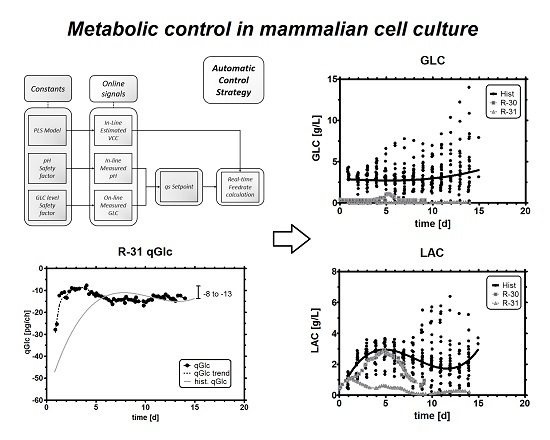
Metabolic Control in Mammalian Fed-Batch Cell Cultures for Reduced Lactic Acid Accumulation and Improved Process Robustness
Abstract
:1. Introduction
1.1. Problem Statement
1.2. Feeding Strategies
1.2.1. Data-Driven Feeding
1.2.2. Adaptive Feeding Strategies
1.3. Challenges in Process Control
| Source | Influence |
|---|---|
| Clones | Metabolic needs may differ greatly, leading to the perpetual development of historical feeding profiles. In adaptive feeding regimes, clone-dependent differences of dielectric properties may complicate biomass estimation when capacitance probes are used, while turbidity probes may detect more or less cell debris in the decline phase, depending on which clone was used. |
| Scales | Especially on-line offgas/kLa–dependent control strategies may become very difficult to transfer because they depend on the aeration and stirrer cascade strategy (i.e., constant or adaptively increasing gas flow to hold pO2). |
| Assumptions | Constant yields (i.e., GLC/OX, X/GLC, X/GLN, etc.) may change over time, leading to stoichiometric over- or under-feeding. |
| Media | Addition of growth-influencing components may change historical feeding profiles completely and make a direct comparison between experiments difficult as these changes have further implications on the process. |
| Process parameters | Changes in temperature, stirrer speed, pO2, pH or pCO2 levels may affect gas solubility, buffer capacity, offgas profiles, cellular stress level, and growth and may change the metabolic requirements for both adaptively or historically calculated feed rates. |
1.4. Goal
2. Experimental Section
2.1. Cell Lines
2.2. Available Dataset
2.3. Media
2.4. Process Setup
2.5. Online Enzymatic Analyzer
2.6. MVDA
2.6.1. Biomass Model
2.6.2. Data Mining
2.7. Process Control
2.7.1. Process Control Scheme
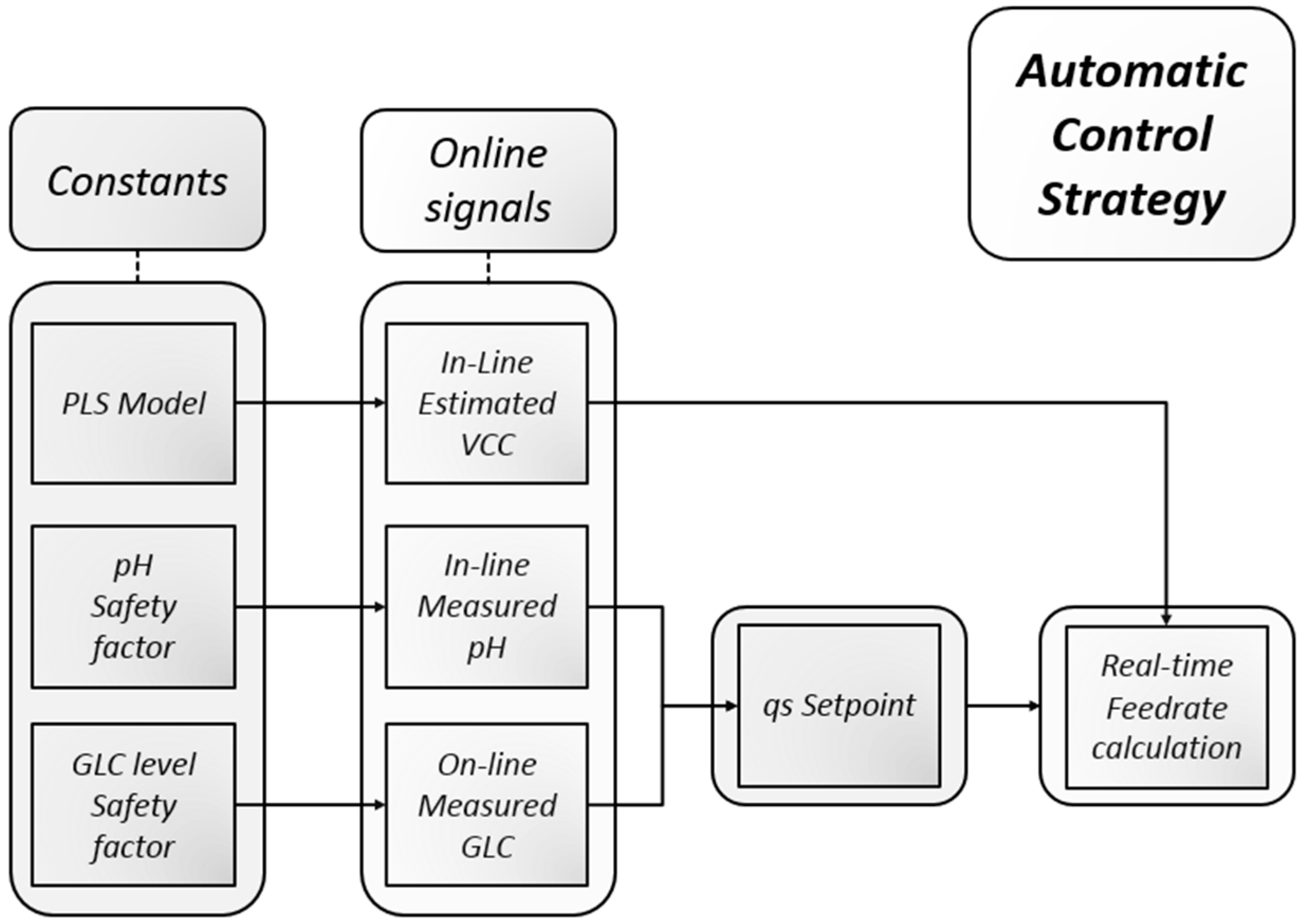
2.7.2. Feed Rate Calculation
| Control Specifications | ||||
|---|---|---|---|---|
| Switch Conditions | Target SP [pg/ch] | |||
| Experiment | R-30 | R-31 | R-30 | R-31 |
| Initialization | Start feed after 2 h | Start feed after 2 h | −15 | −10 |
| pH control † | ||||
| pH high | If pH ≥ 7.1. increase qs by 100% | If pH ≥ 7.4. increase qs by 25% | −30 | −13 |
| pH OK | If pH between 7.1 and 6.9 use the desired qs | If pH between 7.1 and 6.9 use the desired qs | −15 | −10 |
| pH low | If pH ≤ 6.9 reduce qs by 25% | If pH ≤ 6.9 reduce qs by 25% | −11 | −8 |
| Online Analyzer control ‡ | ||||
| GLC low | If Gluc ≤ 0.4 increase qs by 100% | Monitoring | −30 | −10 |
| GLC high ∫ | If Gluc > 0.4 use the desired qs | Monitoring | −15 | −10 |
2.7.3. Operation Window for qs
3. Results and Discussion
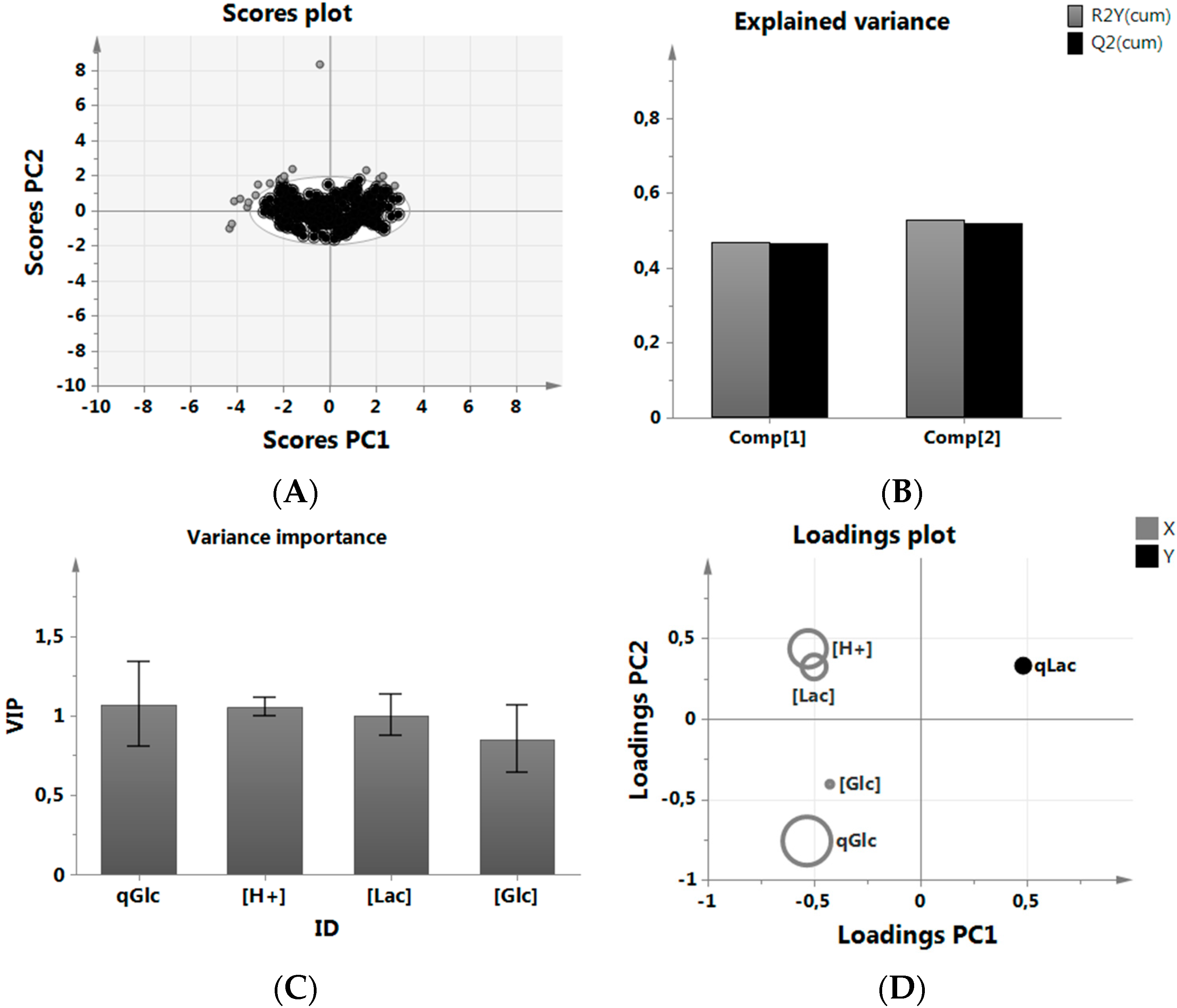
3.1. MVDA for Assessing Lactate Metabolism
- We wanted to be able to set or influence the selected parameters easily, which is the case for the ones we selected, i.e., pH.
- We did not wish to become dependent on other parameters by including them in the analysis, which might then not be frequently analyzed, such as amino acids.
- We did not want to include parameters or define new parameter ranges, which are fixed in a platform/manufacturing process, i.e., pO2 or the pH (acid/base) regulation strategy.
