Optimized Signal Acquisition and Advanced AI for Robust 1D EMG Classification: A Comparative Study of Machine Learning, Deep Learning, and Reinforcement Learning
Abstract
1. Introduction
- Phase 1: Signal Acquisition Optimization. This phase focuses strictly on the foundational hardware-level parameters, specifically evaluating the impact of magnitude quantization (bit depth) and time-domain sampling rates on signal fidelity. By isolating these variables, we empirically determine the optimal operating point called the ‘tipping point’, where data efficiency is maximized without compromising the reconstruction of critical physiological features. This phase provides the rigorous data quality baseline required for valid downstream analysis.
- Phase 2: Model Comparative Analysis. Building directly upon the optimized acquisition parameters established in Phase 1, the second phase benchmarks the efficacy of three distinct AI paradigms: classical Machine Learning (ML), state-of-the-art Deep Learning (DL), and novel Reinforcement Learning (RL) agents. This phase evaluates how well each architecture generalizes across both controlled synthetic environments and noisy real-world datasets, with comparative results.
- Systematic Analysis of Acquisition Parameters: A detailed evaluation of the effects of magnitude quantization (bit depth) and time-domain sampling rates on 1D EMG signal reconstruction error and their direct implications for classification accuracy.
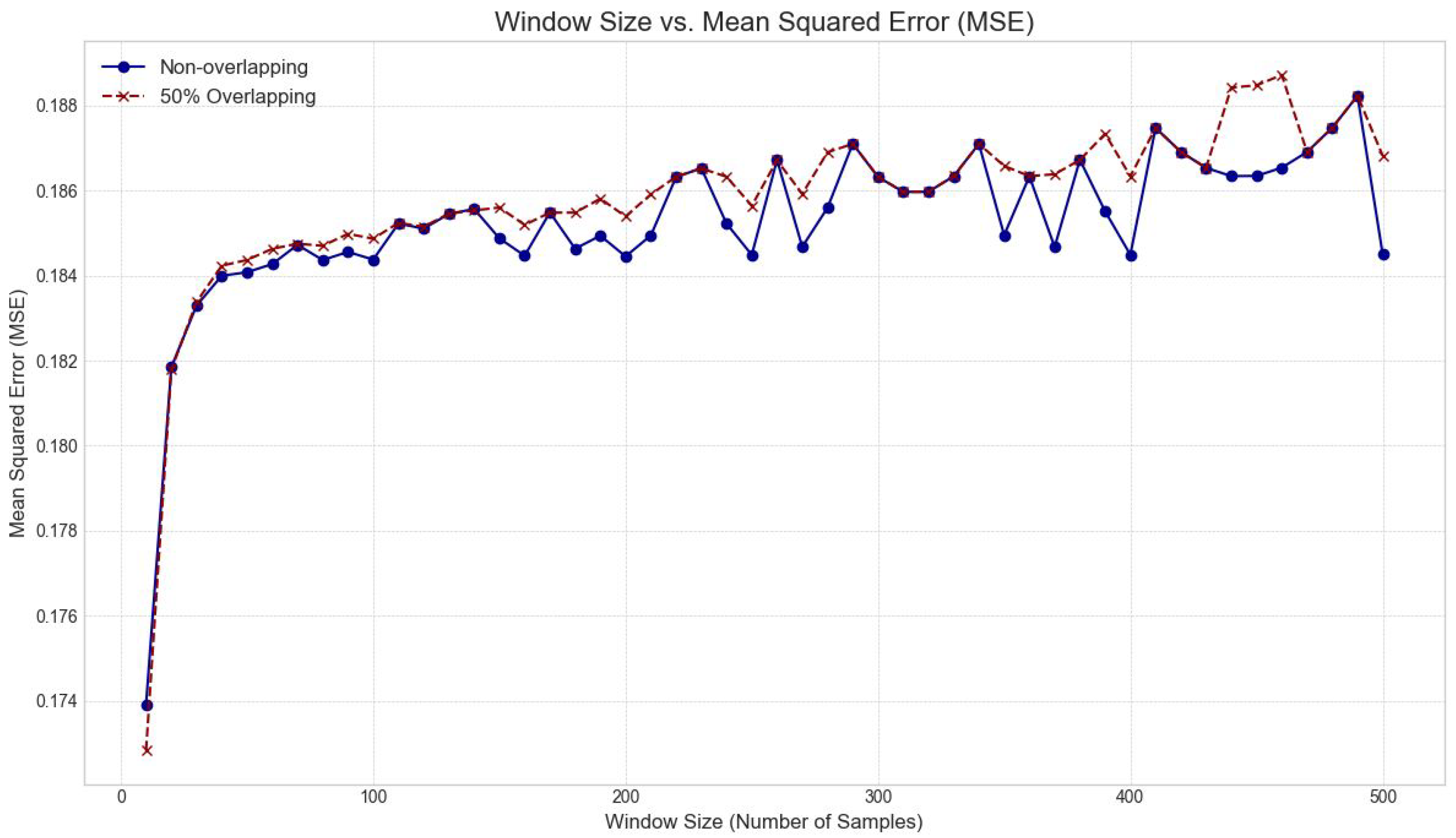
- Optimized Preprocessing Strategies: An in-depth investigation into optimal windowing parameters (window size and overlap percentage) for effective feature extraction and signal segmentation.
- Comprehensive AI Model Comparison (ML & DL): A rigorous comparative study of a diverse suite of traditional machine learning algorithms (Decision Trees, Random Forests, Gradient Boosting, SVM, KNN, LDA, QDA, Extended Associative Memories) and advanced deep learning architectures (1D CNNs, LSTMs, GRUs, Transformers, GANs-discriminator).
- Exploration of Reinforcement Learning for Classification: Presentation of a novel application and performance assessment of Deep Reinforcement Learning agents (DQN, DDPG, Actor-Critic) adapted for the classification of EMG signals, thereby broadening the scope of AI solutions for this domain.
- Performance Insights on Synthetic and Real Data: Practical insights into the most effective methodologies for robust EMG signal interpretation, validated through extensive experimentation on both synthetically generated and real-world EMG datasets.
