Engineering a Quantitative Organ-on-a-Chip Platform for Myogenic Mechanobiology
Abstract
1. Introduction
2. Methods and Materials
2.1. Design of the Organ-on-a-Chip Device
2.2. Finite Element Modeling
2.2.1. Governing Equations
2.2.2. Boundary Conditions
2.3. Fabrication of the Organ-on-a-Chip Molds
2.4. Chip Fabrication and Assembly
2.5. Fabrication of Fluorescent Bead-Embedded Flexible Membranes
2.6. Pneumatic Actuation Platform for Mechanical Stimulation
2.7. Measurement of Membrane Strain and Stress
2.8. Cell Culture
2.9. Preparation of PDMS-Coated Culture Dishes
2.10. Type I Collagen Coating of PDMS Substrates
2.11. Fluorescence Staining
2.12. Statistical Analysis
3. Results
3.1. Design and Structural Optimization of an Organ-on-a-Chip for Myogenic Mechanobiology
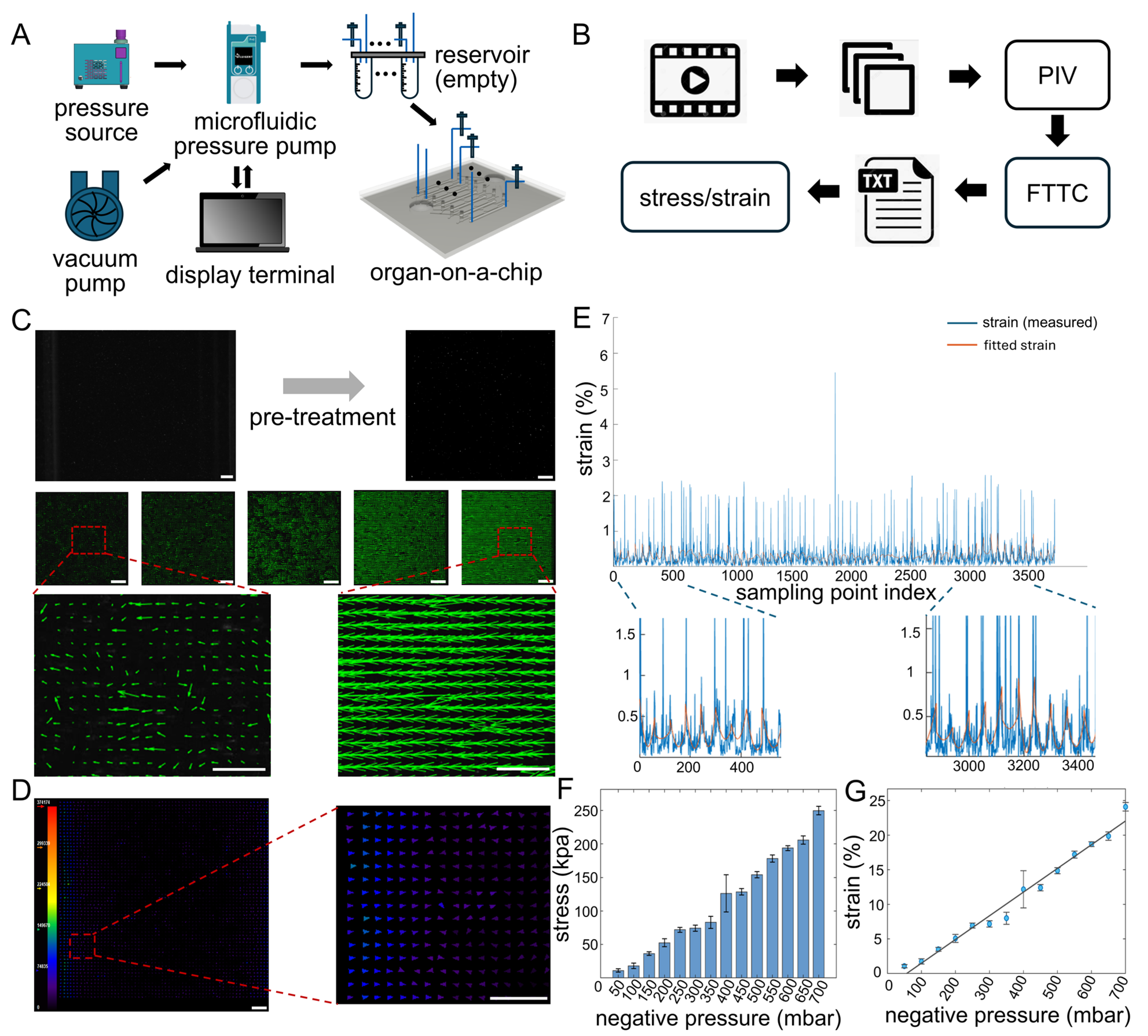
3.2. Fabrication of a Fluorescent Bead-Embedded Flexible Membrane for Quantitative Myogenic Traction Analysis
3.3. Quantitative Characterization of Controllable Cyclic Strain in the Myogenic Microenvironment
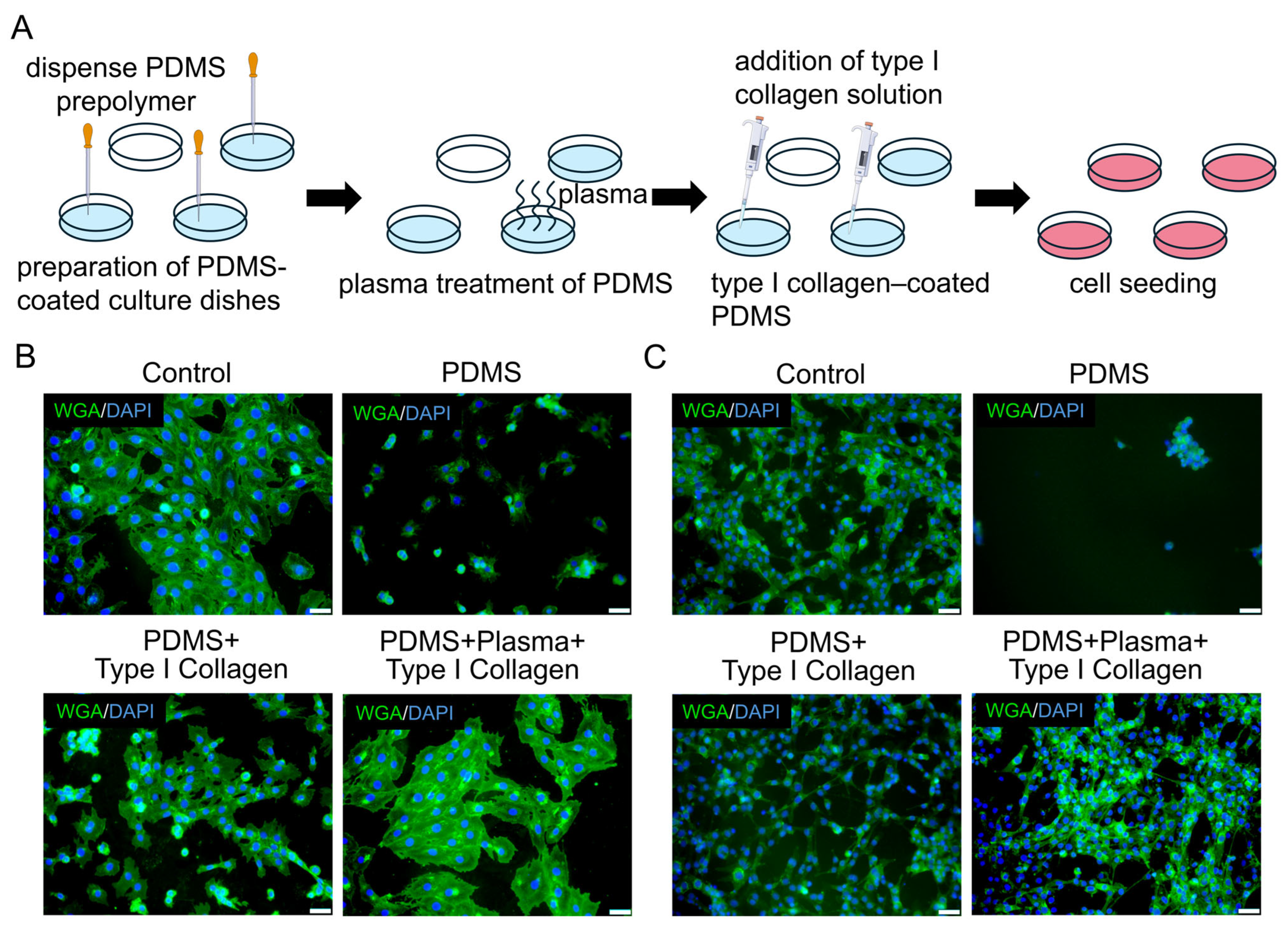
3.4. Surface Modification of PDMS to Support Myogenic Cell Adhesion and Growth
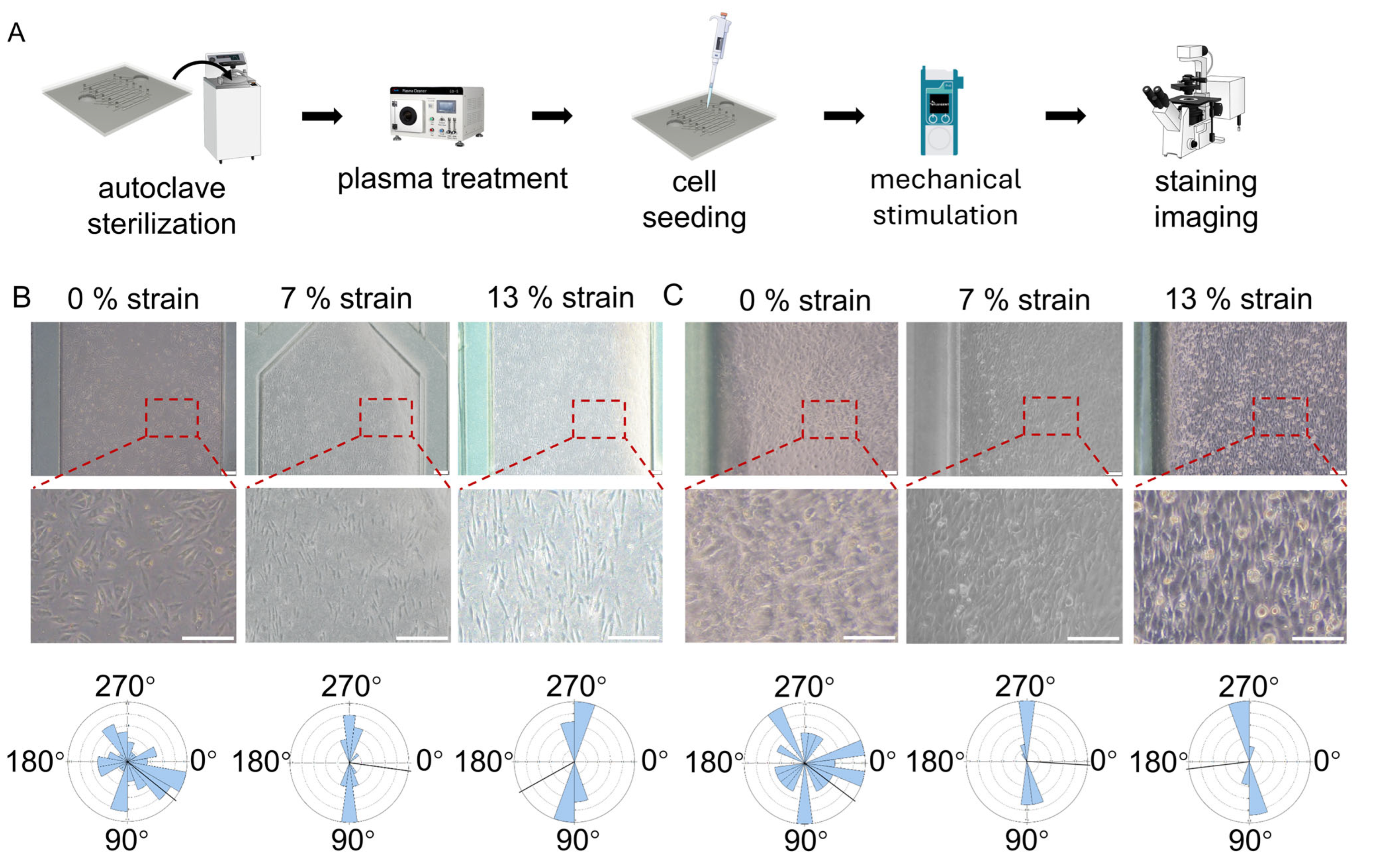
3.5. Cyclic Mechanical Stimulation Coordinately Regulates Alignment of Myogenic Cells
3.6. Cyclic Strain Enhances the Contractile Maturation of Myogenic Phenotypes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bravo-Olín, J.; Martínez-Carreón, S.A.; Francisco-Solano, E.; Lara, A.R.; Beltran-Vargas, N.E. Analysis of the role of perfusion, mechanical, and electrical stimulation in bioreactors for cardiac tissue engineering. Bioprocess Biosyst. Eng. 2024, 47, 767–839. [Google Scholar] [CrossRef]
