and Nuclei Quantification
3. Results
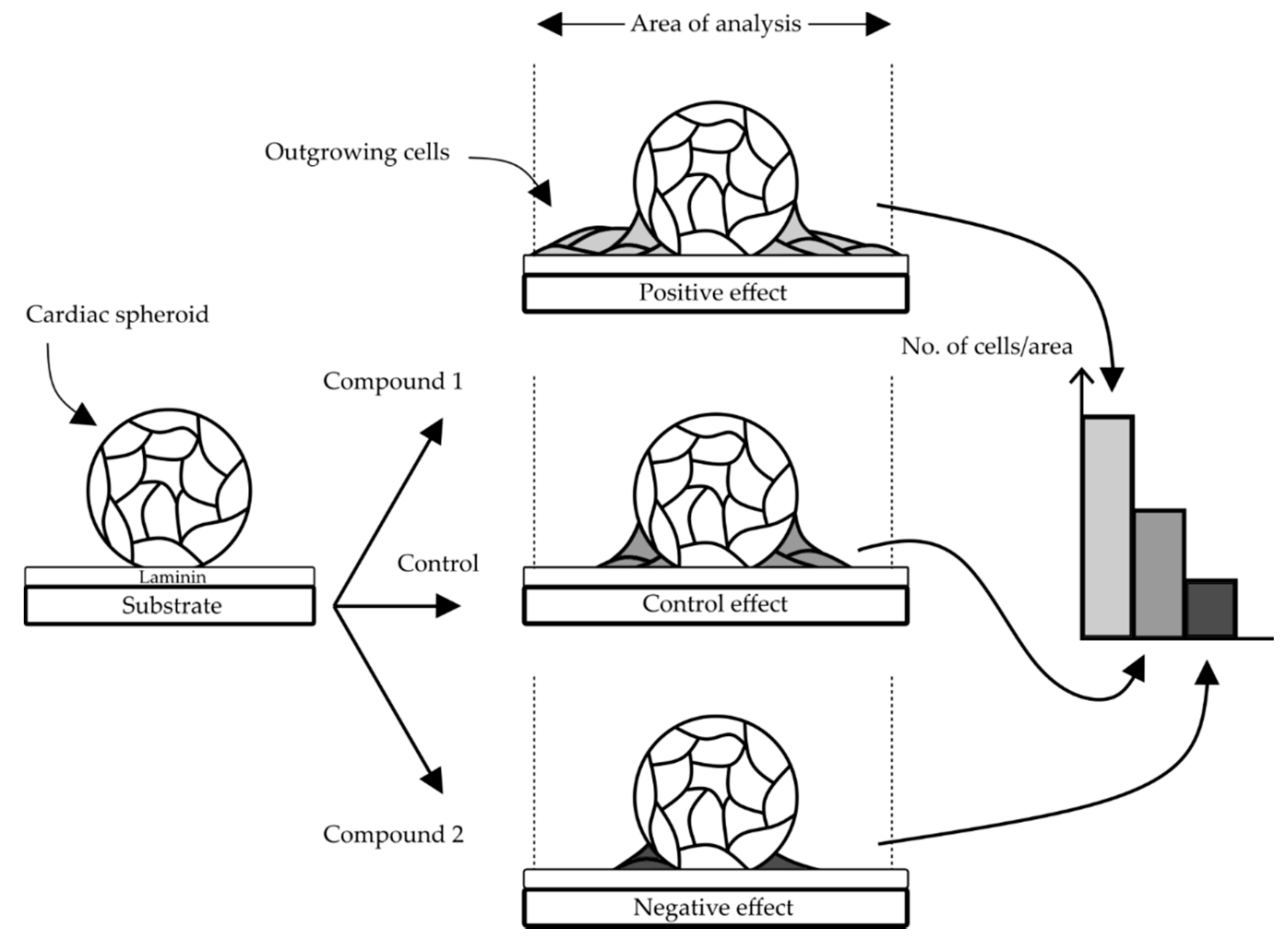
3.1. Design of the Cell Outgrowth Assay
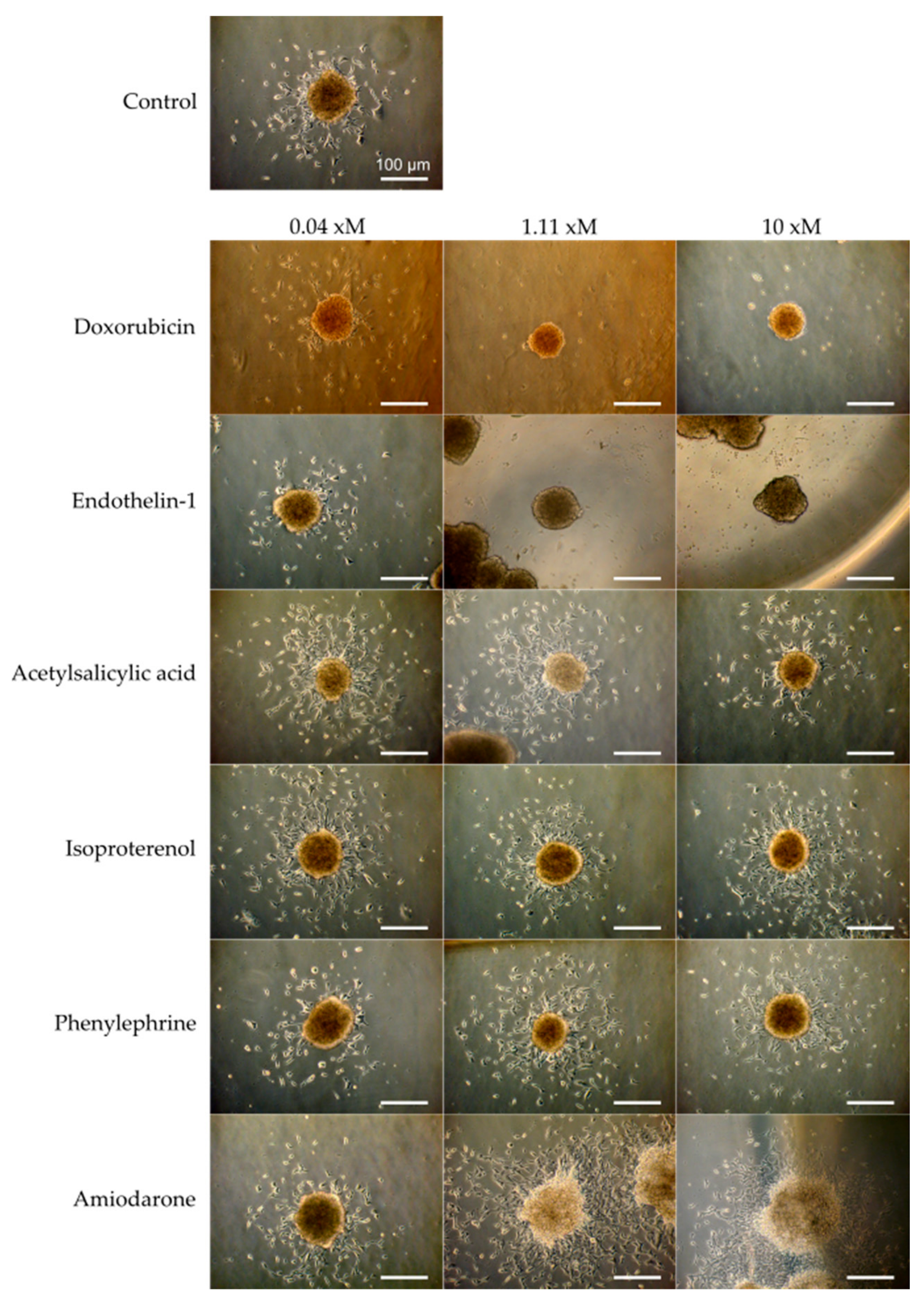
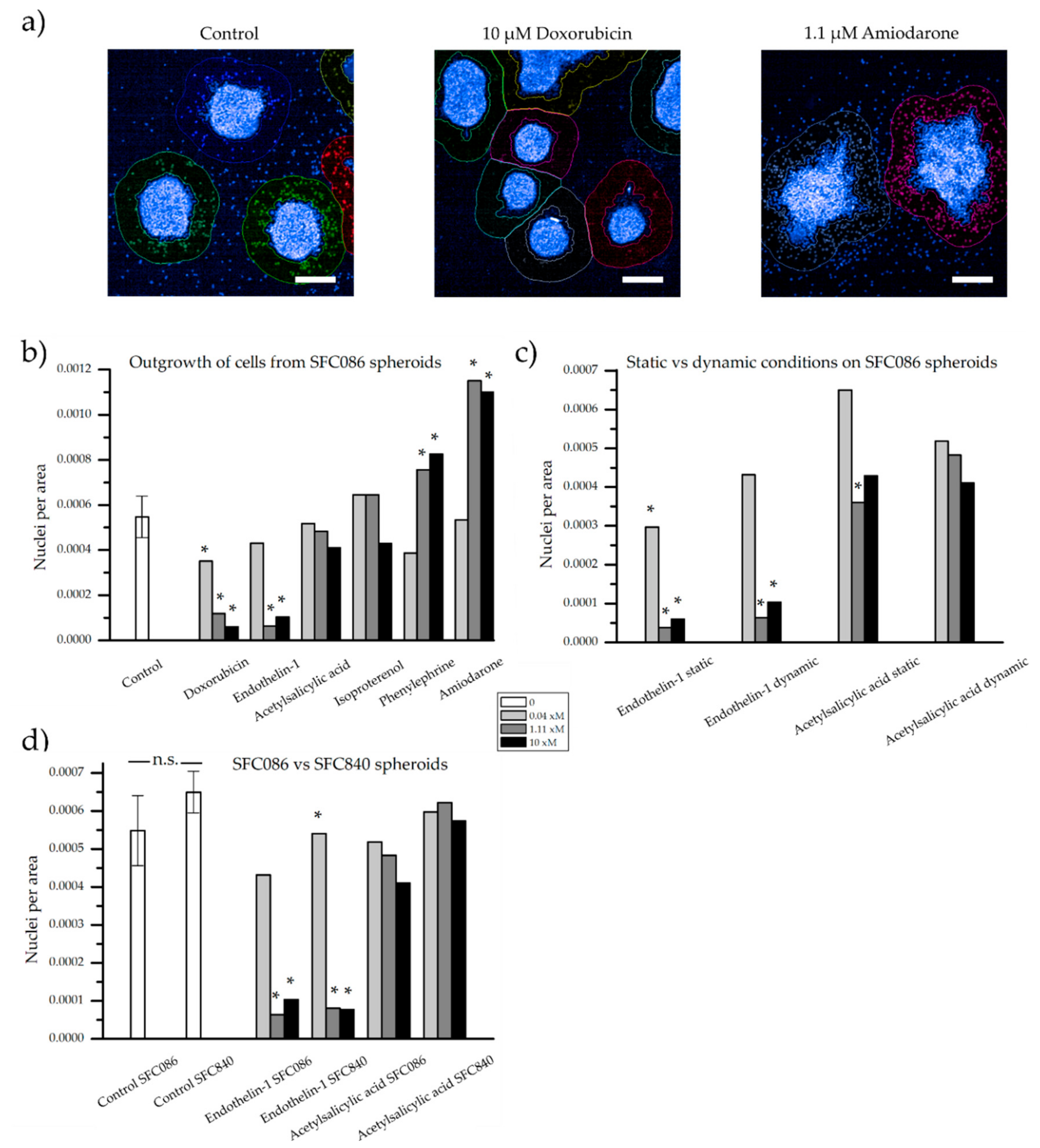
3.2. Verification of the Assay Performance with Six Compounds
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Bhatia, S.N.; Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 2014, 32, 760–772. [Google Scholar] [CrossRef] [PubMed]
- Sung, J.H.; Esch, M.B.; Prot, J.-M.; Long, C.J.; Smith, A.; Hickman, J.J.; Shuler, M.L. Microfabricated mammalian organ systems and their integration into models of whole animals and humans. Lab Chip 2013, 13, 1201–1212. [Google Scholar] [CrossRef] [PubMed]
- Prodanov, L.; Jindal, R.; Bale, S.; Hegde, M.; McCarty, W.J.; Golberg, I.; Bhushan, A.; Yarmush, M.L.; Usta, O.B. Long-term maintenance of a microfluidic 3D human liver sinusoid. Biotechnol. Bioeng. 2016, 113, 241–246. [Google Scholar] [CrossRef] [PubMed]
- Lanz, H.L.; Saleh, A.; Kramer, B.; Cains, J.; Ng, C.P.; Trietsch, S.J.; Hankemeier, T.; Joore, J.; Vulto, P.; Weinshilboum, R.; et al. Therapy response testing of breast cancer in a 3D high-throughput perfused microfluidic platform. BMC Cancer 2017, 17, 709. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, E.; Patch, C.; Aigner, S.; Graf, M.; Iacone, R.; Freskgård, P.O. Multicellular self-assembled spheroidal model of the blood brain barrier. Sci. Rep. 2013, 3, 1500. [Google Scholar] [CrossRef] [PubMed]
- Bell, C.C.; Lauschke, V.M.; Vorrink, S.U.; Palmgren, H.; Duffin, R.; Andersson, T.B.; Ingelman-Sundberg, M. Transcriptional, functional, and mechanistic comparisons of stem cell-derived hepatocytes, HepaRG cells, and three-dimensional human hepatocyte spheroids as predictive in vitro systems for drug-induced liver injury. Drug Metab. Dispos. 2017, 45, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Mueller, D.; Krämer, L.; Hoffmann, E.; Klein, S.; Noor, F. 3D organotypic HepaRG cultures as in vitro model for acute and repeated dose toxicity studies. Toxicol. In Vitro 2014, 28, 104–112. [Google Scholar] [CrossRef] [PubMed]
