MyoNet: Deep Learning-Based Myocardial Strain Quantification from Cine Cardiac MRI
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population and Data Preprocessing
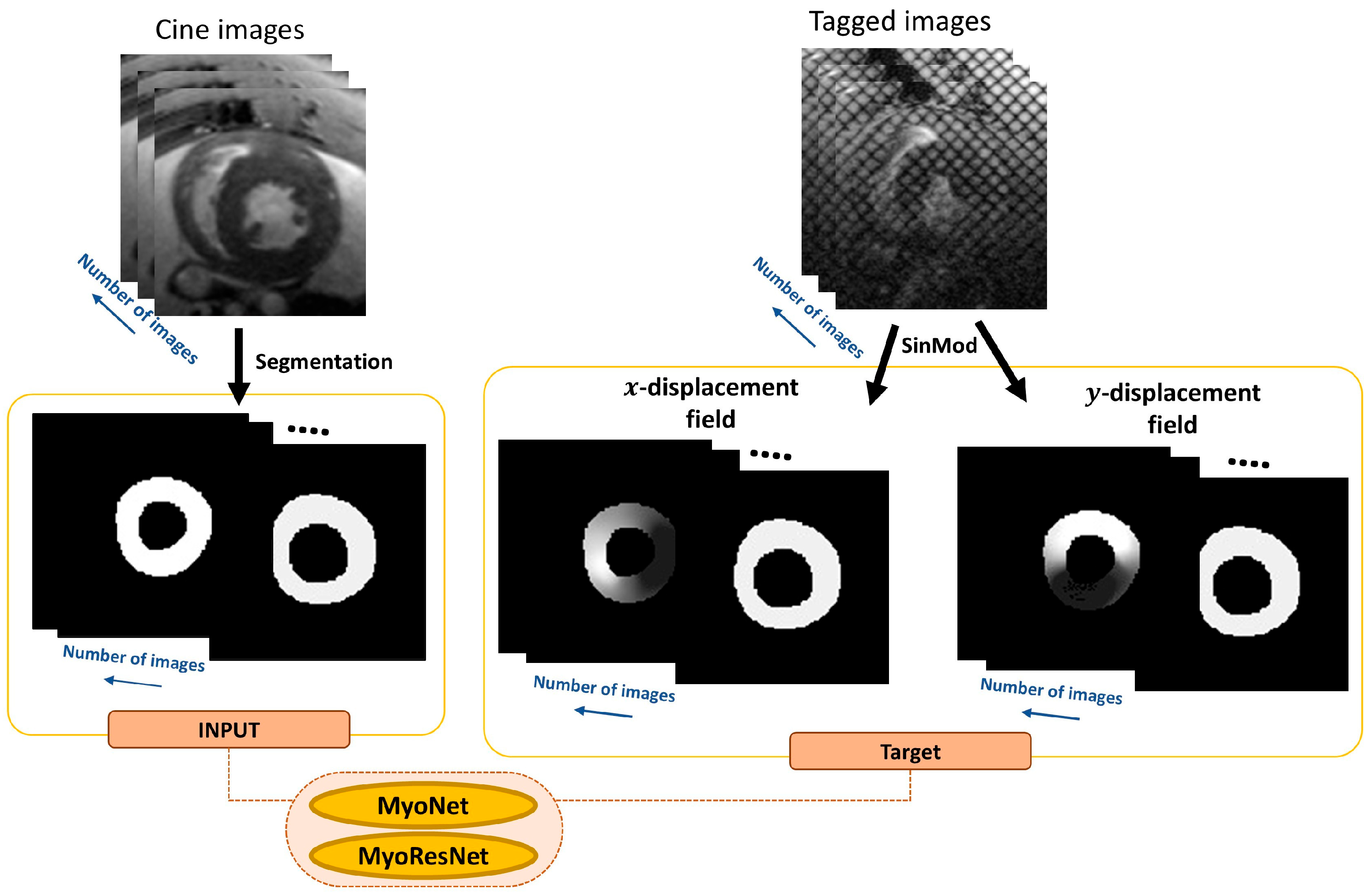
2.2. MyoNet
2.3. ResMyoNet
2.4. Experimental Setup
2.5. Optimization and Loss Functions
2.6. Statistical Analysis
2.7. Strain Computation
3. Results
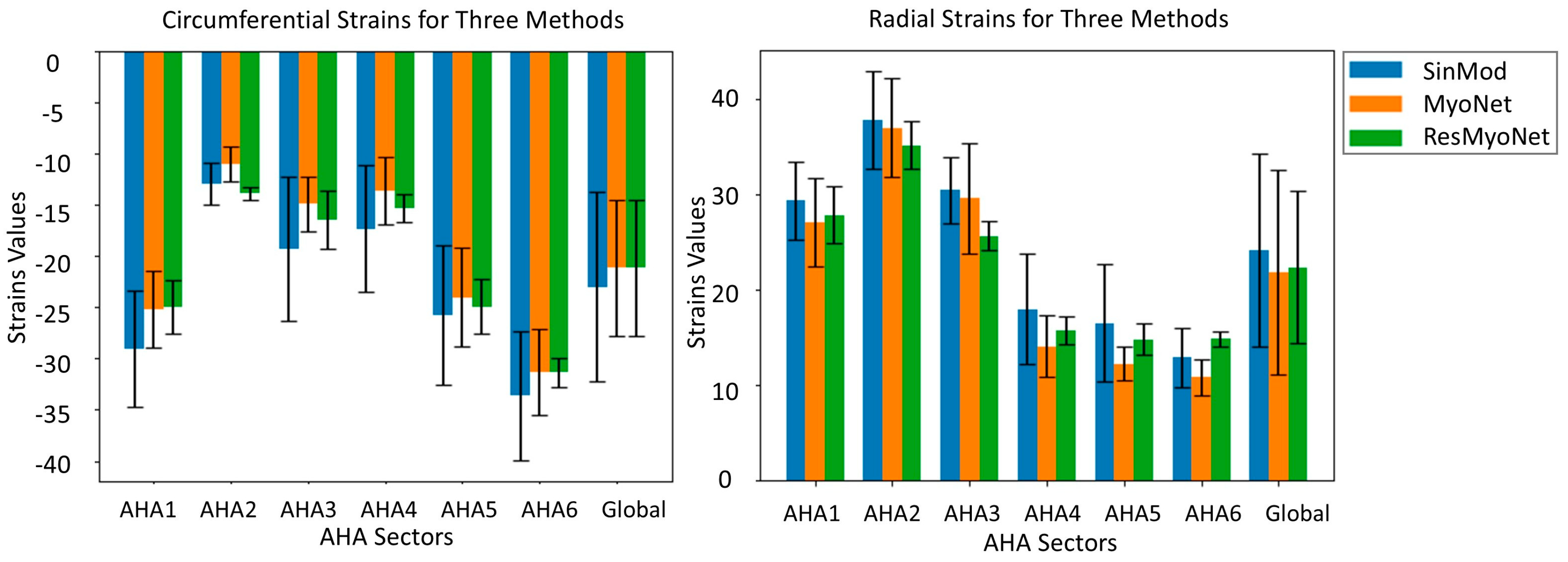
3.1. Strain Analysis by MyoNet and ResMyoNet
3.2. Model Performance and Computational Efficiency
3.3. Performance Metrics of MyoNet and ResMyoNet
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| DL | Deep learning |
| CMR | Cardiac magnetic resonance |
| RT | Radiation therapy |
| Ecc | Circumferential strain |
| Err | Radial strain |
| LVEF | Left ventricular ejection fraction |
| SS | Salt-sensitive |
| CMR-FT | CMR feature tracking |
| LV | Left ventricle |
| RMSE | Root mean squared error |
| MSE | Mean squared error |
| SSIM | Structural similarity index |
| ICC | Intraclass correlation coefficient |
| CV | Coefficient of variation |
| AHA | American Heart Association |
| SD | Standard deviation |
References
- Kongbundansuk, S.; Hundley, W.G. Noninvasive imaging of cardiovascular injury related to the treatment of cancer. JACC Cardiovasc. Imaging 2014, 7, 824–838. [Google Scholar] [CrossRef]
- Anthony, F.Y.; Ky, B. Roadmap for biomarkers of cancer therapy cardiotoxicity. Heart 2016, 102, 425–430. [Google Scholar]
- Henson, K.; McGale, P.; Taylor, C.; Darby, S. Radiation-related mortality from heart disease and lung cancer more than 20 years after radiotherapy for breast cancer. Br. J. Cancer 2013, 108, 179–182. [Google Scholar] [CrossRef]
- Carlson, L.E.; Watt, G.P.; Tonorezos, E.S.; Chow, E.J.; Yu, A.F.; Woods, M.; Lynch, C.F.; John, E.M.; Mellemkjӕr, L.; Brooks, J.D. Coronary artery disease in young women after radiation therapy for breast cancer: The WECARE study. Cardio Oncol. 2021, 3, 381–392. [Google Scholar] [CrossRef]
- Darby, S.C.; Ewertz, M.; McGale, P.; Bennet, A.M.; Blom-Goldman, U.; Brønnum, D.; Correa, C.; Cutter, D.; Gagliardi, G.; Gigante, B. Risk of ischemic heart disease in women after radiotherapy for breast cancer. N. Engl. J. Med. 2013, 368, 987–998. [Google Scholar] [CrossRef]