- Calero-Castro, F.J.; Perez-Puyana, V.M.; Laga, I.; Padillo Ruiz, J.; Romero, A.; de la Portilla de Juan, F. Mechanical Stimulation and Aligned Poly(ε-caprolactone)–Gelatin Electrospun Scaffolds Promote Skeletal Muscle Regeneration. ACS Appl. Bio Mater. 2024, 7, 6430–6440. [Google Scholar] [CrossRef]
- Wang, C.; Ramahdita, G.; Genin, G.; Huebsch, N.; Ma, Z. Dynamic mechanobiology of cardiac cells and tissues: Current status and future perspective. Biophys. Rev. 2023, 4, 011314. [Google Scholar] [CrossRef] [PubMed]
- Pesce, M.; Duda, G.N.; Forte, G.; Girao, H.; Raya, A.; Roca-Cusachs, P.; Sluijter, J.P.G.; Tschöpe, C.; Van Linthout, S. Cardiac fibroblasts and mechanosensation in heart development, health and disease. Nat. Rev. Cardiol. 2023, 20, 309–324. [Google Scholar] [CrossRef] [PubMed]
- Stephanie, G.A.; Vander Roest, A.S. Cardiac disease mechanobiology: Advances using hiPSC-CMs. Front. Cardiovasc. Med. 2025, 12, 1642931. [Google Scholar] [CrossRef]
- Veltrop, R.J.A.; Kukk, M.M.; Topouzidou, K.; Didden, L.; Muchir, A.; van Steenbeek, F.G.; Schurgers, L.J.; Harakalova, M. From gene to mechanics: A comprehensive insight into the mechanobiology of LMNA mutations in cardiomyopathy. Cell Commun. Signal. 2024, 22, 197. [Google Scholar] [CrossRef]
- Visone, R.; Paoletti, C.; Cordiale, A.; Nicoletti, L.; Divieto, C.; Rasponi, M.; Chiono, V.; Occhetta, P. In Vitro Mechanical Stimulation to Reproduce the Pathological Hallmarks of Human Cardiac Fibrosis on a Beating Chip and Predict The Efficacy of Drugs and Advanced Therapies. Adv. Healthc. Mater. 2024, 13, 2301481. [Google Scholar] [CrossRef]
- Mascharak, S.; Guo, J.L.; Griffin, M.E.; Berry, C.C.; Wan, D.T.; Longaker, M. Modelling and targeting mechanical forces in organ fibrosis. Nat. Rev. Bioeng. 2024, 2, 305–323. [Google Scholar] [CrossRef]
