Machine Learning-Based Estimation of Knee Joint Mechanics from Kinematic and Neuromuscular Inputs: A Proof-of-Concept Using the CAMS-Knee Datasets
Abstract
1. Introduction
2. Materials and Methods
3. Results
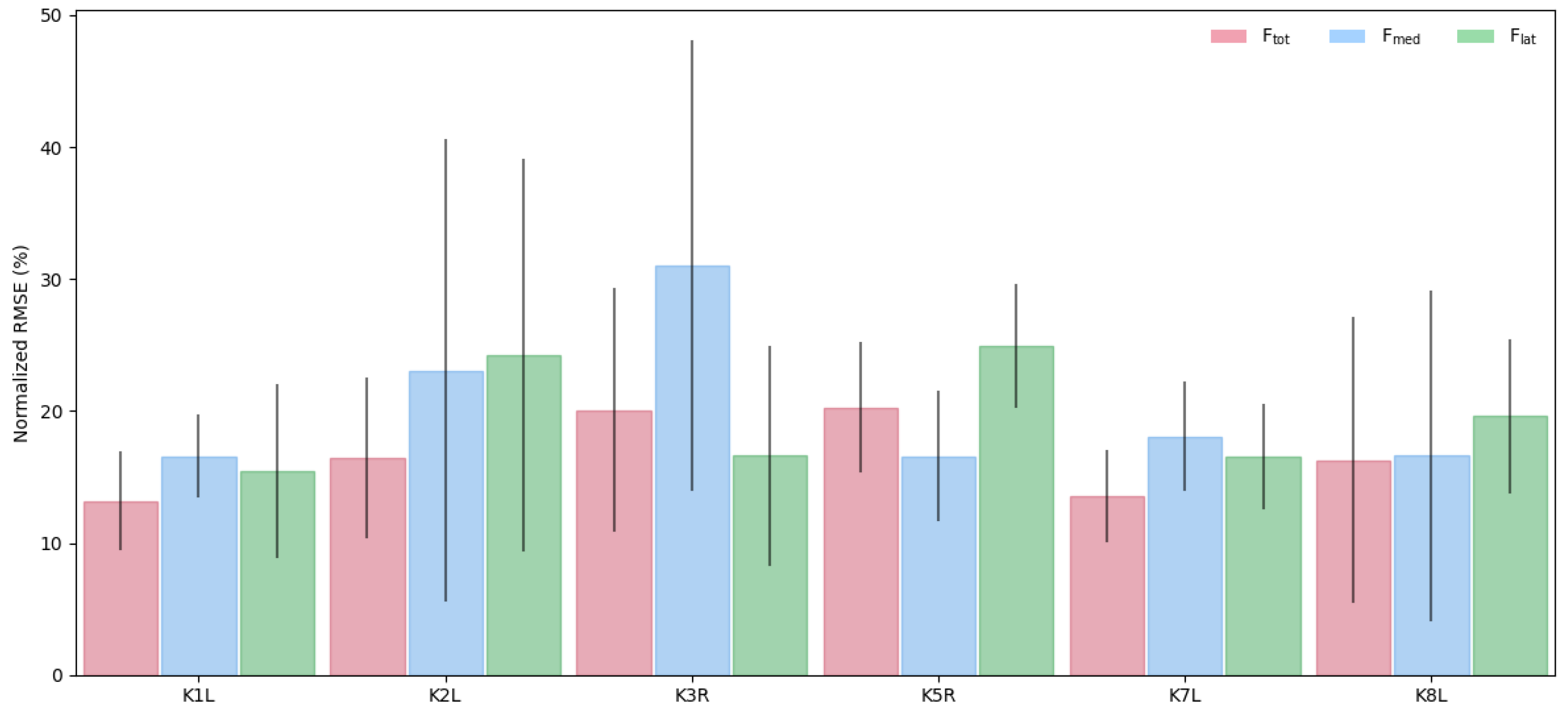
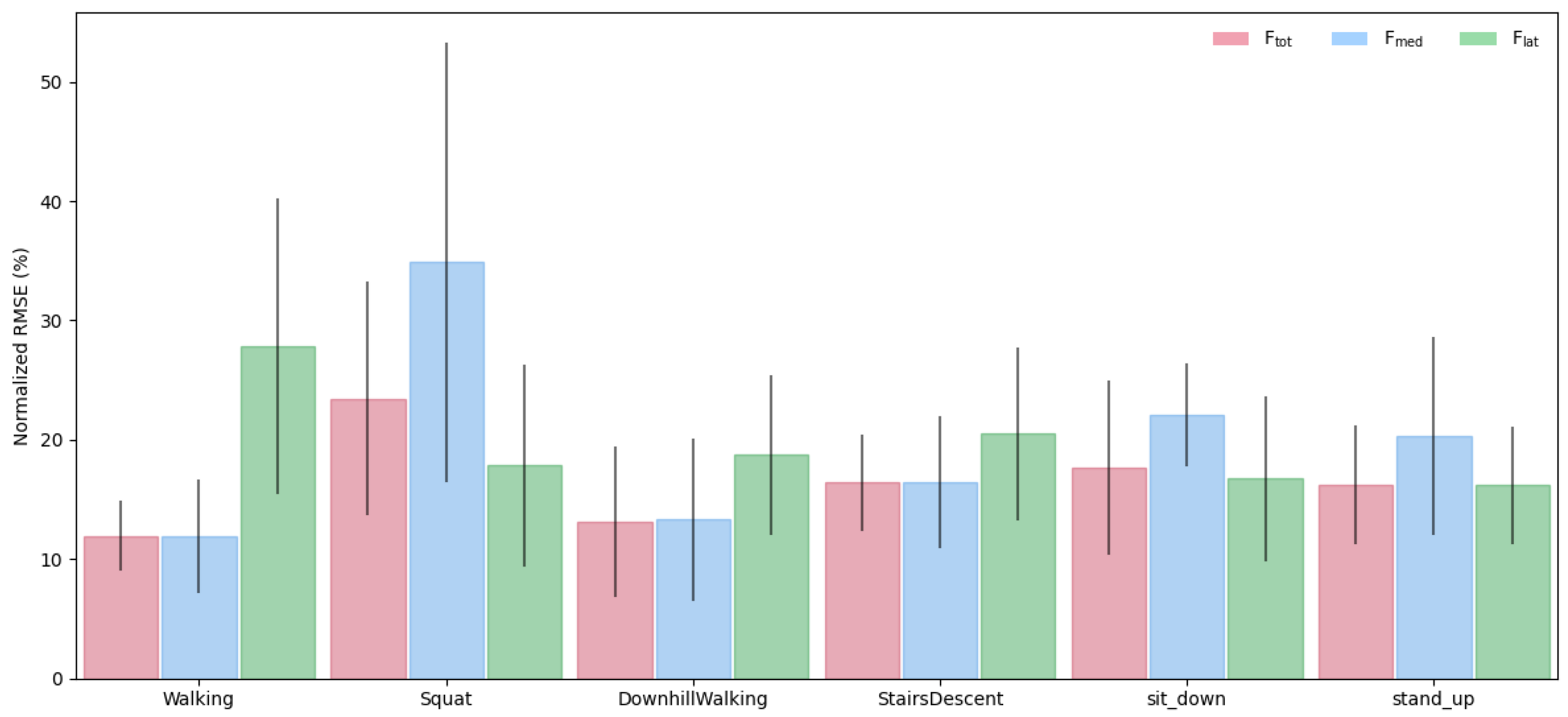
3.1. LSTM Model: Intra-Subject Predictions (LOTO)
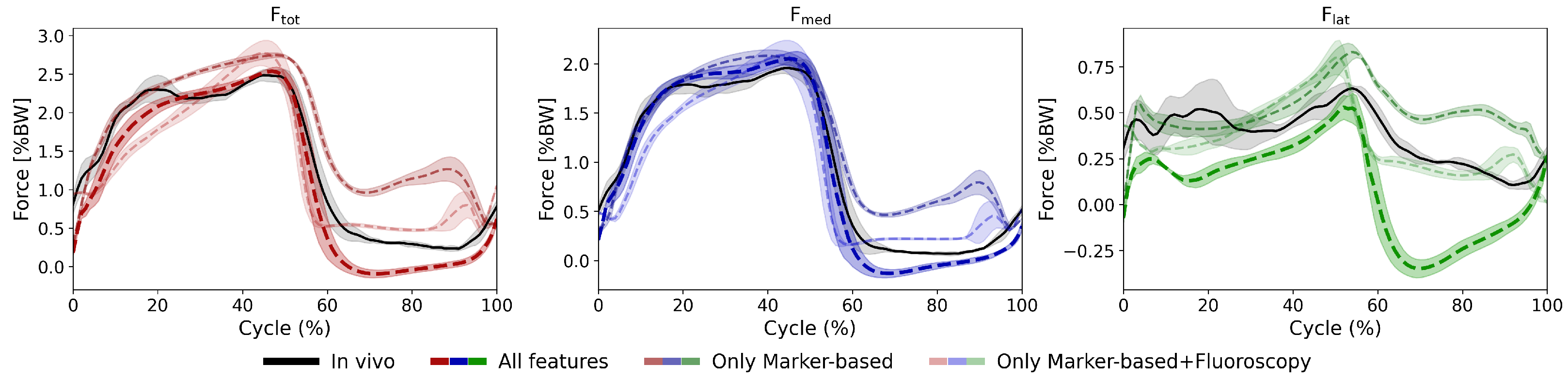
3.2. LSTM Model: Feature Importance Analysis
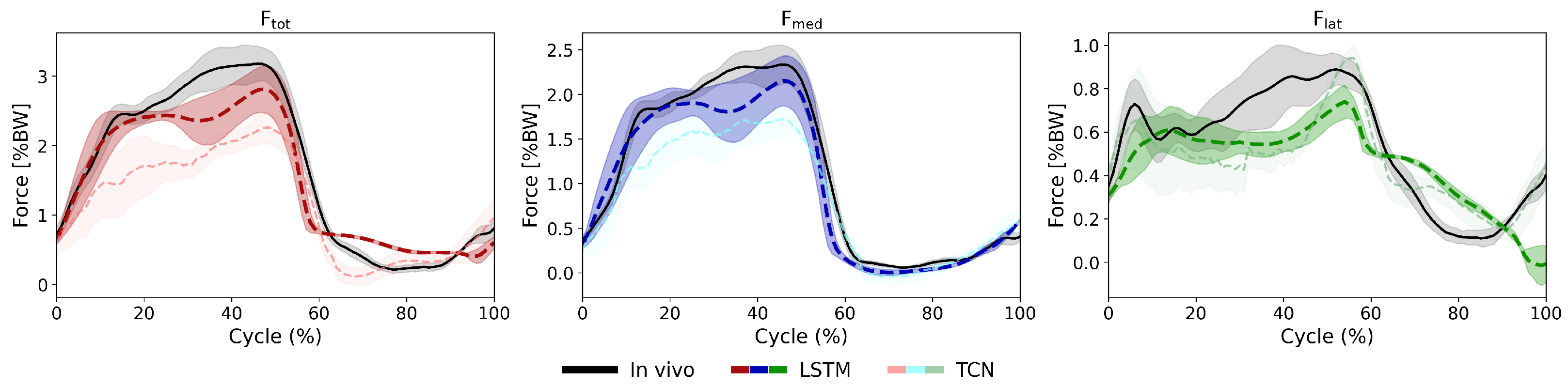
3.3. LSTM vs. TCN
4. Discussion
4.1. Model Performance and Validation
4.2. Comparison with Existing Approaches
4.3. Feature Importance and Input Contributions
4.4. Clinical Applications and Future Directions
4.5. Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| biLSTM-MLP | bidirectional Long Short-Term-Memory Network with a Multi-Layer Perceptron |
| TCN | Temporal Convolutional Network |
| KCF | Knee Contact Force |
| RMSE | Root Mean Squared Error |
| BW | Body Weight |
| EMG | Electromyography |
