Neuroscience-Inspired Deep Learning Brain–Machine Interface Decoder
Abstract
1. Introduction
- 1.
- We propose a DNN-based decoder (CNN-LSTM) for motor decoding, offering a novel formulation that bridges neuroscience mechanisms with deep learning approaches for prosthetic and rehabilitation applications.
- 2.
- We introduce the Single-Direction CNN-LSTM, which decodes joint variables independently across directions, thereby improving task-level generalizability.
2. Materials and Methods
2.1. Data Preparation
2.2. Data Preprocessing
2.3. Proposed Model
2.3.1. Convolutional Layers
2.3.2. LSTM Layer
2.4. Single-Direction CNN-LSTM Decoder
2.5. Baseline Model
2.5.1. Conventional CNN-LSTM Decoder
2.5.2. Linear Decoder
2.6. Fine-Tuned and Generalizability Test
2.7. Environment and Hyperparameter
3. Results
3.1. Validation on All Data
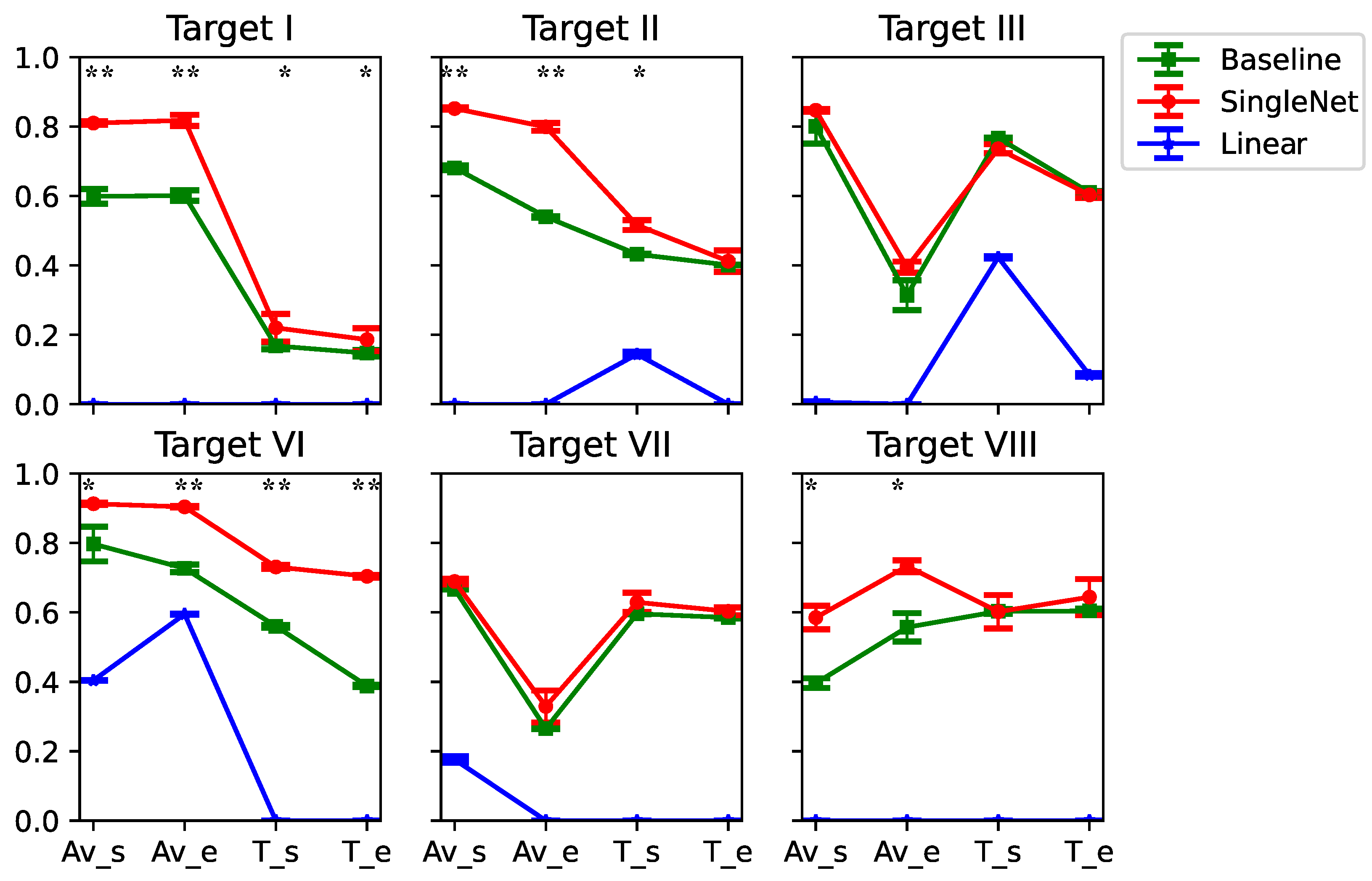
3.2. Generlizability
3.3. Ablation Study
3.4. Features Extracted by the Decoder
3.5. Estimation of Co-Contraction Increase the Generalizability
4. Discussion
4.1. Limitation of Fine-Tuning
4.2. Limitation of Data
4.3. Musculoskeletal Model
4.4. Co-Contraction Judgment
5. Future and Limitation
5.1. Subject-Specific Bias
5.2. Scalability to Human BMI
5.3. Musculoskeletal Model Sensitivity
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lebedev, M.A.; Nicolelis, M.A. Brain–machine interfaces: Past, present and future. TRENDS Neurosci. 2006, 29, 536–546. [Google Scholar] [CrossRef]
- Gao, X.; Wang, Y.; Chen, X.; Gao, S. Interface, interaction, and intelligence in generalized brain–computer interfaces. Trends Cogn. Sci. 2021, 25, 671–684. [Google Scholar] [CrossRef]
- Dong, Y.; Wang, S.; Huang, Q.; Berg, R.W.; Li, G.; He, J. Neural decoding for intracortical brain–computer interfaces. Cyborg Bionic Syst. 2023, 4, 0044. [Google Scholar] [CrossRef]
- Orban, M.; Elsamanty, M.; Guo, K.; Zhang, S.; Yang, H. A review of brain activity and EEG-based brain–computer interfaces for rehabilitation application. Bioengineering 2022, 9, 768. [Google Scholar] [CrossRef]
- Abdullah; Faye, I.; Islam, M.R. EEG channel selection techniques in motor imagery applications: A review and new perspectives. Bioengineering 2022, 9, 726. [Google Scholar] [CrossRef]
- Wu, X.; Metcalfe, B.; He, S.; Tan, H.; Zhang, D. A review of Motor Brain-Computer interfaces using Intracranial Electroencephalography based on surface electrodes and depth electrodes. IEEE Trans. Neural Syst. Rehabil. Eng. 2024, 32, 2408–2431. [Google Scholar] [CrossRef]
