In Vitro Evaluation of Glycoengineered RSV-F in the Human Artificial Lymph Node Reactor
Abstract
:1. Introduction
2. Materials and Methods
2.1. Production and Analysis of Recombinant RSV-F Protein
2.2. Stimulation Experiments
2.3. Cytokine Analysis
2.4. RNA Preparation and NanoString Measurement
3. Results and Discussion
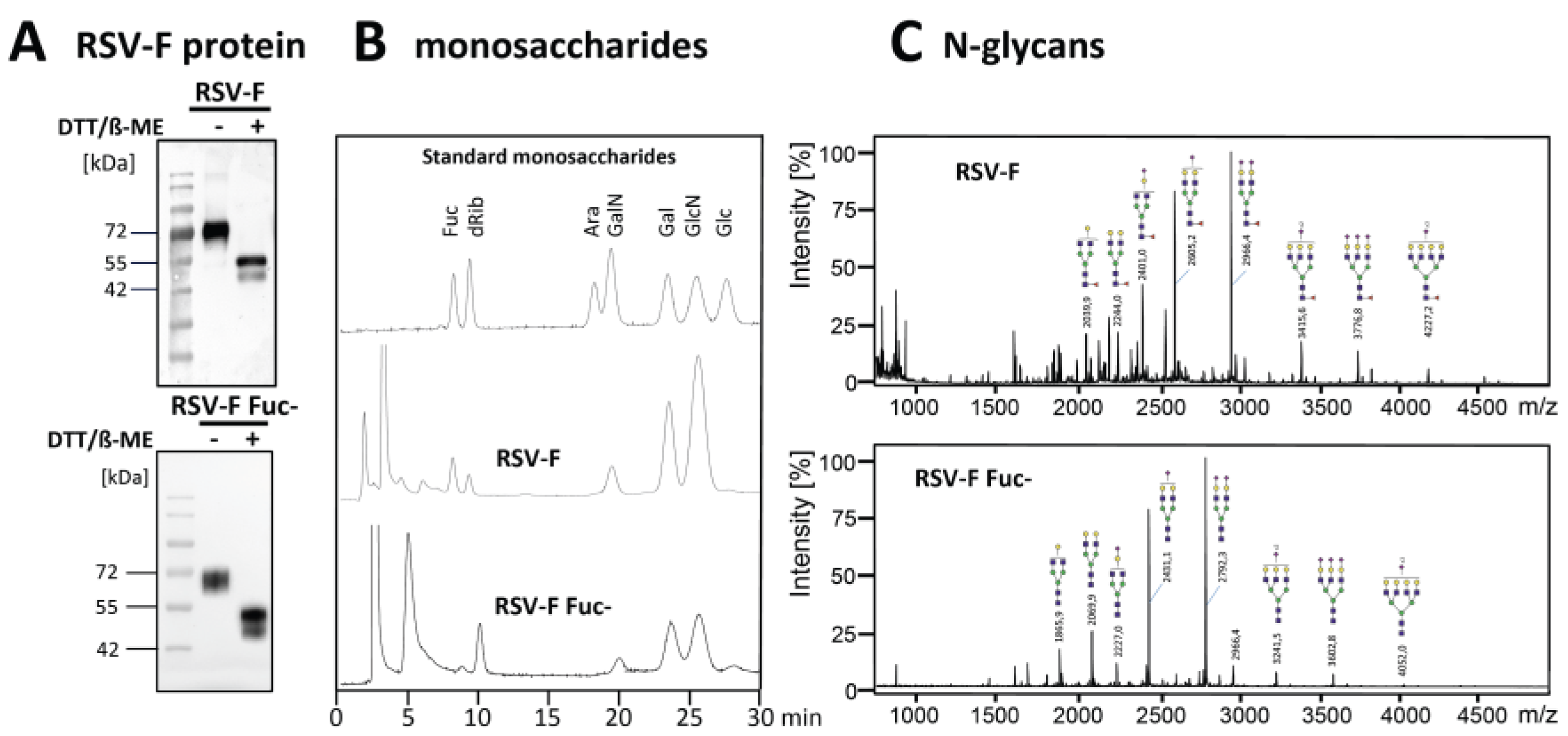
3.1. Production and Analysis of RSV-F Proteins
3.2. Cytokine Analysis
3.3. Quality of RNA Preparations and Nanostring Assays
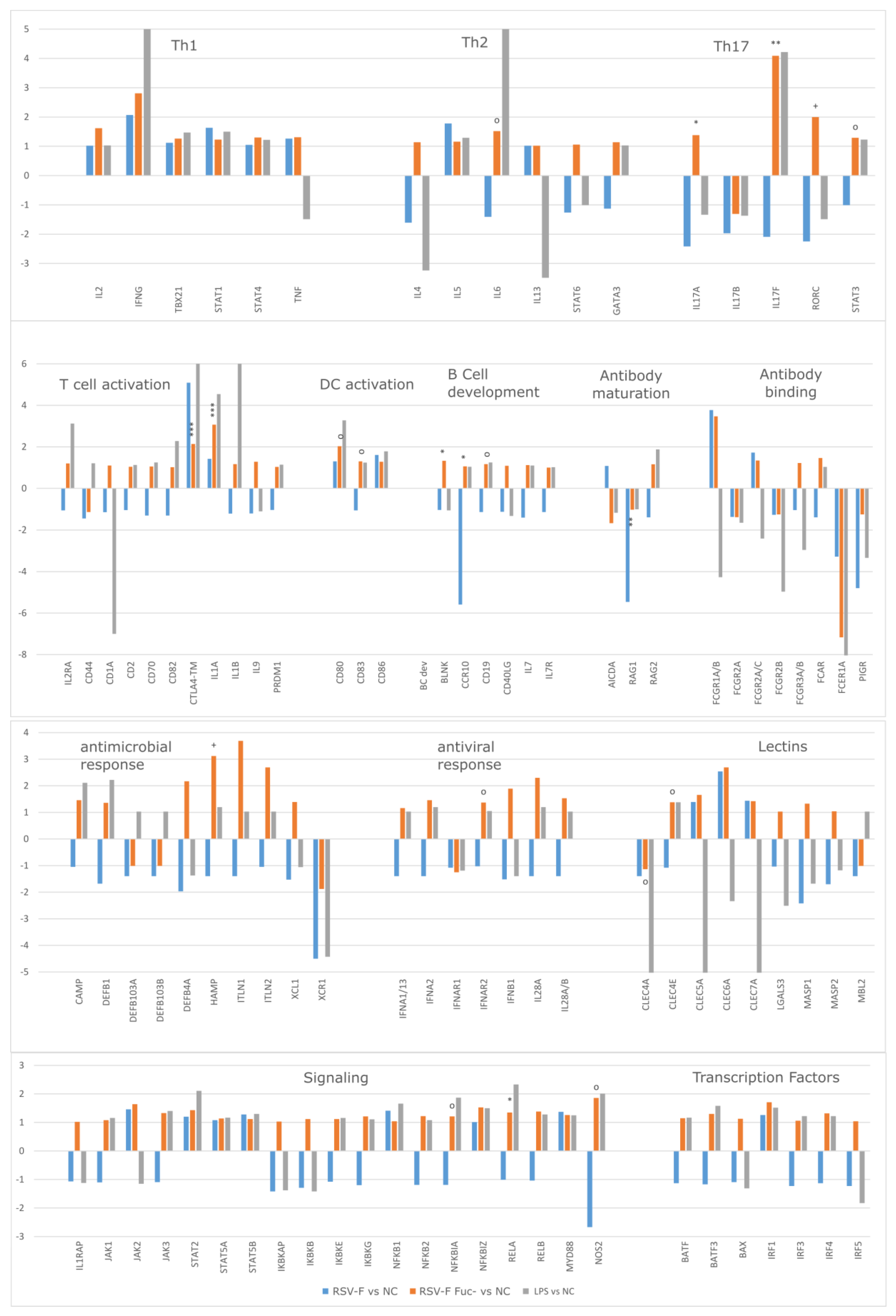
3.4. Gene Expression Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Strugnell, R.; Zepp, F.; Cummingham, A.; Tantawichien, T. Vaccine antigens. Perspect. Vaccinol. 2011, 1, 61–88. [Google Scholar] [CrossRef]
- Israeli, E.; Agmon-Levin, N.; Blank, M.; Shoenfeld, Y. Adjuvants and Autoimmunity. Lupus 2009, 18, 1217–1225. [Google Scholar] [CrossRef] [PubMed]
- Masoudi, S.; Ploen, D.; Kunz, K.; Hildt, E. The adjuvant component α-tocopherol triggers via modulation of Nrf2 the expression and turnover of hypocretin in vitro and its implication to the development of narcolepsy. Vaccine 2014, 32, 2980–2988. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, S.; Lepenies, B. Glycans as Vaccine Antigens and Adjuvants: Immunological Considerations. Methods Mol. Biol. 2015, 1331, 11–26. [Google Scholar] [PubMed]
- Sandig, G.; von Horsten, H.H.; Radke, L.; Blanchard, V.; Frohme, M.; Giese, C.; Sandig, V.; Hinderlich, S. Engineering of CHO Cells for the production of recombinant glycoprotein vaccines with xylosylated N-glycans. Bioengineering 2017, 4, 38. [Google Scholar] [CrossRef]
- Brzezicka, K.; Vogel, V.; Serna, S.; Johannssen, T.; Lepenies, B.; Reichardt, N.C. Influence of Core β-1,2-Xylosylation on Glycoprotein Recognition by Murine C-type Lectin Receptors and Its Impact on Dendritic Cell Targeting. ACS Chem. Biol. 2016, 11, 2347–2356. [Google Scholar] [CrossRef] [PubMed]
- Glezen, W.P.; Taber, L.H.; Frank, A.L.; Kasel, J.A. Risk of primary infection and reinfection with respiratory syncytial virus. Am. J. Dis. Child. 1986, 140, 543–546. [Google Scholar] [CrossRef] [PubMed]
- Thorburn, K. Pre-existing disease is associated with a significantly higher risk of death in severe respiratory syncytial virus infection. Arch. Dis. Child. 2009, 94, 99–103. [Google Scholar] [CrossRef] [PubMed]