- Andersson, H.; Steel, D.; Asp, J.; Dahlenborg, K.; Jonsson, M.; Jeppsson, A.; Lindahl, A.; Kågedal, B.; Sartipy, P.; Mandenius, C.-F. Assaying cardiac biomarkers for toxicity testing using biosensing and cardiomyocytes derived from human embryonic stem cells. J. Biotechnol. 2010, 150, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Bistola, V.; Nikolopoulou, M.; Derventzi, A.; Kataki, A.; Sfyras, N.; Nikou, N.; Toutouza, M.; Toutouzas, P.; Stefanadis, C.; Konstadoulakis, M.M. Long-term primary cultures of human adult atrial cardiac myocytes: Cell viability, structural properties and bnp secretion in vitro. Int. J. Cardiol. 2008, 131, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, A.; Stancescu, M.; Dhir, V.; Armstrong, C.; Sommerhage, F.; Hickman, J.J.; Molnar, P. Patterned cardiomyocytes on microelectrode arrays as a functional, high information content drug screening platform. Biomaterials 2011, 32, 4267–4274. [Google Scholar] [CrossRef] [PubMed]
- Bergström, G.; Christoffersson, J.; Schwanke, K.; Zweigerdt, R.; Mandenius, C.-F. Stem cell derived in vivo-like human cardiac bodies in a microfluidic device for toxicity testing by beating frequency imaging. Lab Chip 2015, 15, 3242–3249. [Google Scholar] [CrossRef] [PubMed]
- Frimat, J.-P.; Sisnaiske, J.; Subbiah, S.; Menne, H.; Godoy, P.; Lampen, P.; Leist, M.; Franzke, J.; Hengstler, J.G.; van Thriel, C.; et al. The network formation assay: A spatially standardized neurite outgrowth analytical display for neurotoxicity screening. Lab Chip 2010, 10, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Van der Meer, A.D.; Vermeul, K.; Poot, A.A.; Feijen, J.; Vermes, I. A microfluidic wound-healing assay for quantifying endothelial cell migration. Am. J. Physiol. Heart Circ. Physiol. 