- Sunadome, K.; Erickson, A.G.; Kah, D.; Fabry, B.; Adori, C.; Kameneva, P.; Faure, L.; Kanatani, S.; Kaucka, M.; Dehnisch Ellström, I.; et al. Directionality of developing skeletal muscles is set by mechanical forces. Nat. Commun. 2023, 14, 3060. [Google Scholar] [CrossRef]
- Hu, W.-W.; Chen, Y.-C.; Tsao, C.-W.; Chen, S.-L.; Tzeng, C.-Y. Development of a multifunctional bioreactor to evaluate the promotion effects of cyclic stretching and electrical stimulation on muscle differentiation. Bioeng. Transl. Med. 2024, 9, e10633. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Shuai, J.; Qiu, H.; Zhang, X.; Li, K.; Zhou, J.; Li, H.; Fu, L. Tri-layered immune-competent intestinal chip with synchronous peristaltic-mucosal flow dynamic for Allergenicity assessment of food proteins. Chem. Eng. J. 2025, 520, 165891. [Google Scholar] [CrossRef]
- Rios, B.; Bu, A.; Sheehan, T.; Kobeissi, H.; Kohli, S.; Shah, K.; Lejeune, E.; Raman, R. Mechanically programming anisotropy in engineered muscle with actuating extracellular matrices. Device 2023, 1, 100097. [Google Scholar] [CrossRef]
- Zhao, Y.; Landau, S.; Okhovatian, S.; Liu, C.; Lu, R.X.Z.; Lai, B.F.L.; Wu, Q.; Kieda, J.; Cheung, K.; Rajasekar, S.; et al. Integrating organoids and organ-on-a-chip devices. Nat. Rev. Bioeng. 2024, 2, 588–608. [Google Scholar] [CrossRef]
- Huang, Y.; Liu, T.; Huang, Q.; Wang, Y. From Organ-on-a-Chip to Human-on-a-Chip: A Review of Research Progress and Latest Applications. ACS Sens. 2024, 9, 3466–3488. [Google Scholar] [CrossRef]
- Wang, Z.; Liu, S.; Han, M.; Xu, J.; Qin, M.; Yang, Q.; Zeng, G.; Long, M.; Li, T.; Yin, J.; et al. 3D Printing-Electrospinning Hybrid Nanofibrous Scaffold as LEGO-Like Bricks for Modular Assembling Skeletal Muscle-on-a-Chip Functional Platform. Adv. Fiber Mater. 2024, 6, 1521–1540. [Google Scholar] [CrossRef]
