ERG-Graph: Graph Signal Processing of the Electroretinogram for Classification of Neurodevelopmental Disorders
Abstract
1. Introduction
- (1)
- To develop and optimize the ERG-Graph construction methodology, including the adaptation of amplitude quantization and graph generation to the ERG waveform, the systematic evaluation of quantization resolution, and the comparison of multiple graph construction strategies (quantization-based, visibility graph, recurrence network, k-nearest neighbor, -ball, and ordinal partition networks).
- (2)
- To extract, evaluate, and compare graph-theoretic features across centrality, topology, clustering, and spectral domains for their ability to differentiate ASD, ADHD, and control groups across multiple classification scenarios (two-group, three-group, and four-group), sex strata, flash strengths, and eye laterality conditions.
- (3)
- To demonstrate that ERG-Graph features provide complementary and superior classification performance compared to traditional time-domain and time–frequency (DWT, VFCDM) features, both independently and in fusion, thereby advancing objective biomarkers for neurodevelopmental disorder screening.
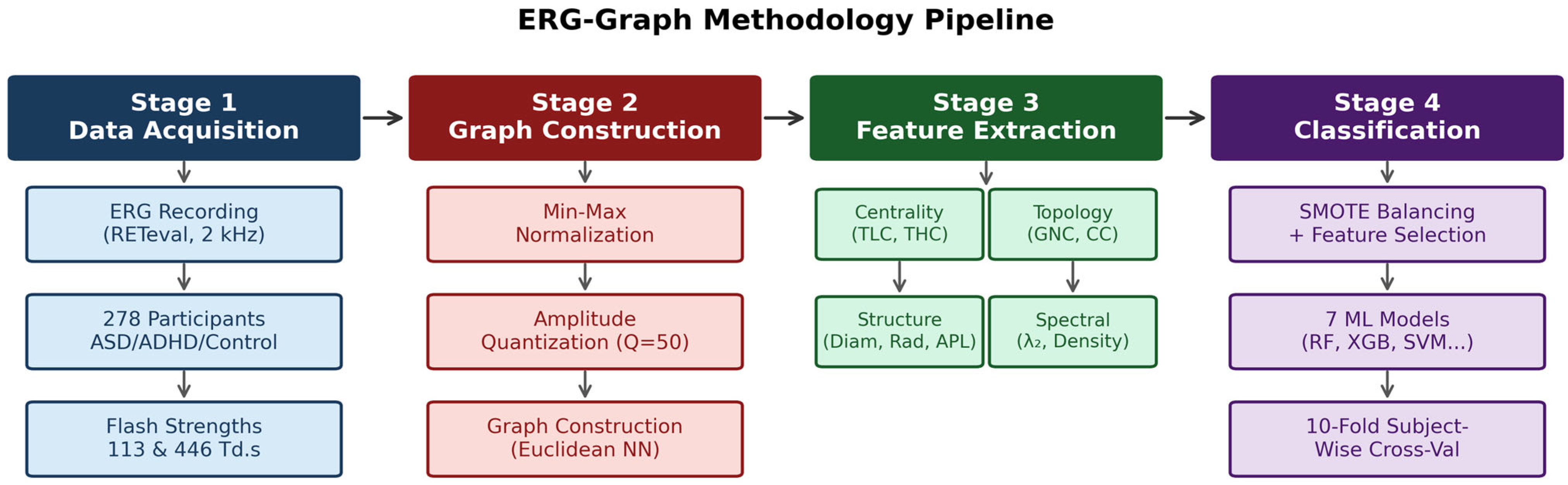
2. Materials and Methods
2.1. Participants and Electrophysiology
2.2. ERG-Graph Construction
2.3. Alternative Graph Construction Methods
2.4. Graph-Theoretic Feature Extraction
- (1)
- Total Load Centrality (TLC). Load centrality quantifies the fraction of shortest paths passing through each node. For a node v, the load centrality is the proportion of all shortest paths between pairs of other nodes that pass through v [59,67]. TLC sums this measure over all nodes:where is the total number of shortest paths from node s to node t, and is the number of those paths passing through v. Higher TLC indicates that more amplitude states serve as obligatory transitions, reflecting waveform complexity.
- (2)
- Total Harmonic Centrality (THC). Harmonic centrality handles disconnected components gracefully by using inverse distances [68]. THC aggregates the harmonic closeness across all nodes:where is the shortest-path distance between nodes v and u. THC reflects global reachability within the graph: signals with diverse amplitude transitions produce graphs where nodes are mutually accessible through short paths.
- (3)
- Number of Cliques (GNC). A clique is a maximally complete subgraph in which every pair of nodes is connected [69]. The number of maximal cliques is:
- (4)
- (5)
- Graph Radius. The radius is the minimum eccentricity over all nodes, where eccentricity = max() for all u [60]:
- (6)
- Average Clustering Coefficient (CC). The local clustering coefficient measures the tendency of a node’s neighbors to be interconnected [70]. The global average is:where is the degree of node v and are edges between neighbors of v. High CC indicates that amplitude levels visited in sequence tend to form closed triangles, reflecting localized oscillatory dwelling.