2009, 298, H719–H725. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.R.; Beh, S.-J.; Bryson-Richardson, R.J.; Doran, P.M. Production of zebrafish cardiospheres and cardiac progenitor cells in vitro and three-dimensional culture of adult zebrafish cardiac tissue in scaffolds. Biotechnol. Bioeng. 2017, 114, 2142–2148. [Google Scholar] [CrossRef] [PubMed]
- Jang, K.-J.; Mehr, A.P.; Hamilton, G.A.; McPartlin, L.A.; Chung, S.; Suh, K.-Y.; Ingber, D.E. Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr. Biol. 2013, 5, 1119–1129. [Google Scholar] [CrossRef] [PubMed]
- Booth, R.; Kim, H. Characterization of a microfluidic in vitro model of the blood-brain barrier (µBBB). Lab Chip 2012, 12, 1784–1792. [Google Scholar] [CrossRef] [PubMed]
- Esch, M.B.; Prot, J.M.; Wang, Y.I.; Miller, P.; Llamas-Vidales, J.R.; Naughton, B.A.; Applegate, D.R.; Shuler, M.L. Multi-cellular 3D human primary liver cell culture elevates metabolic activity under fluidic flow. Lab Chip 2015, 15, 2269–2277. [Google Scholar] [CrossRef] [PubMed]
- Jackman, C.P.; Carlson, A.L.; Bursac, N. Dynamic culture yields engineered myocardium with near-adult functional output. Biomaterials 2016, 111, 66–79. [Google Scholar] [CrossRef] [PubMed]
- Morrison, M.; Klein, C.; Clemann, N.; Collier, D.A.; Hardy, J.; Heiβerer, B.; Cader, M.Z.; Graf, M.; Kaye, J. Stembancc: Governing access to material and data in a large stem cell research consortium. Stem Cell Rev. Rep. 2015, 11, 681–687. [Google Scholar] [CrossRef] [PubMed]
- Kempf, H.; Olmer, R.; Kropp, C.; Ruckert, M.; Jara-Avaca, M.; Robles-Diaz, D.; Franke, A.; Elliott, D.A.; Wojciechowski, D.; Fischer, M.; et al. Controlling expansion and cardiomyogenic differentiation of human pluripotent stem cells in scalable suspension culture. Stem Cell Rep. 2014, 3, 1132–1146. [Google Scholar] [CrossRef] [PubMed]