| KOA | Knee Osteoarthritis |
| TKA | Total Knee Arthroplasty |
| MSK | Musculoskeletal |
| ML | Machine Learning |
| LSTM | Long Short-Term-Memory |
| GRF | Ground Reaction Force |
| MLP | Multi-Layer Perceptron |
| ReLU | Rectified Linear Unit |
| Medial Knee Contact Force | |
| Lateral Knee Contact Force | |
| Total Knee Contact Force | |
| LOSO | Leave-One-Subject-Out |
| PCC | Pearson Correlation Coefficient |
| nRMSE | Normalized Root Mean Squared Error |
| LOTO | Leave-One-Trial-Out |
| LOFO | Leave-One-Feature-Out |
| CCI | Co-Contraction Index |
Appendix A
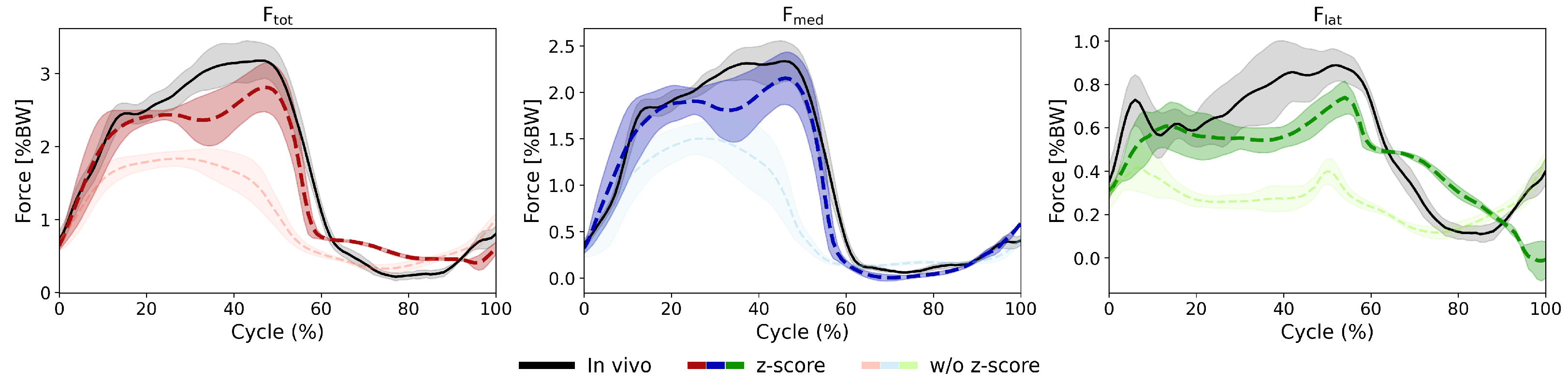
Appendix A.1. LSTM Model: Effect of Input Normalization
| Average Across Subjects (z-Score Normalized Inputs) | Average Across Subjects (Inputs Without z-Score Normalization) | ||
|---|---|---|---|
| Walking | 11.9% (0.97) | 31.2% (0.80) | |
| 11.9% (0.98) | 34.7% (0.81) | ||
| 27.8% (0.66) | 27.8% (0.27) | ||
| Squat | 23.4% (0.84) | 42.3% (0.09) | |