- DeVincenzo, J.P.; Whitley, R.J.; Mackman, R.L.; Scaglioni-Weinlich, C.; Harrison, L.; Farrell, E.; McBride, S.; Lambkin-Williams, R.; Jordan, R.; Xin, Y.; et al. Oral GS-5806 Activity in a Respiratory Syncytial Virus Challenge Study. N. Engl. J. Med. 2014, 371, 711–722. [Google Scholar] [CrossRef] [PubMed]
- Press Release—Novavax. Available online: http://ir.novavax.com/phoenix.zhtml?c=71178&p=irol-newsArticle&ID=2202271 (accessed on 30 March 2017).
- Groothuis, J.R.; King, S.J.; Hogerman, D.A.; Paradiso, P.R.; Simoes, E.A. Safety and immunogenicity of a purified F protein respiratory syncytial virus (PFP-2) vaccine in seropositive children with bronchopulmonary dysplasia. J. Infect. Dis. 1998, 177, 467–469. [Google Scholar] [CrossRef] [PubMed]
- Power, U.F.; Plotnicky, H.; Blaecke, A.; Nguyen, T.N. The immunogenicity, protective efficacy and safety of BBG2Na, a subunit respiratory syncytial virus (RSV) vaccine candidate, against RSV-B. Vaccine 2003, 22, 168–176. [Google Scholar] [CrossRef]
- McDonald, T.P.; Jeffree, C.E.; Li, P.; Rixon, H.W.; Brown, G.; Aitken, J.D.; MacLellan, K.; Sugrue, R.J. Evidence that maturation of the N-linked glycans of the respiratory syncytial virus (RSV) glycoproteins is required for virus-mediated cell fusion: The effect of α-mannosidase inhibitors on RSV infectivity. Virology 2006, 350, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Leist, M.; Hartung, T. Inflammatory findings on species extrapolations: Humans are definitely no 70-kg mice. Arch. Toxicol. 2013, 87, 563–567. [Google Scholar] [CrossRef] [PubMed]
- Hurwitz, J.L. Respiratory syncytial virus vaccine development. Expert Rev. Vaccines 2011, 10, 1415–1433. [Google Scholar] [CrossRef] [PubMed]
- Karron, R.A.; Wright, P.F.; Crowe, J.E., Jr.; Clements-Mann, M.L.; Thompson, J.; Makhene, M.; Casey, R.; Murphy, B.R. Evaluation of Two Live, Cold-Passaged, Temperature-Sensitive Respiratory Syncytial Virus Vaccines in Chimpanzees and in Human Adults, Infants, and Children. J. Infect. Dis. 1997, 176, 1428–1436. [Google Scholar] [CrossRef] [PubMed]
- Parker, J.; Sarlang, S.; Thavagnanam, S.; Williamson, G.; O’donoghue, D.; Villenave, R.; Power, U.; Shields, M.; Heaney, L.; Skibinski, G. A 3-D well-differentiated model of pediatric bronchial epithelium demonstrates unstimulated morphological differences between asthmatic and nonasthmatic cells. Pediatr. Res. 2010, 67, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.F.; Ikizler, M.R.; Gonzales, R.A. Growth of respiratory syncytial virus in primary epithelial cells from the human respiratory tract. J. Virol. 2005, 79, 8651–8654. [Google Scholar] [CrossRef] [PubMed]