- Xu, F.; Jin, H.; Liu, L.; Yang, Y.; Cen, J.; Wu, Y.; Chen, S.; Sun, D. Architecture design and advanced manufacturing of heart-on-a-chip: Scaffolds, stimulation and sensors. Microsyst. Nanoeng. 2024, 10, 96. [Google Scholar] [CrossRef]
- Mainardi, A.; Börsch, A.; Occhetta, P.; Ivanek, R.; Ehrbar, M.; Krattiger, L.; Oertle, P.; Loparic, M.; Martin, I.; Rasponi, M.; et al. An Organ-on-Chip Platform for Strain-Controlled, Tissue-Specific Compression of Cartilage and Mineralized Osteochondral Interface to Study Mechanical Overloading in Osteoarthritis. Adv. Healthc. Mater. 2025, 14, 2501588. [Google Scholar] [CrossRef]
- Li, J.; Zhang, W.; Liu, A.; Lu, Y.; Yu, L.; Liu, X.; Sun, L.; Zhao, B.; Tong, X.; Liu, T.; et al. An Engineered Anisotropic Skeletal Muscle Organoid-on-a-Chip for Deciphering Muscle Response under Intermittent Hypoxia. Adv. Funct. Mater. 2024, 34, 2401564. [Google Scholar] [CrossRef]
- Kołodziejek, D.; Tadko, O.; Iwoń, Z.; Baranowska, P.; Drozd, M.; Wojasiński, M.; Jastrzębska, E. Heart-on-a-chip: A novel microfluidic approach to organ modelling and cellular mechanobiology. Sens. Actuators A Phys. 2025, 394, 116957. [Google Scholar] [CrossRef]
- Huh, D.; Kim, H.J.; Fraser, J.P.; Shea, D.E.; Khan, M.; Bahinski, A.; Hamilton, G.A.; Ingber, D.E. Microfabrication of human organs-on-chips. Nat. Protoc. 2013, 8, 2135–2157. [Google Scholar] [CrossRef]
- Martiel, J.-L.; Leal, A.; Kurzawa, L.; Balland, M.; Wang, I.; Vignaud, T.; Tseng, Q.; Théry, M. Methods in Cell Biology; Paluch, E.K., Ed.; Academic Press: Cambridge, MA, USA, 2015; Volume 125, pp. 269–287. [Google Scholar] [CrossRef]
- Müller, A.; Wapler, M.C.; Wallrabe, U. A quick and accurate method to determine the Poisson’s ratio and the coefficient of thermal expansion of PDMS. Soft Matter 2019, 15, 779–784. [Google Scholar] [CrossRef]