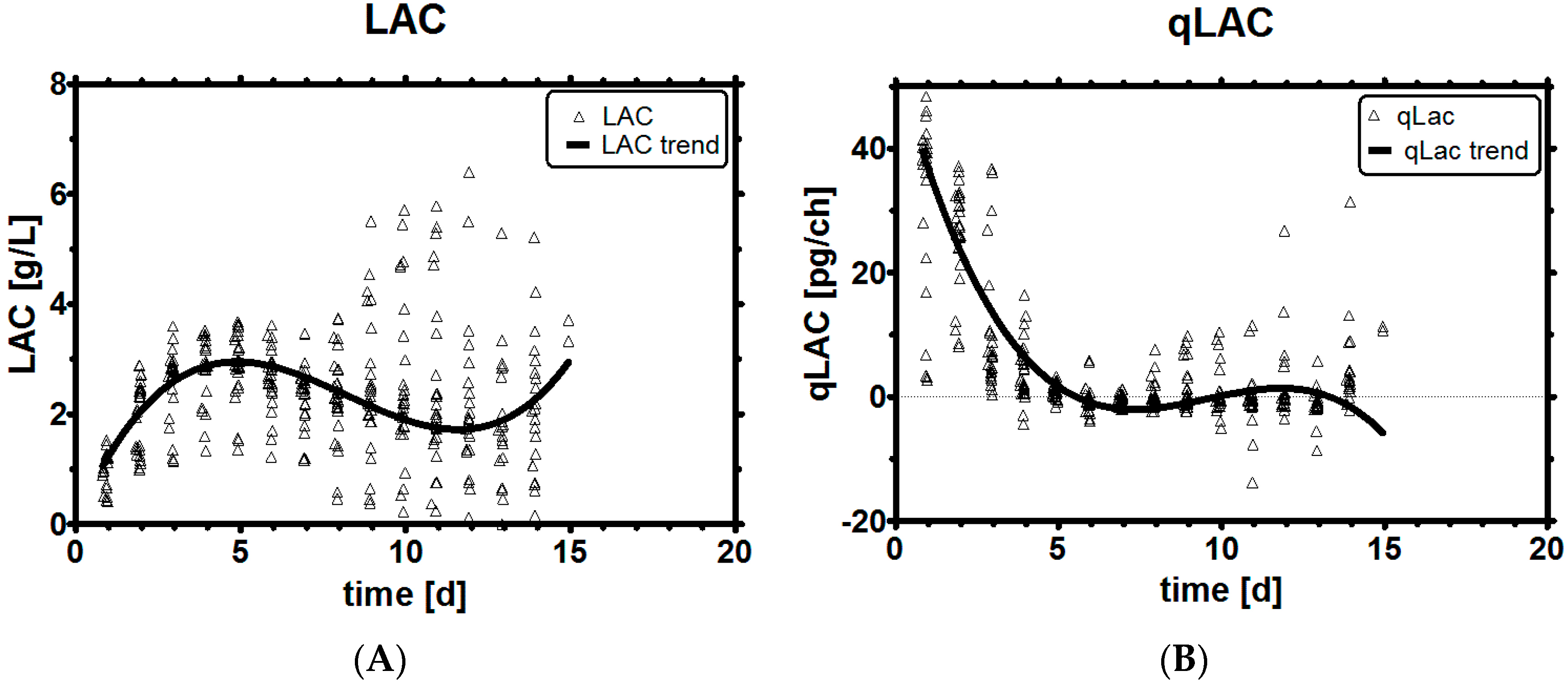
3.2. Lactic Acid Metabolism
3.3. Impact of pH on qGlc
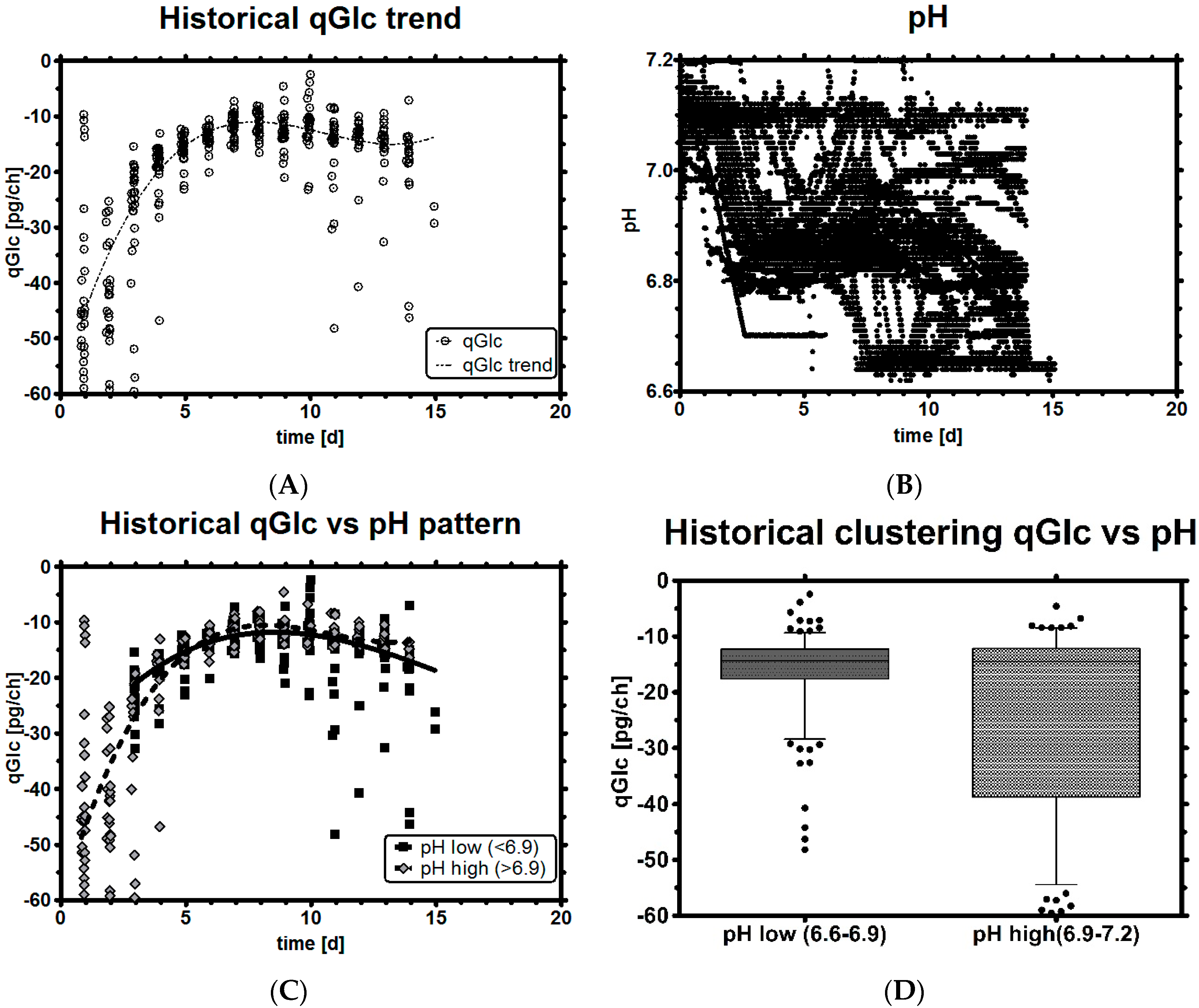
3.4. Set-Point Selection for qGlc
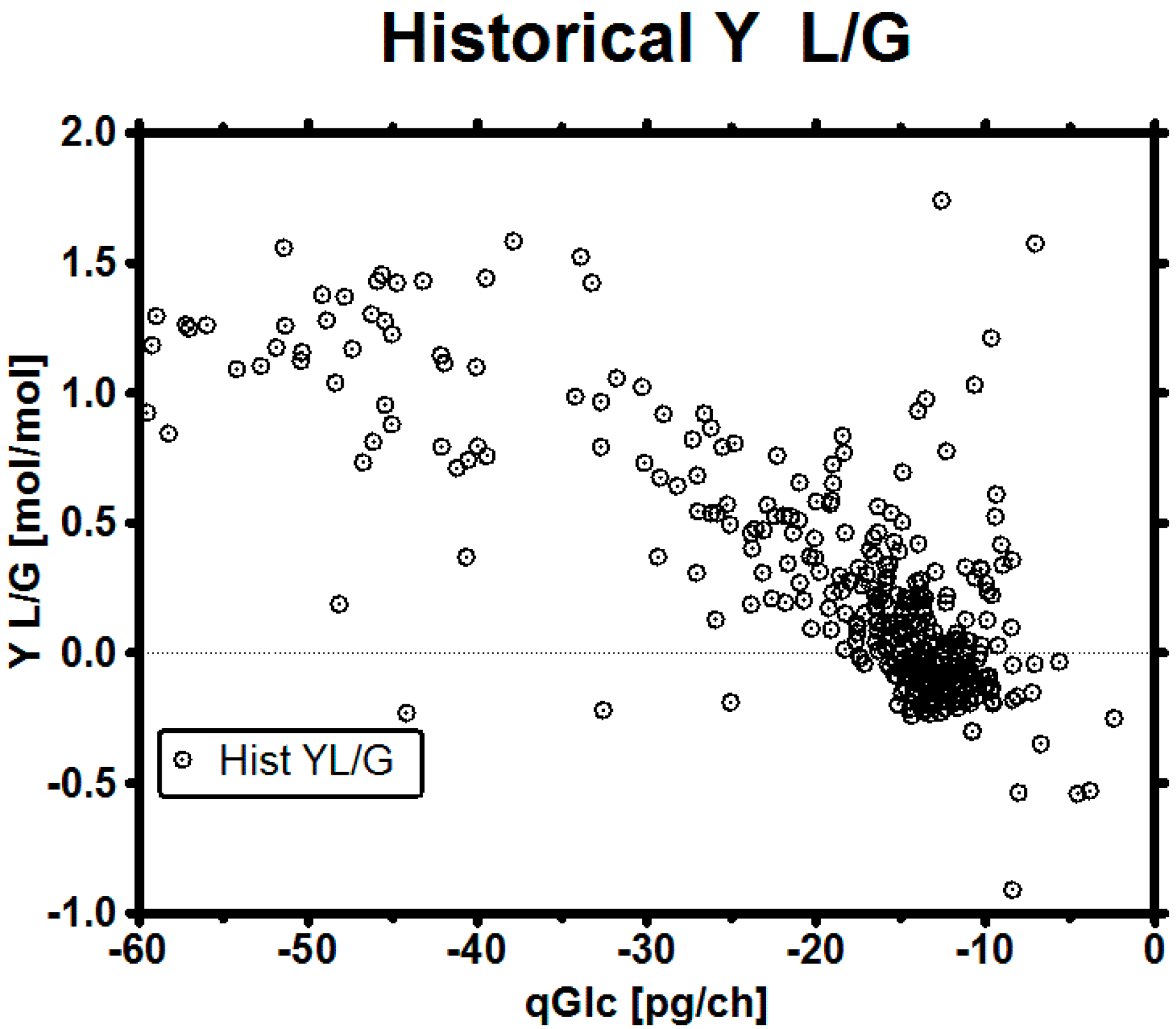
3.5. Impact of a Broad Range as qGlc Set-Point on the Lactic Acid Profile
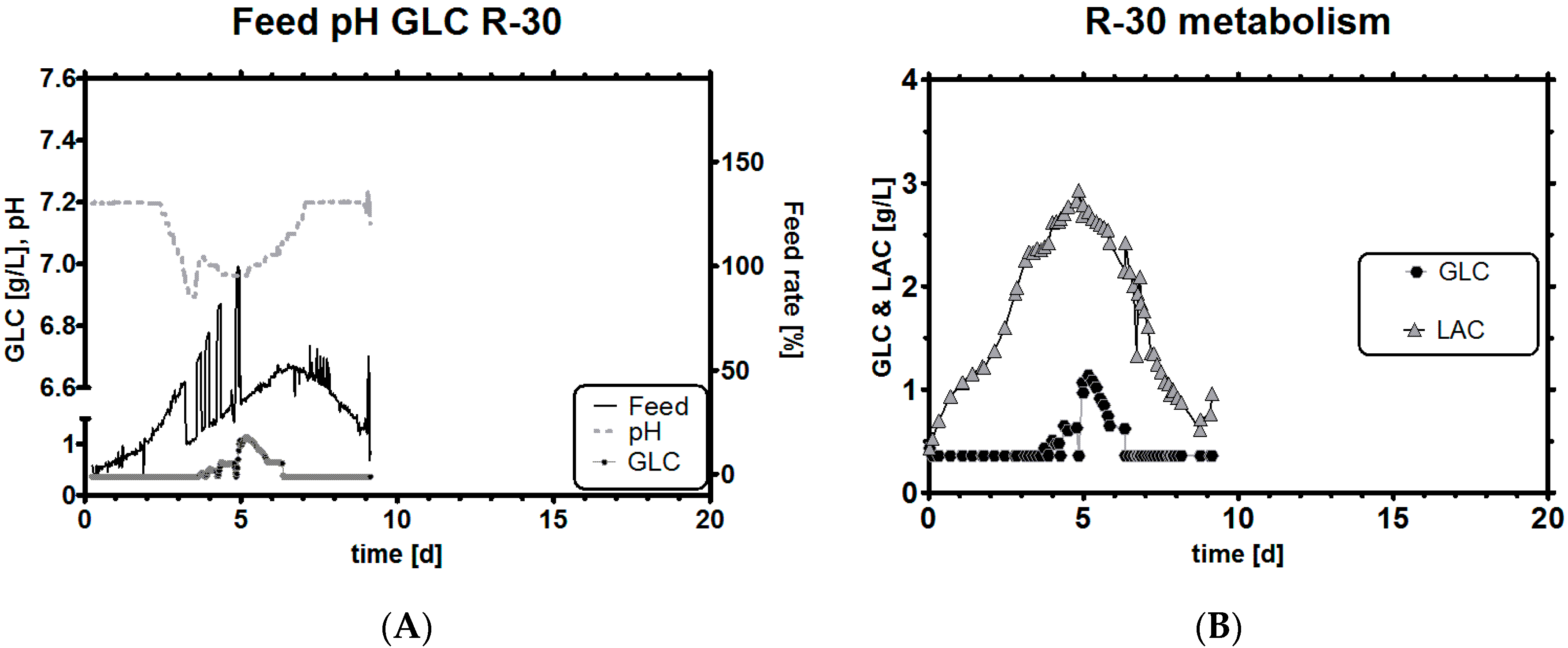
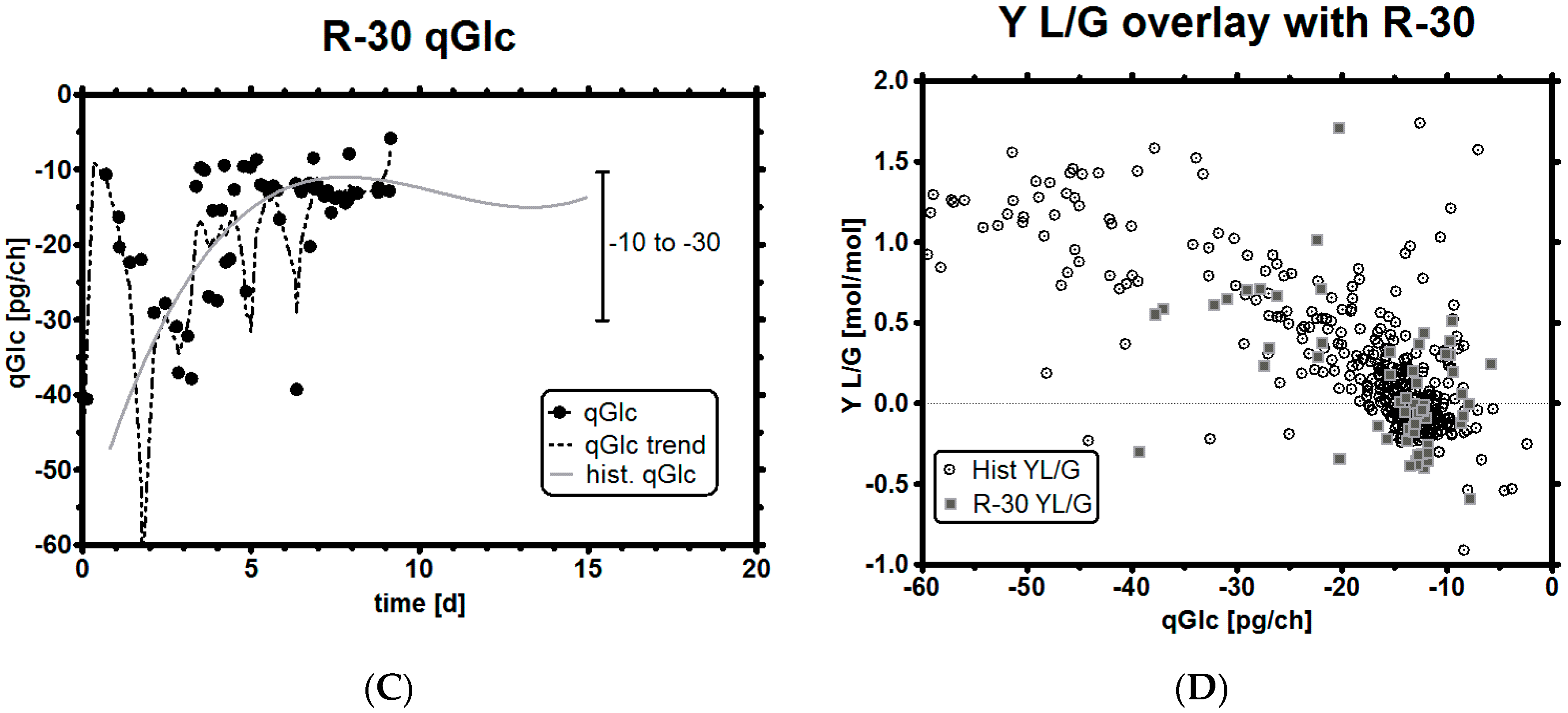
3.6. Impact of a Tight Range as qGlc Set-Point on the Lactic Acid Profile
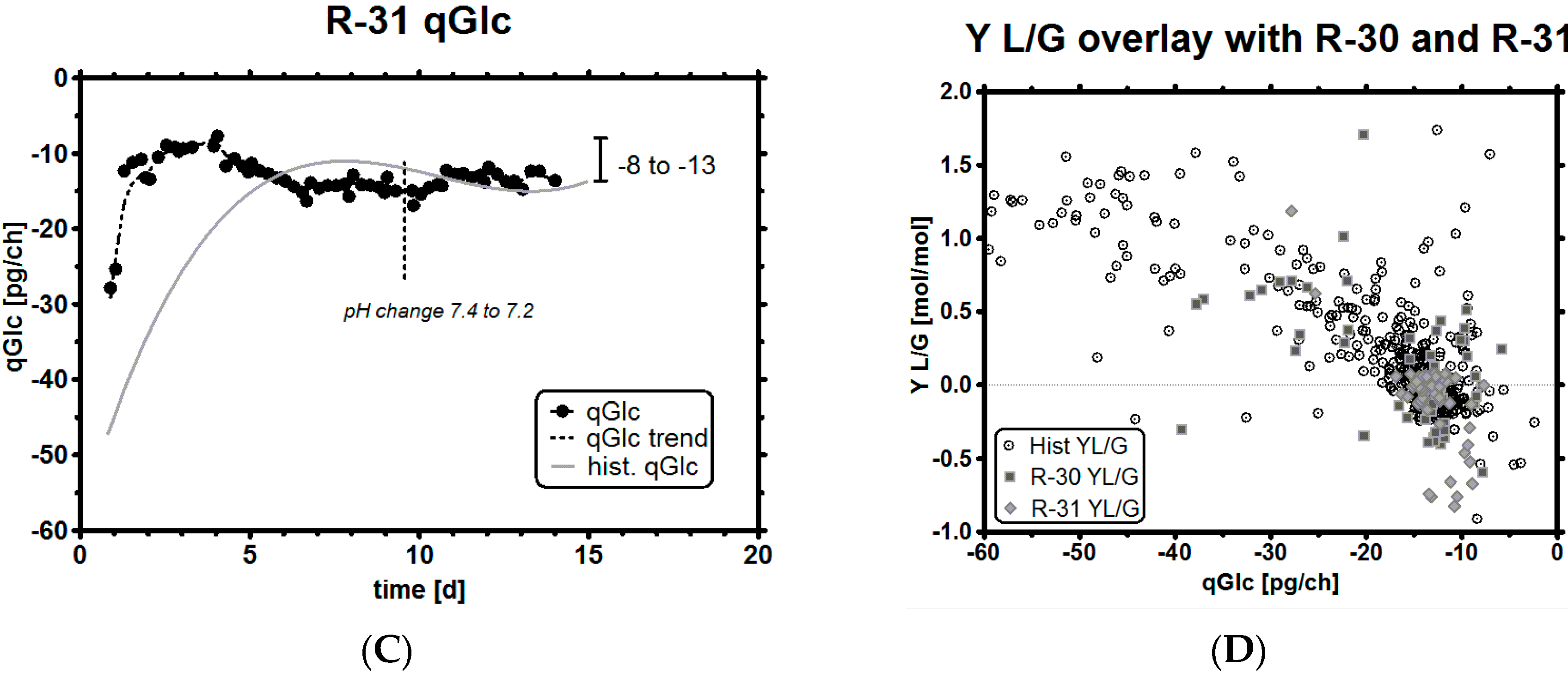
| qGlc [pg/ch] | Y L/G [-] | Y P/G [-] | |||||||
|---|---|---|---|---|---|---|---|---|---|
| ID | Historical | R-30 | R-31 | Historical | R-30 | R-31 | Historical | R-30 | R-31 |
| MEAN | −22.39 | −19.15 | −13.92 | 0.41 | 0.18 | 0.03 | 0.29 | 0.31 | 0.36 |
| SD | 16.53 | 14.62 | 3.64 | 0.63 | 0.49 | 0.31 | 0.12 | 0.05 | 0.1 |
| NAll = 667, NR-30 = 60, NR-31 = 56 | |||||||||
3.7. Adaptive Feeding Using Real-Time Switches
3.7.1. Adaptive Feeding Using pH Correction
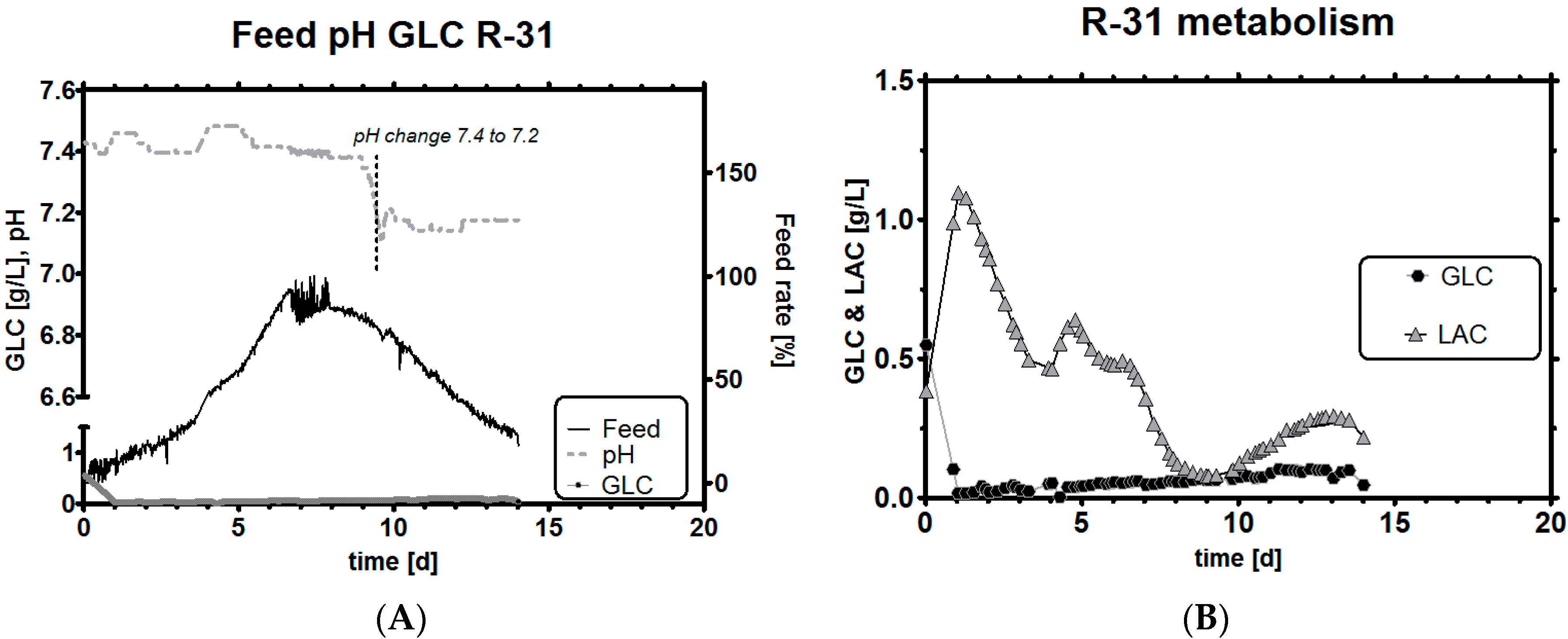
3.7.2. Adaptive Feeding Using an Online Metabolic Analyzer
3.8. Comparison of Experiments with Historical Performance

4. Conclusions
4.1. Eliminating the Root Cause for High Lactic Acid Concentrations
4.2. Directions from MVDA
4.3. Hypothesis for the Positive Effect of pH on Productivity
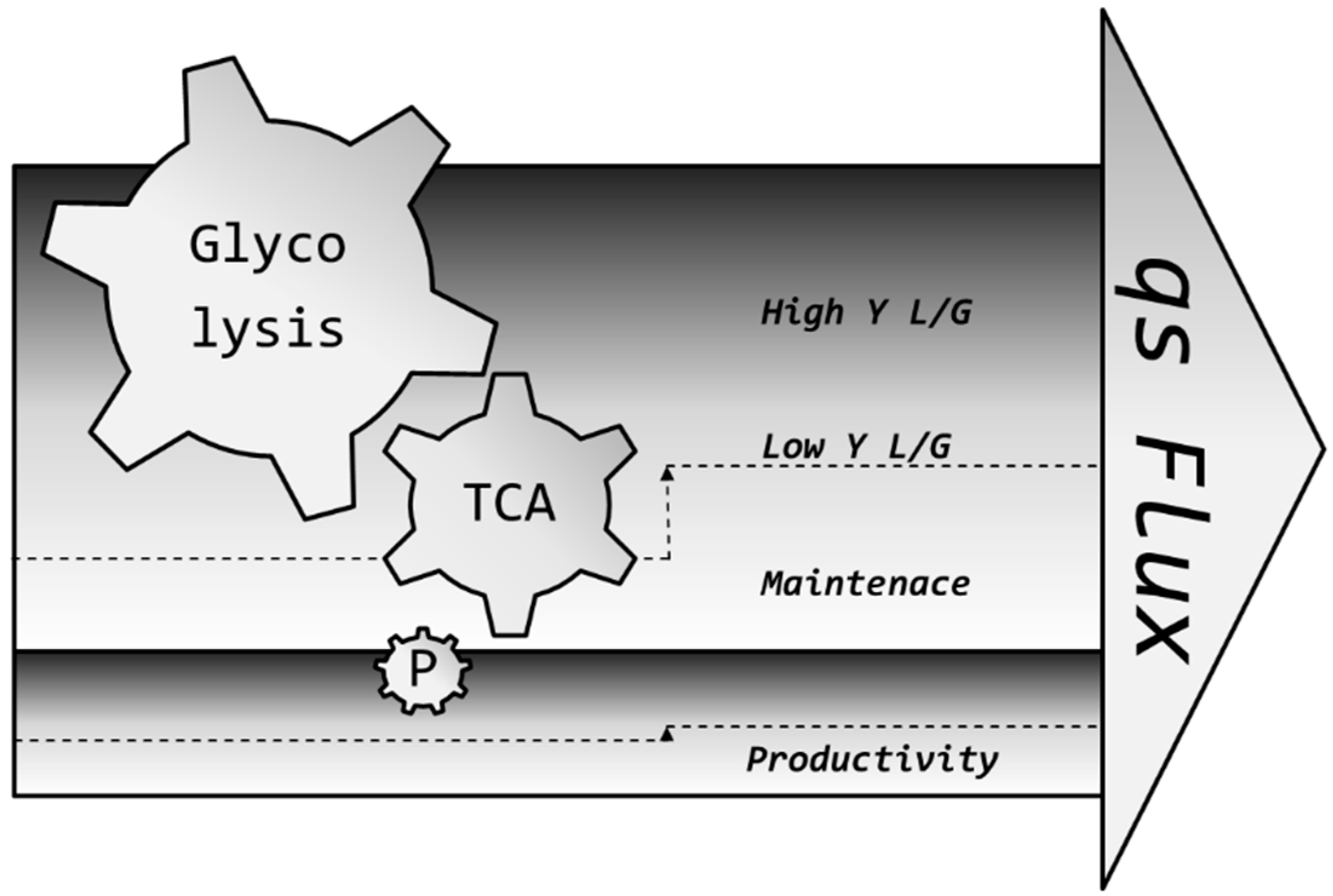
4.4. Living with Uncertainty
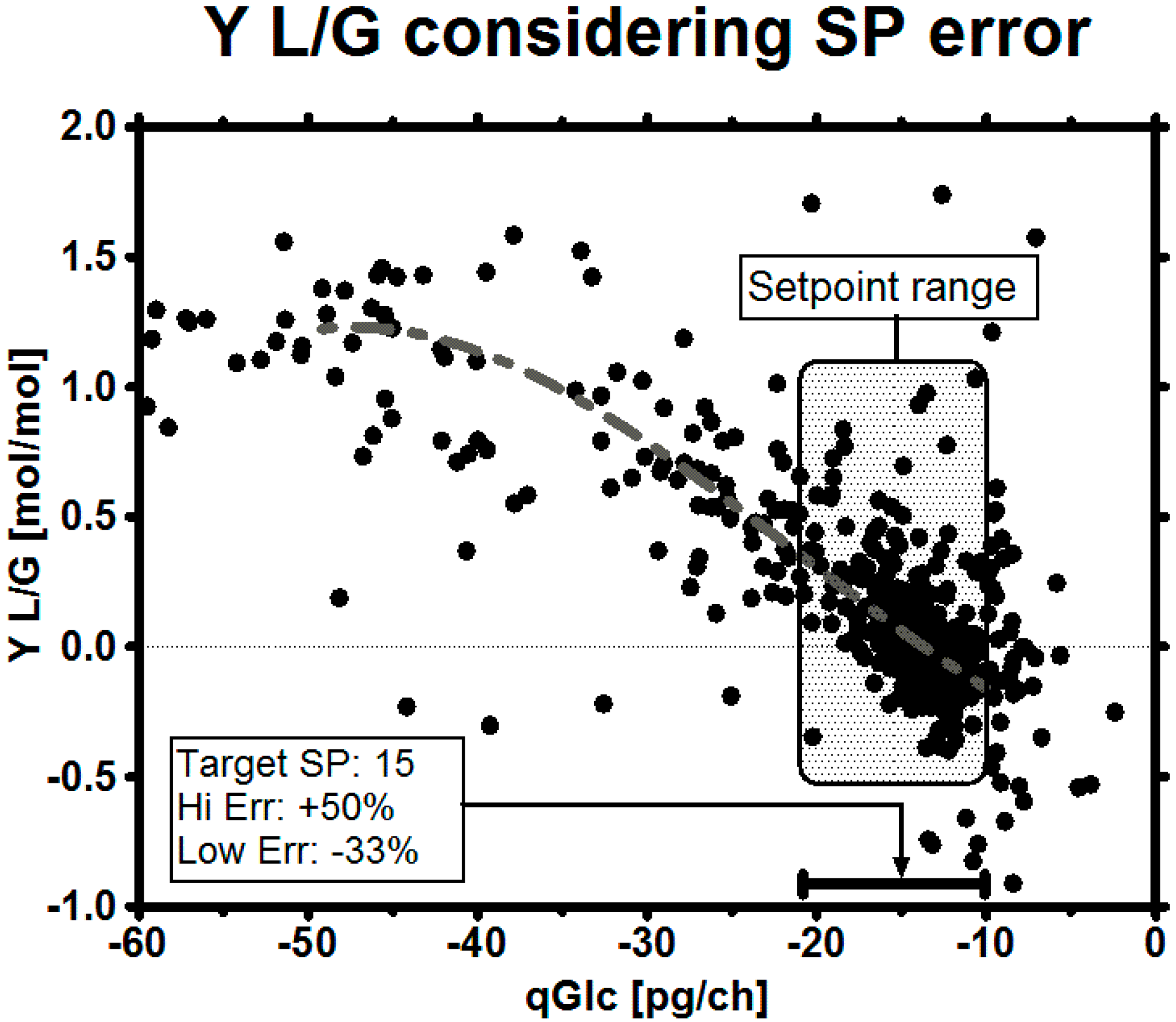
| SP | Lower Range SP | Upper Range SP | |
|---|---|---|---|
| Error | 0% | −33% | +50% |
| Desired set-point considering error | 10 | 6.7 | 15 |
| Robust target set-point | 14.9 | 10 | 22.4 |
4.5. Limitations and Suggested Improvements
4.6. Summary and Outlook
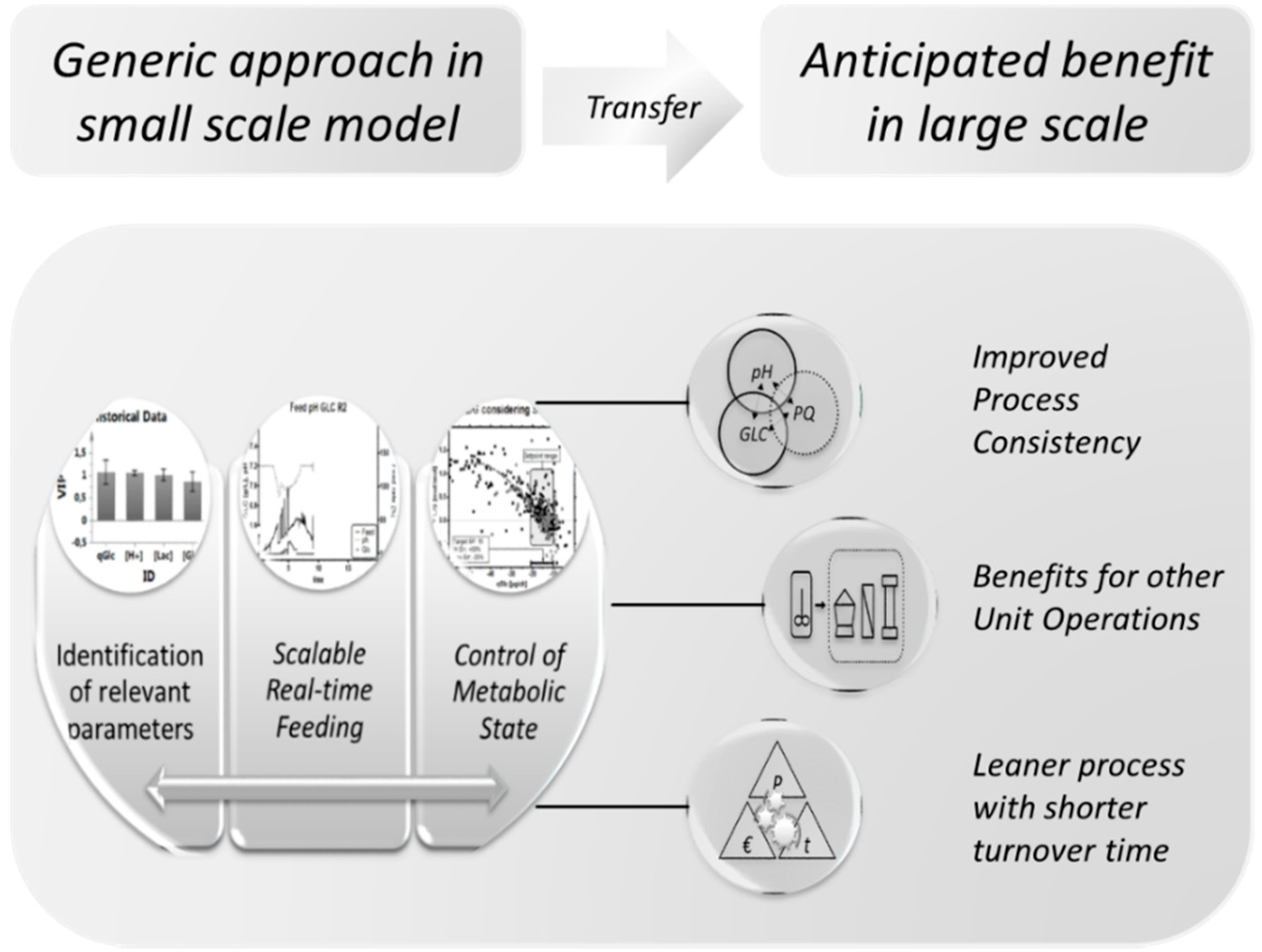
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| CHO | Chinese Hamster Ovary |
| CHO-DG44 | Clone B is derived from this cell line |
| Clone A | Clone in use for a biomass model based on capacitance |
| Clone B | Clone in this contribution (historical and experimental data) |
| CVRMSE | Coefficient of variation of RMSE (Root Mean Square Error) |
| DoE | Design of Experiments |
| Feed concentration, Substrate | Here: Glucose substrate concentration in the feed [mg/mL] |
| GLC | Glucose concentration [g/L] |
| GS-CHO | Glutamine-Synthetase CHO |
| H+ | Hydrogen ion concentration [mol/L] |
| Hist | Historical runs from process development |
| HK | Hexokinase |
| IVCC | Integrated viable cell concentration [c/mL] |
| Km | Clone-dependent Monod constant for glucose affinity (0.1–1 g/L) [g/L] |
| LAC | Lactic acid concentration [g/L] |
| mAbs | Monoclonal Antibodies |
| MVDA | Multivariate Data Analysis |
| N | Number of observations |
| OD | Optical density [-] |
| PAT | Process Analytical Technology |
| pg/ch | Picogram per Cell per hour |
| PFK | 6-Phosphofructo-1-kinase |
| PLS | Partial Least Squares |
| PLS-R | Partial Least Squares Regression |
| Q2 | Variable prediction coefficient |
| QbD | Quality by Design |
| qGlc | Specific glucose uptake rate [pg/ch] |
| qLac | Specific lactic acid uptake/production rate [pg/ch] |
| qOUR | Specific oxygen uptake rate [mmol/ch] |
| qs adapted | New qs set-point after online control action (i.e., pH or online analyzer) |
| qs | General specific uptake rate notation, identical with qGlc as s stands for glucose [pg/ch] |
| R2 | Regression Coefficient [-] |
| R-30 | Experiment R-30, broad range for target qGlc set-point |
| R-31 | Experiment R-31, tight range for target qGlc set-point |
| rlowess | Robust local regression using weighted linear least squares |
| SP | Set-point |
| SD | Standard Deviation |
| VReactorx | Volume of the reactor [mL] |
| VCC | Viable Cell Concentration [cells/mL] |
| VIP | Variance importance of the projection, a measure of relevance of the parameter |
| Y GLC/OX | Yield glucose per oxygen [mol/mol] |
| Y L/G | Yield lactic acid per glucose [mol/mol] |
| Y P/G | Yield product per glucose [mg/g] |
| Y X/GLC | Yield biomass per glucose [cells/g] |
| Y X/GLN | Yield biomass per glutamine [cells/g] |
Appendix A. The Importance of Glucose
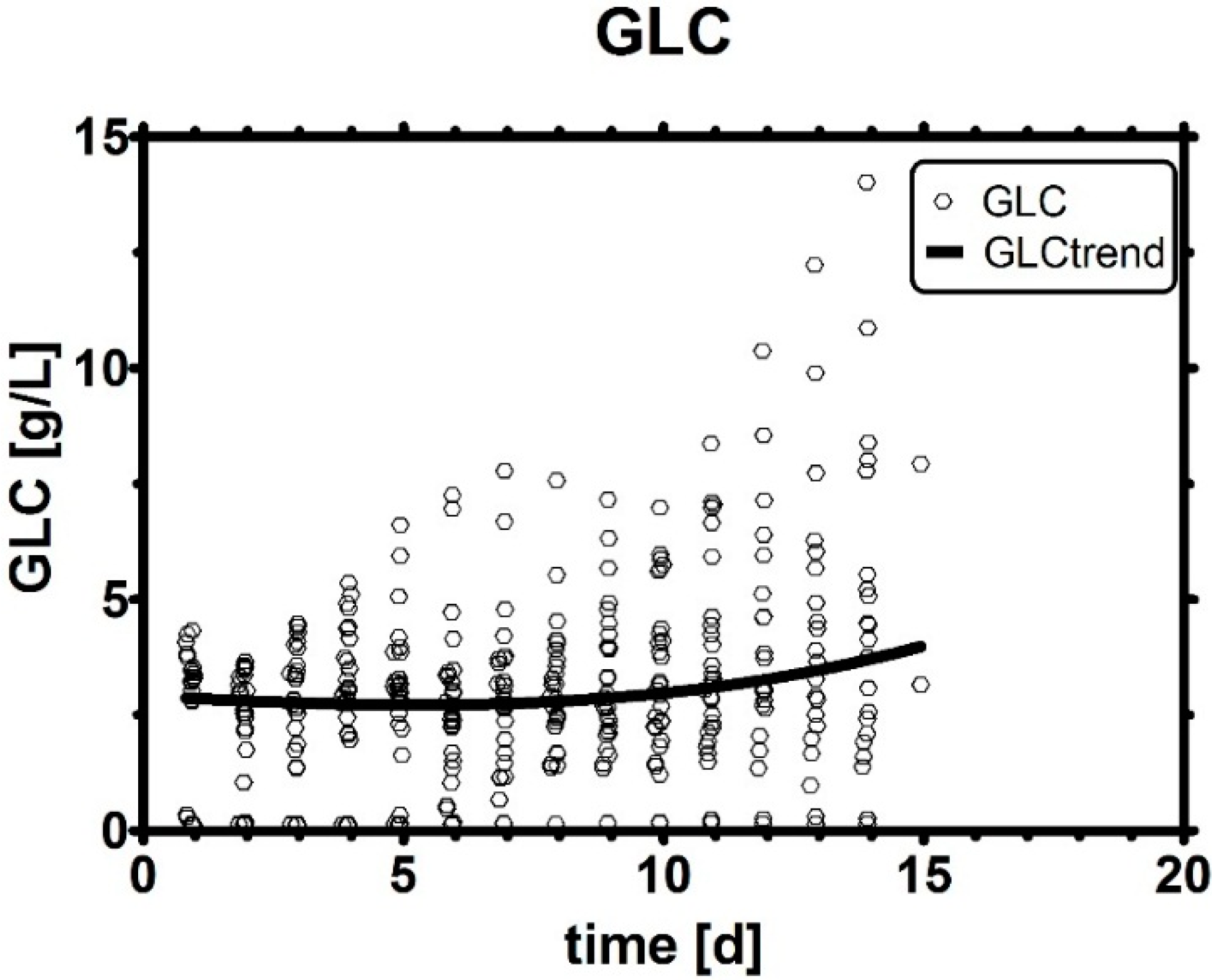
Appendix B. Coefficients in Multivariate Data Analysis (MVDA)
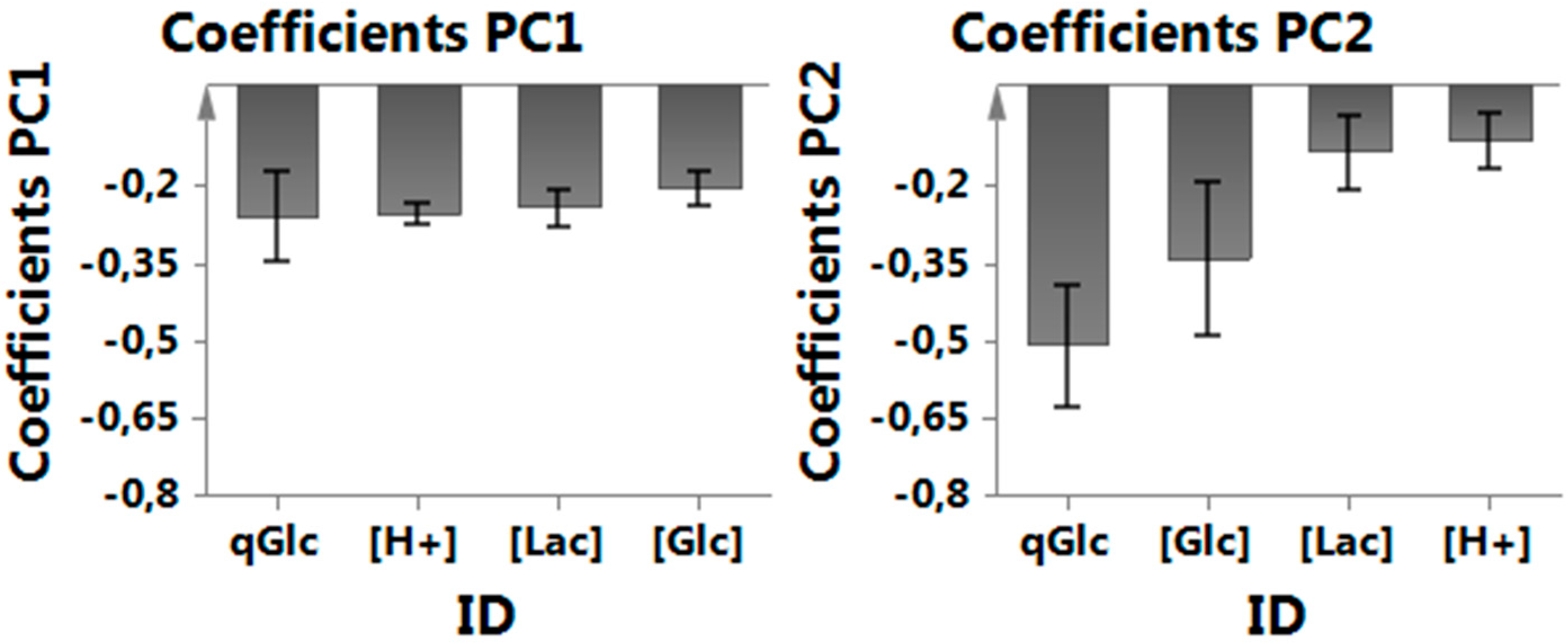
Appendix C. Viability
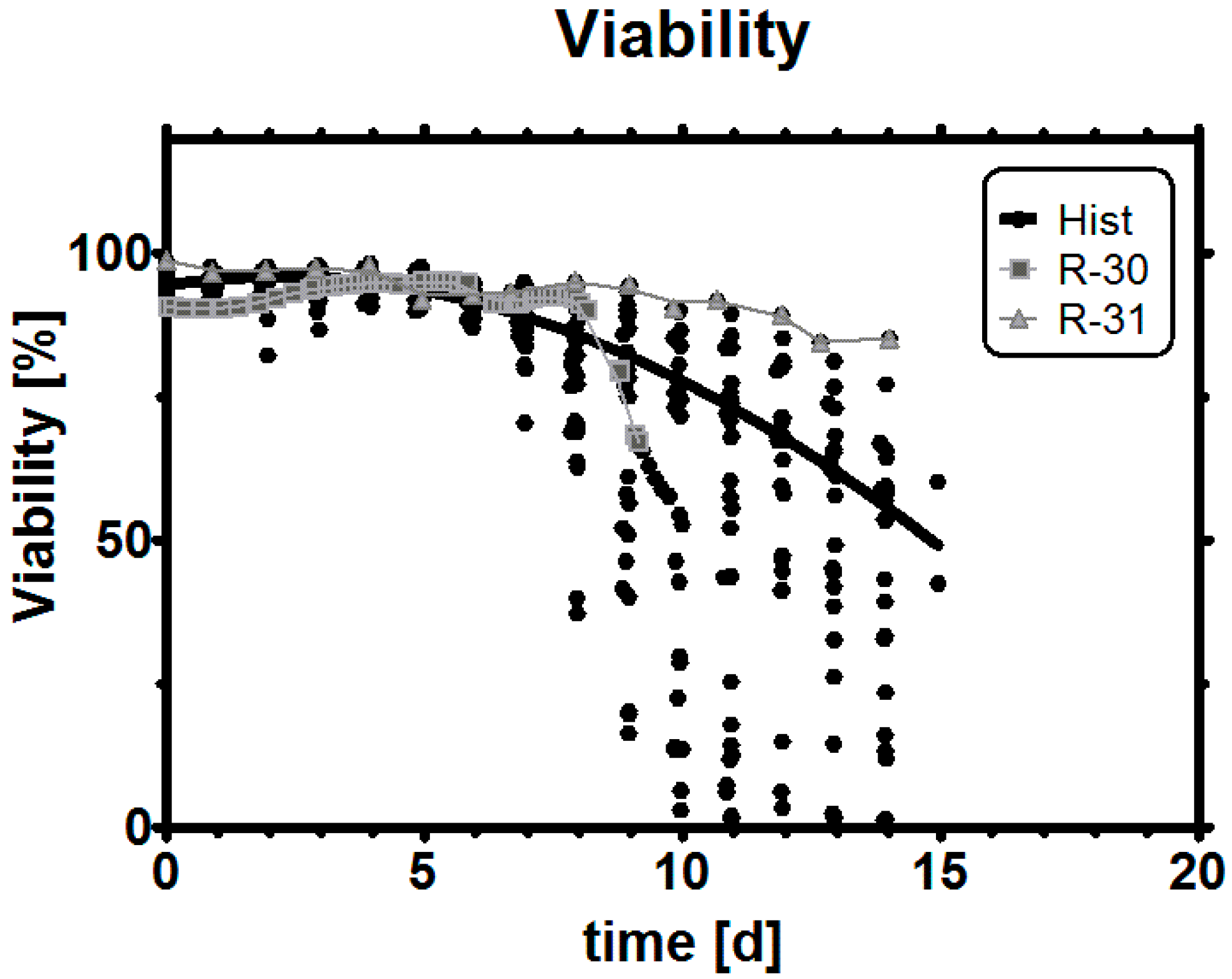
Notes
References
- Rathore, A.S.; Mhatre, R. Quality by Design for Biopharmaceuticals: Principles and Case Studies; Auflage: 1. Wiley-Interscience: Hoboken, NJ, USA, 2011. [Google Scholar]