- Erven, K.; Florian, A.; Slagmolen, P.; Sweldens, C.; Jurcut, R.; Wildiers, H.; Voigt, J.-U.; Weltens, C. Subclinical cardiotoxicity detected by strain rate imaging up to 14 months after breast radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 2013, 85, 1172–1178. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, E.-S.H.; Baruah, D.; Croisille, P.; Stojanovska, J.; Rubenstein, J.C.; Frei, A.; Schlaak, R.A.; Lin, C.-Y.; Pipke, J.L.; Lemke, A. Cardiac magnetic resonance for early detection of radiation therapy-induced cardiotoxicity in a small animal model. Cardio Oncol. 2021, 3, 113–130. [Google Scholar] [CrossRef]
- An, D.; Ibrahim, E.-S. Elucidating Early Radiation-Induced Cardiotoxicity Markers in Preclinical Genetic Models Through Advanced Machine Learning and Cardiac MRI. J. Imaging 2024, 10, 308. [Google Scholar] [CrossRef]
- Lenarczyk, M.; Kronenberg, A.; Mäder, M.; North, P.E.; Komorowski, R.; Cheng, Q.; Little, M.P.; Chiang, I.-H.; LaTessa, C.; Jardine, J. Age at exposure to radiation determines severity of renal and cardiac disease in rats. Radiat. Res. 2019, 192, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Rapp, J. Dahl salt-susceptible and salt-resistant rats. A review. Hypertension 1982, 4, 753–763. [Google Scholar] [CrossRef]
- Axel, L.; Dougherty, L. MR imaging of motion with spatial modulation of magnetization. Radiology 1989, 171, 841–845. [Google Scholar] [CrossRef]
- Hor, K.N.; Baumann, R.; Pedrizzetti, G.; Tonti, G.; Gottliebson, W.M.; Taylor, M.; Benson, D.W.; Mazur, W. Magnetic resonance derived myocardial strain assessment using feature tracking. J. Vis. Exp. JoVE 2011, 48, 2356. [Google Scholar]
- Clarysse, P.; Croisille, P. Cardiac motion analysis in tagged MRI. In Multi-Modality Cardiac Imaging: Processing and Analysis; Wiley: Hoboken, NJ, USA, 2015; pp. 247–255. [Google Scholar]
- Ibrahim, E.-S.H.; Stojanovska, J.; Hassanein, A.; Duvernoy, C.; Croisille, P.; Pop-Busui, R.; Swanson, S.D. Regional cardiac function analysis from tagged MRI images. Comparison of techniques: Harmonic-Phase (HARP) versus Sinusoidal-Modeling (SinMod) analysis. Magn. Reson. Imaging 2018, 54, 271–282. [Google Scholar] [CrossRef]
