A Systematic Review of Contrastive Learning in Medical AI: Foundations, Biomedical Modalities, and Future Directions
Abstract
1. Introduction
- We propose an operational, domain-aware taxonomy of medical CL methods grounded in core design components, including loss functions, positive and negative pairing strategies, augmentation policies, and evaluation regimes.
- We synthesize CL applications across key medical modalities, including medical imaging, electronic health records, physiological time series, genomics and proteomics, and multimodal vision language systems, highlighting modality-specific challenges and transferable design patterns.
- We critically examine methodological limitations in the literature, with emphasis on evaluation heterogeneity, reproducibility constraints, data access limitations, and the need for standardized benchmarking and external validation.
- We provide practical guidance and prioritized future directions for clinically trustworthy CL, focusing on robustness to distribution shift, interpretability, fairness, privacy-preserving training, and deployment considerations.
2. Methodology
2.1. Databases and Search Strategy
- (“contrastive learning” OR “self-supervised”) AND (medical OR healthcare OR clinical)
- (“contrastive learning” OR SimCLR OR MoCo OR BYOL) AND (radiology OR “chest x-ray” OR MRI OR CT)
- (“contrastive learning” OR “self-supervised”) AND (“electronic health record” OR EHR OR “clinical notes”)
- (“contrastive learning” OR “representation learning”) AND (ECG OR EEG OR “physiological signals”)
- (“contrastive learning” OR “self-supervised”) AND (genomics OR proteomics OR “single-cell”)
2.2. Eligibility Criteria
- Peer-reviewed journal articles and full conference papers; influential preprints were included selectively when they introduced widely adopted methods, benchmarks, or were heavily cited in subsequent peer-reviewed work.
- Studies that explicitly employed CL or closely related objectives (e.g., InfoNCE, supervised contrastive loss, MoCo/SimCLR/BYOL-style frameworks, CLIP-style alignment).
- Studies using medical or biomedical data (medical imaging, EHRs, physiological time series, genomics/proteomics, pathology, or multimodal combinations).
- Non-medical studies without biomedical datasets or clinically motivated tasks.
- Abstract-only records, posters lacking sufficient methodological detail, editorials, commentaries, and theses.
- Non-validated technical reports and preprints without experimental results, or preprints superseded by peer-reviewed versions.
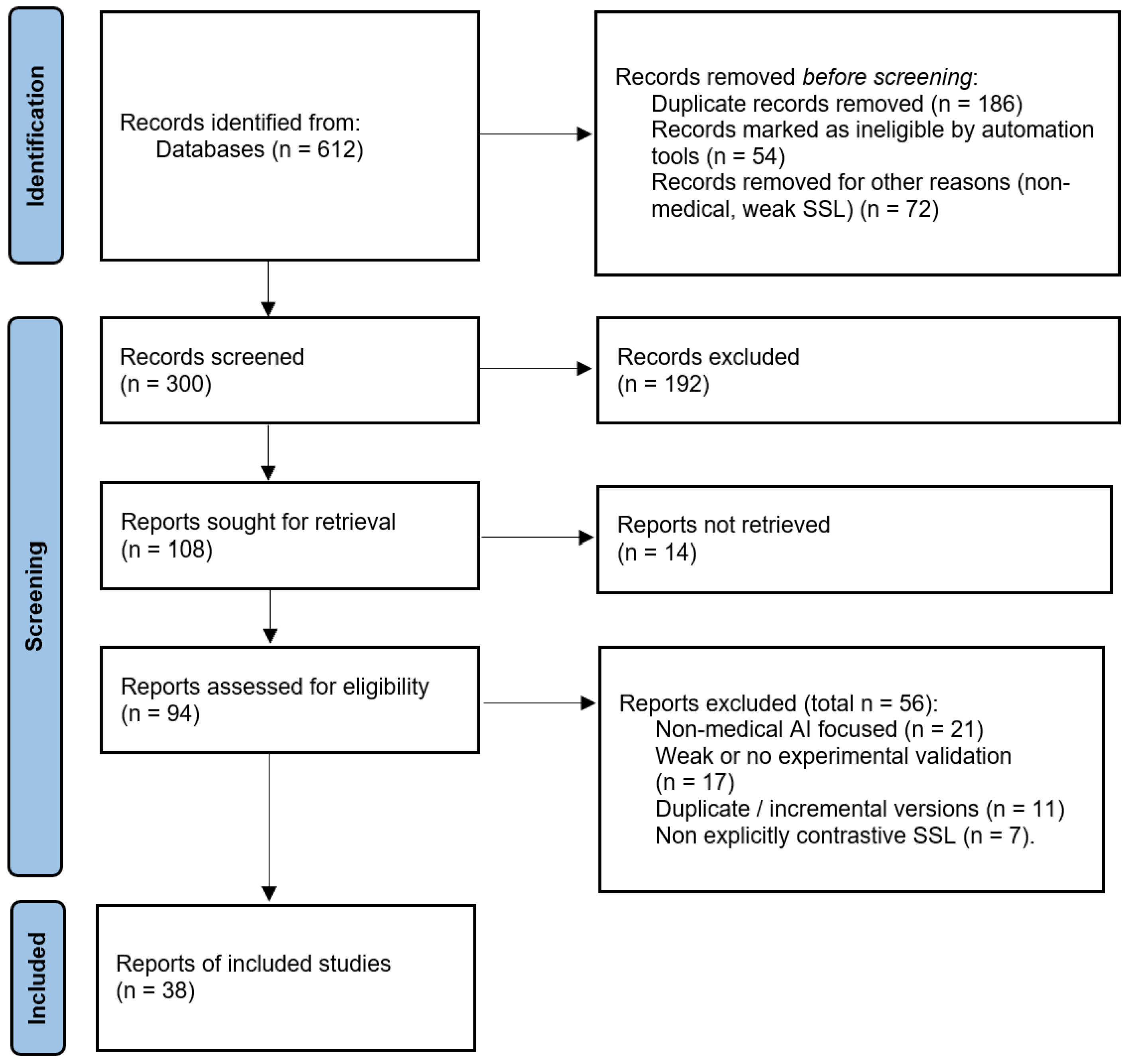
2.3. Study Selection
2.4. Data Extraction
- Modality and dataset(s);
- Contrastive formulation (loss, supervision regime, pairing strategy, negative sampling);
- Encoder architecture and pretraining scale;
- Downstream task(s), evaluation regime (linear probe, fine-tuning, few-shot/zero-shot), and metric(s);
- Reported limitations, external validation, and reproducibility artifacts (e.g., code availability).
2.5. Taxonomy of Medical Contrastive Learning
2.6. Quality and Reporting Appraisal
2.7. Protocol Registration
3. Overview of Contrastive Learning

3.1. Variants and Extensions of Contrastive Learning
3.1.1. InfoNCE-Based Contrastive Learning (SimCLR, MoCo, SupCon)
3.1.2. Negative-Free Self-Distillation (BYOL, DINO, SimSiam)
3.1.3. Clustering-Based Self-Supervised Learning (SwAV)
3.1.4. Multimodal Contrastive Alignment (CLIP, ConVIRT, BioViL)
3.1.5. Temporal and Patient-Aware Objectives (CPC, CLOCS/PCLR)
3.1.6. Optimization Refinements (Hard Negatives, Debiased Objectives)
3.2. Datasets and Benchmarks for Medical Contrastive Learning
| Modality | Representative Datasets/Benchmarks | Typical Downstream Tasks |
|---|---|---|
| Chest imaging | MIMIC-CXR [84]; CheXpert [85]; NIH ChestX-ray14 [86] | Multi-label classification (AUROC); label-scarce transfer; external validation; retrieval and zero-shot classification |
| General medical imaging | BraTS [88,89]; ISIC Archive [90]; ADNI [87] | Segmentation (Dice/HD95); lesion classification; progression prediction; robustness across scanners and time |
| Computational pathology | CAMELYON [91,92]; TCGA-derived slide repositories [98] | WSI classification; region retrieval; weakly supervised detection; cross-site generalization |
| EHR (clinical records) | MIMIC-III [93]; MIMIC-IV [94]; eICU [95] | Mortality and LOS prediction; readmission; phenotyping; temporal outcome modeling; multimodal fusion with notes |
| Physiological signals | PTB-XL [96]; TUH EEG [97] | Diagnosis classification; seizure detection; patient-level transfer; low-label training |
| Genomics and transcriptomics | TCGA [98]; GTEx [99]; UK Biobank [100] | Subtype prediction; survival and risk modeling; representation transfer across cohorts; cross-tissue generalization |
| Single-cell omics | Human Cell Atlas [101] | Cell-type clustering; batch correction; rare cell detection; perturbation response prediction |
4. Applications of Contrastive Learning in Medical AI
4.1. Medical Imaging
4.2. Electronic Health Records
4.3. Genomics and Proteomics
4.4. Multimodal and Cross-Domain Learning
4.5. Time-Series and Physiological Signal Analysis
5. Challenges, Limitations, and Practical Considerations
5.1. Pair Construction and Clinical Semantics
5.2. Augmentation and View Design in Medical Data
5.3. Heterogeneity Across Modalities and Tasks
5.4. Reproducibility and Reporting Gaps
5.5. Multimodal Alignment and Weak Supervision Risks
5.6. Interpretability and Explainability
5.7. Fairness, Bias, and Clinical Validity
6. Discussion and Future Research Directions
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| AUROC | Area Under the Receiver Operating Characteristic Curve |
| AUPRC | Area Under the Precision–Recall Curve |
| BioViL | Biomedical Vision–Language |
| BioViL-T | Biomedical Vision–Language with Temporal Alignment |
| BiomedCLIP | Biomedical CLIP |
| BYOL | Bootstrap Your Own Latent |
| CheXpert | CheXpert dataset |
| CL | Contrastive Learning |
| CLOCS | Contrastive Learning of Cardiac Signals |
| COMET | Hierarchical contrastive framework |
| CONCH | Histopathology foundation model |
| ConVIRT | Contrastive Learning of Visual Representations from Text |
| CPC | Contrastive Predictive Coding |
| CT | Computed Tomography |
| CXR-CLIP | Chest X-Ray CLIP |
| Dice | Dice coefficient |
| DINO | Self-Distillation with No Labels |
| ECG | Electrocardiogram |
| EEG | Electroencephalogram |
| EHR(s) | Electronic Health Record(s) |
| F1 | F1 score |
| GLoRIA | Global–Local image–text alignment in radiology |
| HD95 | 95th percentile Hausdorff Distance |
| ICU | Intensive Care Unit |
| IID | Independent and Identically Distributed |
| InfoNCE | Info Noise-Contrastive Estimation (loss) |
| MaCo | Masked Contrastive Learning |
| MBSL | Multi-Scale and Multimodal Contrastive Learning |
| MedCLIP | Medical CLIP |
| mIoU | mean Intersection over Union |
| MIMIC-CXR | Medical Information Mart for Intensive Care–Chest X-Ray |
| MoCo | Momentum Contrast |
| MRI | Magnetic Resonance Imaging |
| MSCL | Multi-Scale Contrastive Learning |
| OOD | Out-of-Distribution |
| PCLR | Patient Contrastive Learning of Representations |
| PLIP | Pathology Language–Image Pretraining |
| PMQ | Patient Memory Queue |
| PPI(s) | Protein–Protein Interaction(s) |
| PMC-CLIP | PubMed Central CLIP |
| RSNA | Radiological Society of North America |
| SAM | Segment Anything Model |
| scRNA-seq | single-cell RNA sequencing |
| SimCLR | Simple Framework for Contrastive Learning of Visual Representations |
| SSL | Self-Supervised Learning |