2. Literature Review
3. Materials and Methods
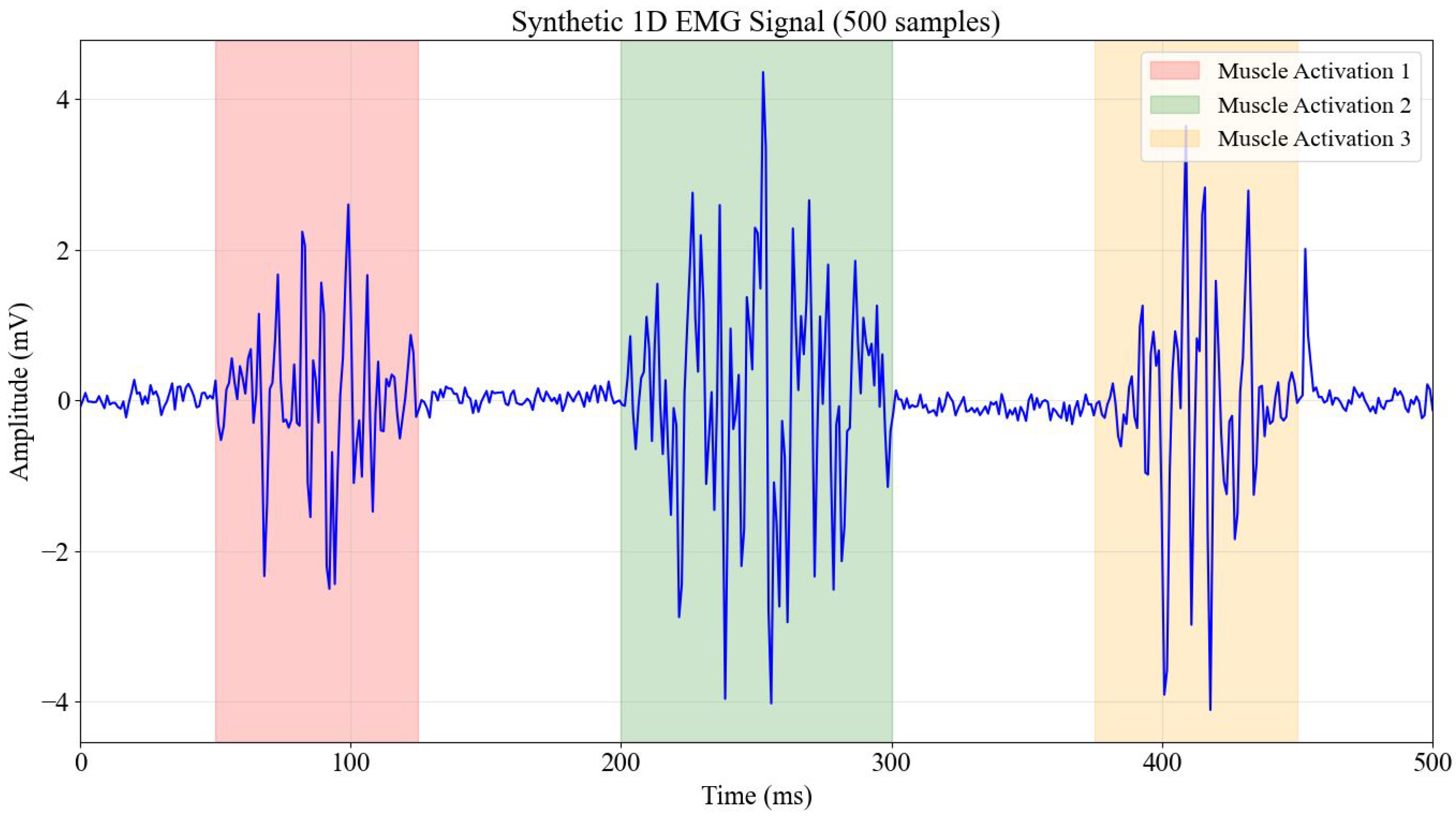
3.1. EMG Signal Generation and Characteristics
- Muscle Activation Bursts: Three distinct periods of muscle activity modulated by Gaussian envelopes simulated the transient nature of muscle contractions, featuring high-frequency content (20–450 Hz) typical of actual EMG signals. This allowed for the evaluation of models’ ability to discern intricate frequency components.
- Baseline Noise: Integrated as random noise to simulate inherent biological noise and artifacts arising from the electrode-skin interface.
- Powerline Interference: A 60 Hz sinusoidal component deliberately introduced to represent common electrical interference.
- Motion Artifacts: Low-frequency components included to simulate artifacts induced by subtle movements, which often confound real EMG recordings.
- Random Spikes: Occasional, sharp random spikes added to mimic artifacts caused by sudden electrode movements or external transient disturbances.
Noise and Artifact Parametrization
- Baseline Noise: White Gaussian noise (sigma = 0.05 mV) was added to simulate the thermal noise floor of the electrode-skin interface and instrumentation amplifier.
- Powerline Interference: A sinusoidal component at 60 Hz with a fixed amplitude of 0.2 mV was introduced to mimic mains hum, a ubiquitous artifact in unshielded acquisition environments.
- Motion Artifacts & Spikes: Low-frequency drift (less than 20 Hz) was modeled using a slow sinusoidal wander (0.1 mV at 1.5 Hz), and random impulsive spikes (amplitude 3.0 mV, probability p = 0.001) were injected to replicate sudden electrode displacements or cable motion.
3.2. Synthetic EMG Data Generation Model
- represents white Gaussian noise with zero mean.
- is the impulse response of a 4th-order Butterworth bandpass filter (20–450 Hz), shaping the noise to match the power spectral density of physiological muscle fiber action potentials.
- is the burst envelope function, modeled as a Gaussian window to simulate the recruitment and de-recruitment phases of muscle contraction.
- and represent the randomized amplitude (2–4 mV) and onset time of the k-th muscle activation burst.
- represents additive artifacts, including 60 Hz powerline interference and baseline thermal noise.