- Attili, A.K.; Schuster, A.; Nagel, E.; Reiber, J.H.; Van der Geest, R.J. Quantification in cardiac MRI: Advances in image acquisition and processing. Int. J. Cardiovasc. Imaging 2010, 26, 27–40. [Google Scholar] [CrossRef]
- Morton, G.; Schuster, A.; Jogiya, R.; Kutty, S.; Beerbaum, P.; Nagel, E. Inter-study reproducibility of cardiovascular magnetic resonance myocardial feature tracking. J. Cardiovasc. Magn. Reson. 2012, 14, 34. [Google Scholar] [CrossRef]
- Schuster, A.; Morton, G.; Hussain, S.T.; Jogiya, R.; Kutty, S.; Asrress, K.N.; Makowski, M.R.; Bigalke, B.; Perera, D.; Beerbaum, P. The intra-observer reproducibility of cardiovascular magnetic resonance myocardial feature tracking strain assessment is independent of field strength. Eur. J. Radiol. 2013, 82, 296–301. [Google Scholar] [CrossRef]
- Kraff, O.; Quick, H.H. 7T: Physics, safety, and potential clinical applications. J. Magn. Reson. Imaging 2017, 46, 1573–1589. [Google Scholar] [CrossRef] [PubMed]
- Soher, B.J.; Dale, B.M.; Merkle, E.M. A review of MR physics: 3T versus 1.5T. Magn. Reson. Imaging Clin. N. Am. 2007, 15, 277–290. [Google Scholar] [CrossRef] [PubMed]
- Schlaak, R.A.; SenthilKumar, G.; Boerma, M.; Bergom, C. Advances in preclinical research models of radiation-induced cardiac toxicity. Cancers 2020, 12, 415. [Google Scholar] [CrossRef]
- Ibrahim, E.-S.H.; Baruah, D.; Budde, M.; Rubenstein, J.; Frei, A.; Schlaak, R.; Gore, E.; Bergom, C. Optimized cardiac functional MRI of small-animal models of cancer radiation therapy. Magn. Reson. Imaging 2020, 73, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Arts, T.; Prinzen, F.W.; Delhaas, T.; Milles, J.R.; Rossi, A.C.; Clarysse, P. Mapping displacement and deformation of the heart with local sine-wave modeling. IEEE Trans. Med. Imaging 2010, 29, 1114–1123. [Google Scholar] [CrossRef] [PubMed]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: Learning dense volumetric segmentation from sparse annotation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2016; Springer: Cham, Switzerland, 2016. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2015. [Google Scholar]
- Han, J.; Moraga, C. The influence of the sigmoid function parameters on the speed of backpropagation learning. In From Natural to Artificial Neural Computation—International Workshop on Artificial Neural Networks; Springer: Berlin/Heidelberg, Germany, 1995. [Google Scholar]