- Mozneb, M.; Jenkins, A.; Sances, S.; Pohlman, S.; Workman, M.J.; West, D.; Ondatje, B.; El-Ghazawi, K.; Woodbury, A.; Garcia, V.J.; et al. Multi-lineage heart-chip models drug cardiotoxicity and enhances maturation of human stem cell-derived cardiovascular cells. Lab Chip 2024, 24, 869–881. [Google Scholar] [CrossRef]
- Shang, Y.; Xu, D.; Sun, L.; Zhao, Y.; Sun, L. A Biomimetic Optical Cardiac Fibrosis-on-a-Chip for High-Throughput Anti-Fibrotic Drug Screening. Research 2024, 7, 0471. [Google Scholar] [CrossRef]
- Lee, L.-W.; Lee, G.-H.; Su, I.H.; Lu, C.-H.; Lin, K.-H.; Wen, F.-L.; Tang, M.-J. Mechanobiological mechanism of cyclic stretch-induced cell columnarization. Cell Rep. 2025, 44, 115662. [Google Scholar] [CrossRef]
- Shi, Y.; Li, D.; Yi, B.; Tang, H.; Xu, T.; Zhang, Y. Physiological cyclic stretching potentiates the cell–cell junctions in vascular endothelial layer formed on aligned fiber substrate. Biomater. Adv. 2024, 157, 213751. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Huang, W.; Li, Z.; Wang, W.; Lei, X.; Chen, J.; Song, X.; Lu, F.; Fan, S.; Zhang, L. Transcriptome Analysis of Differentially Expressed Genes and Molecular Pathways Involved in C2C12 Cells Myogenic Differentiation. Mol. Biotechnol. 2025, 67, 3640–3655. [Google Scholar] [CrossRef]
- Yang, Y.; Dashi, A.; Soong, P.L.; Lin, K.H.; Tan, W.L.W.; Pan, B.; Autio, M.I.; Tiang, Z.; Hartman, R.J.G.; Wei, H.; et al. Long noncoding RNA VENTHEART is required for ventricular cardiomyocyte specification and function. J. Mol. Cell. Cardiol. 2024, 197, 90–102. [Google Scholar] [CrossRef] [PubMed]