- Anjum, M.; Sakib, N.; Islam, M.K. Effect of artifact removal on EEG based motor imagery BCI applications. In Proceedings of the Fourth International Conference on Computer Vision and Information Technology (CVIT 2023); SPIE: Bellingham, WA, USA, 2024; Volume 12984, pp. 69–78. [Google Scholar]
- Abu-Rmileh, A.; Zakkay, E.; Shmuelof, L.; Shriki, O. Co-adaptive training improves efficacy of a multi-day EEG-based motor imagery BCI training. Front. Hum. Neurosci. 2019, 13, 362. [Google Scholar] [CrossRef] [PubMed]
- Altuwaijri, G.A.; Muhammad, G. Electroencephalogram-based motor imagery signals classification using a multi-branch convolutional neural network model with attention blocks. Bioengineering 2022, 9, 323. [Google Scholar] [CrossRef]
- Zhu, D.; Bieger, J.; Garcia Molina, G.; Aarts, R.M. A survey of stimulation methods used in SSVEP-based BCIs. Comput. Intell. Neurosci. 2010, 2010, 702357. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Zhang, C.; Wu, W.; Gao, X. Frequency recognition based on canonical correlation analysis for SSVEP-based BCIs. IEEE Trans. Biomed. Eng. 2006, 53, 2610–2614. [Google Scholar] [CrossRef] [PubMed]
- Gaur, P.; Pachori, R.B.; Wang, H.; Prasad, G. An empirical mode decomposition based filtering method for classification of motor-imagery EEG signals for enhancing brain-computer interface. In Proceedings of the 2015 International Joint Conference on Neural Networks (IJCNN); IEEE: New York, NY, USA, 2015; pp. 1–7. [Google Scholar]
- Mushtaq, F.; Welke, D.; Gallagher, A.; Pavlov, Y.G.; Kouara, L.; Bosch-Bayard, J.; van den Bosch, J.J.; Arvaneh, M.; Bland, A.R.; Chaumon, M.; et al. One hundred years of EEG for brain and behaviour research. Nat. Hum. Behav. 2024, 8, 1437–1443. [Google Scholar] [CrossRef]
- Grobbelaar, M.; Phadikar, S.; Ghaderpour, E.; Struck, A.F.; Sinha, N.; Ghosh, R.; Ahmed, M.Z.I. A survey on denoising techniques of electroencephalogram signals using wavelet transform. Signals 2022, 3, 577–586. [Google Scholar] [CrossRef]
- Buzsáki, G.; Anastassiou, C.A.; Koch, C. The origin of extracellular fields and currents—EEG, ECoG, LFP and spikes. Nat. Rev. Neurosci. 2012, 13, 407–420. [Google Scholar] [CrossRef]
- Georgopoulos, A.P.; Schwartz, A.B.; Kettner, R.E. Neuronal population coding of movement direction. Science 1986, 233, 1416–1419. [Google Scholar] [CrossRef]
- Wu, W.; Gao, Y.; Bienenstock, E.; Donoghue, J.P.; Black, M.J. Bayesian population decoding of motor cortical activity using a Kalman filter. Neural Comput. 2006, 18, 80–118. [Google Scholar] [CrossRef] [PubMed]