- Kempf, H.; Kropp, C.; Olmer, R.; Martin, U.; Zweigerdt, R. Cardiac differentiation of human pluripotent stem cells in scalable suspension culture. Nat. Protoc. 2015, 10, 1345–1361. [Google Scholar] [CrossRef] [PubMed]
- Kempf, H.; Olmer, R.; Haase, A.; Franke, A.; Bolesani, E.; Schwanke, K.; Robles-Diaz, D.; Coffee, M.; Göhring, G.; Dräger, G.; et al. Bulk cell density and Wnt/TGFbeta signalling regulate mesendodermal patterning of human pluripotent stem cells. Nat. Commun. 2016, 7, 13602. [Google Scholar] [CrossRef] [PubMed]
- Maillet, A.; Tan, K.; Chai, X.; Sadananda, S.N.; Mehta, A.; Ooi, J.; Hayden, M.R.; Pouladi, M.A.; Ghosh, S.; Shim, W.; et al. Modeling doxorubicin-induced cardiotoxicity in human pluripotent stem cell derived-cardiomyocytes. Sci. Rep. 2016, 6, 25333. [Google Scholar] [CrossRef] [PubMed]
- Carlson, C.; Koonce, C.; Aoyama, N.; Einhorn, S.; Fiene, S.; Arne, T.; Swanson, B.; Anson, B.; Kattman, S. Phenotypic Screening with Human iPS Cell–Derived Cardiomyocytes: HTS-Compatible Assays for Interrogating Cardiac Hypertrophy. J. Biomol. Screen. 2013, 18, 1203–1211. [Google Scholar] [CrossRef] [PubMed]
- Vane, J.R.; Botting, R.M. The mechanism of action of aspirin. Thromb. Res. 2003, 110, 255–258. [Google Scholar] [CrossRef]
- Schäfer, M.; Frischkopf, K.; Taimor, G.; Piper, H.M.; Schlüter, K.-D. Hypertrophic effect of selective β1-adrenoceptor stimulation on ventricular cardiomyocytes from adult rat. Am. J. Physiol.-Cell Physiol. 2000, 279, C495–C503. [Google Scholar] [CrossRef] [PubMed]
- Ogoh, S.; Sato, K.; Fisher, J.P.; Seifert, T.; Overgaard, M.; Secher, N.H. The effect of phenylephrine on arterial and venous cerebral blood flow in healthy subjects. Clin. Physiol. Funct. Imaging 2011, 31, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Lalevée, N.; Barrère-lemaire, S.; Gautier, P.; Nargeot, J.; Richard, S. Effects of amiodarone and dronedarone on voltage-dependent sodium current in human cardiomyocytes. J. Cardiovasc. Electrophysiol. 