- (7)
- (8)
- Algebraic Connectivity (λ2). The second-smallest eigenvalue of the graph Laplacian L, known as the Fiedler value, measures the robustness of graph connectivity [45,71,72]:where is the first (constant) eigenvector. A higher indicates that the graph is tightly connected with few structural bottlenecks separating amplitude regions. In the ERG context, ASD graphs exhibit elevated because the signal remains within a compact amplitude range, producing redundant transitions between nearby levels. Conversely, ADHD graphs show low because the signal traverses a broader amplitude range, creating a more fragmented trajectory through amplitude space.
- (9)
- Graph Density (ρ). The ratio of observed edges to the maximum possible (Equation (5)). Dense graphs indicate that many different amplitude transitions occur, while sparse graphs indicate a constrained trajectory [60].
2.5. Classification Scenarios
2.6. Machine Learning Pipeline and Statistical Analysis
3. Results
3.1. Quantization Resolution Optimization
3.2. Graph Structure Analysis
3.3. Two-Group Classification Performance
3.4. Three-Group Classification
3.5. Four-Group Classification
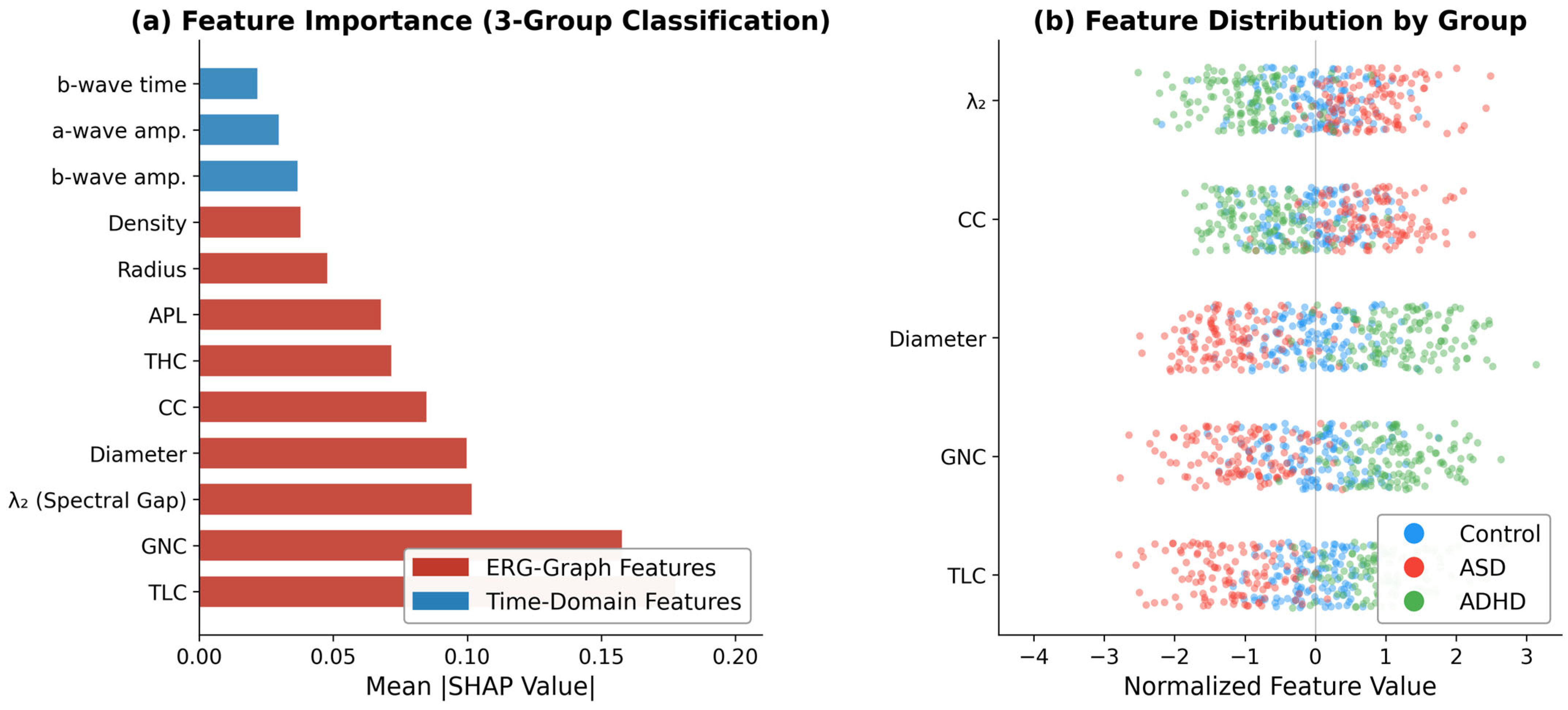
3.6. Feature Importance
3.7. Graph Construction Variant Comparison
| Study | Features | Classification Scenario | Best BA | n | Notes |
|---|---|---|---|---|---|
| Posada-Quintero et al. [10,11] | VFCDM spectral | ASD vs. Control (binary) | Sens. 0.85 /Spec. 0.78 | 278 | Sensitivity/specificity reported |
| Constable et al. [3] | TD + DWT | ASD/ADHD vs. Control | — | 278 | Wavelet energy bands; BA not reported |
| Manjur et al. [13] | TD + DWT + VFCDM | Three-group (ASD/ADHD/Ctrl) | 0.70 | 278 | Previous three-group benchmark |
| Constable et al. [12] | TD + DWT + VFCDM | ASD vs. Ctrl (males)/ADHD vs. Ctrl (females) | 0.87/0.84 | 278 | Four-group BA = 0.53 |
| This study (ERG-Graph only) | GSP graph features | ASD vs. Ctrl (males)/ADHD vs. Ctrl (females) | 0.91/0.88 | 278 | 9 topological + spectral features |
| This study (ERG-Graph + TD) | GSP + TD fusion | Three-group (ASD/ADHD/Ctrl) | 0.81 | 278 | +11 pp vs. prior benchmark; four-group BA = 0.67 |
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Constable, P.A.; Ritvo, E.R.; Ritvo, A.R.; Lee, I.O.; McNair, M.L.; Stahl, D.; Sowden, J.; Quinn, S.; Skuse, D.H.; Thompson, D.A.; et al. Light-adapted electroretinogram differences in autism spectrum disorder. J. Autism Dev. Disord. 2020, 50, 2874–2885. [Google Scholar] [CrossRef]
- Lee, I.O.; Skuse, D.H.; Constable, P.A.; Marmolejo-Ramos, F.; Olsen, L.R.; Thompson, D.A. The electroretinogram b-wave amplitude: A differential measure for neurodevelopmental disorders. J. Neurodev. Disord. 2022, 14, 30. [Google Scholar] [CrossRef] [PubMed]
- Constable, P.A.; Marmolejo-Ramos, F.; Gauthier, M.; Lee, I.O.; Skuse, D.H.; Thompson, D.A. Discrete wavelet transform analysis of the electroretinogram in autism spectrum disorder and attention deficit hyperactivity disorder. Front. Neurosci. 2022, 16, 890461. [Google Scholar] [CrossRef] [PubMed]
- Lavoie, J.; Illiano, P.; Sotnikova, T.D.; Gainetdinov, R.R.; Beaulieu, J.M.; Hébert, M. The electroretinogram as a biomarker of central dopamine and serotonin. Biol. Psychiatry 2014, 75, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Youssef, P.; Nath, S.; Chaimowitz, G.A.; Prat, S.S. Electroretinography in psychiatry: A systematic literature review. Eur. Psychiatry 2019, 62, 97–106. [Google Scholar] [CrossRef]