- Autthasan, P.; Chaisaen, R.; Sudhawiyangkul, T.; Rangpong, P.; Kiatthaveephong, S.; Dilokthanakul, N.; Bhakdisongkhram, G.; Phan, H.; Guan, C.; Wilaiprasitporn, T. MIN2Net: End-to-end multi-task learning for subject-independent motor imagery EEG classification. IEEE Trans. Biomed. Eng. 2021, 69, 2105–2118. [Google Scholar] [CrossRef] [PubMed]
- Lawhern, V.J.; Solon, A.J.; Waytowich, N.R.; Gordon, S.M.; Hung, C.P.; Lance, B.J. EEGNet: A compact convolutional neural network for EEG-based brain–computer interfaces. J. Neural Eng. 2018, 15, 056013. [Google Scholar] [CrossRef] [PubMed]
- Schirrmeister, R.T.; Springenberg, J.T.; Fiederer, L.D.J.; Glasstetter, M.; Eggensperger, K.; Tangermann, M.; Hutter, F.; Burgard, W.; Ball, T. Deep learning with convolutional neural networks for EEG decoding and visualization. Hum. Brain Mapp. 2017, 38, 5391–5420. [Google Scholar] [CrossRef] [PubMed]
- Glaser, J.I.; Benjamin, A.S.; Chowdhury, R.H.; Perich, M.G.; Miller, L.E.; Kording, K.P. Machine learning for neural decoding. eneuro 2020, 7. [Google Scholar] [CrossRef]
- Willett, F.R.; Kunz, E.M.; Fan, C.; Avansino, D.T.; Wilson, G.H.; Choi, E.Y.; Kamdar, F.; Glasser, M.F.; Hochberg, L.R.; Druckmann, S.; et al. A high-performance speech neuroprosthesis. Nature 2023, 620, 1031–1036. [Google Scholar] [CrossRef]
- Chandrasekaran, S.; Wandelt, S.K.; Jangam, A.; Elias, Z.; Ibroci, E.; Maffei, C.; Rosenthal, I.A.; Ramdeo, R.; Kim, J.W.; Xu, J.; et al. Restoring Cortically Mediated Movement and Sensation in Complete Tetraplegia. medRxiv 2025. [Google Scholar] [CrossRef]
- Śliwowski, M.; Martin, M.; Souloumiac, A.; Blanchart, P.; Aksenova, T. Decoding ECoG signal into 3D hand translation using deep learning. J. Neural Eng. 2022, 19, 026023. [Google Scholar] [CrossRef]
- Ji, C. Explainable mst-ecognet decode visual information from ecog signal. arXiv 2024, arXiv:2411.16165. [Google Scholar]
- Xie, Z.; Schwartz, O.; Prasad, A. Decoding of finger trajectory from ECoG using deep learning. J. Neural Eng. 2018, 15, 036009. [Google Scholar] [CrossRef]
- Scott, S.H.; Kalaska, J.F. Reaching movements with similar hand paths but different arm orientations. I. Activity of individual cells in motor cortex. J. Neurophysiol. 1997, 77, 826–852. [Google Scholar] [CrossRef]
- Dey, S.; Yoshida, T.; Foerster, R.H.; Ernst, M.; Schmalz, T.; Carnier, R.M.; Schilling, A.F. A hybrid approach for dynamically training a torque prediction model for devising a human-machine interface control strategy. arXiv 2021, arXiv:2110.03085. [Google Scholar] [CrossRef]