2003, 14, 885–890. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, A.; Yuasa, S.; Mearini, G.; Egashira, T.; Seki, T.; Kodaira, M.; Kusumoto, D.; Kuroda, Y.; Okata, S.; Suzuki, T.; et al. Endothelin-1 induces myofibrillar disarray and contractile vector variability in hypertrophic cardiomyopathy–induced pluripotent stem cell–derived cardiomyocytes. J. Am. Heart Assoc. 2014, 3, e001263. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Miyauchi, T. A novel pharmacological action of ET-1 to prevent the cytotoxicity of doxorubicin in cardiomyocytes. Am. J. Physiol.-Regul. Integr. Comp. Physiol. 2001, 280, R1399–R1406. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.M.; Rovin, J.D.; Parsons, J.T. A role for focal adhesion kinase in phenylephrine-induced hypertrophy of rat ventricular cardiomyocytes. J. Biol. Chem. 2000, 275, 19250–19257. [Google Scholar] [CrossRef] [PubMed]
- Dempsey, G.T.; Chaudhary, K.W.; Atwater, N.; Nguyen, C.; Brown, B.S.; McNeish, J.D.; Cohen, A.E.; Kralj, J.M. Cardiotoxicity screening with simultaneous optogenetic pacing, voltage imaging and calcium imaging. J. Pharmacol. Toxicol. Methods 2016, 81, 240–250. [Google Scholar] [CrossRef] [PubMed]
- Yokoo, N.; Baba, S.; Kaichi, S.; Niwa, A.; Mima, T.; Doi, H.; Yamanaka, S.; Nakahata, T.; Heike, T. The effects of cardioactive drugs on cardiomyocytes derived from human induced pluripotent stem cells. Biochem. Biophys. Res. Commun. 2009, 387, 482–488. [Google Scholar] [CrossRef] [PubMed]
- Mehta, A.; Chung, Y.; Sequiera, G.L.; Wong, P.; Liew, R.; Shim, W. Pharmacoelectrophysiology of viral-free induced pluripotent stem cell–derived human cardiomyocytes. Toxicol. Sci. 2013, 131, 458–469. [Google Scholar] [CrossRef] [PubMed]
- Zweigerdt, R. The art of cobbling a running pump—Will human embryonic stem cells mend broken hearts? Semin. Cell Dev. Biol. 2007, 18, 794–804. [Google Scholar] [CrossRef] [PubMed]