| SupCon | Supervised Contrastive Learning |
| SwAV | Swapping Assignments between Multiple Views |
| VQA | Visual Question Answering |
| 1D CNN | One-dimensional Convolutional Neural Network |
Appendix A. Database-Specific Search Strings
Appendix A.1. PubMed
| ( (``contrastive learning’’[Title/Abstract] OR ``self-supervised’’[Title/Abstract] OR ``self supervised’’[Title/Abstract] OR ``representation learning’’[Title/Abstract] OR SimCLR[Title/Abstract] OR MoCo[Title/Abstract] OR BYOL[Title/Abstract] OR ``supervised contrastive’’[Title/Abstract] OR InfoNCE[Title/Abstract] OR CLIP[Title/Abstract]) AND (medical[Title/Abstract] OR healthcare[Title/Abstract] OR clinical[Title/Abstract] OR radiology[Title/Abstract] OR ``chest x-ray’’[Title/Abstract] OR CXR[Title/Abstract] OR MRI[Title/Abstract] OR CT[Title/Abstract] OR pathology[Title/Abstract] OR ``electronic health record’’[Title/Abstract] OR EHR[Title/Abstract] OR ``clinical notes’’[Title/Abstract] OR ECG[Title/Abstract] OR EEG[Title/Abstract] OR ``physiological signals’’[Title/Abstract] OR genomics[Title/Abstract] OR proteomics[Title/Abstract] OR ``single-cell’’[Title/Abstract] OR ``multi-omics’’[Title/Abstract]) ) |
- Filters: Publication dates: 1 January 2019–31 October 2025; Article types: journal articles and conference papers where applicable; Language: English.
Appendix A.2. IEEE Xplore
| (``contrastive learning’’ OR ``self-supervised’’ OR ``self supervised’’ OR ``representation learning’’ OR SimCLR OR MoCo OR BYOL OR ``supervised contrastive’’ OR InfoNCE OR CLIP) AND (medical OR healthcare OR clinical OR radiology OR ``chest x-ray’’ OR CXR OR MRI OR CT OR pathology OR ``electronic health record’’ OR EHR OR ``clinical notes’’ OR ECG OR EEG OR ``physiological signals’’ OR genomics OR proteomics OR ``single-cell’’ OR ``multi-omics’’) |
- Filters: Years: 2019–2025; Content type: Journals and Conferences; Fields searched: All Metadata.
Appendix A.3. ACM Digital Library
| (``contrastive learning’’ OR ``self-supervised’’ OR ``representation learning’’ OR SimCLR OR MoCo OR BYOL OR ``supervised contrastive’’ OR InfoNCE OR CLIP) AND (medical OR healthcare OR clinical OR radiology OR ``chest x-ray’’ OR MRI OR CT OR pathology OR ``electronic health record’’ OR EHR OR ``clinical notes’’ OR ECG OR EEG OR ``physiological signals’’ OR genomics OR proteomics OR ``single-cell’’ OR ``multi-omics’’) |
- Filters: Publication years: 2019–2025.
Appendix A.4. Web of Science
| TS=( (``contrastive learning’’ OR ``self-supervised’’ OR ``self supervised’’ OR ``representation learning’’ OR SimCLR OR MoCo OR BYOL OR ``supervised contrastive’’ OR InfoNCE OR CLIP) AND (medical OR healthcare OR clinical OR radiology OR ``chest x-ray’’ OR CXR OR MRI OR CT OR pathology OR ``electronic health record’’ OR EHR OR ``clinical notes’’ OR ECG OR EEG OR ``physiological signals’’ OR genomics OR proteomics OR ``single-cell’’ OR ``multi-omics’’) ) |
- Filters: Timespan: 2019–2025; Document types: Article, Proceedings Paper; Language: English.
Appendix A.5. Scopus
| TITLE-ABS-KEY( (``contrastive learning’’ OR ``self-supervised’’ OR ``self supervised’’ OR ``representation learning’’ OR SimCLR OR MoCo OR BYOL OR ``supervised contrastive’’ OR InfoNCE OR CLIP) AND (medical OR healthcare OR clinical OR radiology OR ``chest x-ray’’ OR CXR OR MRI OR CT OR pathology OR ``electronic health record’’ OR EHR OR ``clinical notes’’ OR ECG OR EEG OR ``physiological signals’’ OR genomics OR proteomics OR ``single-cell’’ OR ``multi-omics’’) ) |
- Filters: Year: 2019–2025; Document type: Article, Conference Paper; Language: English.
Appendix A.6. arXiv
| (``contrastive learning’’ OR ``self-supervised’’ OR ``representation learning’’ OR SimCLR OR MoCo OR BYOL OR ``supervised contrastive’’ OR InfoNCE OR CLIP) AND (medical OR healthcare OR clinical OR radiology OR ``chest x-ray’’ OR MRI OR CT OR pathology OR ``electronic health record’’ OR EHR OR ``clinical notes’’ OR ECG OR EEG OR ``physiological signals’’ OR genomics OR proteomics OR ``single-cell’’ OR ``multi-omics’’) |
- Filters: Years: 2019–2025; Categories considered: cs.LG, cs.CV, cs.AI, eess.IV, q-bio.QM.
Appendix B. Reporting Checklist
| Checklist Item | Rationale for Medical CL |
|---|---|
| Patient-level split reported (when applicable) | Reduces leakage from repeated exams/segments across train and test. |
| External validation (cross-site/device/cohort) | Supports generalization beyond in-domain evaluation. |
| Pretraining corpus described (size, source, pairing) | Enables reproducibility and interpretation of representation quality. |
| Pairing strategy specified (what defines positives/negatives) | Central determinant of clinical semantics and false-negative risk. |
| Augmentation policy specified and clinically justified | Prevents invariances that erase subtle pathology or amplify artifacts. |
| Evaluation regime clearly stated (linear probe, fine-tune, few-shot, zero-shot) | Needed for fair comparison across studies. |
| Primary metric(s) justified for task (e.g., AUROC vs F1) | Metrics have different clinical meanings under imbalance and thresholding. |
| Baselines appropriate (supervised, SSL, ImageNet, CL variants) | Prevents inflated claims due to weak comparisons. |
| Code/model released or sufficient implementation detail provided | Enables replication and reuse. |
| Reproducibility constraints discussed (private data, restricted access) | Helps interpret evidence strength and limitations. |
Appendix C. Included Studies Summary Table
| Modality | Study | Year | Dataset(s) | Task | Contrastive Formulation | Label Regime | Eval Protocol | Key Metric |
|---|---|---|---|---|---|---|---|---|
| CXR | Azizi et al. [16] | 2021 | Large-scale chest X-rays (multi-source CXR) | Classification/transfer | Image–image CL (SimCLR-style): augmented views positives; others negatives | SSL pretrain; supervised transfer | IID and external transfer evaluation | AUROC/Acc |
| MRI | Chaitanya et al. [105] | 2020 | Cardiac/brain MRI segmentation benchmarks | Segmentation (limited labels) | Semi-supervised CL: consistency + image-view positives/negatives | SS (few labels + unlabeled) | CV or held-out split | Dice/mIoU |
| Histopathology | Ciga et al. [106] | 2022 | WSI patch datasets (public pathology cohorts) | Cancer/tissue classification | Patch–patch SSL CL (augmentation-invariant representations) | SSL pretrain; supervised fine-tune | Patient-level IID split (where applicable) | AUROC/F1 |
| Cardiac MRI | Guo et al. [107] | 2023 | Cardiac MRI segmentation dataset(s) | Segmentation (myocardium/ventricles) | Multi-scale CL loss (global/local scale alignment) | Supervised + auxiliary CL | Train/val/test or CV | Dice |
| Brain MRI | Luo et al. [108] | 2023 | Brain MRI anomaly datasets | Anomaly detection/localization | Patch-level CL (normality representation learning) | SSL/weak supervision | IID split; lesion localization eval | AUROC/ AUPRC |
| EHR | Krishnan et al. [109] | 2022 | ICU/hospital EHR cohort(s) | Mortality/HF prediction | Patient view–view CL: time-window crops, code dropout/shuffle | SSL + supervised downstream | IID and/or temporal split | AUROC |
| EHR | Pick et al. [110] | 2024 | Hospital/ICU EHR cohort(s) | Mortality + length-of-stay | Patient embedding CL (same-patient/clinical-views positives) | SSL/weak sup + supervised eval | IID or temporal split | AUROC/ MAE |
| EHR (codes + notes) | Sun et al. [114] | 2024 | Paired structured EHR + clinical notes | Progression/ complications | Cross-modal CL (align code-based and note-based embeddings) | Weak supervision (paired modalities) | IID; optional cross-site test | AUROC |
