Synthetic Melanoma Image Generation and Evaluation Using Generative Adversarial Networks
Abstract
1. Introduction
2. Related Work
2.1. GANs in Medical Imaging
2.2. Emerging Alternatives: Diffusion Models
2.3. Synthetic Data Generation for Melanoma Imaging
3. Methods
3.1. Generative Models
3.2. Experimental Setup
3.2.1. Compute Cluster
3.2.2. Datasets
3.2.3. Data Preprocessing
3.2.4. Model Parameter Exploration
3.3. Model Evaluation
4. Results
4.1. FID and FMD Performance Analysis
4.2. Image Generation
4.3. Computational Cost and Parameter Size
5. Downstream Evaluation
5.1. Evaluation Model: External Skin Lesion Classifier
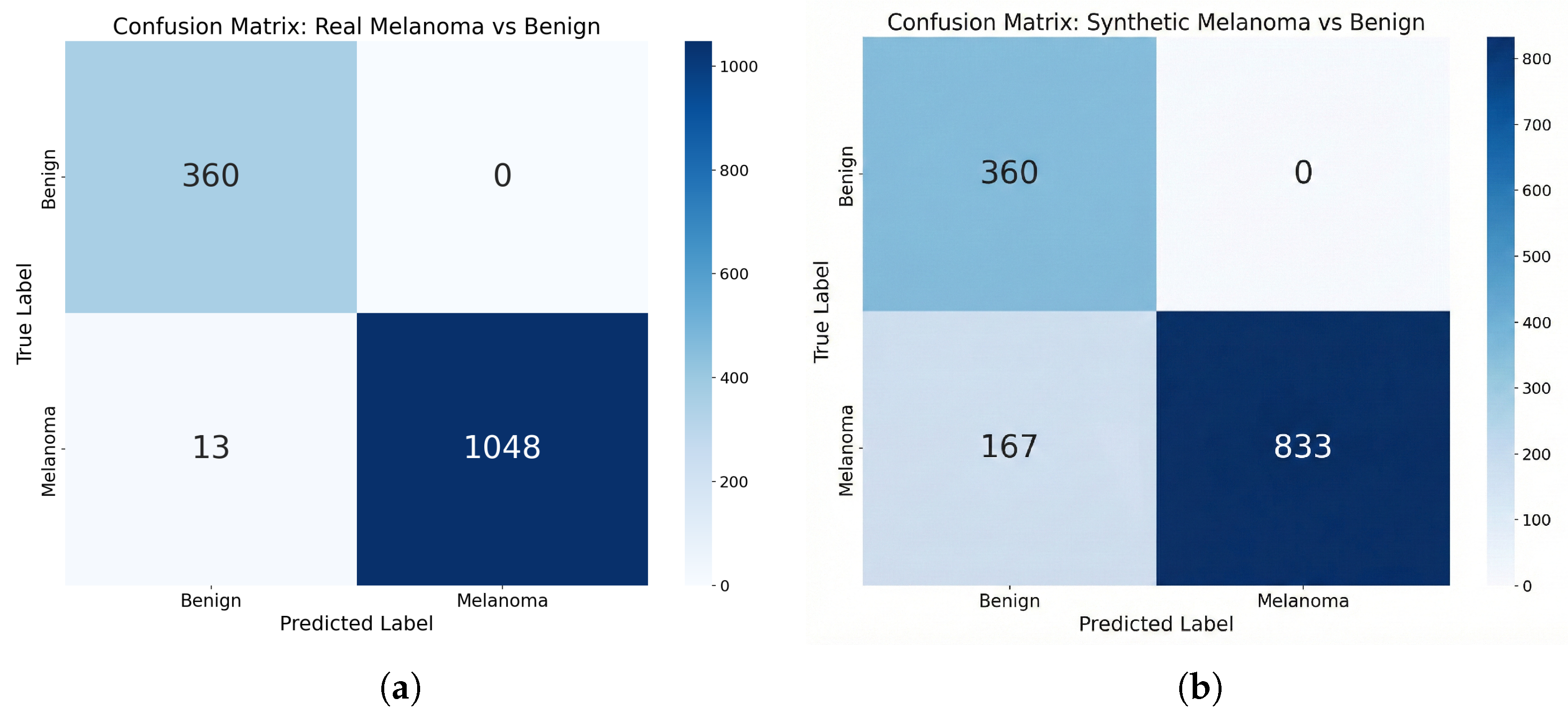
5.2. Downstream Evaluation I: Recognizability Under a Frozen Classifier
5.3. Downstream Evaluation II: Augmentation Utility
- Real-only: The model is trained exclusively on real images (), resulting in a highly imbalanced class distribution with a benign-to-melanoma ratio of approximately 98:2.