- Charaniya, S.P. Systems Analysis of Complex Biological Data for Bioprocess Enhancement. Ph.D. Dissertation, University of Minnesota, Minneapolis, MN, USA, 2008. [Google Scholar]
- Croughan, M.S.; Freund, N.W. Strategy to Reduce Lactic Acid Production and Control PH in Animal Cell Culture. U.S. Patent US8470552 B2, 25 June 2013. Available online: http://www.google.com/patents/US8470552 (accessed on 6 January 2016). [Google Scholar]
- Le, H.; Kabbur, S.; Pollastrini, L.; Sun, Z.; Mills, K.; Johnson, K.; Karypis, G.; Hu, W.-S. Multivariate analysis of cell culture bioprocess data—Lactate consumption as process indicator. J. Biotechnol. 2012, 162, 210–223. [Google Scholar] [CrossRef] [PubMed]
- Le, H.; Castro-Melchor, M.; Hakemeyer, C.; Jung, C.; Szperalski, B.; Karypis, G.; Hu, W.-S. Discerning key parameters influencing high productivity and quality through recognition of patterns in process data. BMC Proc. 2011, 5. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Crawford, Y.; Ng, D.; Tung, J.; Pynn, A.F.J.; Meier, A.; Yuk, I.H.; Vijayasankaran, N.; Leach, K.; Joly, J.; et al. Decreasing lactate level and increasing antibody production in Chinese Hamster Ovary cells (CHO) by reducing the expression of lactate dehydrogenase and pyruvate dehydrogenase kinases. J. Biotechnol. 2011, 153, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.-S.; Kantardjieff, A.; Mulukutla, B.C. Cell Lines that Overexpress Lacatate Dehydrogenase c. Patent WO2012075124 A2, 7 June 2012. [Google Scholar]
- Brown, N.J.; Higham, S.E.; Perunovic, B.; Arafa, M.; Balasubramanian, S.; Rehman, I. Lactate Dehydrogenase-B Is Silenced by Promoter Methylation in a High Frequency of Human Breast Cancers. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Legmann, R.; Melito, J.; Belzer, I.; Ferrick, D. Analysis of glycolytic flux as a rapid screen to identify low lactate producing CHO cell lines with desirable monoclonal antibody yield and glycan profile. BMC Proc. 2011. [Google Scholar] [CrossRef] [PubMed]
- Ma, N.; Ellet, J.; Okediadi, C.; Hermes, P.; McCormick, E.; Casnocha, S. A single nutrient feed supports both chemically defined NS0 and CHO fed-batch processes: Improved productivity and lactate metabolism. Biotechnol. Prog. 2009, 25, 1353–1363. [Google Scholar] [CrossRef] [PubMed]
- Altamirano, C.; Paredes, C.; Illanes, A.; Cairó, J.J.; Gòdia, F. Strategies for fed-batch cultivation of t-PA producing CHO cells: Substitution of glucose and glutamine and rational design of culture medium. J. Biotechnol. 2004, 110, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Wahrheit, J.; Nicolae, A.; Heinzle, E. Dynamics of growth and metabolism controlled by glutamine availability in Chinese hamster ovary cells. Appl. Microbiol. Biotechnol. 2014, 98, 1771–1783. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Lee, G.M. Down-regulation of lactate dehydrogenase-A by siRNAs for reduced lactic acid formation of Chinese hamster ovary cells producing thrombopoietin. Appl. Microbiol. Biotechnol. 2007, 74, 152–159. [Google Scholar] [CrossRef] [PubMed]
- Müller, D.; Katinger, H.; Grillari, J. MicroRNAs as targets for engineering of CHO cell factories. Trends Biotechnol. 2008, 26, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Luan, Y.-T.; Stanek, T.C.; Drapeau, D. Controlling Lactic Acid Production in Fed-Batch Cell Cultures via Variation in Glucose Concentration; Bioreactors and Heterologous Gene Expression. U.S. Patent US7429491 B2, 30 September 2008. Available online: http://www.google.com/patents/US7429491 (accessed on 6 January 2016). [Google Scholar]
- Drapeau, D.; Luan, Y.-T.; Stanek, T.C. Restricted Glucose Feed for Animal Cell Culture. Patent WO2004104186 A1, 2 December 2004. Available online: http://www.google.com/patents/WO2004104186A1 (accessed on 6 January 2016). [Google Scholar]
- Basch, J.O.; Gangloff, S.; Joosten, C.E.; Kothari, D.; Lee, S.S.; Leister, K.; Matlock, L.; Sakhamuri, S.; Schilling, B.M.; Zegarelli, S.G. Product Quality Enhancement in Mammalian Cell Culture Processes for Protein Production. Patent WO2004058944 A2, 15 July 2004. Available online: http://www.google.com/patents/WO2004058944A2 (accessed on 6 January 2016). [Google Scholar]
- Sauer, P.W.; Burky, J.E.; Wesson, M.C.; Sternard, H.D.; Qu, L. A high-yielding, generic fed-batch cell culture process for production of recombinant antibodies. Biotechnol. Bioeng. 2000, 67, 585–597. [Google Scholar] [CrossRef]