- Laflamme, M.A.; Chen, K.Y.; Naumova, A.V.; Muskheli, V.; Fugate, J.A.; Dupras, S.K.; Reinecke, H.; Xu, C.; Hassanipour, M.; Police, S.; et al. Cardiomyocytes derived from human embryonic stem cells in pro-survival factors enhance function of infarcted rat hearts. Nat. Biotechnol. 2007, 25, 1015. [Google Scholar] [CrossRef] [PubMed]
- Rojas, S.V.; Martens, A.; Zweigerdt, R.; Baraki, H.; Rathert, C.; Schecker, N.; Rojas-Hernandez, S.; Schwanke, K.; Martin, U.; Haverich, A.; et al. Transplantation effectiveness of induced pluripotent stem cells is improved by a fibrinogen biomatrix in an experimental model of ischemic heart failure. Tissue Eng. Part A 2015, 21, 1991–2000. [Google Scholar] [CrossRef] [PubMed]
- Van den Akker, F.; Feyen, D.A.M.; van den Hoogen, P.; van Laake, L.W.; van Eeuwijk, E.C.M.; Hoefer, I.; Pasterkamp, G.; Chamuleau, S.A.J.; Grundeman, P.F.; Doevendans, P.A.; et al. Intramyocardial stem cell injection: Go(ne) with the flow. Eur. Heart J. 2017, 38, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Lian, X.; Hsiao, C.; Wilson, G.; Zhu, K.; Hazeltine, L.B.; Azarin, S.M.; Raval, K.K.; Zhang, J.; Kamp, T.J.; Palecek, S.P. Robust cardiomyocyte differentiation from human pluripotent stem cells via temporal modulation of canonical Wnt signaling. Proc. Natl. Acad. Sci. USA 2012, 109, E1848–E1857. [Google Scholar] [CrossRef] [PubMed]
- Weber, N.; Schwanke, K.; Greten, S.; Wendland, M.; Iorga, B.; Fischer, M.; Geers-Knörr, C.; Hegermann, J.; Wrede, C.; Fiedler, J.; et al. Stiff matrix induces switch to pure β-cardiac myosin heavy chain expression in human ESC-derived cardiomyocytes. Basic Res. Cardiol. 2016, 111, 68. [Google Scholar] [CrossRef] [PubMed]
- Iorga, B.; Schwanke, K.; Weber, N.; Wendland, M.; Greten, S.; Piep, B.; dos Remedios, C.G.; Martin, U.; Zweigerdt, R.; Kraft, T.; et al. Differences in contractile function of myofibrils within human embryonic stem cell-derived cardiomyocytes vs. adult ventricular myofibrils are related to distinct sarcomeric protein isoforms. Front. Physiol. 2018, 8, 1111. [Google Scholar] [CrossRef] [PubMed]