- Miyashita, E.; Sakaguchi, Y. State variables of the arm may be encoded by single neuron activity in the monkey motor cortex. IEEE Trans. Ind. Electron. 2015, 63, 1943–1952. [Google Scholar] [CrossRef]
- Tian, K.; Zhao, S.; Zhang, Y.; Yu, S. Multi-dimensional Neural Decoding with Orthogonal Representations for Brain-Computer Interfaces. arXiv 2025, arXiv:2508.08681. [Google Scholar] [CrossRef]
- Georgopoulos, A.P.; Kalaska, J.F.; Caminiti, R.; Massey, J.T. On the relations between the direction of two-dimensional arm movements and cell discharge in primate motor cortex. J. Neurosci. 1982, 2, 1527–1537. [Google Scholar] [CrossRef]
- Ahmadi, N.; Constandinou, T.G.; Bouganis, C.S. Robust and accurate decoding of hand kinematics from entire spiking activity using deep learning. J. Neural Eng. 2021, 18, 026011. [Google Scholar] [CrossRef] [PubMed]
- Meattini, R.; Chiaravalli, D.; Biagiotti, L.; Palli, G.; Melchiorri, C. Combining unsupervised muscle co-contraction estimation with bio-feedback allows augmented kinesthetic teaching. IEEE Robot. Autom. Lett. 2021, 6, 6180–6187. [Google Scholar] [CrossRef]
- Nijhawan, R. Neural delays, visual motion and the flash-lag effect. Trends Cogn. Sci. 2002, 6, 387–393. [Google Scholar] [CrossRef]
- Awasthi, P.; Lin, T.H.; Bae, J.; Miller, L.E.; Danziger, Z.C. Validation of a non-invasive, real-time, human-in-the-loop model of intracortical brain-computer interfaces. J. Neural Eng. 2022, 19, 056038. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long short-term memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Noh, S.H. Analysis of gradient vanishing of RNNs and performance comparison. Information 2021, 12, 442. [Google Scholar] [CrossRef]
- Schmidt-Hieber, J. Nonparametric regression using deep neural networks with ReLU activation function. Ann. Statist. 2020, 48, 1875–1897. [Google Scholar]
- Bhanja, S.; Das, A. Impact of data normalization on deep neural network for time series forecasting. arXiv 2018, arXiv:1812.05519. [Google Scholar]
- Haghi, B.; Aflalo, T.; Kellis, S.; Guan, C.; Gamez de Leon, J.A.; Huang, A.Y.; Pouratian, N.; Andersen, R.A.; Emami, A. Enhanced control of a brain–computer interface by tetraplegic participants via neural-network-mediated feature extraction. Nat. Biomed. Eng. 2024, 9, 917–934. [Google Scholar] [CrossRef]
- Pandarinath, C.; O’Shea, D.J.; Collins, J.; Jozefowicz, R.; Stavisky, S.D.; Kao, J.C.; Trautmann, E.M.; Kaufman, M.T.; Ryu, S.I.; Hochberg, L.R.; et al. Inferring single-trial neural population dynamics using sequential auto-encoders. Nat. Methods 2018, 15, 805–815. [Google Scholar] [CrossRef]