| 34.9% (0.61) | 34.9% (0.24) | ||
| 17.8% (0.92) | 47.7% (0.16) | ||
| Downhill Walking | 13.1% (0.98) | 34.8% (0.77) | |
| 13.3% (0.97) | 35.3% (0.83) | ||
| 18.7% (0.90) | 32.4% (0.44) | ||
| Stairs Descent | 16.4% (0.96) | 36.8% (0.75) | |
| 16.5% (0.95) | 37.4% (0.71) | ||
| 20.5% (0.91) | 35.2% (0.48) | ||
| Sit Down | 17.6% (0.90) | 35.3% (0.65) | |
| 22.1% (0.83) | 31.8% (0.72) | ||
| 16.8% (0.91) | 40.6% (0.61) | ||
| Stand Up | 16.2% (0.96) | 33.5% (0.44) | |
| 20.3% (0.94) | 30.6% (0.51) | ||
| 16.2% (0.95) | 39.7% (0.48) |
Appendix A.2. Impact of Kinematic Input Modalities
| With All Inputs | Without Kinematics | Without Fluoroscopic Kinematics | Without Skin Marker Kinematics | ||
|---|---|---|---|---|---|
| Walking | 11.9% (0.97) | 13.3% (0.96) | 12.6% (0.96) | 12.2% (0.96) | |
| 11.9% (0.98) | 15.1% (0.96) | 12.3% (0.97) | 13.7% (0.97) | ||
| 27.8% (0.66) | 20.7% (0.72) | 22.3% (0.62) | 21.0% (0.71) | ||