- Real + Synthetic: The training set combines all real images with 6500 StyleGAN2-generated synthetic melanoma images, yielding a more balanced benign-to-melanoma ratio of approximately 65:35.
6. Dermatologist Evaluation
6.1. Machine Baseline: StyleGAN2 Discriminator
6.2. Independent Dermatologist Assessment
6.3. Inter-Rater Reliability
6.4. Summary
7. Limitations and Future Work
8. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Kratzer, T.B.; Wagle, N.S.; Sung, H.; Jemal, A. Cancer statistics, 2026. CA Cancer J. Clin. 2026, 76, e70043. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef]
- Luke, J.J.; Flaherty, K.T.; Ribas, A.; Long, G.V. Targeted agents and immunotherapies: Optimizing outcomes in melanoma. Nat. Rev. Clin. Oncol. 2017, 14, 463–482. [Google Scholar] [CrossRef]
- Wadhawan, T.; Situ, N.; Rui, H.; Lancaster, K.; Yuan, X.; Zouridakis, G. Implementation of the 7-point checklist for melanoma detection on smart handheld devices. In Proceedings of the 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society; IEEE: New York, NY, USA, 2011; pp. 3180–3183. [Google Scholar]
- Situ, N.; Wadhawan, T.; Yuan, X.; Zouridakis, G. Modeling spatial relation in skin lesion images by the graph walk kernel. In Proceedings of the 2010 Annual International Conference of the IEEE Engineering in Medicine and Biology; IEEE: New York, NY, USA, 2010; pp. 6130–6133. [Google Scholar]
- Zouridakis, G. Methods for Screening and Diagnosing a Skin Condition. US Patent 10,593,040, 17 March 2020. [Google Scholar]
- Zareen, S.S.; Hossain, M.S.; Wang, J.; Kang, Y. Recent innovations in machine learning for skin cancer lesion analysis and classification: A comprehensive analysis of computer-aided diagnosis. Precis. Med. Sci. 2025, 14, 15–40. [Google Scholar] [CrossRef]
- Yao, P.; Shen, S.; Xu, M.; Liu, P.; Zhang, F.; Xing, J.; Shao, P.; Kaffenberger, B.; Xu, R.X. Single Model Deep Learning on Imbalanced Small Datasets for Skin Lesion Classification. IEEE Trans. Med. Imaging 2022, 41, 1242–1254. [Google Scholar] [CrossRef]
- Raza, W.H.; Shah, A.B.; Wen, Y.; Shen, Y.; Lemus, J.D.M.; Schiess, M.C.; Ellmore, T.M.; Hu, R.; Fu, X. NeuroMoE: A Transformer-Based Mixture-of-Experts Framework for Multi-Modal Neurological Disorder Classification. arXiv 2025, arXiv:2506.14970. [Google Scholar]
- Innani, S.; Dutande, P.; Baid, U.; Pokuri, V.; Bakas, S.; Talbar, S.; Baheti, B.; Guntuku, S.C. Generative adversarial networks based skin lesion segmentation. Sci. Rep. 2023, 13, 13467. [Google Scholar] [CrossRef] [PubMed]
- La Salvia, M.; Torti, E.; Leon, R.; Fabelo, H.; Ortega, S.; Martinez-Vega, B.; Callico, G.M.; Leporati, F. Deep Convolutional Generative Adversarial Networks to Enhance Artificial Intelligence in Healthcare: A Skin Cancer Application. Sensors 2022, 22, 6145. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez Alonso, A.; Sanchez Diez, A.; Cancho Galán, G.; Ibarrola Altuna, R.; Irigoyen Miró, G.; Penas Lago, C.; Boyano López, M.D.; Izu Belloso, R. Enhancing Melanoma Diagnosis in Histopathology with Deep Learning and Synthetic Data Augmentation. Bioengineering 2025, 12, 1001. [Google Scholar] [CrossRef] [PubMed]
- Fumagal-González, G.A.; Garza-Abdala, J.A.; Pérez, A.A.P.; Tamez-Pena, J.G. Assessing a Proposed Dynamic Ratio in Dataset Class Imbalance with GAN’s Generated Melanoma Images. In Proceedings of the 2025 IEEE 38th International Symposium on Computer-Based Medical Systems (CBMS), Madrid, Spain, 18–20 June 2025; pp. 500–503. [Google Scholar] [CrossRef]