- Ritvo, E.R.; Creel, D.; Realmuto, G.; Crandall, A.S.; Freeman, B.J.; Bateman, J.B.; Barr, R.; Pingree, C.; Coleman, M.; Purple, R. Electroretinograms in autism: A pilot study of b-wave amplitudes. Am. J. Psychiatry 1988, 145, 229–232. [Google Scholar] [CrossRef]
- Constable, P.A.; Gaigg, S.B.; Bowler, D.M.; Jägle, H.; Thompson, D.A. Full-field electroretinogram in autism spectrum disorder. Doc. Ophthalmol. 2016, 132, 83–99. [Google Scholar] [CrossRef]
- Friedel, E.B.N.; Schäfer, M.; Endres, D.; Maier, S.; Runge, K.; Bach, M.; Heinrich, S.P.; Ebert, D.; Domschke, K.; Tebartz van Elst, L.; et al. Electroretinography in adults with high-functioning autism spectrum disorder. Autism Res. 2022, 15, 2026–2037. [Google Scholar] [CrossRef]
- Constable, P.A.; Lee, I.O.; Marmolejo-Ramos, F.; Skuse, D.H.; Thompson, D.A. The photopic negative response in autism spectrum disorder. Clin. Exp. Optom. 2021, 104, 841–847. [Google Scholar] [CrossRef]
- Posada-Quintero, H.F.; Manjur, S.M.; Hossain, M.B.; Marmolejo-Ramos, F.; Lee, I.O.; Skuse, D.H.; Thompson, D.A.; Constable, P.A. Autism spectrum disorder detection using variable-frequency complex demodulation of the electroretinogram. Res. Autism Spectr. Disord. 2023, 109, 102258. [Google Scholar] [CrossRef]
- Mohammad-Manjur, S.; Hossain, M.-B.; Constable, P.A.; Thompson, D.A.; Marmolejo-Ramos, F.; Lee, I.O.; Skuse, D.; Posada-Quintero, H.F. Detecting autism spectrum disorder using spectral analysis of electroretinogram and machine learning. In Proceedings of the 44th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Glasgow, Scotland, UK; Institute of Electrical and Electronics Engineers (IEEE): New York, NY, USA, 2022; pp. 3435–3438. [Google Scholar]
- Constable, P.A.; Pinzon-Arenas, J.O.; Mercado Diaz, L.R.; Lee, I.O.; Marmolejo-Ramos, F.; Loh, L.; Posada-Quintero, H. Spectral analysis of light-adapted electroretinograms in neurodevelopmental disorders: Classification with machine learning. Bioengineering 2024, 12, 15. [Google Scholar] [CrossRef]
- Manjur, S.M.; Diaz, L.R.M.; Lee, I.O.; Skuse, D.H.; Thompson, D.A.; Marmolejos-Ramos, F.; Constable, P.A.; Posada-Quintero, H.F. Detecting autism spectrum disorder and attention deficit hyperactivity disorder using multimodal time-frequency analysis with machine learning using the electroretinogram. J. Autism Dev. Disord. 2025, 55, 1365–1378. [Google Scholar] [CrossRef]
- Micai, M.; Fatta, L.M.; Gila, L.; Caruso, A.; Salvitti, T.; Fulceri, F.; Ciaramella, A.; D’Amico, R.; Del Giovane, C.; Bertelli, M.; et al. Prevalence of co-occurring conditions in children and adults with autism spectrum disorder: A systematic review and meta-analysis. Neurosci. Biobehav. Rev. 2023, 155, 105436. [Google Scholar] [CrossRef]
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 5th ed.; American Psychiatric Publishing: Arlington, VA, USA, 2013. [Google Scholar]
- Faraone, S.V.; Asherson, P.; Banaschewski, T.; Biederman, J.; Buitelaar, J.K.; Ramos-Quiroga, J.A.; Rohde, L.A.; Sonuga-Barke, E.J.; Tannock, R.; Franke, B. Attention-deficit/hyperactivity disorder. Nat. Rev. Dis. Primers 2015, 1, 15020. [Google Scholar] [CrossRef]
- Zeidan, J.; Fombonne, E.; Scorah, J.; Ibrahim, A.; Durkin, M.S.; Saxena, S.; Yusuf, A.; Shih, A.; Elsabbagh, M. Global prevalence of autism: A systematic review update. Autism Res. 2022, 15, 778–790. [Google Scholar] [CrossRef]
- Polanczyk, G.; de Lima, M.S.; Horta, B.L.; Biederman, J.; Rohde, L.A. The worldwide prevalence of ADHD: A systematic review and metaregression analysis. Am. J. Psychiatry 2007, 164, 942–948. [Google Scholar] [CrossRef]