- Diakogiannis, F.I.; Waldner, F.; Caccetta, P.; Wu, C. ResUNet-a: A deep learning framework for semantic segmentation of remotely sensed data. ISPRS J. Photogramm. Remote Sens. 2020, 162, 94–114. [Google Scholar] [CrossRef]
- Nwankpa, C.; Ijomah, W.; Gachagan, A.; Marshall, S. Activation functions: Comparison of trends in practice and research for deep learning. arXiv 2018, arXiv:1811.03378. [Google Scholar] [CrossRef]
- Marquez, E.S.; Hare, J.S.; Niranjan, M. Deep cascade learning. IEEE Trans. Neural Netw. Learn. Syst. 2018, 29, 5475–5485. [Google Scholar] [CrossRef]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar] [CrossRef]
- Koch, G.G. Intraclass correlation coefficient. In Encyclopedia of Statistical Sciences; John Wiley & Sons: Hoboken, NJ, USA, 2004. [Google Scholar]
- Pearson, K., VII. Note on regression and inheritance in the case of two parents. Proc. R. Soc. Lond. 1895, 58, 240–242. [Google Scholar] [CrossRef]
- Bland, J.M.; Altman, D. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 1986, 327, 307–310. [Google Scholar] [CrossRef]
- Ibrahim, E.-S.H. Heart Mechanics: Magnetic Resonance Imaging—Mathematical Modeling, Pulse Sequences, and Image Analysis; CRC Press: Boca Raton, FL, USA, 2017. [Google Scholar]
- Cerqueira, M.D.; Weissman, N.J.; Dilsizian, V.; Jacobs, A.K.; Kaul, S.; Laskey, W.K.; Pennell, D.J.; Rumberger, J.A.; Ryan, T.; Verani, M.S.; et al. Standardized myocardial segmentation and nomenclature for tomographic imaging of the heart: A statement for healthcare professionals from the Cardiac Imaging Committee of the Council on Clinical Cardiology of the American Heart Association. Circulation 2002, 105, 539–542. [Google Scholar]
- Hammouda, K.; Khalifa, F.; Abdeltawab, H.; Elnakib, A.; Giridharan, G.; Zhu, M.; Ng, C.; Dassanayaka, S.; Kong, M.; Darwish, H. A new framework for performing cardiac strain analysis from cine MRI imaging in mice. Sci. Rep. 2020, 10, 7725. [Google Scholar] [CrossRef] [PubMed]
- Graves, C.V.; Rebelo, M.F.; Moreno, R.A.; Dantas, R.N., Jr.; Assunção, A.N., Jr.; Nomura, C.H.; Gutierrez, M.A. Siamese pyramidal deep learning network for strain estimation in 3D cardiac cine-MR. Comput. Med. Imaging Graph. 2023, 108, 102283. [Google Scholar] [CrossRef]