- Bissoto, A.; Perez, F.; Valle, E.; Avila, S. Skin Lesion Synthesis with Generative Adversarial Networks. In Proceedings of the IEEE International Symposium on Biomedical Imaging (ISBI), Venice, Italy, 8–11 April 2019; pp. 1–4. [Google Scholar]
- Behara, K.; Bhero, E.; Agee, J.T. Skin Lesion Synthesis and Classification Using an Improved DCGAN Classifier. Diagnostics 2023, 13, 2635. [Google Scholar] [CrossRef]
- Ali, Z.; Naz, S.; Zaffar, H.; Choi, J.; Kim, Y. An IoMT-Based Melanoma Lesion Segmentation Using Conditional Generative Adversarial Networks. Sensors 2023, 23, 3548. [Google Scholar] [CrossRef] [PubMed]
- Frid-Adar, M.; Diamant, I.; Klang, E.; Amitai, M.; Goldberger, J.; Greenspan, H. GAN-Based Synthetic Medical Image Augmentation for Increased CNN Performance in Liver Lesion Classification. Neurocomputing 2018, 321, 321–331. [Google Scholar] [CrossRef]
- Karras, T.; Aila, T.; Laine, S.; Lehtinen, J. Progressive Growing of GANs for Improved Quality, Stability, and Variation. In Proceedings of the International Conference on Learning Representations (ICLR), Vancouver, BC, Canada, 30 April–3 May 2018. [Google Scholar]
- Bissoto, A.; Fornaciali, M.; Valle, E.; Avila, S. (De)constructing Bias on Skin Lesion Datasets. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), Long Beach, CA, USA, 16–17 June 2019. [Google Scholar]
- Karras, T.; Laine, S.; Aila, T. A Style-Based Generator Architecture for Generative Adversarial Networks. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 16–17 June 2019; pp. 4401–4410. [Google Scholar]
- Karras, T.; Aittala, M.; Laine, S.; Härkönen, E.; Hellsten, J.; Lehtinen, J.; Aila, T. Training Generative Adversarial Networks with Limited Data. In Proceedings of the Advances in Neural Information Processing Systems (NeurIPS), Virtual, 6–12 December 2020. [Google Scholar]
- Costa, P.; Galdran, A.; Smailagic, A.; Campilho, A. A Weakly-Supervised Framework for Interpretability of Retinal Disease Classification. IEEE Trans. Med. Imaging 2018, 37, 2539–2551. [Google Scholar]
- Dhariwal, P.; Nichol, A. Diffusion Models Beat GANs on Image Synthesis. In Proceedings of the Advances in Neural Information Processing Systems, Online, 6–14 December 2021; Volume 34, pp. 8780–8794. [Google Scholar]
- Rombach, R.; Blattmann, A.; Lorenz, D.; Esser, P.; Ommer, B. High-Resolution Image Synthesis with Latent Diffusion Models. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA, 19–20 June 2022; pp. 10684–10695. [Google Scholar]
- Akrout, M.; Gyepesi, B.; Holló, P.; Poór, A.; Kincső, B.; Solis, S.; Cirone, K.; Kawahara, J.; Slade, D.; Abid, L.; et al. Diffusion-Based Data Augmentation for Skin Disease Classification: Impact Across Original Medical Datasets to Fully Synthetic Images. In Proceedings of the Deep Generative Models, DGM4MICCAI 2023; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2024; Volume 14533, pp. 99–109. [Google Scholar] [CrossRef]
- Farooq, M.A.; Yao, W.; Schukat, M.; Little, M.A.; Corcoran, P. Derm-T2IM: Harnessing Synthetic Skin Lesion Data via Stable Diffusion Models for Enhanced Skin Disease Classification. arXiv 2024, arXiv:2401.05159. [Google Scholar] [CrossRef]
- Wang, J.; Chung, Y.; Ding, Z.; Hamm, J. From Majority to Minority: A Diffusion-based Augmentation for Underrepresented Groups in Skin Lesion Analysis. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer Nature: Cham, Switzerland, 2024; pp. 14–23. [Google Scholar]
- Abbasi, S.F.; Bilal, M.; Mukherjee, T.; Islam, S.u.; Pournik, O.; Arvanitis, T.N. Synthetic Image Generation for Skin Lesion Analysis Using Stable Diffusion Models. In Proceedings of the Studies in Health Technology and Informatics; IOS Press: Amsterdam, The Netherlands, 2025; Volume 323, pp. 81–85. [Google Scholar] [CrossRef]