- Leitner, Y. The co-occurrence of autism and attention deficit hyperactivity disorder in children. Front. Hum. Neurosci. 2014, 8, 268. [Google Scholar] [CrossRef]
- Rommelse, N.N.; Franke, B.; Geurts, H.M.; Hartman, C.A.; Buitelaar, J.K. Shared heritability of attention-deficit/hyperactivity disorder and autism spectrum disorder. Eur. Child Adolesc. Psychiatry 2010, 19, 281–295. [Google Scholar] [CrossRef]
- Lord, C.; Elsabbagh, M.; Baird, G.; Veenstra-Vanderweele, J. Autism spectrum disorder. Lancet 2018, 392, 508–520. [Google Scholar] [CrossRef]
- Thapar, A.; Cooper, M. Attention deficit hyperactivity disorder. Lancet 2016, 387, 1240–1250. [Google Scholar] [CrossRef]
- Zwaigenbaum, L.; Bauman, M.L.; Stone, W.L.; Yirmiya, N.; Estes, A.; Hansen, R.L.; McPartland, J.C.; Natowicz, M.R.; Choueiri, R.; Fein, D.; et al. Early identification of autism spectrum disorder: Recommendations for practice and research. Pediatrics 2015, 136, S10–S40. [Google Scholar] [CrossRef]
- Dawson, G. Early behavioral intervention, brain plasticity, and the prevention of autism spectrum disorder. Dev. Psychopathol. 2008, 20, 775–803. [Google Scholar] [CrossRef]
- Constable, P.A.; Lim, J.K.H.; Thompson, D.A. Retinal electrophysiology in central nervous system disorders: A review of human and mouse studies. Front. Neurosci. 2023, 17, 1215097. [Google Scholar] [CrossRef]
- Robson, A.G.; Frishman, L.J.; Grigg, J.; Hamilton, R.; Jeffrey, B.G.; Kondo, M.; Li, S.; McCulloch, D.L. ISCEV standard for full-field clinical electroretinography (2022 update). Doc. Ophthalmol. 2022, 144, 165–177. [Google Scholar] [CrossRef] [PubMed]
- London, A.; Benhar, I.; Schwartz, M. The retina as a window to the brain—From eye research to CNS disorders. Nat. Rev. Neurol. 2013, 9, 44–53. [Google Scholar] [CrossRef]
- Dowling, J. The Retina: An Approachable Part of the Brain; Harvard University Press: Cambridge, MA, USA, 2012. [Google Scholar]
- Schwitzer, T.; Le Cam, S.; Cosker, E.; Vinsard, H.; Leguay, A.; Angioi-Duprez, K.; Laprevote, V.; Ranta, R.; Schwan, R.; Dorr, V.L. Retinal electroretinogram features can detect depression state and treatment response in adults: A machine learning approach. J. Affect. Disord. 2022, 306, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Nowacka, B.; Lubiński, W.; Honczarenko, K.; Potemkowski, A.; Safranow, K. Bioelectrical function and structural assessment of the retina in patients with early stages of Parkinson’s disease (PD). Doc. Ophthalmol. 2015, 131, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Ellis, C.L.; Leo, S.M.; Velthuis, H.; Pereira, A.C.; Dimitrov, M.; Ponteduro, F.M.; Wong, N.M.L.; Daly, E.; Murphy, D.G.M.; et al. Retinal GABAergic Alterations in Adults with Autism Spectrum Disorder. J. Neurosci. 2024, 44, e1218232024. [Google Scholar] [CrossRef] [PubMed]
- Dubois, M.A.; Pelletier, C.A.; Mérette, C.; Jomphe, V.; Turgeon, R.; Bélanger, R.E.; Grondin, S.; Hébert, M. Evaluation of electroretinography (ERG) parameters as a biomarker for ADHD. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2023, 127, 110807. [Google Scholar] [CrossRef]
- Constable, P.A.; Skuse, D.H.; Thompson, D.A.; Lee, I.O. Brief report: Effects of methylphenidate on the light adapted electroretinogram. Doc. Ophthalmol. 2025, 150, 25–32. [Google Scholar] [CrossRef]
- Gauvin, M.; Little, J.M.; Lina, J.M.; Lachapelle, P. Functional decomposition of the human electroretinogram based on the discrete wavelet transform. J. Vis. 2014, 14, 15. [Google Scholar]