- Ruan, J.-L.; Tulloch, N.L.; Saiget, M.; Paige, S.L.; Razumova, M.V.; Regnier, M.; Tung, K.C.; Keller, G.; Pabon, L.; Reinecke, H.; et al. Mechanical Stress Promotes Maturation of Human Myocardium From Pluripotent Stem Cell-Derived Progenitors. Stem Cells 2015, 33, 2148–2157. [Google Scholar] [CrossRef]
- Shimkunas, R.; Hegyi, B.; Jian, Z.; Shaw, J.A.; Kazemi-Lari, M.A.; Mitra, D.; Leach, J.K.; Li, X.; Jaradeh, M.; Balardi, N.; et al. Mechanical Load Regulates Excitation-Ca2+ Signaling-Contraction in Cardiomyocyte. Circ. Res. 2021, 128, 772–774. [Google Scholar] [CrossRef] [PubMed]
- Alghannam, F.; Alayed, M.; Alfihed, S.; Sakr, M.A.; Almutairi, D.; Alshamrani, N.; Al Fayez, N. Recent Progress in PDMS-Based Microfluidics Toward Integrated Organ-on-a-Chip Biosensors and Personalized Medicine. Biosensors 2025, 15, 76. [Google Scholar] [CrossRef]
- Guimaraes, A.P.P.; Calori, I.R.; Stilhano, R.S.; Tedesco, A.C. Renal proximal tubule-on-a-chip in PDMS: Fabrication, functionalization, and RPTEC:HUVEC co-culture evaluation. Biofabrication 2024, 16, 025024. [Google Scholar] [CrossRef]
- Greaney, A.J.; McCarthy, C.M.; Vethil, J.P.; Abubaker, M.; Reardon, E.C.; Crowley, F.D.; Cunnane, E.M.; Mulvihill, J.J.E. A comprehensive protocol for PDMS fabrication for use in cell culture. PLoS ONE 2025, 20, e0323283. [Google Scholar] [CrossRef]
- Regagnon, T.; Raynaud, F.; Subra, G.; Carnac, G.; Hugon, G.; Flatres, A.; Humblot, V.; Raymond, L.; Martin, J.; Carretero, E.; et al. A new biofunctionalized and micropatterned PDMS is able to promote stretching induced human myotube maturation. Lab Chip 2025, 25, 1586–1599. [Google Scholar] [CrossRef]
- Kefallinou, D.; Grigoriou, M.; Constantoudis, V.; Raptis, I.; Xenogiannopoulou, E.; Dimoulas, A.; Boumpas, D.T.; Tserepi, A. Enhanced and stabilized mesenchymal stem cell growth inside plasma pre-treated and collagen-coated PDMS-based microfluidic chambers. Surf. Coat. Technol. 2023, 466, 129658. [Google Scholar] [CrossRef]