- Templeton, N.; Dean, J.; Reddy, P.; Young, J.D. Peak antibody production is associated with increased oxidative metabolism in an industrially relevant fed-batch CHO cell culture. Biotechnol. Bioeng. 2013, 110, 2013–2024. [Google Scholar] [CrossRef] [PubMed]
- Mulukutla, B.C.; Khan, S.; Lange, A.; Hu, W.-S. Glucose metabolism in mammalian cell culture: New insights for tweaking vintage pathways. Trends Biotechnol. 2010, 28, 476–484. [Google Scholar] [CrossRef] [PubMed]
- Yongky, A.; Lee, J.; Le, T.; Mulukutla, B.C.; Daoutidis, P.; Hu, W.-S. Mechanism for multiplicity of steady states with distinct cell concentration in continuous culture of mammalian cells. Biotechnol. Bioeng. 2015, 112, 1437–1445. [Google Scholar] [CrossRef] [PubMed]
- Mulukutla, B.C.; Yongky, A.; Daoutidis, P.; Hu, W.-S. Bistability in Glycolysis Pathway as a Physiological Switch in Energy Metabolism. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Wahrheit, J.; Niklas, J.; Heinzle, E. Metabolic control at the cytosol-mitochondria interface in different growth phases of CHO cells. Metab. Eng. 2014, 23, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.M.; Blanch, H.W.; Wilke, C.R. A kinetic analysis of hybridoma growth and metabolism in batch and continuous suspension culture: Effect of nutrient concentration, dilution rate, and pH. Biotechnol. Bioeng. 1988, 32, 947–965. [Google Scholar] [CrossRef] [PubMed]
- Ivarsson, M.; Noh, H.; Morbidelli, M.; Soos, M. Insights into pH-induced metabolic switch by flux balance analysis. Biotechnol. Prog. 2015, 31, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Trummer, E.; Fauland, K.; Seidinger, S.; Schriebl, K.; Lattenmayer, C.; Kunert, R.; Vorauer-Uhl, K.; Weik, R.; Borth, N.; Katinger, H.; et al. Process parameter shifting: Part I. Effect of DOT, pH, and temperature on the performance of Epo-Fc expressing CHO cells cultivated in controlled batch bioreactors. Biotechnol. Bioeng. 2006, 94, 1033–1044. [Google Scholar] [CrossRef] [PubMed]
- Osman, J.J.; Birch, J.; Varley, J. The response of GS-NS0 myeloma cells to pH shifts and pH perturbations. Biotechnol. Bioeng. 2001, 75, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Mulukutla, B.C.; Gramer, M.; Hu, W.-S. On metabolic shift to lactate consumption in fed-batch culture of mammalian cells. Metab. Eng. 2012, 14, 138–149. [Google Scholar] [CrossRef] [PubMed]
- Joosten, C.E.; Leist, C.; Schmidt, J. Cell Cultivation Process. U.S. Patent US8765413 B2, 1 July 2014. Available online: http://www.google.com/patents/US8765413 (accessed on 6 January 2016). [Google Scholar]
- Ozturk, S.S.; Palsson, B.O. Growth, metabolic, and antibody production kinetics of hybridoma cell culture: 2. Effects of serum concentration, dissolved oxygen concentration, and medium pH in a batch reactor. Biotechnol. Prog. 1991, 7, 481–494. [Google Scholar] [CrossRef] [PubMed]
- Patel, S.D.; Papoutsakis, E.T.; Winter, J.N.; Miller, W.M. The lactate issue revisited: Novel feeding protocols to examine inhibition of cell proliferation and glucose metabolism in hematopoietic cell cultures. Biotechnol. Prog. 2000, 16, 885–892. [Google Scholar] [CrossRef] [PubMed]
- Ivarsson, M.; Villiger, T.K.; Morbidelli, M.; Soos, M. Evaluating the impact of cell culture process parameters on monoclonal antibody N-glycosylation. J. Biotechnol. 2014, 188, 88–96. [Google Scholar] [CrossRef] [PubMed]
- L’Allemain, G.; Paris, S.; Pouysségur, J. Growth factor action and intracellular pH regulation in fibroblasts. Evidence for a major role of the Na+/H+ antiport. J. Biol. Chem. 1984, 259, 5809–5815. [Google Scholar] [PubMed]
- Rathore, A.S.; Bhambure, R.; Ghare, V. Process analytical technology (PAT) for biopharmaceutical products. Anal. Bioanal. Chem. 2010, 398, 137–154. [Google Scholar] [CrossRef] [PubMed]
- Luttmann, R.; Bracewell, D.G.; Cornelissen, G.; Gernaey, K.V.; Glassey, J.; Hass, V.C.; Kaiser, C.; Preusse, C.; Striedner, G.; Mandenius, C.-F. Soft sensors in bioprocessing: A status report and recommendations. Biotechnol. J. 2012, 7, 1040–1048. [Google Scholar] [CrossRef] [PubMed]
- Wechselberger, P.; Sagmeister, P.; Herwig, C. Real-time estimation of biomass and specific growth rate in physiologically variable recombinant fed-batch processes. Bioprocess Biosyst. Eng. 2013, 36, 1205–1218. [Google Scholar] [CrossRef] [PubMed]
- Lu, F.; Toh, P.C.; Burnett, I.; Li, F.; Hudson, T.; Amanullah, A.; Li, J. Automated dynamic fed-batch process and media optimization for high productivity cell culture process development. Biotechnol. Bioeng. 2013, 110, 191–205. [Google Scholar] [CrossRef] [PubMed]