Parameter Selection and Window Optimization
3.3. Signal Acquisition and Pre-Processing Analysis
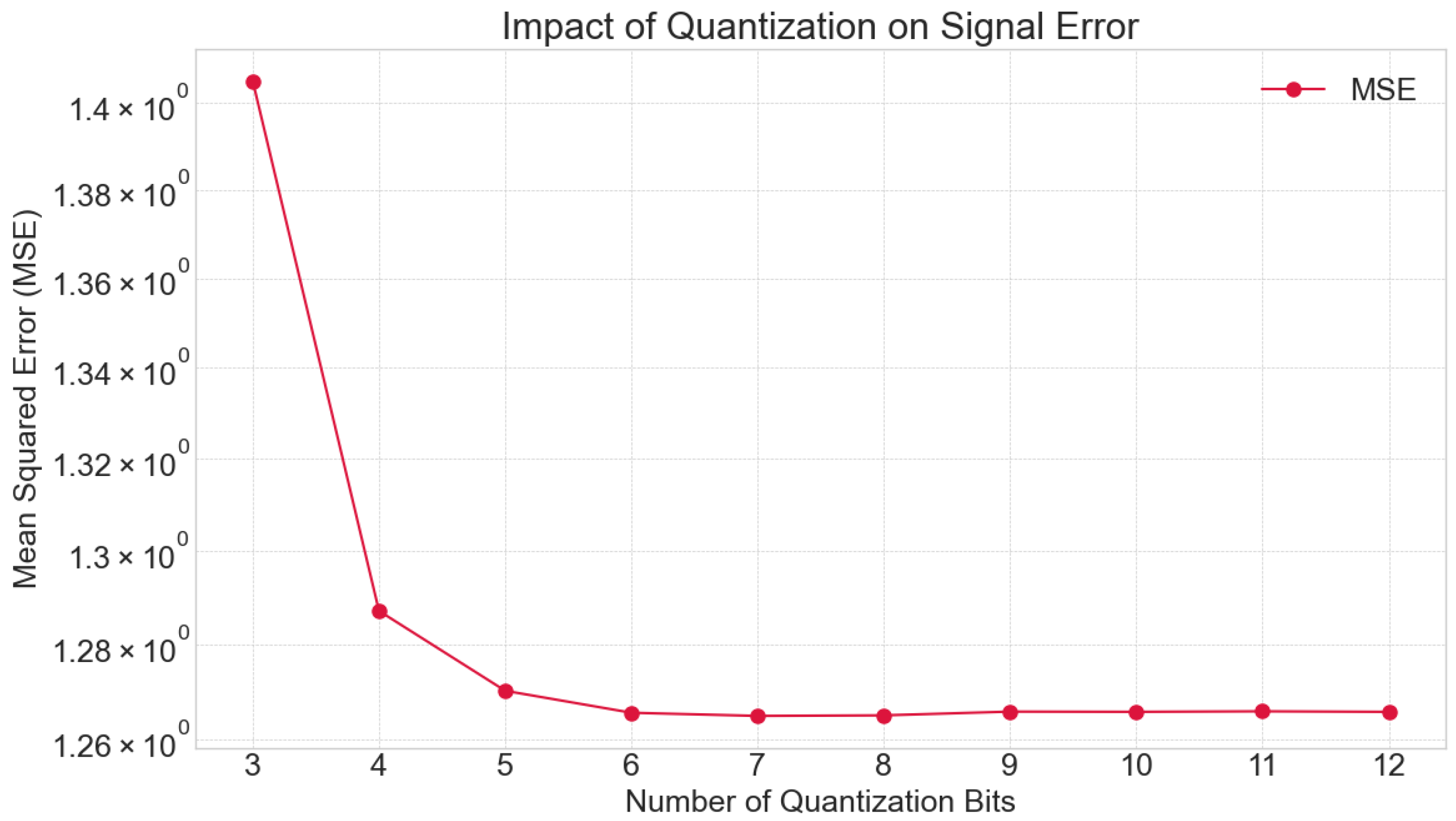
3.3.1. Magnitude Quantization (Bit Depth) Analysis
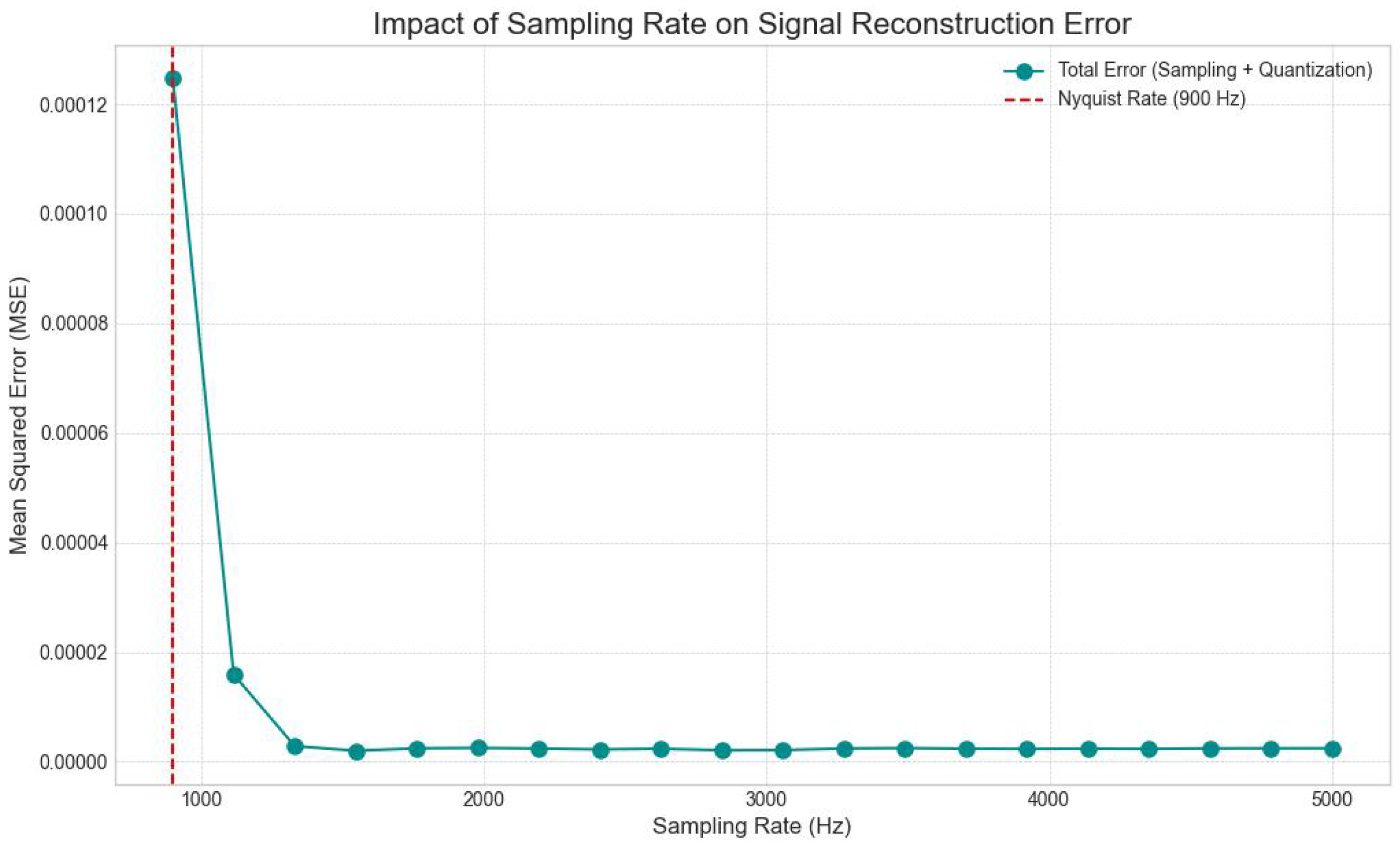
3.3.2. Time Domain Sampling Rate Analysis
3.3.3. Windowing Strategy Optimization
3.4. Final Signal Preprocessing Configuration
3.5. Feature Extraction and Automatic Labeling
- Statistical Features: Mean (average amplitude), Standard Deviation (variability), Max (peak positive amplitude), Min (peak negative amplitude), Variance (spread of data), and Range (difference between max and min).