| EHR (large-scale) | Cai et al. [115] | 2024 | Large EHR warehouse(s) | Generalizable patient modeling | Distributed CL pretraining (large memory bank/shards) | SSL pretrain; supervised fine-tune | Cross-population/cross-site when available | AUROC |
| EHR survival | Kerdabadi et al. [111] | 2023 | Large longitudinal EHR cohort for AKI risk forecasting | Survival risk prediction (AKI) | Temporal distinctiveness CL: hardness-aware negative sampling; time-aware pairs | Supervised survival + CL auxiliary loss | Patient-level split; temporal evaluation | C-index/ AUROC |
| EHR (missing/irregular) | Liu et al. [112] | 2023 | Two real-world EHR datasets (ICU/hospital cohorts) | In-hospital mortality prediction (with imputation) | CL-imputation–prediction network: patient stratification + contrastive repr. learning | Supervised prediction + CL auxiliary | IID split; patient-level where applicable | AUROC |
| EHR | Zang and Wang [113] | 2021 | Longitudinal EHR cohorts (risk prediction benchmarks) | Mortality + phenotyping (multi-label) | Supervised CL loss: contrastive cross-entropy + supervised CL regularizer | Supervised | IID split | AUROC/ micro-F1 |
| Genomics | Zhong et al. [116] | 2024 | Bulk genomics datasets (expression/pathways) | Biomarker discovery/prediction | Multi-scale CL (gene-level + pathway-level alignment) | Supervised + CL regularization or SSL + FT | IID; CV common | AUROC/F1 |
| Proteins | Bepler and Berger [119] | 2021 | Protein sequence corpora (+ structures) | Function/structure prediction | Contrastive protein sequence representations | SSL pretrain; supervised downstream | IID; family holdout possible | Acc/AUROC |
| Multi-omics | Liu et al. [117] | 2022 | Multi-omics cohorts (paired omics) | Disease susceptibility/outcome prediction | Cross-omics CL: align patient embeddings across modalities | Weak sup (paired omics) + supervised task | IID; CV | AUROC |
| scRNA-seq | Li et al. [118] | 2024 | Multiple scRNA-seq datasets | Clustering/rare cell discovery | Cell–cell CL with augmented count views; batch robustness | SSL | Cross-dataset/batch-aware evaluation | ARI/NMI |
| Seq + structure | Zhang et al. [120] | 2024 | Paired peptide sequence–structure datasets | PPI prediction | Sequence–structure CL alignment | Weak sup (paired modalities) + supervised | IID + cold-start split | AUROC |
| CXR + report | Zhang et al. [18] | 2022 | Paired CXR–report corpora (e.g., MIMIC-CXR) | CXR repr. learning; classification/retrieval | Image–text CL (CLIP-style) | Weak sup (paired) | IID; external transfer | AUROC/ Recall@K |
| CXR + report | Huang et al. [65] | 2021 | MIMIC-CXR | Classification/retrieval/ grounding | Global–local image–text CL (region–phrase + global) | Weak sup (paired) | IID; external optional | AUROC/ Recall@K |
| CXR + report | Boecking et al. [66] | 2022 | Large CXR-report corpora | ZS/FS radiology benchmarks | BioViL vision–language CL | Weak sup (paired) + ZS eval | IID; external | AUROC |
| CXR (temporal) | Bannur et al. [67] | 2023 | MIMIC-CXR (longitudinal pairs) | Disease progression tracking | Temporal CL across patient studies/time | Weak sup (paired + temporal IDs) | Patient split; temporal test | AUROC |
| CXR + prompt | You et al. [68] | 2023 | CheXpert/MIMIC-CXR | Prompt-based recognition | Prompt/label-text alignment CL | Weak sup + supervised finetune | IID; external | AUROC |
| CXR (ZS) | Tiu et al. [69] | 2022 | Pretrain paired CXR–report; eval CheXpert | Zero-shot multi-label classification | CLIP-style image–text CL; prompt inference | Weak sup + ZS eval | External benchmark | AUROC |
| Radiology VLP | Wang et al. [70] | 2022 | Paired + unpaired image/text corpora | Radiology classification robustness | Decoupled/knowledge-aware CL matching | Weak supervision | IID; external validation | AUROC |
| Pathology VLP | Huang et al. [71] | 2023 | Pathology image–text corpora | Zero-shot pathology transfer | CLIP-style PLIP | Weak sup (paired) | Cross-dataset transfer | AUROC |
| Histo VLP | Lu et al. [72] | 2024 | 1.17 M histopathology image–caption pairs | Retrieval/segmentation transfer | Large-scale image–text CL | Weak sup | Multi-task transfer evaluation | AUROC/ Dice |
| Biomedical VLP | Zhang et al. [73] | 2023 | PubMed-scale biomedical image–text | Zero/few-shot across tasks | CLIP-style biomedical CL | Weak sup | Benchmark suite | AUROC/ Recall@K |
| PMC VLP | Lin et al. [74] | 2023 | 1.6 M PMC figure–caption pairs | VQA/retrieval | Figure–caption image–text CL | Weak sup | Benchmark evaluation | VQA Acc/ Recall@K |
| CXR masked CL | Huang et al. [121] | 2024 | MIMIC-CXR/CheXpert | ZS + localization | Masked CL + image–text alignment | Weak sup | IID + external evaluation | AUROC |
| Segmentation (multi) | Koleilat et al. [122] | 2024 | Ultrasound/MRI/CT segmentation sets | Text-driven segmentation | MedCLIP + SAM; text prompts guide masks | Weak sup (prompts) | Multi-dataset transfer | Dice/mIoU |
| EHR (structured) | Liu et al. [112] | 2023 | MIMIC-III; eICU | Clinical time-series imputation + in-hospital mortality prediction | Imputation: unsupervised CL (patient vs augmented view) | Hybrid: SSL (imputation) + supervised (mortality labels) | Random split 70/15/15 train/val/test; repeated 10 runs | Imputation: MAE/MRE; Prediction: AUROC/AUPRC |
| ECG | Diamant et al. [19] | 2022 | Repeated ECG per patient cohorts | Cardiac disease prediction | Patient-level CL (same-patient positives) | SSL + supervised downstream | Patient split | AUROC |
| ECG | Yuan et al. [124] | 2025 | ECG recordings (windowed) | Classification repr. learning | Poly-window CL (overlap-aware positives) | SSL | IID/patient split | AUROC |
| ECG/EEG | Wang et al. [125] | 2023 | ECG and/or EEG datasets | Few-label classification | Hierarchical CL (instance/segment/patient) | SSL/SS + supervised FT | Few-label protocol | AUROC/F1 |
| ECG | Chen et al. [76] | 2025 | Multi-lead ECG datasets | Robust classification (lead/time shift) | Spatiotemporal CL (lead-wise + time-wise invariance) | SSL | Patient split; robustness tests | AUROC |
| Physio + labs | Raghu et al. [126] | 2022 | Multimodal ICU time-series | Outcome prediction | Multimodal temporal CL (align modalities/time contexts) | Weak sup (paired) | Temporal or IID split | AUROC |
| Wearables | Guo et al. [127] | 2025 | Respiration/HR/ motion datasets | Multi-task inference | Multi-scale multimodal CL | SSL + supervised tasks | IID/subject split | AUROC/F1 |
| ECG | Sun et al. [128] | 2025 | Repeated ECG per patient datasets | Reduce false negatives in CL pretraining | Patient memory queue (intra-patient positives) | SSL + supervised downstream | Patient split | AUROC |
References
- Parvin, N.; Joo, S.W.; Jung, J.H.; Mandal, T.K. Multimodal AI in Biomedicine: Pioneering the Future of Biomaterials, Diagnostics, and Personalized Healthcare. Nanomaterials 2025, 15, 895. [Google Scholar] [CrossRef] [PubMed]
- Nazir, A.; Hussain, A.; Singh, M.; Assad, A. Deep learning in medicine: Advancing healthcare with intelligent solutions and the future of holography imaging in early diagnosis. Multimed. Tools Appl. 2025, 84, 17677–17740. [Google Scholar] [CrossRef]
- Mienye, I.D.; Swart, T.G.; Obaido, G.; Jordan, M.; Ilono, P. Deep convolutional neural networks in medical image analysis: A review. Information 2025, 16, 195. [Google Scholar] [CrossRef]
- Mienye, I.D.; Jere, N.; Obaido, G.; Ogunruku, O.O.; Esenogho, E.; Modisane, C. Large language models: An overview of foundational architectures, recent trends, and a new taxonomy. Discov. Appl. Sci. 2025, 7, 1027. [Google Scholar] [CrossRef]