- Ozturk, S.S.; Thrift, J.C.; Blackie, J.D.; Naveh, D. Real-time monitoring and control of glucose and lactate concentrations in a mammalian cell perfusion reactor. Biotechnol. Bioeng. 1997, 53, 372–378. [Google Scholar] [CrossRef]
- Franze, R.; Link, T.; Takuma, S.; Takagi, Y.; Hirashima, C.; Tsuda, Y. Method for the Production of a Glycosylated Immunoglobulin. U.S. Patent US20110117087 A1, 19 May 2011. Available online: https://www.google.com/patents/US20110117087 (accessed on 6 January 2016). [Google Scholar]
- Lenas, P.; Kitade, T.; Watanabe, H.; Honda, H.; Kobayashi, T. Adaptive fuzzy control of nutrients concentration in fed-batch culture of mammalian cells. Cytotechnology 1997, 25, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Luan, Y.; Stanek, T.C.; Drapeau, D. Restricted Glucose Feed for Animal Cell Culture. U.S. Patent US7429491. Available online: http://www.sumobrain.com/patents/us/Restricted-glucose-feed-animal-cell/US7429491.html (accessed on 6 January 2016).
- Gagnon, M.; Hiller, G.; Luan, Y.-T.; Kittredge, A.; DeFelice, J.; Drapeau, D. High-End pH-controlled delivery of glucose effectively suppresses lactate accumulation in CHO Fed-batch cultures. Biotechnol. Bioeng. 2011, 108, 1328–1337. [Google Scholar] [CrossRef] [PubMed]
- KWlaschin, F.; Hu, W.-S. Fedbatch culture and dynamic nutrient feeding. Adv. Biochem. Eng. Biotechnol. 2006, 101, 43–74. [Google Scholar]
- Aehle, M.; Schaepe, S.; Kuprijanov, A.; Simutis, R.; Lübbert, A. Simple and efficient control of CHO cell cultures. J. Biotechnol. 2011, 153, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Bezaire, J. Pre-Programmed Non-Feedback Controlled Continuous Feeding of Cell Cultures. Patent WO2013040444 A1, 21 March 2013. [Google Scholar]
- Zhou, W.; Hu, W.-S. On-line characterization of a hybridoma cell culture process. Biotechnol. Bioeng. 1994, 44, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Noll, T.; Biselli, M. Dielectric spectroscopy in the cultivation of suspended and immobilized hybridoma cells. J. Biotechnol. 1998, 63, 187–198. [Google Scholar] [CrossRef]
- JDowd, E.; Jubb, A.; Kwok, K.E.; Piret, J.M. Optimization and control of perfusion cultures using a viable cell probe and cell specific perfusion rates. Cytotechnology 2003, 42, 35–45. [Google Scholar]
- Zhou, W.; Rehm, J.; Hu, W.-S. High viable cell concentration fed-batch cultures of hybridoma cells through on-line nutrient feeding. Biotechnol. Bioeng. 1995, 46, 579–587. [Google Scholar] [CrossRef] [PubMed]
- Europa, A.F.; Gambhir, A.; Fu, P.-C.; Hu, W.-S. Multiple steady states with distinct cellular metabolism in continuous culture of mammalian cells. Biotechnol. Bioeng. 2000, 67, 25–34. [Google Scholar] [CrossRef]
- Mi, L.; Feng, Q.; Li, L.; Wang, X.H. Method for Parameter Control of the Process for Culturing Serum-Suspension Free Animal Cell. Patent CN1557948 A, 29 December 2004. Available online: http://www.google.com/patents/CN1557948A (accessed on 6 January 2016). [Google Scholar]
- Konakovsky, V.; Yagtu, A.C.; Clemens, C.; Müller, M.M.; Berger, M.; Schlatter, S.; Herwig, C. Universal Capacitance Model for Real-Time Biomass in Cell Culture. Sensors 2015, 15, 22128–22150. [Google Scholar] [CrossRef] [PubMed]
- Dietzsch, C.; Spadiut, O.; Herwig, C. On-line multiple component analysis for efficient quantitative bioprocess development. J. Biotechnol. 2013, 163, 362–370. [Google Scholar] [CrossRef] [PubMed]
- Lohninger, H. Datalab 3.5, A Programme for Statistical Analysis. 2000. Available online: http://datalab.epina.at.
- MATLAB, Inc. Robust Local Regression Using Weighted Linear Least Squares in Matlab (RLOWESS). Available online: http://de.mathworks.com/help/curvefit/smoothing-data.html#bq_6ys3–3 (accessed on 6 January 2016).
- Jobé, A.M.; Herwig, C.; Surzyn, M.; Walker, B.; Marison, I.; von Stockar, U. Generally applicable fed-batch culture concept based on the detection of metabolic state by on-line balancing. Biotechnol. Bioeng. 2003, 82, 627–639. [Google Scholar] [CrossRef] [PubMed]
- Wold, S.; Sjöström, M.; Eriksson, L. PLS-regression: A basic tool of chemometrics. Chemom. Intell. Lab. Syst. 2001, 58, 109–130. [Google Scholar] [CrossRef]
- Glassey, J.; Gernaey, K.V.; Clemens, C.; Schulz, T.W.; Oliveira, R.; Striedner, G.; Mandenius, C.-F. Process analytical technology (PAT) for biopharmaceuticals. Biotechnol. J. 2011, 6, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Haenlein, M.; Kaplan, A.M. A Beginner’s Guide to Partial Least Squares Analysis. Underst. Stat. 2004, 3, 283–297. [Google Scholar] [CrossRef]