- Signal Processing Features: Root Mean Square (RMS) (overall signal strength), Energy (sum of squared amplitudes, indicating signal power), Mean Absolute Value (MAV) (average rectified value), and Waveform Length (cumulative length of the waveform, indicating complexity and firing rate).
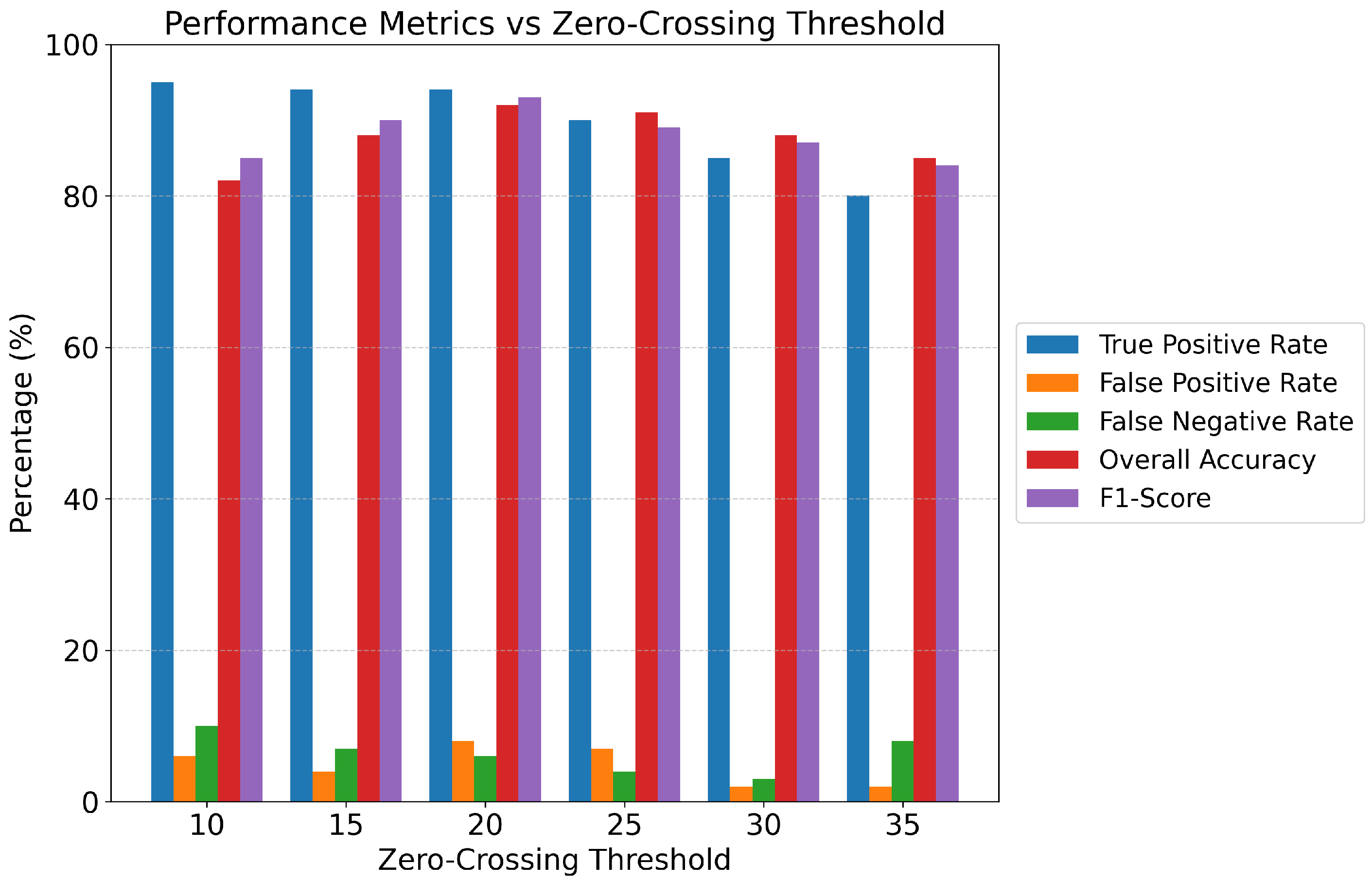
- Activity Indicators: Zero Crossings (number of times the signal crosses the zero amplitude axis, reflecting frequency content and neural drive), and Skewness (asymmetry of the amplitude distribution).
3.6. Machine Learning Model Implementations
- Decision Tree Classifier: A tree-based model that partitions the feature space based on feature values, optimized with a maximum depth of 10 and a minimum of 5 samples required to split an internal node, incorporating built-in overfitting prevention mechanisms.
- Random Forest Classifier: An ensemble method that constructs multiple decision trees during training and outputs the mode of the classes (for classification) of the individual trees, leveraging bagging to reduce variance.
- Gradient Boosting Classifier: Another ensemble method that builds trees sequentially, with each new tree correcting errors made by previous ones, optimizing a differentiable loss function; LightGBM was specifically tested as an efficient implementation of gradient boosting.