- Gauvin, M.; Lina, J.M.; Lachapelle, P. Advance in ERG analysis: From peak time and amplitude to frequency, power, and energy. BioMed Res. Int. 2014, 2014, 246096. [Google Scholar] [CrossRef]
- Gauvin, M.; Sustar, M.; Little, J.M.; Brecelj, J.; Lina, J.M.; Lachapelle, P. Quantifying the ON and OFF contributions to the flash ERG with the discrete wavelet transform. Transl. Vis. Sci. Technol. 2017, 6, 3. [Google Scholar] [CrossRef]
- Shwetar, Y.J.; Lalush, D.S.; Zhang, A.Y.; McAnany, J.J.; Jeffrey, B.G.; Haendel, M.A. A practical introduction to wavelet analysis in electroretinography. Doc. Ophthalmol. 2026, 152, 23–31. [Google Scholar] [CrossRef]
- Albasu, F.; Kulyabin, M.; Zhdanov, A.; Dolganov, A.; Ronkin, M.; Borisov, V.; Dorosinsky, L.; Constable, P.A.; Al-Masni, M.A.; Maier, A. Electroretinogram analysis using short-time Fourier transform and machine learning techniques. Bioengineering 2024, 11, 866. [Google Scholar] [CrossRef]
- Kulyabin, M.; Constable, P.A.; Zhdanov, A.; Lee, I.O.; Thompson, D.A.; Maier, A. Attention to the electroretinogram: Gated multilayer perceptron for ASD classification. IEEE Access 2024, 12, 52352–52362. [Google Scholar] [CrossRef]
- Chistiakov, S.; Dolganov, A.; Constable, P.A.; Zhdanov, A.; Kulyabin, M.; Thompson, D.A.; Lee, I.O.; Albasu, F.; Borisov, V.; Ronkin, M. Time series classification of autism spectrum disorder using the light-adapted electroretinogram. Bioengineering 2025, 12, 951. [Google Scholar] [CrossRef] [PubMed]
- Ortega, A.; Frossard, P.; Kovačević, J.; Moura, J.M.F.; Vandergheynst, P. Graph signal processing: Overview, challenges, and applications. Proc. IEEE 2018, 106, 808–828. [Google Scholar] [CrossRef]
- Shuman, D.I.; Narang, S.K.; Frossard, P.; Ortega, A.; Vandergheynst, P. The emerging field of signal processing on graphs: Extending high-dimensional data analysis to networks and other irregular domains. IEEE Signal Process. Mag. 2013, 30, 83–98. [Google Scholar] [CrossRef]
- Sandryhaila, A.; Moura, J.M.F. Discrete signal processing on graphs. IEEE Trans. Signal Process. 2013, 61, 1644–1656. [Google Scholar] [CrossRef]
- Sandryhaila, A.; Moura, J.M.F. Discrete signal processing on graphs: Frequency analysis. IEEE Trans. Signal Process. 2014, 62, 3042–3054. [Google Scholar] [CrossRef]
- Chung, F. Spectral Graph Theory; American Mathematical Society: Providence, RI, USA, 1997. [Google Scholar]
- Calazans, M.A.A.; Ferreira, F.A.B.S.; Santos, F.A.N.; Madeiro, F.; Lima, J.B. Machine learning and graph signal processing applied to healthcare: A review. Bioengineering 2024, 11, 671. [Google Scholar] [CrossRef]
- Bullmore, E.; Sporns, O. Complex brain networks: Graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 2009, 10, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Rubinov, M.; Sporns, O. Complex network measures of brain connectivity: Uses and interpretations. NeuroImage 2010, 52, 1059–1069. [Google Scholar] [CrossRef]
- Huang, W.; Bolton, T.A.W.; Medaglia, J.D.; Bassett, D.S.; Ribeiro, A.; Ville, D.V.D. A graph signal processing perspective on functional brain imaging. Proc. IEEE 2018, 106, 868–885. [Google Scholar] [CrossRef]
- Stam, C. Modern network science of neurological disorders. Nat. Rev. Neurosci. 2014, 15, 683–695. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Yuan, X.; Radfar, M.; Marendy, P.; Ni, W.; O’Brien, T.J.; Casillas-Espinosa, P. Graph signal processing, graph neural networks, and graph learning on biological data: A systematic review. IEEE Rev. Biomed. Eng. 2023, 16, 109–135. [Google Scholar] [CrossRef]