- Nichyporuk, B.; Cardinell, J.; Szeto, J.; Mehta, R.; Falet, J.P.R.; Arnold, D.L.; Tsaftaris, S.A.; Arbel, T. Rethinking generalization: The impact of annotation style on medical image segmentation. arXiv 2022, arXiv:2210.17398. [Google Scholar] [CrossRef]
- Daneshjou, R.; Yuksekgonul, M.; Cai, Z.R.; Novoa, R.; Zou, J.Y. Skincon: A skin disease dataset densely annotated by domain experts for fine-grained debugging and analysis. Adv. Neural Inf. Process. Syst. 2022, 35, 18157–18167. [Google Scholar]
- Krenzer, A.; Makowski, K.; Hekalo, A.; Fitting, D.; Troya, J.; Zoller, W.G.; Hann, A.; Puppe, F. Fast machine learning annotation in the medical domain: A semi-automated video annotation tool for gastroenterologists. Biomed. Eng. Online 2022, 21, 33. [Google Scholar] [CrossRef]
- Chen, H.; Gouin-Vallerand, C.; Bouchard, K.; Gaboury, S.; Couture, M.; Bier, N.; Giroux, S. Contrastive Self-Supervised Learning for Sensor-Based Human Activity Recognition: A Review. IEEE Access 2024, 12, 152511–152531. [Google Scholar] [CrossRef]
- Liu, S.; Zhao, L.; Chen, D.; Song, Z. Contrastive learning for image complexity representation. arXiv 2024, arXiv:2408.03230. [Google Scholar] [CrossRef]
- Ren, X.; Wei, W.; Xia, L.; Huang, C. A comprehensive survey on self-supervised learning for recommendation. ACM Comput. Surv. 2025, 58, 1–38. [Google Scholar] [CrossRef]
- Prince, J.S.; Alvarez, G.A.; Konkle, T. Contrastive learning explains the emergence and function of visual category-selective regions. Sci. Adv. 2024, 10, eadl1776. [Google Scholar] [CrossRef]
- Albelwi, S. Survey on self-supervised learning: Auxiliary pretext tasks and contrastive learning methods in imaging. Entropy 2022, 24, 551. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Asmatullah, L.; Malik, A.; Khan, S.; Asif, H. A Survey on Self-supervised Contrastive Learning for Multimodal Text-Image Analysis. arXiv 2025, arXiv:2503.11101. [Google Scholar]
- Zeng, D.; Wu, Y.; Hu, X.; Xu, X.; Shi, Y. Contrastive learning with synthetic positives. In Proceedings of the European Conference on Computer Vision, Milan, Italy, 29 September–4 October 2024; Springer: Berlin/Heidelberg, Germany, 2024; pp. 430–447. [Google Scholar]
- Xu, Z.; Dai, Y.; Liu, F.; Wu, B.; Chen, W.; Shi, L. Swin MoCo: Improving parotid gland MRI segmentation using contrastive learning. Med. Phys. 2024, 51, 5295–5307. [Google Scholar] [CrossRef] [PubMed]
- Azizi, S.; Mustafa, B.; Ryan, F.; Beaver, Z.; Freyberg, J.; Deaton, J.; Loh, A.; Karthikesalingam, A.; Kornblith, S.; Chen, T.; et al. Big self-supervised models advance medical image classification. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 11–17 October 2021; pp. 3478–3488. [Google Scholar]
- Sowrirajan, H.; Yang, J.; Ng, A.Y.; Rajpurkar, P. Moco pretraining improves representation and transferability of chest x-ray models. In Proceedings of the Medical Imaging with Deep Learning, Lübeck, Germany, 7–9 July 2021; PMLR: New York, NY, USA, 2021; pp. 728–744. [Google Scholar]
- Zhang, Y.; Jiang, H.; Miura, Y.; Manning, C.D.; Langlotz, C.P. Contrastive learning of medical visual representations from paired images and text. In Proceedings of the Machine Learning for Healthcare Conference, Durham, NC, USA, 5–6 August 2022; PMLR: New York, NY, USA, 2022; pp. 2–25. [Google Scholar]
- Diamant, N.; Reinertsen, E.; Song, S.; Aguirre, A.D.; Stultz, C.M.; Batra, P. Patient contrastive learning: A performant, expressive, and practical approach to electrocardiogram modeling. PLoS Comput. Biol. 2022, 18, e1009862. [Google Scholar] [CrossRef]
- Jaiswal, A.; Babu, A.R.; Zadeh, M.Z.; Banerjee, D.; Makedon, F. A survey on contrastive self-supervised learning. Technologies 2020, 9, 2. [Google Scholar] [CrossRef]
- Gui, J.; Chen, T.; Zhang, J.; Cao, Q.; Sun, Z.; Luo, H.; Tao, D. A Survey on Self-supervised Learning: Algorithms, Applications, and Future Trends. IEEE Trans. Pattern Anal. Mach. Intell. 2024, 46, 9052–9071. [Google Scholar] [CrossRef]
- Hu, H.; Wang, X.; Zhang, Y.; Chen, Q.; Guan, Q. A comprehensive survey on contrastive learning. Neurocomputing 2024, 610, 128645. [Google Scholar] [CrossRef]
- Liu, R. Understand and improve contrastive learning methods for visual representation: A review. arXiv 2021, arXiv:2106.03259. [Google Scholar] [CrossRef]
- Shurrab, S.; Duwairi, R. Self-supervised learning methods and applications in medical imaging analysis: A survey. PeerJ Comput. Sci. 2022, 8, e1045. [Google Scholar] [CrossRef]
- Wang, W.C.; Ahn, E.; Feng, D.; Kim, J. A review of predictive and contrastive self-supervised learning for medical images. Mach. Intell. Res. 2023, 20, 483–513. [Google Scholar] [CrossRef]
- Huang, S.C.; Pareek, A.; Jensen, M.; Lungren, M.P.; Yeung, S.; Chaudhari, A.S. Self-supervised learning for medical image classification: A systematic review and implementation guidelines. NPJ Digit. Med. 2023, 6, 74. [Google Scholar] [CrossRef] [PubMed]
- VanBerlo, B.; Hoey, J.; Wong, A. A survey of the impact of self-supervised pretraining for diagnostic tasks in medical X-ray, CT, MRI, and ultrasound. BMC Med. Imaging 2024, 24, 79. [Google Scholar] [CrossRef] [PubMed]
- Yeh, C.H.; Hong, C.Y.; Hsu, Y.C.; Liu, T.L.; Chen, Y.; LeCun, Y. Decoupled contrastive learning. In Proceedings of the European Conference on Computer Vision, Tel Aviv, Israel, 23–27 October 2022; Springer: Berlin/Heidelberg, Germany, 2022; pp. 668–684. [Google Scholar]
- Khosla, P.; Teterwak, P.; Wang, C.; Sarna, A.; Tian, Y.; Isola, P.; Maschinot, A.; Liu, C.; Krishnan, D. Supervised contrastive learning. Adv. Neural Inf. Process. Syst. 2020, 33, 18661–18673. [Google Scholar]
- Tian, Y.; Sun, C.; Poole, B.; Krishnan, D.; Schmid, C.; Isola, P. What makes for good views for contrastive learning? Adv. Neural Inf. Process. Syst. 2020, 33, 6827–6839. [Google Scholar]
- Kalantidis, Y.; Sariyildiz, M.B.; Pion, N.; Weinzaepfel, P.; Larlus, D. Hard negative mixing for contrastive learning. Adv. Neural Inf. Process. Syst. 2020, 33, 21798–21809. [Google Scholar]
- Wu, J.; Chen, J.; Wu, J.; Shi, W.; Wang, X.; He, X. Understanding contrastive learning via distributionally robust optimization. Adv. Neural Inf. Process. Syst. 2024, 36, 23297–23320. [Google Scholar]
- Le-Khac, P.H.; Healy, G.; Smeaton, A.F. Contrastive representation learning: A framework and review. IEEE Access 2020, 8, 193907–193934. [Google Scholar] [CrossRef]
- Falcon, W.; Cho, K. A framework for contrastive self-supervised learning and designing a new approach. arXiv 2020, arXiv:2009.00104. [Google Scholar] [CrossRef]
- Peng, X.; Wang, K.; Zhu, Z.; Wang, M.; You, Y. Crafting better contrastive views for siamese representation learning. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA, 18–24 June 2022; pp. 16031–16040. [Google Scholar]
- Kim, B.; Ye, J.C. Energy-based contrastive learning of visual representations. Adv. Neural Inf. Process. Syst. 2022, 35, 4358–4369. [Google Scholar]
- Wu, L.; Zhuang, J.; Chen, H. Voco: A simple-yet-effective volume contrastive learning framework for 3d medical image analysis. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 16–22 June 2024; pp. 22873–22882. [Google Scholar]
- Tang, C.; Zeng, X.; Zhou, L.; Zhou, Q.; Wang, P.; Wu, X.; Ren, H.; Zhou, J.; Wang, Y. Semi-supervised medical image segmentation via hard positives oriented contrastive learning. Pattern Recognit. 2024, 146, 110020. [Google Scholar] [CrossRef]
- Kundu, R. The Beginner’s Guide to Contrastive Learning. 2022. Available online: https://www.v7labs.com/blog/contrastive-learning-guide (accessed on 10 October 2025).