- Support Vector Machine (SVM): A powerful discriminative classifier that finds an optimal hyperplane to separate data points into classes, with various kernel functions (e.g., RBF) explored to handle non-linear separability.
- K-Nearest Neighbors (KNN): A non-parametric, instance-based learning algorithm that classifies a data point based on the majority class among its k nearest neighbors in the feature space; optimized parameters for KNN included , using the euclidean distance metric, and uniform weighting for neighbors.
- Linear Discriminant Analysis (LDA): A linear classification algorithm that finds a linear combination of features that characterizes or separates two or more classes, projecting data onto a lower-dimensional space to maximize class separability.
- Quadratic Discriminant Analysis (QDA): Similar to LDA but assumes that the covariance matrices for each class are different, leading to a quadratic decision boundary.
- Extended Associative Memories (EAM): A neural network-inspired model that leverages principles of associative memory for pattern recognition.
3.7. Deep Learning Model Implementations
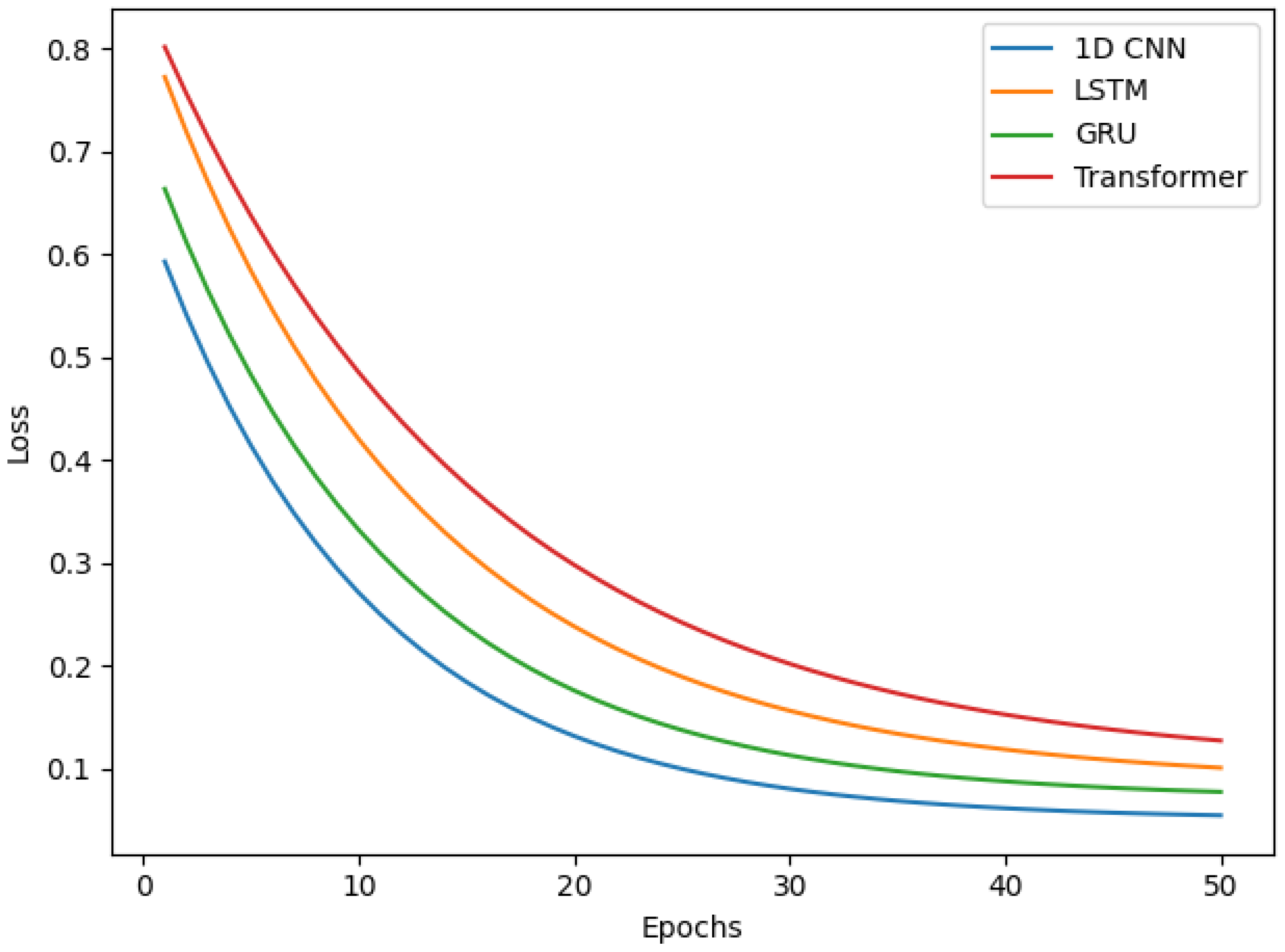
- 1D Convolutional Neural Network (1D CNN): Designed for processing sequential data like 1D EMG signals, its architecture consisted of a Conv1D layer with an output shape of (None, 600, 32) (192 parameters), followed by MaxPooling1D (None, 300, 32) (0 parameters) and Dropout (None, 300, 32) (0 parameters). This was succeeded by another Conv1D (None, 300, 64) (10,304 parameters), MaxPooling1D (None, 150, 64) (0 parameters), and Dropout (None, 150, 64) (0 parameters). A Flatten layer (None, 9600) (0 parameters) then connected to a Dense layer (None, 100) (960,100 parameters), culminating in a final Dense layer for binary classification (None, 1) (101 parameters) with sigmoid activation, or (None, NumClasses) with softmax activation for multiclass tasks.
- Recurrent Neural Networks (RNNs)-LSTM and GRU: Employed for their ability to capture long-range temporal dependencies in sequential data.
- –
- LSTM Architecture: Comprised an initial LSTM layer (None, 600, 64) (16,896 parameters), followed by Dropout (None, 600, 64) (0 parameters), another LSTM layer (None, 64) (33,024 parameters), Dropout (None, 64) (0 parameters), a Dense layer (None, 100) (6500 parameters), and a final Dense layer (None, 1) (101 parameters).
- –
- GRU Architecture: Similarly featured an initial GRU layer (None, 600, 64) (12,864 parameters), Dropout (None, 600, 64) (0 parameters), a second GRU layer (None, 64) (24,960 parameters), Dropout (None, 64) (0 parameters), a Dense layer (None, 100) (6500 parameters), and a final Dense layer (None, 1) (101 parameters).