- Lacasa, L.; Luque, B.; Ballesteros, F.; Luque, J.; Nuño, J.C. From time series to complex networks: The visibility graph. Proc. Natl. Acad. Sci. USA 2008, 105, 4972–4975. [Google Scholar] [CrossRef] [PubMed]
- Donner, R.V.; Zou, Y.; Donges, J.F.; Marwan, N.; Kurths, J. Recurrence networks—A novel paradigm for nonlinear time series analysis. New J. Phys. 2010, 12, 033025. [Google Scholar] [CrossRef]
- Zou, Y.; Donner, R.V.; Marwan, N.; Donges, J.F.; Kurths, J. Complex network approaches to nonlinear time series analysis. Phys. Rep. 2019, 787, 1–97. [Google Scholar] [CrossRef]
- Ahmadlou, M.; Adeli, H.; Adeli, A. New diagnostic EEG markers of the Alzheimer’s disease using visibility graph. J. Neural Transm. 2010, 117, 1099–1109. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Bian, C.; Ning, X.; Ma, Q.D.Y. Visibility graph analysis on heartbeat dynamics of meditation training. Appl. Phys. Lett. 2013, 102, 253702. [Google Scholar] [CrossRef]
- Marwan, N.; Carmen Romano, M.; Thiel, M.; Kurths, J. Recurrence plots for the analysis of complex systems. Phys. Rep. 2007, 438, 237–329. [Google Scholar] [CrossRef]
- McCullough, M.; Small, M.; Stemler, T.; Iu, H.H.M. Time lagged ordinal partition networks for capturing dynamics of continuous dynamical systems. Chaos 2015, 25, 053101. [Google Scholar] [CrossRef]
- Freeman, L.C. Centrality in social networks: Conceptual clarification. Soc. Netw. 1979, 1, 215–239. [Google Scholar] [CrossRef]
- Newman, M. Networks: An Introduction; Oxford University Press: Oxford, UK, 2010. [Google Scholar]
- Barrat, A.; Barthélemy, M.; Pastor-Satorras, R.; Vespignani, A. The architecture of complex weighted networks. Proc. Natl. Acad. Sci. USA 2004, 101, 3747–3752. [Google Scholar] [CrossRef]
- Mercado-Diaz, L.R.; Veeranki, Y.R.; Marmolejo-Ramos, F.; Posada-Quintero, H.F. EDA-Graph: Graph signal processing of electrodermal activity for emotional state detection. IEEE J. Biomed. Health Inform. 2024, 28, 4599–4612. [Google Scholar] [CrossRef]
- Veeranki, Y.R.; Diaz, L.R.M.; Swaminathan, R.; Posada-Quintero, H.F. Nonlinear signal processing methods for automatic emotion recognition using electrodermal activity. IEEE Sens. J. 2024, 24, 8140–8150. [Google Scholar] [CrossRef]
- Mercado-Diaz, L.R.; Veeranki, Y.R.; Large, E.W.; Posada-Quintero, H.F. Fractal analysis of electrodermal activity for emotion recognition: A novel approach using detrended fluctuation analysis and wavelet entropy. Sensors 2024, 24, 8130. [Google Scholar] [CrossRef]
- Viswanathan, S.; Frishman, L.J.; Robson, J.G.; Walters, J.W. The photopic negative response of the flash electroretinogram in primary open angle glaucoma. Investig. Ophthalmol. Vis. Sci. 2001, 42, 514–522. [Google Scholar]
- Wachtmeister, L. Some aspects of the oscillatory response of the retina. Prog. Brain Res. 2001, 131, 465–474. [Google Scholar] [PubMed]
- Brandes, U. A faster algorithm for betweenness centrality. J. Math. Sociol. 2001, 25, 163–177. [Google Scholar] [CrossRef]
- Boldi, P.; Vigna, S. Axioms for centrality. Internet Math. 2014, 10, 222–262. [Google Scholar] [CrossRef]
- Bron, C.; Kerbosch, J. Algorithm 457: Finding all cliques of an undirected graph. Commun. ACM 1973, 16, 575–577. [Google Scholar] [CrossRef]
- Watts, D.; Strogatz, S. Collective dynamics of ‘small-world’ networks. Nature 1998, 393, 440–442. [Google Scholar] [CrossRef]
- Fiedler, M. Algebraic connectivity of graphs. Czech. Math. J. 1973, 23, 298–305. [Google Scholar] [CrossRef]