- Zhang, C.; Zhang, K.; Pham, T.X.; Niu, A.; Qiao, Z.; Yoo, C.D.; Kweon, I.S. Dual temperature helps contrastive learning without many negative samples: Towards understanding and simplifying moco. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA, 18–24 June 2022; pp. 14441–14450. [Google Scholar]
- Hoffmann, D.T.; Behrmann, N.; Gall, J.; Brox, T.; Noroozi, M. Ranking info noise contrastive estimation: Boosting contrastive learning via ranked positives. In Proceedings of the AAAI Conference on Artificial Intelligence, Online, 22 February–1 March 2022; Volume 36, pp. 897–905. [Google Scholar]
- Xu, L.; Xie, H.; Li, Z.; Wang, F.L.; Wang, W.; Li, Q. Contrastive learning models for sentence representations. ACM Trans. Intell. Syst. Technol. 2023, 14, 1–34. [Google Scholar] [CrossRef]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A simple framework for contrastive learning of visual representations. In Proceedings of the International Conference on Machine Learning, Virtual Event, 13–18 July 2020; PMLR: New York, NY, USA, 2020; pp. 1597–1607. [Google Scholar]
- Zhang, H.; Cao, Y. Understanding the benefits of simclr pre-training in two-layer convolutional neural networks. arXiv 2024, arXiv:2409.18685. [Google Scholar]
- Bunyang, S.; Thedwichienchai, N.; Pintong, K.; Lael, N.; Kunaborimas, W.; Boonrat, P.; Siriborvornratanakul, T. Self-supervised learning advanced plant disease image classification with SimCLR. Adv. Comput. Intell. 2023, 3, 18. [Google Scholar] [CrossRef]
- Fırıldak, K.; Çelik, G.; Talu, M.F. SimCLR-based Self-Supervised Learning Approach for Limited Brain MRI and Unlabeled Images. Bitlis Eren Üniv. Fen Bilim. Derg. 2024, 13, 1304–1313. [Google Scholar] [CrossRef]
- Chen, X.; Fan, H.; Girshick, R.; He, K. Improved baselines with momentum contrastive learning. arXiv 2020, arXiv:2003.04297. [Google Scholar] [CrossRef]
- Li, Y.; Liu, Q.; Zhou, L.; Zhao, W.; Tian, Y.; Zhang, W. Improved contrastive learning with MoCo framework. In Proceedings of the 2023 3rd International Conference on Consumer Electronics and Computer Engineering (ICCECE), Guangzhou, China, 6–8 January 2023; IEEE: Piscataway, NJ, USA, 2023; pp. 729–732. [Google Scholar]
- He, Y.; Wang, X.; Shi, T. Ddpm-moco: Advancing industrial surface defect generation and detection with generative and contrastive learning. In Proceedings of the International Joint Conference on Artificial Intelligence, Jeju, Republic of Korea, 3–9 August 2024; Springer: Berlin/Heidelberg, Germany, 2024; pp. 34–49. [Google Scholar]
- Xie, E.; Ding, J.; Wang, W.; Zhan, X.; Xu, H.; Sun, P.; Li, Z.; Luo, P. Detco: Unsupervised contrastive learning for object detection. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 11–17 October 2021; pp. 8392–8401. [Google Scholar]
- Xiao, T.; Wang, X.; Efros, A.A.; Darrell, T. What should not be contrastive in contrastive learning. arXiv 2020, arXiv:2008.05659. [Google Scholar] [CrossRef]
- Grill, J.B.; Strub, F.; Altché, F.; Tallec, C.; Richemond, P.; Buchatskaya, E.; Doersch, C.; Avila Pires, B.; Guo, Z.; Gheshlaghi Azar, M.; et al. Bootstrap your own latent-a new approach to self-supervised learning. Adv. Neural Inf. Process. Syst. 2020, 33, 21271–21284. [Google Scholar]
- Deng, Z.; Man, J.; Song, Z.; Yang, G. A Few-Shot Anomaly Detection Method Based on BYOL Contrastive Learning Framework. In Proceedings of the 2024 4th International Conference on Robotics, Automation and Intelligent Control (ICRAIC), Changsha, China, 6–9 December 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 439–443. [Google Scholar]
- Richemond, P.H.; Grill, J.B.; Altché, F.; Tallec, C.; Strub, F.; Brock, A.; Smith, S.; De, S.; Pascanu, R.; Piot, B.; et al. Byol works even without batch statistics. arXiv 2020, arXiv:2010.10241. [Google Scholar] [CrossRef]
- Caron, M.; Touvron, H.; Misra, I.; Jégou, H.; Mairal, J.; Bojanowski, P.; Joulin, A. Emerging properties in self-supervised vision transformers. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 10–17 October 2021; pp. 9650–9660. [Google Scholar]
- Takanami, K.; Takahashi, T.; Sakata, A. The effect of optimal self-distillation in noisy gaussian mixture model. arXiv 2025, arXiv:2501.16226. [Google Scholar] [CrossRef]
- Wang, Q.; Mao, Z.; Gao, J.; Zhang, Y. Document-level relation extraction with progressive self-distillation. ACM Trans. Inf. Syst. 2024, 42, 1–34. [Google Scholar] [CrossRef]
- Tong, S.; Xia, Z.; Alahi, A.; He, X.; Shi, Y. GeoDistill: Geometry-Guided Self-Distillation for Weakly Supervised Cross-View Localization. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Honolulu, HI, USA, 19–23 October 2025; pp. 25357–25366. [Google Scholar]
- Chen, X.; He, K. Exploring simple siamese representation learning. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Nashville, TN, USA, 20–25 June 2021; pp. 15750–15758. [Google Scholar]
- Zhang, C.; Zhang, K.; Zhang, C.; Pham, T.X.; Yoo, C.D.; Kweon, I.S. How does simsiam avoid collapse without negative samples? A unified understanding with self-supervised contrastive learning. arXiv 2022, arXiv:2203.16262. [Google Scholar]
- Lu, Y.; Jha, A.; Deng, R.; Huo, Y. Contrastive learning meets transfer learning: A case study in medical image analysis. In Proceedings of the Medical Imaging 2022: Computer-Aided Diagnosis, San Diego, CA, USA, 20–24 February 2022; SPIE: Bellingham, WA, USA, 2022; Volume 12033, pp. 729–736. [Google Scholar]
- Khorram, S.; Kim, J.; Tripathi, A.; Lu, H.; Zhang, Q.; Sak, H. Contrastive siamese network for semi-supervised speech recognition. In Proceedings of the ICASSP 2022—2022 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Singapore, 22–27 May 2022; IEEE: Piscataway, NJ, USA, 2022; pp. 7207–7211. [Google Scholar]
- Caron, M.; Misra, I.; Mairal, J.; Goyal, P.; Bojanowski, P.; Joulin, A. Unsupervised learning of visual features by contrasting cluster assignments. Adv. Neural Inf. Process. Syst. 2020, 33, 9912–9924. [Google Scholar]
- Radford, A.; Kim, J.W.; Hallacy, C.; Ramesh, A.; Goh, G.; Agarwal, S.; Sastry, G.; Askell, A.; Mishkin, P.; Clark, J.; et al. Learning transferable visual models from natural language supervision. In Proceedings of the International Conference on Machine Learning, Virtual, 18–24 July 2021; PMLR: New York, NY, USA, 2021; pp. 8748–8763. [Google Scholar]
- Huang, S.C.; Shen, L.; Lungren, M.P.; Yeung, S. Gloria: A multimodal global-local representation learning framework for label-efficient medical image recognition. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 11–17 October 2021; pp. 3942–3951. [Google Scholar]
- Boecking, B.; Usuyama, N.; Bannur, S.; Castro, D.C.; Schwaighofer, A.; Hyland, S.; Wetscherek, M.; Naumann, T.; Nori, A.; Alvarez-Valle, J.; et al. Making the most of text semantics to improve biomedical vision–language processing. In Proceedings of the European Conference on Computer Vision, Tel Aviv, Israel, 23–27 October 2022; Springer: Berlin/Heidelberg, Germany, 2022; pp. 1–21. [Google Scholar]
- Bannur, S.; Hyland, S.; Liu, Q.; Perez-Garcia, F.; Ilse, M.; Castro, D.C.; Boecking, B.; Sharma, H.; Bouzid, K.; Thieme, A.; et al. Learning to exploit temporal structure for biomedical vision-language processing. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Vancouver, BC, Canada, 17–24 June 2023; pp. 15016–15027. [Google Scholar]
- You, K.; Gu, J.; Ham, J.; Park, B.; Kim, J.; Hong, E.K.; Baek, W.; Roh, B. Cxr-clip: Toward large scale chest x-ray language-image pre-training. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Vancouver, BC, Canada, 8–12 October 2023; Springer: Berlin/Heidelberg, Germany, 2023; pp. 101–111. [Google Scholar]
- Tiu, E.; Talius, E.; Patel, P.; Langlotz, C.P.; Ng, A.Y.; Rajpurkar, P. Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning. Nat. Biomed. Eng. 2022, 6, 1399–1406. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wu, Z.; Agarwal, D.; Sun, J. Medclip: Contrastive learning from unpaired medical images and text. In Proceedings of the Conference on Empirical Methods in Natural Language Processing. Conference on Empirical Methods in Natural Language Processing, Abu Dhabi, United Arab Emirates, 7–11 December 2022; Volume 2022, p. 3876. [Google Scholar]
- Huang, Z.; Bianchi, F.; Yuksekgonul, M.; Montine, T.J.; Zou, J. A visual–language foundation model for pathology image analysis using medical twitter. Nat. Med. 2023, 29, 2307–2316. [Google Scholar] [CrossRef]
- Lu, M.Y.; Chen, B.; Williamson, D.F.; Chen, R.J.; Liang, I.; Ding, T.; Jaume, G.; Odintsov, I.; Le, L.P.; Gerber, G.; et al. A visual-language foundation model for computational pathology. Nat. Med. 2024, 30, 863–874. [Google Scholar] [CrossRef]
- Zhang, S.; Xu, Y.; Usuyama, N.; Xu, H.; Bagga, J.; Tinn, R.; Preston, S.; Rao, R.; Wei, M.; Valluri, N.; et al. Biomedclip: A multimodal biomedical foundation model pretrained from fifteen million scientific image-text pairs. arXiv 2023, arXiv:2303.00915. [Google Scholar]
- Lin, W.; Zhao, Z.; Zhang, X.; Wu, C.; Zhang, Y.; Wang, Y.; Xie, W. Pmc-clip: Contrastive language-image pre-training using biomedical documents. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Vancouver, BC, Canada, 8–12 October 2023; Springer: Berlin/Heidelberg, Germany, 2023; pp. 525–536. [Google Scholar]