- Transformers: Leveraging self-attention mechanisms, they were implemented to evaluate their efficacy in capturing global dependencies within EMG sequences. The specific architecture involved a stacked encoder setup for sequence classification.
- Generative Adversarial Network (GAN) Discriminator for Classification: A GAN framework was explored, specifically utilizing the discriminator component for classification, trained to distinguish between real and synthetically generated EMG signals. Its final layers were adapted for binary classification (EMG Activity vs. No EMG Activity), with the discriminator network architecture consisting of: Conv1D (None, 593, 32) (288 parameters), MaxPooling1D (None, 148, 32) (0 parameters), Conv1D (None, 144, 64) (10,304 parameters), MaxPooling1D (None, 36, 64) (0 parameters), Flatten (None, 2304) (0 parameters), Dense (None, 64) (147,520 parameters), and a final Dense layer (None, 2) (130 parameters) for classification output.
Reinforcement Learning (RL) Agents for Classification
3.8. Machine Learning and Deep Learning Architectures
3.9. Data Splitting and Evaluation Metrics
- Accuracy Score: The proportion of correctly classified instances.
- Classification Report: Providing Precision (proportion of true positive predictions among all positive predictions), Recall (Sensitivity, proportion of true positive predictions among all actual positive instances), and F1-score (harmonic mean of Precision and Recall) for each class, as well as macro and weighted averages.
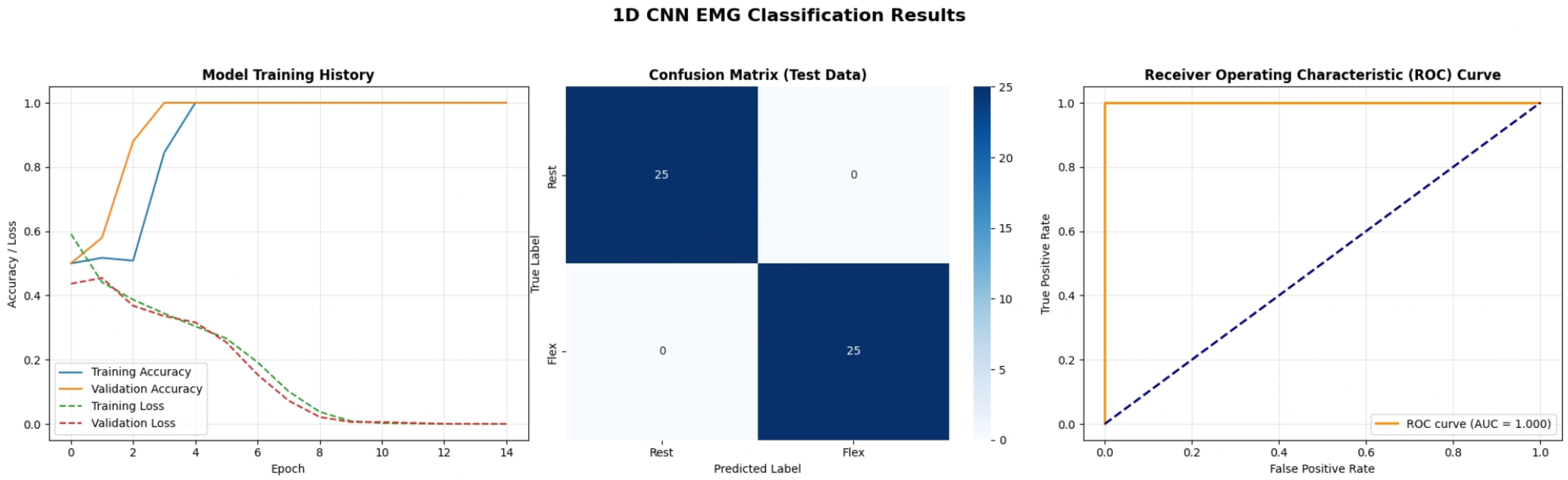
- Confusion Matrix: A table summarizing the number of true positive, true negative, false positive, and false negative predictions, providing a detailed breakdown of classifier performance.
- Receiver Operating Characteristic (ROC) Curve and Area Under the Curve (AUC): Visualizing the trade-off between the True Positive Rate (TPR) and False Positive Rate (FPR) at various threshold settings, with AUC providing an aggregate measure of separability.
3.10. Real EMG Data
4. Results and Discussion
4.1. Impact of Signal Acquisition and Preprocessing Parameters
4.2. Comparative Performance of Machine Learning and Deep Learning Models
4.3. Implications and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Abou-Zahra, Y.; Hassan, M.; Chen, J. Machine Learning- and Deep Learning-Based Myoelectric Control System for Upper Limb Rehabilitation Utilizing EEG and EMG Signals: A Systematic Review. Bioengineering 2025, 12, 144. [Google Scholar] [CrossRef]
- Al-Ayyad, M.; Owida, H.A.; De Fazio, R.; Al-Naami, B.; Visconti, P. Electromyography monitoring systems in rehabilitation: A review of clinical applications, wearable devices and signal acquisition methodologies. Electronics 2023, 12, 1520. [Google Scholar] [CrossRef]
- Ashburner, J.; Klöppel, S. Multivariate models of inter-subject anatomical variability. Neuroimage 2011, 56, 422–439. [Google Scholar] [CrossRef]
- Saleh, A.; Hameed, H.; Najafi, B.; Soleimani, M. Machine Learning-Based Feature Extraction and Classification of EMG Signals for Intuitive Prosthetic Control. Appl. Sci. 2024, 14, 5784. [Google Scholar] [CrossRef]
- Ahmed, M.R.; Newby, S.; Potluri, P.; Mirihanage, W.; Fernando, A. Emerging paradigms in fetal heart rate monitoring: Evaluating the efficacy and application of innovative textile-based wearables. Sensors 2024, 24, 6066. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhang, X.; Cheng, Y.; Xi, N. Deep Learning for EMG-based Human-Machine Interaction: A Review. IEEE CAA J. Autom. Sin. 2021, 8, 777–796. [Google Scholar]
- Hassan, H.F.; Abou-Loukh, S.J.; Ibraheem, I.K. EMG Pattern Recognition in the Era of Big Data and Deep Learning. Big Data Cogn. Comput. 2018, 2, 21. [Google Scholar] [CrossRef]
- Yousefi, J.; Hamilton-Wright, A. Characterizing EMG data using machine-learning tools. Comput. Biol. Med. 2014, 51, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Rani, G.J.; Hashmi, M.F.; Gupta, A. Surface electromyography and artificial intelligence for human activity recognition—A systematic review on methods, emerging trends applications, challenges, and future implementation. IEEE Access 2023, 11, 105140–105169. [Google Scholar] [CrossRef]
- Yousif, H.A.; Zakaria, A.; Rahim, N.A.; Salleh, A.F.B.; Mahmood, M.; Alfarhan, K.A.; Kamarudin, L.M.; Mamduh, S.M.; Hasan, A.M.; Hussain, M.K. Assessment of muscles fatigue based on surface EMG signals using machine learning and statistical approaches: A review. In IOP Conference Series: Materials Science and Engineering; IOP Publishing: Bristol, UK, 2019; Volume 705, p. 012010. [Google Scholar]