- Giese, C.; Demmler, C.D.; Ammer, R.; Hartmann, S.; Lubitz, A.; Miller, L.; Müller, R.; Marx, U. A human lymph node in vitro—Challenges and progress. Artif. Organs 2006, 30, 803–808. [Google Scholar] [CrossRef] [PubMed]
- Giese, C.; Lubitz, A.; Demmler, C.D.; Reuschel, J.; Bergner, K.; Marx, U. Immunological substance testing on human lymphatic micro-organoids in vitro. J. Biotechnol. 2010, 148, 38–45. [Google Scholar] [CrossRef] [PubMed]
- Radke, L.; López Hemmerling, D.A.; Lubitz, A.; Giese, C.; Frohme, M. Induced cytokine response of human PMBC-cultures: Correlation of gene expression and secretion profiling and the effect of cryopreservation. Cell. Immun. 2012, 272, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Radke, L.; Giese, C.; Lubitz, A.; Hinderlich, S.; Sandig, G.; Hummel, M.; Frohme, M. Reference gene stability in peripheral blood mononuclear cells determined by qPCR and NanoString. Microchim. Acta 2014, 181, 1733–1742. [Google Scholar] [CrossRef]
- Von Horsten, H.H.; Ogorek, C.; Blanchard, V.; Demmler, C.; Giese, C.; Winkler, K.; Kaup, M.; Berger, M.; Jordan, I.; Sandig, V. Production of non-fucosylated antibodies by co-expression of heterologous GDP-6-deoxy-d-lyxo-4-hexulose reductase. Glycobiology 2010, 20, 1607–1618. [Google Scholar] [CrossRef] [PubMed]
- Rosenlöcher, J.; Böhrsch, V.; Sacharjat, M.; Blanchard, V.; Giese, C.; Sandig, V.; Hackenberger, C.P.R.; Hinderlich, S. Applying Acylated Fucose Analogues to Metabolic Glycoengineering. Bioengineering 2015, 2, 213–234. [Google Scholar] [CrossRef]
- Reinke, S.O.; Bayer, M.; Berger, M.; Blanchard, V.; Hinderlich, S. Analysis of cell surface N-glycosylation of the human embryonic kidney 293t cell line. J. Carbohydr. Chem. 2011, 30, 218–232. [Google Scholar] [CrossRef]
- Frisch, E.; Kaup, M.; Egerer, K.; Weimann, A.; Tauber, R.; Berger, M.; Blanchard, V. Profiling of endo H-released serum N-glycans using CE-LIF and MALDI-TOF-MS-application to rheumatoid arthritis. Electrophoresis 2011, 32, 3510–3515. [Google Scholar] [CrossRef] [PubMed]
- Keil, K.; Radke, L.; Tillich, U.; Frohme, M. Automatisierung des Bio-Plex Pro Analyseverfahrens. Wissenschaftliche Beiträge 2015 2015, 19, 15–19. (In German) [Google Scholar] [CrossRef]
- Wang, H.; Horbinski, C.; Wu, H.; Liu, Y.; Stromberg, A.J.; Wang, C. A Negative Binomial Model-Based Method for Differential Expression Analysis Based on NanoString nCounter Data. Nucleic Acids Res. 2016, 44, e151. [Google Scholar] [PubMed]