- Fornito, A.; Zalesky, A.; Bullmore, E. Fundamentals of Brain Network Analysis; Academic Press: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Lundberg, S.; Lee, S.I. A unified approach to interpreting model predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems (NeurIPS), Long Beach, CA, USA; Associates, Inc.: Seattle, WA, USA, 2017; pp. 4765–4774. [Google Scholar]
- Bellman, R. Adaptive Control Processes: A Guided Tour; Princeton University Press: Princeton, NJ, USA, 1961. [Google Scholar]
- Nasrolahzadeh, M.; Mohammadpoory, Z.; Haddadnia, J. A novel method for distinction heart rate variability during meditation using LSTM recurrent neural networks based on visibility graph. Biomed. Signal Process. Control 2024, 90, 105822. [Google Scholar] [CrossRef]
- Patel, S.A.; Smith, R.J.; Yildirim, A. Gershgorin circle theorem-based feature extraction for biomedical signal analysis. Front. Neuroinform. 2024, 18, 1395916. [Google Scholar] [CrossRef]
- Nguyen-Legros, J.; Versaux-Botteri, C.; Vernier, P. Dopamine receptor localization in the mammalian retina. Mol. Neurobiol. 1999, 19, 181–204. [Google Scholar] [CrossRef]
- Choi, H.; Hong, J.; Kang, H.G.; Park, M.H.; Ha, S.; Lee, J.; Yoon, S.; Kim, D.; Park, Y.R.; Cheon, K.A. Retinal fundus imaging as a biomarker for ADHD using machine learning. npj Digit. Med. 2025, 8, 164. [Google Scholar] [CrossRef]
- Wang, J.; Wang, Y.X.; Zeng, D.; Zhu, Z.; Li, D.; Liu, Y.; Sheng, B.; Grzybowski, A.; Wong, T.Y. AI-enhanced retinal imaging as a biomarker for systemic diseases. Theranostics 2025, 15, 3223–3233. [Google Scholar] [CrossRef]
- Kulyabin, M.; Zhdanov, A.; Lee, I.O.; Skuse, D.H.; Thompson, D.A.; Maier, A.; Constable, P.A. Synthetic electroretinogram signal generation using a conditional generative adversarial network. Doc. Ophthalmol. 2025, 151, 161–177. [Google Scholar] [CrossRef]
- Sporns, O. Graph theory methods: Applications in brain networks. Dialogues Clin. Neurosci. 2018, 20, 111–121. [Google Scholar] [CrossRef]
- Jain, A.K. Data clustering: 50 years beyond K-means. Pattern Recognit. Lett. 2010, 31, 651–666. [Google Scholar] [CrossRef]
- von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 2017, 17, 395–416. [Google Scholar] [CrossRef]
- Fortunato, S. Community detection in graphs. Phys. Rep. 2010, 486, 75–174. [Google Scholar] [CrossRef]
- Constable, P.A.; Gaigg, S.B.; Bowler, D.M.; Thompson, D.A. Motion and pattern cortical potentials in adults with high-functioning autism spectrum disorder. Doc. Ophthalmol. 2012, 125, 219–227. [Google Scholar] [CrossRef]
- Lee, I.O.; Fritsch, D.M.; Kerz, M.; Sowden, J.C.; Constable, P.A.; Skuse, D.H.; Thompson, D.A. Global motion coherent deficits in individuals with autism spectrum disorder and their family members are associated with retinal function. Sci. Rep. 2025, 15, 28249. [Google Scholar] [CrossRef]
- Brabec, M.; Constable, P.A.; Thompson, D.A.; Marmolejo-Ramos, F. Group comparisons of the individual electroretinogram time trajectories for the ascending limb of the b-wave using a raw and registered time series. BMC Res. Notes 2023, 16, 238. [Google Scholar] [CrossRef] [PubMed]
- Brabec, M.; Marmolejo-Ramos, F.; Loh, L.; Lee, I.O.; Kulyabin, M.; Zhdanov, A.; Posada-Quintero, H.; Thompson, D.A.; Constable, P.A. Remodeling the light-adapted electroretinogram using a bayesian statistical approach. BMC Res. Notes 2025, 18, 33. [Google Scholar] [CrossRef] [PubMed]
- Brabec, M.; Loh, L.; Lee, I.O.; Marmolejo-Ramos, F.; Skuse, D.H.; Thompson, D.A.; Constable, P.A. Technical note: Contour plot visualization of the light adapted electroretinogram using a generalized additive model. Doc. Ophthalmol. 2026, 152, 1–11. [Google Scholar] [CrossRef] [PubMed]