- Di Nardo, F.; Nocera, A.; Cucchiarelli, A.; Fioretti, S.; Morbidoni, C. Machine learning for detection of muscular activity from surface EMG signals. Sensors 2022, 22, 3393. [Google Scholar] [CrossRef]
- Yaman, E.; Subasi, A. Comparison of bagging and boosting ensemble machine learning methods for automated EMG signal classification. BioMed Res. Int. 2019, 2019, 9152506. [Google Scholar] [CrossRef]
- Khan, M.U.; Khan, H.; Muneeb, M.; Abbasi, Z.; Abbasi, U.B.; Baloch, N.K. Supervised machine learning based fast hand gesture recognition and classification using electromyography (EMG) signals. In Proceedings of the 2021 International Conference on Applied and Engineering Mathematics (ICAEM), Taxila, Pakistan, 30–31 August 2021; IEEE: New York, NY, USA, 2021; pp. 81–86. [Google Scholar]
- Zhou, Y.; Chen, C.; Ni, J.; Ni, G.; Li, M.; Xu, G.; Cavanaugh, J.; Cheng, M.; Lemos, S. EMG signal processing for hand motion pattern recognition using machine learning algorithms. Arch. Orthop. 2020, 1, 17–26. [Google Scholar]
- Mokri, C.; Bamdad, M.; Abolghasemi, V. Muscle force estimation from lower limb EMG signals using novel optimised machine learning techniques. Med. Biol. Eng. Comput. 2022, 60, 683–699. [Google Scholar] [CrossRef]
- Zhou, Y.; Chen, C.; Cheng, M.; Alshahrani, Y.; Franovic, S.; Lau, E.; Xu, G.; Ni, G.; Cavanaugh, J.M.; Muh, S.; et al. Comparison of machine learning methods in sEMG signal processing for shoulder motion recognition. Biomed. Signal Process. Control 2021, 68, 102577. [Google Scholar] [CrossRef]
- Jaramillo, A.G.; Benalcazar, M.E. Real-time hand gesture recognition with EMG using machine learning. In Proceedings of the 2017 IEEE Second Ecuador Technical Chapters Meeting (ETCM), Salinas, Ecuador, 16–20 October 2017; IEEE: New York, NY, USA, 2017; pp. 1–5. [Google Scholar]
- Ali, S.; Ahmad, N.; Khan, M.R.; Shah, S.A. EMG Signal Processing for Hand Motion Pattern Recognition Using Machine Learning Algorithms. Sci. Arch. 2024, 5, 125–140. [Google Scholar]
- Haque, F.; Nandy, A.; Zadeh, P.M. Surface EMG Signal Classification by Using WPD and Ensemble Tree Classifiers. In Proceedings of the International Conference on Computational Intelligence and Data Engineering; Springer: Berlin/Heidelberg, Germany, 2017; pp. 475–484. [Google Scholar]
- Mahmood, N.T.; Al-Muifraje, M.H.; Salih, S.K.; Saeed, T.R. Pattern recognition of composite motions based on emg signal via machine learning. Eng. Technol. J. 2021, 39, 295–305. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, R.; Chen, W.; Xiong, C. Advances of Machine Learning in Electromyography (EMG) Signal Classification. Biomed. Signal Process. Control 2021, 68, 102796. [Google Scholar]
- Nazmi, N.; Rahman, M.A.A.; Yamamoto, S.I.; Ahmad, S.A. Processing Surface EMG Signals for Exoskeleton Motion Control. Front. Neurorobotics 2020, 14, 40. [Google Scholar] [CrossRef] [PubMed]
- Phinyomark, A.; Khushaba, R.N.; Ibánez-Marcelo, E.; Petri, A.; St-Onge, S.; Scheme, E. Interpreting Deep Learning Features for Myoelectric Control: A Comparison with Handcrafted Features. Front. Bioeng. Biotechnol. 2020, 8, 158. [Google Scholar] [CrossRef]
- Turgunov, A.; Zohirov, K.; Nasimov, R.; Mirzakhalilov, S. Comparative analysis of the results of EMG signal classification based on machine learning algorithms. In Proceedings of the 2021 International Conference on Information Science and Communications Technologies (ICISCT), Tashkent, Uzbekistan, 3–5 November 2021; IEEE: New York, NY, USA, 2021; pp. 1–4. [Google Scholar]
- Pérez-Reynoso, F.; Farrera-Vazquez, N.; Capetillo, C.; Méndez-Lozano, N.; González-Gutiérrez, C.; López-Neri, E. Pattern recognition of EMG signals by machine learning for the control of a manipulator robot. Sensors 2022, 22, 3424. [Google Scholar] [CrossRef]
- Sonmezocak, T.; Kurt, S. Machine learning and regression analysis for diagnosis of bruxism by using EMG signals of jaw muscles. Biomed. Signal Process. Control 2021, 69, 102905. [Google Scholar] [CrossRef]
- Shankar, H.; Nair, S.; Chow, C.; Daud, W.; Azizan, M. A Machine Learning System for Classification of EMG Signals to Assist Exoskeleton Performance. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Washington, DC, USA, 9–11 October 2018; IEEE: New York, NY, USA, 2019; pp. 3309–3312. [Google Scholar]
- Nurhanim, K.; Elamvazuthi, I.; Izhar, L.; Capi, G.; Su, S. EMG signals classification on human activity recognition using machine learning algorithm. In Proceedings of the 2021 8th NAFOSTED Conference on Information and Computer Science (NICS), Hanoi, Vietnam, 21–22 December 2021; IEEE: New York, NY, USA, 2021; pp. 369–373. [Google Scholar]