- Luschi, A.; Tognetti, L.; Cartocci, A.; Cinotti, E.; Rubegni, G.; Calabrese, L.; D’onghia, M.; Dragotto, M.; Moscarella, E.; Brancaccio, G.; et al. Design and development of a systematic validation protocol for synthetic melanoma images for responsible use in medical artificial intelligence. Biocybern. Biomed. Eng. 2025, 45, 608–616. [Google Scholar] [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Nets. In Proceedings of the Advances in Neural Information Processing Systems (NeurIPS), Montreal, QC, Canada, 8–13 December 2014; pp. 2672–2680. [Google Scholar]
- Bastien, F.; Lamblin, P.; Pascanu, R.; Bergstra, J.; Goodfellow, I.; Bergeron, A.; Bouchard, N.; Warde-Farley, D.; Bengio, Y. Theano: New Features and Speed Improvements. arXiv 2012, arXiv:1211.5590. [Google Scholar] [CrossRef]
- Radford, A.; Metz, L.; Chintala, S. Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks. arXiv 2015, arXiv:1511.06434. [Google Scholar] [CrossRef]
- Karras, T.; Laine, S.; Aittala, M.; Hellsten, J.; Lehtinen, J.; Aila, T. Analyzing and Improving the Image Quality of StyleGAN. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Seattle, WA, USA, 14–19 June 2020; pp. 8110–8119. [Google Scholar]
- Karras, T.; Aittala, M.; Laine, S.; Härkönen, E.; Hellsten, J.; Lehtinen, J.; Aila, T. Alias-Free Generative Adversarial Networks. In Proceedings of the Advances in Neural Information Processing Systems (NeurIPS), Virtual, 6–14 December 2021. [Google Scholar]
- Codella, N.; Rotemberg, V.; Tschandl, P.; Celebi, M.E.; Dusza, S.; Gutman, D.; Helba, B.; Kalloo, A.; Liopyris, K.; Marchetti, M.; et al. Skin Lesion Analysis Toward Melanoma Detection 2018: A Challenge Hosted by the International Skin Imaging Collaboration (ISIC). arXiv 2019, arXiv:1902.03368. [Google Scholar] [CrossRef]
- International Skin Imaging Collaboration (ISIC). SIIM-ISIC 2020 Challenge Dataset, 2020. Available online: https://www.isic-archive.com (accessed on 11 February 2025).
- Tomasi, C.; Manduchi, R. Bilateral Filtering for Gray and Color Images. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Bombay, India, 4–7 January 1998; pp. 839–846. [Google Scholar]
- Gavaskar, R.G.; Chaudhury, K.N. Fast Adaptive Bilateral Filtering. IEEE Trans. Image Process. 2018, 28, 779–790. [Google Scholar] [CrossRef]
- Mutepfe, F.; Kalejahi, B.K.; Meshgini, S.; Danishvar, S. Generative Adversarial Network Image Synthesis Method for Skin Lesion Generation and Classification. J. Med. Signals Sens. 2021, 11, 237–252. [Google Scholar] [CrossRef]
- Heusel, M.; Ramsauer, H.; Unterthiner, T.; Nessler, B.; Hochreiter, S. GANs Trained by a Two Time-Scale Update Rule Converge to a Local Nash Equilibrium. In Proceedings of the Advances in Neural Information Processing Systems (NeurIPS), Long Beach, CA, USA, 4–9 December 2017; pp. 6626–6637. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception Architecture for Computer Vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 2818–2826. [Google Scholar]
- Abdusalomov, A.B.; Nasimov, R.; Nasimova, N.; Muminov, B.; Whangbo, T.K. Evaluating Synthetic Medical Images Using Artificial Intelligence with the GAN Algorithm. Sensors 2023, 23, 3440. [Google Scholar] [CrossRef]
- Argenziano, G.; Fabbrocini, G.; Carli, P.; De Giorgi, V.; Sammarco, E.; Delfino, M. Epiluminescence Microscopy for the Diagnosis of Doubtful Melanocytic Skin Lesions: Comparison of the ABCD Rule of Dermatoscopy and a New 7-Point Checklist. Arch. Dermatol. 1998, 134, 1563–1570. [Google Scholar] [CrossRef]