| Feature | Control | ASD | ADHD | p-Value | δ ASD/Ctrl | δ ADHD/Ctrl | δ ASD/ADHD |
|---|---|---|---|---|---|---|---|
| TLC | 245.3 ± 42.1 | 198.7 ± 38.5 | 278.4 ± 51.2 | <0.001 *** | −0.52(L) | 0.34(M) | −0.61(L) |
| THC | 312.8 ± 55.6 | 289.4 ± 49.3 | 341.2 ± 62.7 | 0.002 ** | −0.22(S) | 0.27(S) | −0.41(M) |
| GNC | 18.4 ± 4.2 | 15.1 ± 3.8 | 21.7 ± 5.1 | <0.001 *** | −0.41(M) | 0.36(M) | −0.58(L) |
| Diam. | 8.2 ± 1.9 | 6.8 ± 1.5 | 9.5 ± 2.3 | <0.001 *** | −0.44(L) | 0.39(M) | −0.56(L) |
| Radius | 4.5 ± 1.1 | 3.9 ± 0.9 | 5.2 ± 1.4 | 0.001 ** | −0.31(M) | 0.28(M) | −0.47(L) |
| CC | 0.42 ± 0.08 | 0.48 ± 0.09 | 0.37 ± 0.07 | <0.001 *** | 0.38(M) | −0.33(M) | 0.59(L) |
| APL | 3.8 ± 0.7 | 3.2 ± 0.6 | 4.3 ± 0.9 | <0.001 *** | −0.47(L) | 0.31(M) | −0.57(L) |
| λ2 | 0.35 ± 0.08 | 0.41 ± 0.09 | 0.29 ± 0.07 | 0.003 ** | 0.35(M) | −0.40(M) | 0.55(L) |
| Density | 0.28 ± 0.05 | 0.32 ± 0.06 | 0.24 ± 0.05 | <0.001 *** | 0.37(M) | −0.42(M) | 0.54(L) |
| Comparison | Sex | Model | Flash/Eye | BA | F1 | Feat. |
|---|---|---|---|---|---|---|
| ASD vs. Ctrl | Male | XGB | 446 Td.s/Right | 0.91 | 0.90 | 7 |
| ASD vs. Ctrl | Female | RF | 446 Td.s/Right | 0.84 | 0.83 | 8 |
| ASD vs. Ctrl | All | XGB | 446 Td.s/R + L | 0.84 | 0.83 | 12 |
| ADHD vs. Ctrl | Male | SVM | 446 Td.s/Right | 0.83 | 0.82 | 6 |
| ADHD vs. Ctrl | Female | RF | 446 Td.s/Right | 0.88 | 0.87 | 7 |
| ADHD vs. Ctrl | All | RF | 446 Td.s/R + L | 0.83 | 0.82 | 11 |
| Feature Set | Model | BA | F1 | Feat. |
|---|---|---|---|---|
| TD only | AdaB | 0.62 | 0.60 | 8 |
| TD + DWT | GradB | 0.67 | 0.65 | 24 |
| TD + VFCDM | XGB | 0.70 | 0.68 | 32 |
| TD + DWT+VFCDM | XGB | 0.70 | 0.69 | 45 |
| ERG-Graph only | XGB | 0.78 | 0.76 | 18 |
| ERG-Graph + TD | XGB | 0.81 | 0.79 | 22 |
| Full fusion | XGB | 0.79 | 0.77 | 38 |
| Feature Set | Model | BA | F1 | #Feat. |
|---|---|---|---|---|
| TD + DWT + VFCDM [12] | XGB | 0.53 | 0.51 | 45 |
| ERG-Graph only | XGB | 0.64 | 0.62 | 18 |
| ERG-Graph + TD | XGB | 0.67 | 0.65 | 22 |
| Full fusion | XGB | 0.65 | 0.63 | 38 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Mercado-Diaz, L.R.; Pinzon-Arenas, J.O.; Constable, P.A.; Lee, I.O.; Loh, L.; Thompson, D.A.; Posada-Quintero, H.F. ERG-Graph: Graph Signal Processing of the Electroretinogram for Classification of Neurodevelopmental Disorders. Bioengineering 2026, 13, 446. https://doi.org/10.3390/bioengineering13040446
Mercado-Diaz LR, Pinzon-Arenas JO, Constable PA, Lee IO, Loh L, Thompson DA, Posada-Quintero HF. ERG-Graph: Graph Signal Processing of the Electroretinogram for Classification of Neurodevelopmental Disorders. Bioengineering. 2026; 13(4):446. https://doi.org/10.3390/bioengineering13040446
Chicago/Turabian StyleMercado-Diaz, Luis Roberto, Javier O. Pinzon-Arenas, Paul A. Constable, Irene O. Lee, Lynne Loh, Dorothy A. Thompson, and Hugo F. Posada-Quintero. 2026. "ERG-Graph: Graph Signal Processing of the Electroretinogram for Classification of Neurodevelopmental Disorders" Bioengineering 13, no. 4: 446. https://doi.org/10.3390/bioengineering13040446
APA StyleMercado-Diaz, L. R., Pinzon-Arenas, J. O., Constable, P. A., Lee, I. O., Loh, L., Thompson, D. A., & Posada-Quintero, H. F. (2026). ERG-Graph: Graph Signal Processing of the Electroretinogram for Classification of Neurodevelopmental Disorders. Bioengineering, 13(4), 446. https://doi.org/10.3390/bioengineering13040446