- Zhang, L.; Soselia, D.; Wang, R.; Gutierrez-Farewik, E.M. Lower-limb joint torque prediction using LSTM neural networks and transfer learning. IEEE Trans. Neural Syst. Rehabil. Eng. 2022, 30, 600–609. [Google Scholar] [CrossRef]
- Shah, S.; Haghi, B.; Kellis, S.; Bashford, L.; Kramer, D.; Lee, B.; Liu, C.; Andersen, R.; Emami, A. Decoding kinematics from human parietal cortex using neural networks. In Proceedings of the 2019 9th International IEEE/EMBS Conference on Neural Engineering (NER); IEEE: New York, NY, USA, 2019; pp. 1138–1141. [Google Scholar]
- De Feo, V.; Boi, F.; Safaai, H.; Onken, A.; Panzeri, S.; Vato, A. State-dependent decoding algorithms improve the performance of a bidirectional bmi in anesthetized rats. Front. Neurosci. 2017, 11, 269. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Rao, C.R.; Rao, C.R.; Statistiker, M.; Rao, C.R.; Rao, C.R. Linear Statistical Inference and Its Applications; Wiley: New York, NY, USA, 1973; Volume 2. [Google Scholar]
- Peterson, S.M.; Steine-Hanson, Z.; Davis, N.; Rao, R.P.; Brunton, B.W. Generalized neural decoders for transfer learning across participants and recording modalities. J. Neural Eng. 2021, 18, 026014. [Google Scholar] [CrossRef]
- Churchland, M.M.; Shenoy, K.V. Temporal complexity and heterogeneity of single-neuron activity in premotor and motor cortex. J. Neurophysiol. 2007, 97, 4235–4257. [Google Scholar] [CrossRef]
- Chen, X.; Fu, Z.; Zhang, P.; Chen, X.; Huang, J. Intracortical Brain-Machine Interfaces with High-Performance Neural Decoding through Efficient Transfer Meta-learning. IEEE Trans. Biomed. Eng. 2025, 73, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Iman, M.; Arabnia, H.R.; Rasheed, K. A review of deep transfer learning and recent advancements. Technologies 2023, 11, 40. [Google Scholar] [CrossRef]
- Hong, X.; Zheng, Q.; Liu, L.; Chen, P.; Ma, K.; Gao, Z.; Zheng, Y. Dynamic joint domain adaptation network for motor imagery classification. IEEE Trans. Neural Syst. Rehabil. Eng. 2021, 29, 556–565. [Google Scholar] [CrossRef]
- Lee, D.Y.; Lee, M.; Lee, S.W. Decoding imagined speech based on deep metric learning for intuitive BCI communication. IEEE Trans. Neural Syst. Rehabil. Eng. 2021, 29, 1363–1374. [Google Scholar] [CrossRef] [PubMed]
- Halmich, C.; Höschler, L.; Schranz, C.; Borgelt, C. Data augmentation of time-series data in human movement biomechanics: A scoping review. PLoS ONE 2025, 20, e0327038. [Google Scholar] [CrossRef]