- Sugrue, R.J.; Brown, C.; Brown, G.; Aitken, J.; McL Rixon, H.W. Furin cleavage of the respiratory syncytial virus fusion protein is not a requirement for its transport to the surface of virus-infected cells. J. Gen. Virol. 2001, 82, 1375–1386. [Google Scholar] [CrossRef] [PubMed]
- Ceroni, A.; Maass, K.; Geyer, H.; Geyer, R.; Dell, A.; Haslam, S.M. GlycoWorkbench: A Tool for the Computer-Assisted Annotation of Mass Spectra of Glycans. J. Proteome Res. 2008, 7, 1650–1659. [Google Scholar] [CrossRef] [PubMed]
- Kutscher, S.; Dembek, C.J.; Deckert, S.; Russo, C.; Körber, N.; Bogner, J.R.; Geisler, F.; Umgelter, A.; Neuenhahn, M.; Albrecht, J.; et al. Overnight Resting of PBMC Changes Functional Signatures of Antigen Specific T- Cell Responses: Impact for Immune Monitoring within Clinical Trials. PLoS ONE 2013, 8, e76215. [Google Scholar] [CrossRef] [PubMed]
- Röth, A.; Schneider, L.; Himmelreich, H.; Baerlocher, G.M.; Dührsen, U. Impact of culture conditions on the proliferative lifespan of human T cells in vitro. Cytotherapy 2007, 9, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Sprent, J. T-cell survival and the role of cytokines. Immunol. Cell Biol. 2001, 79, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Samarajiwa, S.A.; Forster, S.; Auchettl, K.; Hertzog, P.J. INTERFEROME: The database of interferon regulated genes. Nucleic Acids Res. 2009, 37, D852–D857. [Google Scholar] [CrossRef] [PubMed]
- Guglani, L.; Khader, S.A. Th17 cytokines in mucosal immunity and inflammation. Curr. Opin. HIV AIDS 2010, 5, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Crome, S.Q.; Wang, A.Y.; Levings, M.K. Translational Mini-Review Series on Th17 Cells: Function and regulation of human T helper 17 cells in health and disease. Clin. Exp. Immunol. 2009, 159, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Walsh, E.E.; Falsey, A.R. Humoral and mucosal immunity in protection from natural respiratory syncytial virus infection in adults. J. Infect. Dis. 2004, 190, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Rojas, R.; Apodaca, G. Immunoglobulin transport across polarized epithelial cells. Nat. Rev. Mol. Cell Biol. 2012, 3, 944–956. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Ladinsky, M.S.; Huey-Tubman, K.E.; Jensen, G.J.; McIntosh, J.R.; Björkman, P.J. FcRn-mediated antibody transport across epithelial cells revealed by electron tomography. Nature 2008, 455, 542–546. [Google Scholar] [CrossRef] [PubMed]