- van den Oord, A.; Li, Y.; Vinyals, O. Representation learning with contrastive predictive coding. arXiv 2018, arXiv:1807.03748. [Google Scholar]
- Chen, W.; Wang, H.; Zhang, L.; Zhang, M. Temporal and spatial self supervised learning methods for electrocardiograms. Sci. Rep. 2025, 15, 6029. [Google Scholar] [CrossRef]
- Chuang, C.Y.; Robinson, J.; Lin, Y.C.; Torralba, A.; Jegelka, S. Debiased contrastive learning. Adv. Neural Inf. Process. Syst. 2020, 33, 8765–8775. [Google Scholar]
- Robinson, J.; Sun, L.; Yu, K.; Batmanghelich, K.; Jegelka, S.; Sra, S. Can contrastive learning avoid shortcut solutions? Adv. Neural Inf. Process. Syst. 2021, 34, 4974–4986. [Google Scholar] [PubMed]
- Jang, T.; Wang, X. Difficulty-based sampling for debiased contrastive representation learning. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Vancouver, BC, Canada, 17–24 June 2023; pp. 24039–24048. [Google Scholar]
- Biswas, D.; Tešić, J. Unsupervised domain adaptation with debiased contrastive learning and support-set guided pseudolabeling for remote sensing images. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2024, 17, 3197–3210. [Google Scholar] [CrossRef]
- Agarwal, A.; Banerjee, T.; Romine, W.L.; Cajita, M. Debias-clr: A contrastive learning based debiasing method for algorithmic fairness in healthcare applications. In Proceedings of the 2024 IEEE International Conference on Big Data (BigData), Washington DC, USA, 15–18 December 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 6411–6419. [Google Scholar]
- Yun, B.; Zhao, S.; Li, Q.; Kot, A.; Wang, Y. Debiasing Medical Knowledge for Prompting Universal Model in CT Image Segmentation. IEEE Trans. Med. Imaging 2025, 44, 5142–5154. [Google Scholar] [CrossRef] [PubMed]
- Tang, P.; Ouyang, C.; Liu, Y. Debiasing medication recommendation with counterfactual analysis. In Proceedings of the International Conference on Neural Information Processing, Changsha, China, 20–23 November 2023; Springer: Berlin/Heidelberg, Germany, 2023; pp. 426–438. [Google Scholar]
- Johnson, A.E.W.; Pollard, T.J.; Berkowitz, S.J.; Greenbaum, N.R.; Lungren, M.P.; Deng, C.Y.; Mark, R.G.; Horng, S. MIMIC-CXR: A large publicly available database of labeled chest radiographs. arXiv 2019, arXiv:1901.07042. [Google Scholar]
- Irvin, J.; Rajpurkar, P.; Ko, M.; Yu, Y.; Ciurea-Ilinca, M.; Chute, C.; Marklund, H.; Haghgoo, B.; Ball, R.L.; Shpanskaya, K.; et al. CheXpert: A Large Chest Radiograph Dataset with Uncertainty Labels and Expert Comparison. Proc. AAAI Conf. Artif. Intell. 2019, 33, 590–597. [Google Scholar] [CrossRef]
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. ChestX-ray8: Hospital-scale Chest X-ray Database and Benchmarks on Weakly-Supervised Classification and Localization of Common Thorax Diseases. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 2097–2106. [Google Scholar] [CrossRef]
- Weiner, M.W.; Veitch, D.P.; Aisen, P.S.; Beckett, L.A.; Cairns, N.J.; Green, R.C.; Harvey, D.; Jack, C.R.; Jagust, W.; Liu, E.; et al. The Alzheimer’s Disease Neuroimaging Initiative: A review of papers published since its inception. Alzheimer’s Dement. 2015, 11, e1–e120. [Google Scholar] [CrossRef]
- Menze, B.H.; Jakab, A.; Bauer, S.; Kalpathy-Cramer, J.; Farahani, K.; Kirby, J.; Burren, Y.; Porz, N.; Slotboom, J.; Wiest, R.; et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 2014, 34, 1993–2024. [Google Scholar] [CrossRef]
- Baid, U.; Ghodasara, S.; Mohan, S.; Bilello, M.; Calabrese, E.; Colak, E.; Farahani, K.; Kalpathy-Cramer, J.; Kitamura, F.C.; Pati, S.; et al. The rsna-asnr-miccai brats 2021 benchmark on brain tumor segmentation and radiogenomic classification. arXiv 2021, arXiv:2107.02314. [Google Scholar]
- Cassidy, B.; Kendrick, C.; Brodzicki, A.; Jaworek-Korjakowska, J.; Yap, M.H. Analysis of the ISIC image datasets: Usage, benchmarks and recommendations. Med. Image Anal. 2022, 75, 102305. [Google Scholar] [CrossRef]
- Bejnordi, B.E.; Veta, M.; Van Diest, P.J.; Van Ginneken, B.; Karssemeijer, N.; Litjens, G.; Van Der Laak, J.A.; Hermsen, M.; Manson, Q.F.; Balkenhol, M.; et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 2017, 318, 2199–2210. [Google Scholar] [CrossRef]
- Underwood, T. Pan-cancer analysis of whole genomes. Nature 2020, 578, 82–93. [Google Scholar] [CrossRef]
- Johnson, A.E.; Pollard, T.J.; Shen, L.; Lehman, L.w.H.; Feng, M.; Ghassemi, M.; Moody, B.; Szolovits, P.; Anthony Celi, L.; Mark, R.G. MIMIC-III, a freely accessible critical care database. Sci. Data 2016, 3, 160035. [Google Scholar] [CrossRef]
- Johnson, A.E.; Bulgarelli, L.; Shen, L.; Gayles, A.; Shammout, A.; Horng, S.; Pollard, T.J.; Hao, S.; Moody, B.; Gow, B.; et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci. Data 2023, 10, 1. [Google Scholar] [CrossRef]
- Pollard, T.J.; Johnson, A.E.; Raffa, J.D.; Celi, L.A.; Mark, R.G.; Badawi, O. The eICU Collaborative Research Database, a freely available multi-center database for critical care research. Sci. Data 2018, 5, 180178. [Google Scholar] [CrossRef] [PubMed]
- Wagner, P.; Strodthoff, N.; Bousseljot, R.D.; Kreiseler, D.; Lunze, F.I.; Samek, W.; Schaeffter, T. PTB-XL, a large publicly available electrocardiography dataset. Sci. Data 2020, 7, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Obeid, I.; Picone, J. The temple university hospital EEG data corpus. Front. Neurosci. 2016, 10, 196. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The cancer genome atlas pan-cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Lonsdale, J.; Thomas, J.; Salvatore, M.; Phillips, R.; Lo, E.; Shad, S.; Hasz, R.; Walters, G.; Garcia, F.; Young, N.; et al. The genotype-tissue expression (GTEx) project. Nat. Genet. 2013, 45, 580–585. [Google Scholar] [CrossRef]
- Bycroft, C.; Freeman, C.; Petkova, D.; Band, G.; Elliott, L.T.; Sharp, K.; Motyer, A.; Vukcevic, D.; Delaneau, O.; O’Connell, J.; et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 2018, 562, 203–209. [Google Scholar] [CrossRef]
- Regev, A.; Teichmann, S.A.; Lander, E.S.; Amit, I.; Benoist, C.; Birney, E.; Bodenmiller, B.; Campbell, P.; Carninci, P.; Clatworthy, M.; et al. The human cell atlas. elife 2017, 6, e27041. [Google Scholar] [CrossRef]
- Hasanah, U.; Leu, J.S.; Avian, C.; Azmi, I.; Prakosa, S.W. A systematic review of multilabel chest X-ray classification using deep learning. Multimed. Tools Appl. 2025, 84, 26719–26753. [Google Scholar] [CrossRef]
- Bhusal, D.; Panday, S.P. Multi-label classification of thoracic diseases using dense convolutional network on chest radiographs. arXiv 2022, arXiv:2202.03583. [Google Scholar]
- Sammani, F.; Joukovsky, B.; Deligiannis, N. Visualizing and understanding contrastive learning. IEEE Trans. Image Process. 2023, 33, 541–555. [Google Scholar] [CrossRef] [PubMed]
- Chaitanya, K.; Erdil, E.; Karani, N.; Konukoglu, E. Contrastive learning of global and local features for medical image segmentation with limited annotations. Adv. Neural Inf. Process. Syst. 2020, 33, 12546–12558. [Google Scholar]
- Ciga, O.; Xu, T.; Martel, A.L. Self supervised contrastive learning for digital histopathology. Mach. Learn. Appl. 2022, 7, 100198. [Google Scholar] [CrossRef]
- Guo, Z.; Zhang, Y.; Qiu, Z.; Dong, S.; He, S.; Gao, H.; Zhang, J.; Chen, Y.; He, B.; Kong, Z.; et al. An improved contrastive learning network for semi-supervised multi-structure segmentation in echocardiography. Front. Cardiovasc. Med. 2023, 10, 1266260. [Google Scholar] [CrossRef]
- Luo, G.; Xie, W.; Gao, R.; Zheng, T.; Chen, L.; Sun, H. Unsupervised anomaly detection in brain MRI: Learning abstract distribution from massive healthy brains. Comput. Biol. Med. 2023, 154, 106610. [Google Scholar] [CrossRef]
- Krishnan, R.; Rajpurkar, P.; Topol, E.J. Self-supervised learning in medicine and healthcare. Nat. Biomed. Eng. 2022, 6, 1346–1352. [Google Scholar] [CrossRef]
- Pick, F.; Xie, X.; Wu, L.Y. Contrastive Multitask Transformer for Hospital Mortality and Length-of-Stay Prediction. In Proceedings of the International Conference on AI in Healthcare, Laguna Hills, CA, USA, 5–7 February 2024; Springer: Berlin/Heidelberg, Germany, 2024; pp. 134–145. [Google Scholar]
- Nayebi Kerdabadi, M.; Hadizadeh Moghaddam, A.; Liu, B.; Liu, M.; Yao, Z. Contrastive learning of temporal distinctiveness for survival analysis in electronic health records. In Proceedings of the 32nd ACM International Conference on Information and Knowledge Management, Birmingham, UK, 21–25 October 2023; pp. 1897–1906. [Google Scholar]
- Liu, Y.; Zhang, Z.; Qin, S.; Salim, F.D.; Yepes, A.J. Contrastive learning-based imputation-prediction networks for in-hospital mortality risk modeling using ehrs. In Proceedings of the Joint European Conference on Machine Learning and Knowledge Discovery in Databases, Turin, Italy, 18–22 September 2023; Springer: Berlin/Heidelberg, Germany, 2023; pp. 428–443. [Google Scholar]
- Zang, C.; Wang, F. Scehr: Supervised contrastive learning for clinical risk prediction using electronic health records. In Proceedings of the IEEE International Conference on Data Mining, Auckland, New Zealand, 7–10 December 2021; Volume 2021, p. 857. [Google Scholar]