- Ha, Q.; Liu, B.; Liu, F. Identifying Melanoma Images using EfficientNet Ensemble: Winning Solution to the SIIM-ISIC Melanoma Classification Challenge. arXiv 2020, arXiv:2010.05351. [Google Scholar] [CrossRef]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef]
- Lancaster, K.; Zouridakis, G. Asymmetry Measures of Dermoscopic Images for Automated Melanoma Detection. In Proceedings of the 2023 IEEE 7th Portuguese Meeting on Bioengineering (ENBENG); IEEE: New York, NY, USA, 2023; pp. 151–154. [Google Scholar]
- Rotemberg, V.; Kurtansky, N.; Betz-Stablein, B.; Caffery, L.; Chousakos, E.; Codella, N.; Combalia, M.; Dusza, S.; Guitera, P.; Gutman, D.; et al. A patient-centric dataset of images and metadata for identifying melanomas using clinical context. Sci. Data 2021, 8, 34. [Google Scholar] [CrossRef]
- Queen, C.M.; Hu, R.; Zouridakis, G. Towards the development of reliable and economical mHealth solutions: A methodology for accurate detection of Buruli ulcer for hard-to-reach communities. Front. Trop. Dis. 2023, 3, 1031352. [Google Scholar] [CrossRef]






| Metric | DCGAN | StyleGAN2 | StyleGAN3-T | StyleGAN3-R |
|---|---|---|---|---|
| FID ↓ | 66.49 | 31.58 | 246.42 | 26.47 |
| FMD ↓ | 695.93 | 50.08 | 41.37 | 49.21 |
| FID (ISIC 2018) | FID (ISIC 2020) | |
|---|---|---|
| 0.8 | 24.8 | 7.96 |
| 1.6 | 27.4 | 9.48 |
| 8.0 | 31.6 | 9.91 |
| 10.0 | 33.2 | 10.4 |
| DCGAN | StyleGAN2 | StyleGAN3-T | StyleGAN3-R | |
|---|---|---|---|---|
| Parameters (M), | 4 + 1 | 30 + 29 | 25 + 29 | 25 + 29 |
| Training time (hours) | 0.9 | 2.8 | 9.2 | 9.2 |
| Training Data | Accuracy (%) † | Melanoma AUC | Melanoma F1 |
|---|---|---|---|
| Real-only | 85.07 | 0.9252 | 0.1682 |
| Real + Synthetic | 98.27 | 0.9445 | 0.2586 |
| Metric | Dermatologist 1 | Dermatologist 2 | Human Mean | Discriminator |
|---|---|---|---|---|
| Overall Accuracy | 71.0% (p < 0.001) | 62.0% (p < 0.001) | 66.5% | 59.5% |
| Real Accuracy | 51.0% | 70.0% | 60.5% | 35.0% |
| Synthetic Accuracy | 91.0% | 54.0% | 72.5% | 84.0% |
| Accepted as Real † | 9.0% | 46.0% | 27.5% | 16.0% |
| Comparison | Cohen’s | p-Value | Agreement |
|---|---|---|---|
| Dermatologist 1 vs. Dermatologist 2 | 0.173 | 0.009 | Slight |
| Dermatologist 1 vs. Discriminator | 0.042 | 0.482 | Slight |
| Dermatologist 2 vs. Discriminator | 0.082 | 0.187 | Slight |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Lin, P.-Y.; Shen, Y.; Mathew, N.; Hu, R.; Huang, S.; Queen, C.M.; West, C.E.; Ciurea, A.; Zouridakis, G. Synthetic Melanoma Image Generation and Evaluation Using Generative Adversarial Networks. Bioengineering 2026, 13, 245. https://doi.org/10.3390/bioengineering13020245
Lin P-Y, Shen Y, Mathew N, Hu R, Huang S, Queen CM, West CE, Ciurea A, Zouridakis G. Synthetic Melanoma Image Generation and Evaluation Using Generative Adversarial Networks. Bioengineering. 2026; 13(2):245. https://doi.org/10.3390/bioengineering13020245
Chicago/Turabian StyleLin, Pei-Yu, Yidan Shen, Neville Mathew, Renjie Hu, Siyu Huang, Courtney M. Queen, Cameron E. West, Ana Ciurea, and George Zouridakis. 2026. "Synthetic Melanoma Image Generation and Evaluation Using Generative Adversarial Networks" Bioengineering 13, no. 2: 245. https://doi.org/10.3390/bioengineering13020245
APA StyleLin, P.-Y., Shen, Y., Mathew, N., Hu, R., Huang, S., Queen, C. M., West, C. E., Ciurea, A., & Zouridakis, G. (2026). Synthetic Melanoma Image Generation and Evaluation Using Generative Adversarial Networks. Bioengineering, 13(2), 245. https://doi.org/10.3390/bioengineering13020245