- Banks, C.L.; Huang, H.J.; Little, V.L.; Patten, C. Electromyography exposes heterogeneity in muscle co-contraction following stroke. Front. Neurol. 2017, 8, 699. [Google Scholar] [CrossRef]
- Kumar, S.; Alawieh, H.; Racz, F.S.; Fakhreddine, R.; Millán, J.d.R. Transfer learning promotes acquisition of individual BCI skills. PNAS Nexus 2024, 3, pgae076. [Google Scholar] [CrossRef]
- Chen, S.; Chen, M.; Wang, X.; Liu, X.; Liu, B.; Ming, D. Brain–computer interfaces in 2023–2024. Brain-x 2025, 3, e70024. [Google Scholar] [CrossRef]
- Wang, Z.; Li, S.; Wu, D. Canine EEG helps human: Cross-species and cross-modality epileptic seizure detection via multi-space alignment. Natl. Sci. Rev. 2025, 12, nwaf086. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Ma, S.; Zhao, Y.; Shi, C.; Zhang, Z. A physics-informed low-shot adversarial learning for semg-based estimation of muscle force and joint kinematics. IEEE J. Biomed. Health Inform. 2023, 28, 1309–1320. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Kim, C.; Hwangbo, J. Learning forward dynamics model and informed trajectory sampler for safe quadruped navigation. arXiv 2022, arXiv:2204.08647. [Google Scholar] [CrossRef]







| Branch | Layer | Activation Function | Hyperparameter | Value | Output Shape |
|---|---|---|---|---|---|
| Input | N/A | N/A | Input Shape | [32 × 150 × 73 × 30] | [32 × 150 × 73 × 30 × 1] |
| Extension | Conv_2D | ReLU | Number of temporal filters | 8 | [32 × 180 × 73 × 30 × 8] |
| Kernel size | (1, 30) | ||||
| Padding | same | ||||
| stride step | 1 | ||||
| Depthwise Conv_2D | ReLU | Depth multiplier | 2 | [32 × 180 × 1 × 30 × 16] | |
| Kernel size | (73, 1) | ||||
| Padding | valid | ||||
| Stride step | 1 | ||||
| Average Pooling & Flatten | N/A | Pooling kernel size/stride | 4 | [32 × 180 × 112] | |
| LSTM | tanh | Number of layers | 1 | [32 × 180 × 16] | |
| Number of hidden units | 16 | ||||
| Sequence length | 180 | ||||
| FC | ReLU | Number of units | 8 | [32 × 180 × 8] | |
| Flexion | Conv_2D | ReLU | Number of temporal filters | 8 | [32 × 180 × 73 × 30 × 8] |
| Kernel size | (1, 30) | ||||
| Padding | same | ||||
| stride step | 1 | ||||
| Depthwise Conv_2D | ReLU | Depth multiplier | 2 | [32 × 180 × 1 × 30 × 16] | |
| Kernel size | (73, 1) | ||||
| Padding | valid | ||||
| Stride step | 1 | ||||
| Average Pooling & Flatten | N/A | Pooling kernel size/stride | 4 | [32 × 180 × 112] | |
| LSTM | tanh | Number of layers | 1 | ||
| Number of hidden units | 16 | ||||
| Sequence length | 180 | [32 × 180 × 16] | |||