- Sun, M.; Yang, X.; Niu, J.; Gu, Y.; Wang, C.; Zhang, W. A cross-modal clinical prediction system for intensive care unit patient outcome. Knowl.-Based Syst. 2024, 283, 111160. [Google Scholar] [CrossRef]
- Cai, T.; Huang, F.; Nakada, R.; Zhang, L.; Zhou, D. Contrastive Learning on Multimodal Analysis of Electronic Health Records. arXiv 2024, arXiv:2403.14926. [Google Scholar] [CrossRef]
- Zhong, X.; Batmanghelich, K.; Sun, L. Enhancing Biomedical Multi-modal Representation Learning with Multi-scale Pre-training and Perturbed Report Discrimination. In Proceedings of the 2024 IEEE Conference on Artificial Intelligence (CAI), Singapore, 25–27 June 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 480–485. [Google Scholar]
- Liu, X.; Xu, X.; Xu, X.; Li, X.; Xie, G. Representation Learning for Multi-omics Data with Heterogeneous Gene Regulatory Network. In Proceedings of the 2021 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Houston, TX, USA, 9–12 December 2021; pp. 702–705. [Google Scholar] [CrossRef]
- Li, S.; Ma, J.; Zhao, T.; Jia, Y.; Liu, B.; Luo, R.; Huang, Y. CellContrast: Reconstructing spatial relationships in single-cell RNA sequencing data via deep contrastive learning. Patterns 2024, 5, 101022. [Google Scholar] [CrossRef] [PubMed]
- Bepler, T.; Berger, B. Learning the protein language: Evolution, structure, and function. Cell Syst. 2021, 12, 654–669. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Wu, H.; Liu, C.; Li, H.; Wu, Y.; Li, K.; Wang, Y.; Deng, Y.; Chen, J.; Zhou, F.; et al. Pepharmony: A multi-view contrastive learning framework for integrated sequence and structure-based peptide encoding. arXiv 2024, arXiv:2401.11360. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Li, C.; Zhou, H.Y.; Yang, H.; Liu, J.; Liang, Y.; Zheng, H.; Zhang, S.; Wang, S. Enhancing representation in radiography-reports foundation model: A granular alignment algorithm using masked contrastive learning. Nat. Commun. 2024, 15, 7620. [Google Scholar] [CrossRef]
- Koleilat, T.; Asgariandehkordi, H.; Rivaz, H.; Xiao, Y. Medclip-sam: Bridging text and image towards universal medical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Marrakesh, Morocco, 6–10 October 2024; Springer: Berlin/Heidelberg, Germany, 2024; pp. 643–653. [Google Scholar]
- Liu, Z.; Alavi, A.; Li, M.; Zhang, X. Self-supervised contrastive learning for medical time series: A systematic review. Sensors 2023, 23, 4221. [Google Scholar] [CrossRef]
- Yuan, Y.; Van Duyn, J.; Yan, R.; Huang, Z.; Vesal, S.; Plis, S.; Hu, X.; Kwak, G.H.; Xiao, R.; Fedorov, A. Learning ECG Representations via Poly-Window Contrastive Learning. arXiv 2025, arXiv:2508.15225. [Google Scholar] [CrossRef]
- Wang, Y.; Han, Y.; Wang, H.; Zhang, X. Contrast everything: A hierarchical contrastive framework for medical time-series. Adv. Neural Inf. Process. Syst. 2023, 36, 55694–55717. [Google Scholar]
- Raghu, A.; Chandak, P.; Alam, R.; Guttag, J.; Stultz, C. Contrastive pre-training for multimodal medical time series. In Proceedings of the NeurIPS 2022 Workshop on Learning from Time Series for Health, New Orleans, LA, USA, 2 December 2022. [Google Scholar]
- Guo, H.; Xu, X.; Wu, H.; Liu, B.; Xia, J.; Cheng, Y.; Guo, Q.; Chen, Y.; Xu, T.; Wang, J.; et al. Multi-scale and multi-modal contrastive learning network for biomedical time series. Biomed. Signal Process. Control 2025, 106, 107697. [Google Scholar] [CrossRef]
- Sun, X.; Yang, Y.; Dong, X. Enhancing Contrastive Learning-based Electrocardiogram Pretrained Model with Patient Memory Queue. arXiv 2025, arXiv:2506.06310. [Google Scholar]
- Zech, J.R.; Badgeley, M.A.; Liu, M.; Costa, A.B.; Titano, J.J.; Oermann, E.K. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: A cross-sectional study. PLoS Med. 2018, 15, e1002683. [Google Scholar] [CrossRef]
- Oakden-Rayner, L.; Dunnmon, J.; Carneiro, G.; Ré, C. Hidden stratification causes clinically meaningful failures in machine learning for medical imaging. In Proceedings of the ACM Conference on Health, Inference, and Learning, Toronto, ON, Canada, 2–4 April 2020; pp. 151–159. [Google Scholar]
- Guo, C.; Pleiss, G.; Sun, Y.; Weinberger, K.Q. On calibration of modern neural networks. In Proceedings of the International Conference on Machine Learning, Sydney, Australia, 6–11 August 2017; PMLR: New York, NY, USA, 2017; pp. 1321–1330. [Google Scholar]
- Mehrabi, N.; Morstatter, F.; Saxena, N.; Lerman, K.; Galstyan, A. A survey on bias and fairness in machine learning. ACM Comput. Surv. (CSUR) 2021, 54, 1–35. [Google Scholar] [CrossRef]
- Samek, W.; Wiegand, T.; Müller, K.R. Explainable Artificial Intelligence: Understanding, Visualizing and Interpreting Deep Learning Models. arXiv 2017, arXiv:1708.08296. [Google Scholar] [CrossRef]
- Ribeiro, M.T.; Singh, S.; Guestrin, C. “Why should I trust you?” Explaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 1135–1144. [Google Scholar]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]
- Finlayson, S.G.; Bowers, J.D.; Ito, J.; Zittrain, J.L.; Beam, A.L.; Kohane, I.S. Adversarial attacks on medical machine learning. Science 2019, 363, 1287–1289. [Google Scholar] [CrossRef]
- Arjovsky, M.; Bottou, L.; Gulrajani, I.; Lopez-Paz, D. Invariant Risk Minimization. arXiv 2019, arXiv:1907.02893. [Google Scholar]
- Schölkopf, B.; Locatello, F.; Bauer, S.; Ke, N.R.; Kalchbrenner, N.; Goyal, A.; Bengio, Y. Toward Causal Representation Learning. Proc. IEEE 2021, 109, 612–634. [Google Scholar] [CrossRef]
- McMahan, H.B.; Moore, E.; Ramage, D.; Hampson, S.; y Arcas, B.A. Communication-Efficient Learning of Deep Networks from Decentralized Data. In Proceedings of the 20th International Conference on Artificial Intelligence and Statistics (AISTATS), Ft. Lauderdale, FL, USA, 20–22 April 2017; PMLR: New York, NY, USA, 2017; pp. 1273–1282. [Google Scholar]
- Abadi, M.; Chu, A.; Goodfellow, I.; McMahan, H.B.; Mironov, I.; Talwar, K.; Zhang, L. Deep Learning with Differential Privacy. In Proceedings of the 2016 ACM SIGSAC Conference on Computer and Communications Security (CCS), Vienna, Austria, 24–28 October 2016; ACM: New York, NY, USA, 2016; pp. 308–318. [Google Scholar]
- Rajpurkar, P.; Chen, E.; Banerjee, O.; Topol, E.J. AI in health and medicine. Nat. Med. 2022, 28, 31–38. [Google Scholar] [CrossRef]

| Review | Year | Primary Scope | Medical Coverage | Notes for Positioning in This Paper |
|---|---|---|---|---|
| Jaiswal et al. [20] | 2020 | General CL and SSL overview | Limited | Early overview of CL principles and variants. |
| Gui et al. [21] | 2024 | Broad SSL survey | Limited | Includes contrastive, generative, and clustering-based SSL methods. |
| Hu et al. [22] | 2024 | General CL survey | Indirect | Presents a universal CL framework and component-level advances. |
| Liu [23] | 2021 | CL for visual representation | Indirect | Vision-focused synthesis of CL components, limitations, and improvements. |
| Shurrab et al. [24] | 2022 | SSL in medical imaging | Imaging only | Focuses on imaging applications with limited clinical data coverage. |
| Wang et al. [25] | 2023 | Medical imaging SSL and CL | Imaging only | Emphasizes adaptation of natural image SSL methods to medical imaging. |
| Huang et al. [26] | 2023 | Systematic review of SSL for medical imaging | Imaging only | Systematic evidence synthesis and reporting trends for image classification studies. |
| VanBerlo et al. [27] | 2024 | Evidence review of SSL pretraining in imaging | Imaging only | Highlights comparisons against supervised baselines and transfer protocols. |
| This review | 2025 | CL in medical AI | Imaging, EHR, signals, omics, multimodal | Cross modality synthesis with operational taxonomy and evaluation guidance. |
| Dimension | Definition | Common Categories/Options |
|---|---|---|
| Loss family/ objective | The contrastive or self-supervised learning signal is used to shape representations. | InfoNCE/NT-Xent; supervised contrastive; distillation/negative-free; clustering consistency; cross-modal retrieval loss; temporal predictive losses (e.g., CPC). |
| Pairing strategy (positives) | How positives are constructed, i.e., what the method enforces to be similar. | Augmented views of the same instance; same-class positives; patient-aware positives; longitudinal (same patient across time); cross-modal paired (image–text, ECG–EHR); region–text phrase. |
| Augmentations/ views | Transformations are used to generate alternative views and define invariances. | Imaging: mild geometry/intensity, modality-specific; signals: masking/jitter/windowing; EHR: time masking/cropping; text: entity-aware processing; multi-view sampling (slices/windows). |
| Label regime | How labels are used during pretraining (if at all) and during evaluation. | Unsupervised (no labels); weak/self-labels; supervised contrastive; semi-supervised (mix of labeled/unlabeled); label-scarce learning curves. |
| Evaluation protocol | How the downstream benefit is tested, including whether shift and calibration are considered. | Linear probe; full fine-tuning; few-shot/label fractions; external validation (site/device/time); temporal split; retrieval evaluation (Recall@K); calibration (ECE/Brier) for risk models. |