- Svetlova, A.; Law, H.T.J.; Kim, D.; Enriquez, N.; Soleimani, A.; Al-Shami, A.; Kohan, S.; Peck, R.; Eremina, O.E.; Zavaleta, C.; et al. Benchtop Fabrication and Integration of Laser-Induced Graphene Strain Gauges and Stimulation Electrodes in Muscle on a Chip Devices. Adv. Funct. Mater. 2025, 35, 2417184. [Google Scholar] [CrossRef]
- Su, W.; Yang, Q.; Li, T.; Xu, J.; Yin, P.; Han, M.; Lin, Z.; Deng, Y.; Wu, Y.; Huang, W.; et al. Electrospun Aligned Nanofiber Yarns Constructed Biomimetic M-Type Interface Integrated into Precise Co-Culture System as Muscle-Tendon Junction-on-a-Chip for Drug Development. Small Methods 2024, 8, 2301754. [Google Scholar] [CrossRef] [PubMed]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhou, Z.; Chen, Z.; Bai, Z.; Chen, F.; Huang, Y.; Guo, Y. Engineering a Quantitative Organ-on-a-Chip Platform for Myogenic Mechanobiology. Bioengineering 2026, 13, 371. https://doi.org/10.3390/bioengineering13030371
Zhou Z, Chen Z, Bai Z, Chen F, Huang Y, Guo Y. Engineering a Quantitative Organ-on-a-Chip Platform for Myogenic Mechanobiology. Bioengineering. 2026; 13(3):371. https://doi.org/10.3390/bioengineering13030371
Chicago/Turabian StyleZhou, Zepeng, Zhu Chen, Zhuojun Bai, Fengling Chen, Yujuan Huang, and Yuan Guo. 2026. "Engineering a Quantitative Organ-on-a-Chip Platform for Myogenic Mechanobiology" Bioengineering 13, no. 3: 371. https://doi.org/10.3390/bioengineering13030371
APA StyleZhou, Z., Chen, Z., Bai, Z., Chen, F., Huang, Y., & Guo, Y. (2026). Engineering a Quantitative Organ-on-a-Chip Platform for Myogenic Mechanobiology. Bioengineering, 13(3), 371. https://doi.org/10.3390/bioengineering13030371