| FC | ReLU | Number of units | 8 | [32 × 180 × 8] |
| Av_s | Av_e | T_s | T_e | |
|---|---|---|---|---|
| Target I II | 0.775 | 0.534 | 0.645 | 0.612 |
| Target VI VIII | 0.827 | 0.740 | 0.486 | 0.542 |
| Target IV V | 0.827 | 0.710 | 0.604 | 0.587 |
| I | II | III | VI | VII | VIII | ||
|---|---|---|---|---|---|---|---|
| SingleNet | 0.818 ± 0.016 | 0.852 ± 0.003 | 0.847 ± 0.004 ↑ | 0.913 ± 0.003 ↑ | 0.689 ± 0.008 | 0.773 ± 0.017 | |
| 0.810 ± 0.005 ↑ | 0.799 ± 0.011 | 0.395 ± 0.016 ↑ | 0.904 ± 0.002 ↑ | 0.329 ± 0.046 | 0.585 ± 0.034 ↑ | ||
| 0.220 ± 0.040 | 0.516 ± 0.014 | 0.736 ± 0.013 | 0.731 ± 0.006 ↑ | 0.629 ± 0.028 | 0.602 ± 0.048 | ||
| 0.186 ± 0.033 | 0.412 ± 0.031 | 0.603 ± 0.009 | 0.704 ± 0.004 ↑ | 0.603 ± 0.011 | 0.644 ± 0.052 | ||
| LinearNet | 0.848 ± 0.005 ↑ | 0.856 ± 0.006 ↑ | 0.831 ± 0.008 | 0.897 ± 0.002 | 0.724 ± 0.014 ↑ | 0.500 ± 0.063 | |
| 0.803 ± 0.013 | 0.816 ± 0.013 | 0.394 ± 0.029 | 0.828 ± 0.043 | 0.360 ± 0.028 | 0.767 ± 0.029 | ||
| 0.209 ± 0.016 | 0.558 ± 0.014 ↑ | 0.745 ± 0.004 ↑ | 0.688 ± 0.006 | 0.666 ± 0.013 ↑ | 0.611 ± 0.026 | ||
| 0.226 ± 0.056 ↑ | 0.400 ± 0.040 | 0.595 ± 0.029 | 0.569 ± 0.022 | 0.604 ± 0.013 | 0.670 ± 0.032 ↑ | ||
| SharedNet | 0.827 ± 0.007 | 0.833 ± 0.005 | 0.842 ± 0.007 | 0.875 ± 0.002 | 0.344 ± 0.015 | 0.801 ± 0.010 ↑ | |
| 0.778 ± 0.019 | 0.863 ± 0.003 ↑ | 0.371 ± 0.051 | 0.893 ± 0.004 | 0.703 ± 0.004 ↑ | 0.544 ± 0.036 | ||
| 0.226 ± 0.051 ↑ | 0.463 ± 0.008 | 0.602 ± 0.021 | 0.279 ± 0.016 | 0.620 ± 0.012 | 0.698 ± 0.012 ↑ | ||
| 0.209 ± 0.010 | 0.571 ± 0.013 ↑ | 0.741 ± 0.008 ↑ | 0.694 ± 0.009 | 0.605 ± 0.065 ↑ | 0.632 ± 0.054 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ou, H.-Y.; Hasegawa, T.; Fukayama, O.; Miyashita, E. Neuroscience-Inspired Deep Learning Brain–Machine Interface Decoder. Bioengineering 2026, 13, 440. https://doi.org/10.3390/bioengineering13040440
Ou H-Y, Hasegawa T, Fukayama O, Miyashita E. Neuroscience-Inspired Deep Learning Brain–Machine Interface Decoder. Bioengineering. 2026; 13(4):440. https://doi.org/10.3390/bioengineering13040440
Chicago/Turabian StyleOu, Hong-Yun, Takahiro Hasegawa, Osamu Fukayama, and Eizo Miyashita. 2026. "Neuroscience-Inspired Deep Learning Brain–Machine Interface Decoder" Bioengineering 13, no. 4: 440. https://doi.org/10.3390/bioengineering13040440
APA StyleOu, H.-Y., Hasegawa, T., Fukayama, O., & Miyashita, E. (2026). Neuroscience-Inspired Deep Learning Brain–Machine Interface Decoder. Bioengineering, 13(4), 440. https://doi.org/10.3390/bioengineering13040440