| Task family | Clinical endpoint and decision context used for evaluation. | Classification (diagnosis/risk); survival/prognosis; segmentation; detection; retrieval/triage; phenotype discovery/clustering; zero-shot/VLM prompting. |
| Protocol | What It Tests | Typical Metrics (Examples) |
|---|---|---|
| Linear probe | Representation quality under minimal supervision | AUROC/AUPRC (multi-label); accuracy; macro-F1 (class imbalance); calibration metrics when reported |
| Full fine-tuning | End-task performance with task-specific adaptation | AUROC/AUPRC; Dice/HD95 for segmentation; patient-level metrics for clinical outcomes |
| Few-shot and label-scarce | Label efficiency and robustness when annotation is limited | AUROC/F1 vs label fraction; confidence intervals across seeds; sensitivity to class imbalance |
| External validation and site shift | Generalization across institutions, devices, protocols, and time | Performance drop under shift; subgroup robustness; calibration under shift; domain-wise reporting |
| Retrieval and alignment (vision–language) | Cross-modal grounding and matching fidelity | Recall@K; median rank; phrase grounding scores; zero-shot AUROC using text prompts |
| Application Domain | Author(s) | Year | Method | Application |
|---|---|---|---|---|
| Medical Imaging | Azizi et al. [16] | 2021 | SimCLR-based pretraining on unlabeled chest X-rays | Learned transferable visual representations for medical image classification and segmentation using unlabeled data. |
| Chaitanya et al. [105] | 2020 | Semi-supervised contrastive framework for MRI | Improved segmentation and classification in MRI with limited labels using augmented positive and negative pairs. | |
| Ciga et al. [106] | 2022 | Self-supervised contrastive learning for histopathology | Enhanced cancer detection and tissue differentiation in biopsy samples using augmentation-invariant representations. | |
| Guo et al. [107] | 2023 | Multi-scale contrastive loss for cardiac MRI segmentation | Captured both global and local structures, improving accuracy of myocardium and ventricle segmentation. | |
| Luo et al. [108] | 2023 | Self-supervised anomaly detection using contrastive loss | Detected abnormal regions in brain MRI scans by distinguishing normal and pathological patches. | |
| Electronic Health Records | Krishnan et al. [109] | 2022 | Self-supervised contrastive learning on augmented EHR views | Modeled temporal and clinical correlations for mortality and heart failure prediction. |
| Pick et al. [110] | 2024 | Contrastive patient representation learning | Improved prediction of hospital mortality and length-of-stay through patient-level embeddings. | |
| Sun et al. [114] | 2024 | Cross-modal contrastive framework for EHR integration | Aligned structured and unstructured EHR data to predict disease progression and complications. | |
| Cai et al. [115] | 2024 | Distributed large-scale contrastive learning | Scalable training on large EHR datasets for improved generalization across patient populations. | |
| Kerdabadi et al. [111] | 2023 | ontology-aware temporal contrastive survival | Learns temporally distinctive EHR embeddings with hardness-aware negatives for AKI survival risk prediction. | |
| Liu et al. [112] | 2023 | Contrastive imputation–prediction network (CL-IPN) | Contrastive-enhanced imputation with patient stratification improves in-hospital mortality prediction under missing/irregular EHRs. | |
| Zang and Wang [113] | 2021 | Supervised Contrastive framework using longitudinal EHR | Unified supervised contrastive loss improves EHR classification outcomes. | |
| Genomics and Proteomics | Zhong et al. [116] | 2024 | Multi-scale contrastive learning (MSCL) for genomics | Identified disease-associated genetic markers by modeling gene and pathway-level interactions. |
| Bepler and Berger [119] | 2021 | Contrastive protein sequence representation learning | Learned structural and functional protein embeddings for improved function prediction and drug discovery. | |
| Liu et al. [117] | 2022 | Multi-omics contrastive learning (MoHeG/GenCL) | Integrated genomics, transcriptomics, and epigenomics to predict disease susceptibility and treatment outcomes. | |
| Li et al. [118] | 2024 | CellContrast for single-cell RNA sequencing | Enhanced clustering and identification of rare cell types in scRNA-seq data. | |
| Zhang et al. [120] | 2024 | Pepharmony: sequence–structure contrastive learning | Predicted protein–protein interactions with improved accuracy using multimodal peptide representations. | |
| Multimodal and Cross-Domain Learning | Zhang et al. [18] | 2022 | ConVIRT (image–text alignment) | Learned chest X-ray representations by aligning images and radiology reports with bidirectional contrastive loss. |
| Huang et al. [65] | 2021 | GLoRIA (global–local image–text alignment) | Improved retrieval and classification on MIMIC-CXR through local region–phrase alignment. | |
| Boecking et al. [66] | 2022 | BioViL (biomedical vision–language model) | Enhanced zero-shot radiology performance using domain-specific text pretraining. | |
| Bannur et al. [67] | 2023 | BioViL-T (temporal alignment) | Improved disease progression tracking in chest X-rays via temporal contrastive learning. | |
| You et al. [68] | 2023 | CXR-CLIP (prompt-based multimodal CL) | Combined image–label and image–text supervision for robust chest X-ray recognition. | |
| Tiu et al. [69] | 2022 | CheXzero (CLIP-style vision–language model) | Achieved radiologist-level zero-shot classification on the CheXpert benchmark. | |
| Wang et al. [70] | 2022 | MedCLIP (knowledge-aware matching loss) | Reduced false negatives in radiology by decoupling image–text corpora for efficient pretraining. | |
| Huang et al. [71] | 2023 | PLIP (pathology vision–language foundation model) | Achieved state-of-the-art performance in pathology classification and zero-shot transfer. | |
| Lu et al. [72] | 2024 | CONCH (large-scale histopathology pretraining) | Trained on 1.17 M image–caption pairs for generalizable pathology retrieval and segmentation. | |
| Zhang et al. [73] | 2023 | BiomedCLIP (PubMed multimodal foundation model) | Pretrained on 15 M image–text pairs for broad biomedical zero/few-shot applications. | |
| Lin et al. [74] | 2023 | PMC-CLIP (literature-derived pretraining) | Improved biomedical VQA and retrieval from 1.6M figure–caption pairs. | |
| Huang et al. [121] | 2024 | MaCo (masked contrastive learning) | Applied to chest X-rays for enhanced zero-shot and localized recognition. | |
| Koleilat et al. [122] | 2024 | MedCLIP + SAM (text-driven segmentation) | Enabled multimodal segmentation across ultrasound, MRI, and CT without explicit labels. | |
| Time-Series and Physiological Signals | Liu et al. [123] | 2023 | Systematic review of contrastive time-series methods | Identified key design trends in self-supervised ECG/EEG contrastive learning. |
| Diamant et al. [19] | 2022 | PCLR (patient-level contrastive learning) | Leveraged same-patient ECGs to improve cardiac disease prediction tasks. | |
| Yuan et al. [124] | 2025 | Poly-window contrastive learning | Modeled temporal overlap in ECGs to enhance representation efficiency. | |
| Wang et al. [125] | 2023 | COMET (hierarchical contrastive framework) | Applied multi-level contrastive learning for ECG and EEG classification with few labels. | |
| Chen et al. [76] | 2025 | CLOCS (spatiotemporal contrastive model) | Improved robustness in cardiac signals under lead and time variation. | |
| Raghu et al. [126] | 2022 | Multimodal temporal contrastive pretraining | Integrated physiological signals with lab and vitals data for outcome prediction. | |
| Guo et al. [127] | 2025 | MBSL (multi-scale multimodal contrastive learning) | Combined respiration, heart rate, and motion signals for multi-task biomedical inference. | |
| Sun et al. [128] | 2025 | PMQ (patient memory queue) | Mitigated false negatives in ECG pretraining by leveraging intra-patient memory banks. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Obaido, G.; Mienye, I.D.; Aruleba, K.; Chukwu, C.W.; Esenogho, E.; Modisane, C. A Systematic Review of Contrastive Learning in Medical AI: Foundations, Biomedical Modalities, and Future Directions. Bioengineering 2026, 13, 176. https://doi.org/10.3390/bioengineering13020176
Obaido G, Mienye ID, Aruleba K, Chukwu CW, Esenogho E, Modisane C. A Systematic Review of Contrastive Learning in Medical AI: Foundations, Biomedical Modalities, and Future Directions. Bioengineering. 2026; 13(2):176. https://doi.org/10.3390/bioengineering13020176
Chicago/Turabian StyleObaido, George, Ibomoiye Domor Mienye, Kehinde Aruleba, Chidozie Williams Chukwu, Ebenezer Esenogho, and Cameron Modisane. 2026. "A Systematic Review of Contrastive Learning in Medical AI: Foundations, Biomedical Modalities, and Future Directions" Bioengineering 13, no. 2: 176. https://doi.org/10.3390/bioengineering13020176
APA StyleObaido, G., Mienye, I. D., Aruleba, K., Chukwu, C. W., Esenogho, E., & Modisane, C. (2026). A Systematic Review of Contrastive Learning in Medical AI: Foundations, Biomedical Modalities, and Future Directions. Bioengineering, 13(2), 176. https://doi.org/10.3390/bioengineering13020176