- Halestrap, A.P.; Price, N.T. The proton-linked monocarboxylate transporter (MCT) family: structure, function and regulation. Biochem. J. 1999, 343, 281–299. [Google Scholar] [CrossRef] [PubMed]
- Smerilli, M.; Neureiter, M.; Wurz, S.; Haas, C.; Frühauf, S.; Fuchs, W. Direct fermentation of potato starch and potato residues to lactic acid by Geobacillus stearothermophilus under non-sterile conditions. J. Chem. Technol. Biotechnol. 2015, 90, 648–657. [Google Scholar] [CrossRef] [PubMed]
- Kurano, N.; Leist, C.; Messi, F.; Kurano, S.; Fiechter, A. Growth behavior of Chinese hamster ovary cells in a compact loop bioreactor: 1. Effects of physical and chemical environments. J. Biotechnol. 1990, 15, 101–111. [Google Scholar] [CrossRef]
- Ansorge, S.; Esteban, G.; Ghommidh, C.; Schmid, G. Monitoring Nutrient Limitations by Online Capacitance Measurements in Batch & Fed-batch CHO Fermentations. In Cell Technology for Cell Products; Smith, R., Ed.; Springer: Houten, Netherlands, 2007; pp. 723–726. [Google Scholar]
- Yongky, A. Analysis of Central Metabolic Pathways in Cultured Mammalian Cells. Ph.D. Dissertation, University of Minnesota, Minneapolis, MN, USA, 2014. [Google Scholar]
- Ündey, C.; Ertunç, S.; Mistretta, T.; Looze, B. Applied advanced process analytics in biopharmaceutical manufacturing: Challenges and prospects in real-time monitoring and control. J. Process Control 2010, 20, 1009–1018. [Google Scholar] [CrossRef]
- Spadiut, O.; Rittmann, S.; Dietzsch, C.; Herwig, C. Dynamic process conditions in bioprocess development. Eng. Life Sci. 2013, 13, 88–101. [Google Scholar] [CrossRef]
- Kantardjieff, A. Transcriptome Analysis in Mammalian Cell Culture: Applications in Process Development and Characterization. Ph.D. Dissertation, University of Minnesota, Minneapolis, MN, USA, 2009. [Google Scholar]
- Fadok, V.A.; Bratton, D.L.; Guthrie, L.; Henson, P.M. Differential effects of apoptotic versus lysed cells on macrophage production of cytokines: role of proteases. J. Immunol. (Baltim. Md.: 1950) 2001, 166, 6847–6854. [Google Scholar] [CrossRef]
- Rathmell, J.C.; Fox, C.J.; Plas, D.R.; Hammerman, P.S.; Cinalli, R.M.; Thompson, C.B. Akt-directed glucose metabolism can prevent Bax conformation change and promote growth factor-independent survival. Mol. Cell. Biol. 2003, 23, 7315–7328. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Dengler, T.J.; Kluger, M.S.; Madge, L.A.; Schechner, J.S.; Maher, S.E.; Pober, J.S.; Bothwell, A.L. Cytoprotection of human umbilical vein endothelial cells against apoptosis and CTL-mediated lysis provided by caspase-resistant Bcl-2 without alterations in growth or activation responses. J. Immunol. (Baltim. Md.: 1950) 2000, 164, 4665–4671. [Google Scholar] [CrossRef]
- Ansorge, S.; Esteban, G.; Schmid, G. On-line monitoring of responses to nutrient feed additions by multi-frequency permittivity measurements in fed-batch cultivations of CHO cells. Cytotechnology 2010, 62, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Ansorge, S.; Esteban, G.; Schmid, G. On-line monitoring of infected Sf-9 insect cell cultures by scanning permittivity measurements and comparison with off-line biovolume measurements. Cytotechnology 2007, 55, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Dabros, M.; Dennewald, D.; Currie, D.J.; Lee, M.H.; Todd, R.W.; Marison, I.W.; Stockar, U. Cole–Cole, linear and multivariate modeling of capacitance data for on-line monitoring of biomass. Bioprocess Biosyst. Eng. 2008, 32, 161–173. [Google Scholar] [CrossRef] [PubMed]
- Cannizzaro, C.; Gügerli, R.; Marison, I.; von Stockar, U. On-line biomass monitoring of CHO perfusion culture with scanning dielectric spectroscopy. Biotechnol. Bioeng. 2003, 84, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Carvell, J.P.; Dowd, J.E. On-line Measurements and Control of Viable Cell Density in Cell Culture Manufacturing Processes using Radio-frequency Impedance. Cytotechnology 2006, 50, 35–48. [Google Scholar] [CrossRef] [PubMed]
- De Jesus, M.J.; Bourgeois, M.; Baumgartner, G.; Tromba, P.; Jordan, D.M.; Amstutz, M.H.; Wurm, P.F.M. The Influence of pH on Cell Growth and Specific Productivity of Two CHO Cell Lines Producing Human Anti Rh D IgG. In Animal Cell Technology: From Target to Market; Lindner-Olsson, D.E., Chatzissavidou, M.N., Lüllau, D.E., Eds.; Springer: Houten, Netherlands, 2001; pp. 197–199. [Google Scholar]