- Amamcherla, N.; Turlapaty, A.; Gokaraju, B. A machine learning system for classification of emg signals to assist exoskeleton performance. In Proceedings of the 2018 IEEE Applied Imagery Pattern Recognition Workshop (AIPR), Washington, DC, USA, 9–11 October 2018; IEEE: New York, NY, USA, 2018; pp. 1–4. [Google Scholar]
- Khairuddin, I.M.; Sidek, S.N.; Majeed, A.P.A.; Razman, M.A.M.; Puzi, A.A.; Yusof, H.M. The classification of movement intention through machine learning models: The identification of significant time-domain EMG features. PeerJ Comput. Sci. 2021, 7, e379. [Google Scholar] [CrossRef] [PubMed]
- Tsinganos, P.; Cornelis, B.; Cornelis, J.; Jansen, B.; Skodras, A. Multiday EMG-Based Classification of Hand Motions with Deep Learning Techniques. Sensors 2018, 18, 2497. [Google Scholar] [CrossRef] [PubMed]














| Model Family | Algorithm/Architecture | Key Hyperparameters & Configuration |
|---|---|---|
| Ensemble ML | Random Forest | 50 Trees, Gini impurity, Bootstrap sampling |
| Gradient Boosting (LightGBM) | Learning rate = 0.1, Max depth = 3, Seq. correction | |
| Classical ML | Decision Tree | Max depth = 10, Min samples split = 5 |
| SVM | Kernel = RBF, , Gamma = Scale | |
| KNN | , Metric = Euclidean, Weights = Uniform | |
| LDA/QDA | SVD solver/SVD solver (no priors) | |
| 1D CNN | Input(600) → Conv1D(32) → MaxPool → Conv1D(64) → MaxPool → Dense(100) → Output | Filters = 32/64, Kernel = 3, Pool = 2, Activation = ReLU, Dropout = 0.3 |
| RNN (LSTM) | Input(600) → LSTM(64) → Dropout → LSTM(64) → Dense(100) → Output | Units = 64, Return Sequences = True/False, Activation = Tanh, Dropout = 0.3 |
| RNN (GRU) | Input(600) → GRU(64) → Dropout → GRU(64) → Dense(100) → Output | Units = 64, Reset_after = True, Activation = Tanh, Dropout = 0.3 |
| Transformer | Input → Pos. Encoding → Multi-Head Attn → Add & Norm → Feed Forward → Output | Heads = 4, Key_dim = 64, FF_dim = 128, Dropout = 0.1 |
| Model | Params | Model Size (FP32) | Inf. Time (CPU) | Inf. Time (GPU) | Train Time | Memory Footprint | Suitability |
|---|---|---|---|---|---|---|---|
| Decision Tree | N/A | ∼15 KB | 0.8 ms | N/A | <1 min | Low | Microcontroller |
| Random Forest | N/A | ∼850 KB | 4.2 ms | N/A | 2 min | Moderate | Edge Device |
| Gradient Boost | N/A | ∼1.2 MB | 5.8 ms | N/A | 3.5 min | Moderate | Edge Device |
| SVM | N/A | ∼180 KB | 2.1 ms | N/A | 1.5 min | Low | Microcontroller |
| KNN | N/A | ∼2.5 MB | 12.3 ms | N/A | <1 min | High | Impractical |
| 1D CNN | 970,697 | 3.7 MB | 8.5 ms | 2.1 ms | 3 min | Moderate | Edge/GPU |
| LSTM | 56,421 | 220 KB | 6.3 ms | 1.8 ms | 5 min | Moderate | Edge Device |
| GRU | 44,425 | 175 KB | 5.1 ms | 1.5 ms | 4.5 min | Moderate | Edge Device |
| Transformer | 1,800,000 | 7.2 MB | 18.7 ms | 4.2 ms | 8 min | High | Cloud/Server |
| DQN (RL) | 304,386 | 1.2 MB | 9.8 ms | 2.5 ms | 45 min | Moderate | Edge Device |
| Study/Architecture | Dataset | Classes | Freq. (Hz) | Res. | Acc. (%) |
|---|---|---|---|---|---|
| Proposed (Voting Ensemble) | Real EMG | 3 | 2000 | 10-bit | 100.0 |
| Proposed (LSTM) | Real EMG | 3 | 2000 | 10-bit | 96.3 |
| Saleh et al. [4] (CNN-Voting) | Ninapro DB4 | 52 | 2000 | 12-bit | 89.5 |
| Tsinganos et al. [31] (CNN) | Myo Dataset | 7 | 200 | 8-bit | 97.8 |
| Baspinar et al. [12] (Ensemble) | Ninapro DB1 | 10 | 100 | 12-bit | 94.8 |
| Nazmi et al. [22] (Review) | Exoskeleton | Various | 1000 | – | 90–95 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Shinde, A.; Shete, V.; Mehendale, N. Optimized Signal Acquisition and Advanced AI for Robust 1D EMG Classification: A Comparative Study of Machine Learning, Deep Learning, and Reinforcement Learning. Bioengineering 2026, 13, 463. https://doi.org/10.3390/bioengineering13040463
Shinde A, Shete V, Mehendale N. Optimized Signal Acquisition and Advanced AI for Robust 1D EMG Classification: A Comparative Study of Machine Learning, Deep Learning, and Reinforcement Learning. Bioengineering. 2026; 13(4):463. https://doi.org/10.3390/bioengineering13040463
Chicago/Turabian StyleShinde, Anagha, Virendra Shete, and Ninad Mehendale. 2026. "Optimized Signal Acquisition and Advanced AI for Robust 1D EMG Classification: A Comparative Study of Machine Learning, Deep Learning, and Reinforcement Learning" Bioengineering 13, no. 4: 463. https://doi.org/10.3390/bioengineering13040463
APA StyleShinde, A., Shete, V., & Mehendale, N. (2026). Optimized Signal Acquisition and Advanced AI for Robust 1D EMG Classification: A Comparative Study of Machine Learning, Deep Learning, and Reinforcement Learning. Bioengineering, 13(4), 463. https://doi.org/10.3390/bioengineering13040463