- Sandino, C.M.; Lai, P.; Vasanawala, S.S.; Cheng, J.Y. Accelerating cardiac cine MRI using a deep learning-based ESPIRiT reconstruction. Magn. Reson. Med. 2021, 85, 152–167. [Google Scholar] [CrossRef] [PubMed]





| MyoNet | ResMyoNet | |||
|---|---|---|---|---|
| Ecc | Err | Ecc | Err | |
| AHA 1 | t = 1.45 | t = 3.16 | t = 1.80 | t = 1.42 |
| p = 0.20 | p = 0.02 * | p = 0.12 | p = 0.20 | |
| AHA 2 | t = 1.65 | t = 0.37 | t = 0.96 | t = 1.55 |
| p = 0.15 | p = 0.72 | p = 0.38 | p = 0.17 | |
| AHA 3 | t = 1.89 | t = 0.30 | t = 1.34 | t = 2.88 |
| p = 0.11 | p = 0.77 | p = 0.46 | p = 0.03 * | |
| AHA 4 | t = 1.14 | t = 1.28 | t = 0.79 | t = 1.05 |
| p = 0.30 | p = 0.25 | p = 0.80 | p = 0.49 | |
| AHA 5 | t = 0.46 | t = 1.58 | t = 0.26 | t = 0.74 |
| p = 0.66 | p = 0.17 | p = 0.80 | p = 0.49 | |
| AHA 6 | t = 0.75 | t = 1.74 | t = 0.95 | t = −1.53 |
| p = 0.48 | p = 0.13 | p = 0.38 | p = 0.18 | |
| MyoNet | ResMyoNet | |||
|---|---|---|---|---|
| Ecc | Err | Ecc | Err | |
| SSIM | 0.961 | 0.960 | 0.937 | 0.934 |
| ICC | 0.973 | 0.975 | 0.955 | 0.955 |
| Pearson CC | 0.973 | 0.975 | 0.956 | 0.955 |
| CV | 32.447 | 34.445 | 21.749 | 22.116 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
An, D.; Nencka, A.; Clarysse, P.; Croisille, P.; Bergom, C.; Ibrahim, E.-S. MyoNet: Deep Learning-Based Myocardial Strain Quantification from Cine Cardiac MRI. Bioengineering 2026, 13, 310. https://doi.org/10.3390/bioengineering13030310
An D, Nencka A, Clarysse P, Croisille P, Bergom C, Ibrahim E-S. MyoNet: Deep Learning-Based Myocardial Strain Quantification from Cine Cardiac MRI. Bioengineering. 2026; 13(3):310. https://doi.org/10.3390/bioengineering13030310
Chicago/Turabian StyleAn, Dayeong, Andrew Nencka, Patrick Clarysse, Pierre Croisille, Carmen Bergom, and El-Sayed Ibrahim. 2026. "MyoNet: Deep Learning-Based Myocardial Strain Quantification from Cine Cardiac MRI" Bioengineering 13, no. 3: 310. https://doi.org/10.3390/bioengineering13030310
APA StyleAn, D., Nencka, A., Clarysse, P., Croisille, P., Bergom, C., & Ibrahim, E.-S. (2026). MyoNet: Deep Learning-Based Myocardial Strain Quantification from Cine Cardiac MRI. Bioengineering, 13(3), 310. https://doi.org/10.3390/bioengineering13030310