- Gambhir, A.; Korke, R.; Lee, J.; Fu, P.-C.; Europa, A.; Hu, W.-S. Analysis of cellular metabolism of hybridoma cells at distinct physiological states. J. Biosci. Bioeng. 2003, 95, 317–327. [Google Scholar] [CrossRef]
- Eyal, A.M.; Starr, J.N.; Fisher, R.; Hazan, B.; Canari, R.; Witzke, D.R.; Gruber, P.R.; Kolstad, J.J. Lactic Acid Processing; Methods; Arrangements; and, Product. U.S. Patent US6320077 B1, 20 November 2001. Available online: http://www.google.com/patents/US6320077 (accessed on 6 January 2016). [Google Scholar]
- Abdel-Rahman, M.A.; Tashiro, Y.; Sonomoto, K. Recent advances in lactic acid production by microbial fermentation processes. Biotechnol. Adv. 2013, 31, 877–902. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Huang, J.; Zhou, R. Progress in engineering acid stress resistance of lactic acid bacteria. Appl. Microbiol. Biotechnol. 2014, 98, 1055–1063. [Google Scholar] [CrossRef] [PubMed]
- Klein, T.; Heinzel, N.; Kroll, P.; Brunner, M.; Herwig, C.; Neutsch, L. Quantification of cell lysis during CHO bioprocesses: Impact on cell count, growth kinetics and productivity. J. Biotechnol. 2015, 207, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Boron, W.F. Regulation of intracellular pH. Adv. Physiol. Educ. 2004, 28, 160–179. [Google Scholar] [CrossRef] [PubMed]
- Dechant, R.; Binda, M.; Lee, S.S.; Pelet, S.; Winderickx, J.; Peter, M. Cytosolic pH is a second messenger for glucose and regulates the PKA pathway through V-ATPase. EMBO J. 2010, 29, 2515–2526. [Google Scholar] [CrossRef] [PubMed]
- Boyer, M.J.; Tannock, I.F. Regulation of intracellular pH in tumor cell lines: Influence of microenvironmental conditions. Cancer Res. 1992, 52, 4441–4447. [Google Scholar] [PubMed]
- Olsnes, S.; Tønnessen, T.I.; Sandvig, K. pH-regulated anion antiport in nucleated mammalian cells. J. Cell Biol. 1986, 102, 967–971. [Google Scholar] [CrossRef] [PubMed]
- Pilkis, S.J.; Claus, T.H. Hepatic Gluconeogenesis/Glycolysis: Regulation and Structure/Function Relationships of Substrate Cycle Enzymes. Annu. Rev. Nutr. 1991, 11, 465–515. [Google Scholar] [CrossRef] [PubMed]
- Okar, D.A.; Wu, C.; Lange, A.J. Regulation of the regulatory enzyme, 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase. Adv. Enzym. Regul. 2004, 44, 123–154. [Google Scholar] [CrossRef] [PubMed]
- Somberg, E.W.; Mehlman, M.A. Regulation of gluconeogenesis and lipogenesis. The regulation of mitochondrial pyruvate metabolism in guinea-pig liver synthesizing precursors for gluconeogenesis. Biochem. J. 1969, 112, 435–447. [Google Scholar] [CrossRef] [PubMed]
- Mulquiney, P.J.; Kuchel, P.W. Model of 2,3-bisphosphoglycerate metabolism in the human erythrocyte based on detailed enzyme kinetic equations: Equations and parameter refinement. Biochem. J. 1999, 342, 581–596. [Google Scholar] [CrossRef] [PubMed]
- Hutkins, R.W.; Nannen, N.L. pH Homeostasis in Lactic Acid Bacteria. J. Dairy Sci. 1993, 76, 2354–2365. [Google Scholar] [CrossRef]
- Nolan, R.P.; Lee, K. Dynamic model of CHO cell metabolism. Metab. Eng. 2011, 13, 108–124. [Google Scholar] [CrossRef] [PubMed]
- Shivhare, M.; McCreath, G. Practical Considerations for DoE Implementation in Quality by Design. BioProcess Int. 2010, 22–30. Available online: http://www.bioprocessintl.com/wp-content/uploads/bpi-content/BPI_A_100806AR03_O_98037a.pdf (accessed on 6 January 2016). [Google Scholar]
- Carrondo, M.J.T.; Alves, P.M.; Carinhas, N.; Glassey, J.; Hesse, F.; Merten, O.-W.; Micheletti, M.; Noll, T.; Oliveira, R.; Reichl, U.; et al. How can measurement, monitoring, modeling and control advance cell culture in industrial biotechnology? Biotechnol. J. 2012, 7, 1522–1529. [Google Scholar] [CrossRef] [PubMed]
- Oeggerli, A.; Eyer, K.; Heinzle, E. On-line gas analysis in animal cell cultivation: I. Control of dissolved oxygen and pH. Biotechnol. Bioeng. 1995, 45, 42–53. [Google Scholar] [CrossRef] [PubMed]
- Åström, K.J.; Murray, R.M. Feedback Systems Web Site. 5 April 2008. Available online: http://www.cds.caltech.edu/~murray/amwiki. (accessed on 29 May 2015).
- Dumont, G. EECE Courses Prof. Guy Dumont: Lecture Notes. Available online: http://www.phoneoximeter.org/ece-courses/eece-460/lecture-notes/ (accessed on 29 May 2015).
- Press, W.H.; Teukolsky, S.A.; Vetterling, W.T.; Flannery, B.P. Numerical Recipes 3rd Edition: The Art of Scientific Computing, 3rd ed.; Cambridge University Press: Cambridge, UK; New York, NY, USA, 2007. [Google Scholar]