- Sardi, M.; Lubitz, A.; Giese, C. Modeling Human Immunity In Vitro: Improving Artificial Lymph Node Physiology by Stromal Cells. Appl. In Vitro Toxicol. 2016, 3, 143–150. [Google Scholar] [CrossRef]
- Mahan, A.E.; Mattoo, H.; Dionne, K.; Tedesco, J.; Pillai, S.; Alter, G. IgG Glycosylation Is Programmed and Remembered after Immunization with TLR Stimulating Adjuvants. AIDS Res. Hum. Retrovir. 2014, 30, A65. [Google Scholar] [CrossRef]
- Shields, R.L.; Lai, J.; Keck, R.; O’Connell, L.Y.; Hong, K.; Meng, Y.G.; Weikert, S.H.A.; Presta, L.G. Lack of Fucose on Human IgG1 N-Linked Oligosaccharide Improves Binding to Human FcγRIII and Antibody-dependent Cellular Toxicity. J. Biol. Chem. 2002, 277, 26733–26740. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, G.; Rohn, M.; McGregor, G.P.; Schemann, M.; Conzelmann, K.K.; Herrler, G. Virokinin, a bioactive peptide of the tachykinin family, is released from the fusion protein of bovine respiratory syncytial virus. J. Biol. Chem. 2003, 278, 46854–46861. [Google Scholar] [CrossRef] [PubMed]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Radke, L.; Sandig, G.; Lubitz, A.; Schließer, U.; Von Horsten, H.H.; Blanchard, V.; Keil, K.; Sandig, V.; Giese, C.; Hummel, M.; et al. In Vitro Evaluation of Glycoengineered RSV-F in the Human Artificial Lymph Node Reactor. Bioengineering 2017, 4, 70. https://doi.org/10.3390/bioengineering4030070
Radke L, Sandig G, Lubitz A, Schließer U, Von Horsten HH, Blanchard V, Keil K, Sandig V, Giese C, Hummel M, et al. In Vitro Evaluation of Glycoengineered RSV-F in the Human Artificial Lymph Node Reactor. Bioengineering. 2017; 4(3):70. https://doi.org/10.3390/bioengineering4030070
Chicago/Turabian StyleRadke, Lars, Grit Sandig, Annika Lubitz, Ulrike Schließer, Hans Henning Von Horsten, Veronique Blanchard, Karolin Keil, Volker Sandig, Christoph Giese, Michael Hummel, and et al. 2017. "In Vitro Evaluation of Glycoengineered RSV-F in the Human Artificial Lymph Node Reactor" Bioengineering 4, no. 3: 70. https://doi.org/10.3390/bioengineering4030070
APA StyleRadke, L., Sandig, G., Lubitz, A., Schließer, U., Von Horsten, H. H., Blanchard, V., Keil, K., Sandig, V., Giese, C., Hummel, M., Hinderlich, S., & Frohme, M. (2017). In Vitro Evaluation of Glycoengineered RSV-F in the Human Artificial Lymph Node Reactor. Bioengineering, 4(3), 70. https://doi.org/10.3390/bioengineering4030070