| Compound | Type | Mechanism | Expected Effect (−/\/+) 1 |
|---|---|---|---|
| Doxorubicin | Chemotherapeutic | DNA intercalator | − |
| Endothelin-1 | Vasoconstrictor | Hypertrophy inducer | − |
| Acetylsalicylic acid | Analgesic | Anti-inflammation | \ |
| Isoproterenol | Positive inotrope | β-adrenergic receptor agonist | + |
| Phenylephrine | Vasoconstrictor | α-adrenergic receptor agonist | + |
| Amiodarone | Antiarrhythmic agent | K+ channel blocker | + |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Christoffersson, J.; Meier, F.; Kempf, H.; Schwanke, K.; Coffee, M.; Beilmann, M.; Zweigerdt, R.; Mandenius, C.-F. A Cardiac Cell Outgrowth Assay for Evaluating Drug Compounds Using a Cardiac Spheroid-on-a-Chip Device. Bioengineering 2018, 5, 36. https://doi.org/10.3390/bioengineering5020036
Christoffersson J, Meier F, Kempf H, Schwanke K, Coffee M, Beilmann M, Zweigerdt R, Mandenius C-F. A Cardiac Cell Outgrowth Assay for Evaluating Drug Compounds Using a Cardiac Spheroid-on-a-Chip Device. Bioengineering. 2018; 5(2):36. https://doi.org/10.3390/bioengineering5020036
Chicago/Turabian StyleChristoffersson, Jonas, Florian Meier, Henning Kempf, Kristin Schwanke, Michelle Coffee, Mario Beilmann, Robert Zweigerdt, and Carl-Fredrik Mandenius. 2018. "A Cardiac Cell Outgrowth Assay for Evaluating Drug Compounds Using a Cardiac Spheroid-on-a-Chip Device" Bioengineering 5, no. 2: 36. https://doi.org/10.3390/bioengineering5020036
APA StyleChristoffersson, J., Meier, F., Kempf, H., Schwanke, K., Coffee, M., Beilmann, M., Zweigerdt, R., & Mandenius, C.-F. (2018). A Cardiac Cell Outgrowth Assay for Evaluating Drug Compounds Using a Cardiac Spheroid-on-a-Chip Device. Bioengineering, 5(2), 36. https://doi.org/10.3390/bioengineering5020036