- Motulsky, H. Intuitive Biostatistics: A Nonmathematical Guide to Statistical Thinking; Oxford University Press: New York, NY, USA, 2013. [Google Scholar]
- Motulsky, H. Fitting Models to Biological Data Using Linear and Nonlinear Regression: A Practical Guide to Curve Fitting; 1. Aufl.; Oxford University Press: New York, NY, USA, 2004. [Google Scholar]
- Aghamohseni, H.; Ohadi, K.; Spearman, M.; Krahn, N.; Moo-Young, M.; Scharer, J.M.; Butler, M.; Budman, H.M. Effects of nutrient levels and average culture pH on the glycosylation pattern of camelid-humanized monoclonal antibody. J. Biotechnol. 2014, 186, 98–109. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Del Val Jimenez, I.; Müller, C.; Wagtberg Sen, J.; Rasmussen, S.K.; Kontoravdi, C.; Weilguny, D.; Andersen, M.R. Amino acid and glucose metabolism in fed-batch CHO cell culture affects antibody production and glycosylation. Biotechnol. Bioeng. 2015, 112, 521–535. [Google Scholar] [CrossRef] [PubMed]
- Wechselberger, P.; Herwig, C. Model-based analysis on the relationship of signal quality to real-time extraction of information in bioprocesses. Biotechnol. Prog. 2012, 28, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Zalai, D.; Dietzsch, C.; Herwig, C. Risk-based Process Development of Biosimilars as Part of the Quality by Design Paradigm. PDA J. Pharm. Sci. Technol. PDA 2013, 67, 569–580. [Google Scholar] [CrossRef] [PubMed]
- Gnoth, S.; Jenzsch, M.; Simutis, R.; Lübbert, A. Process Analytical Technology (PAT): Batch-to-batch reproducibility of fermentation processes by robust process operational design and control. J. Biotechnol. 2007, 132, 180–186. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Vijayasankaran, N.; Shen, A.; Kiss, R.; Amanullah, A. Cell culture processes for monoclonal antibody production. mAbs 2010, 2, 466–477. [Google Scholar] [CrossRef] [PubMed]
- Abu-Absi, S.F.; Yang, L.; Thompson, P.; Jiang, C.; Kandula, S.; Schilling, B.; Shukla, A.A. Defining process design space for monoclonal antibody cell culture. Biotechnol. Bioeng. 2010, 106, 894–905. [Google Scholar] [CrossRef] [PubMed]
- Gronemeyer, P.; Ditz, R.; Strube, J. Trends in Upstream and Downstream Process Development for Antibody Manufacturing. Bioengineering 2014, 1, 188–212. [Google Scholar] [CrossRef]
- Gao, Y.; Kipling, K.; Glassey, J.; Willis, M.; Montague, G.; Zhou, Y.; Titchener-Hooker, N.J. Application of agent-based system for bioprocess description and process improvement. Biotechnol. Prog. 2010, 26, 706–716. [Google Scholar] [CrossRef] [PubMed]
- Green, A.; Glassey, J. Multivariate analysis of the effect of operating conditions on hybridoma cell metabolism and glycosylation of produced antibody: Effect of operating conditions on mAb glycosylation. J. Chem. Technol. Biotechnol. 2015, 90, 303–313. [Google Scholar] [CrossRef]
- Hossler, P.; Mulukutla, B.C.; Hu, W.-S. Systems Analysis of N-Glycan Processing in Mammalian Cells. PLoS ONE 2007, 2, e713. [Google Scholar] [CrossRef] [PubMed]
- Le, H.T.N. Mining High-Dimensional Bioprocess and Gene Expression Data for Enhanced Process Performance. Ph.D. Dissertation, University of Minnesota, Minneapolis, MN, USA, 2012. [Google Scholar]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Konakovsky, V.; Clemens, C.; Müller, M.M.; Bechmann, J.; Berger, M.; Schlatter, S.; Herwig, C. Metabolic Control in Mammalian Fed-Batch Cell Cultures for Reduced Lactic Acid Accumulation and Improved Process Robustness. Bioengineering 2016, 3, 5. https://doi.org/10.3390/bioengineering3010005
Konakovsky V, Clemens C, Müller MM, Bechmann J, Berger M, Schlatter S, Herwig C. Metabolic Control in Mammalian Fed-Batch Cell Cultures for Reduced Lactic Acid Accumulation and Improved Process Robustness. Bioengineering. 2016; 3(1):5. https://doi.org/10.3390/bioengineering3010005
Chicago/Turabian StyleKonakovsky, Viktor, Christoph Clemens, Markus Michael Müller, Jan Bechmann, Martina Berger, Stefan Schlatter, and Christoph Herwig. 2016. "Metabolic Control in Mammalian Fed-Batch Cell Cultures for Reduced Lactic Acid Accumulation and Improved Process Robustness" Bioengineering 3, no. 1: 5. https://doi.org/10.3390/bioengineering3010005
APA StyleKonakovsky, V., Clemens, C., Müller, M. M., Bechmann, J., Berger, M., Schlatter, S., & Herwig, C. (2016). Metabolic Control in Mammalian Fed-Batch Cell Cultures for Reduced Lactic Acid Accumulation and Improved Process Robustness. Bioengineering, 3(1), 5. https://doi.org/10.3390/bioengineering3010005