| Squat | 23.4% (0.84) | 31.0% (0.81) | 21.5% (0.79) | 23.3% (0.82) | |
| 34.9% (0.61) | 37.2% (0.63) | 31.3% (0.50) | 32.5% (0.58) | ||
| 17.8% (0.92) | 30.5% (0.89) | 20.5% (0.88) | 21.0% (0.89) | ||
| Downhill Walking | 13.1% (0.98) | 13.0% (0.97) | 12.1% (0.98) | 11.6% (0.98) | |
| 13.3% (0.97) | 16.6% (0.96) | 12.6% (0.97) | 13.5% (0.97) | ||
| 18.7% (0.90) | 19.1% (0.82) | 18.0% (0.88) | 16.7% (0.90) | ||
| Stairs Descent | 16.4% (0.96) | 17.6% (0.96) | 14.3% (0.97) | 14.7% (0.97) | |
| 16.5% (0.95) | 18.7% (0.94) | 13.9% (0.96) | 16.4% (0.96) | ||
| 20.5% (0.91) | 21.1% (0.84) | 18.5% (0.90) | 17.4% (0.91) | ||
| Sit Down | 17.6% (0.90) | 28.2% (0.87) | 18.8% (0.91) | 20.3% (0.89) | |
| 22.1% (0.83) | 27.0% (0.84) | 27.0% (0.73) | 22.7% (0.83) | ||
| 16.8% (0.91) | 30.5% (0.87) | 20.2% (0.89) | 23.7% (0.88) | ||
| Stand Up | 16.2% (0.96) | 20.3% (0.90) | 18.2% (0.94) | 13.4% (0.96) | |
| 20.3% (0.94) | 22.3% (0.86) | 19.1% (0.88) | 18.1% (0.92) | ||
| 16.2% (0.95) | 23.3% (0.87) | 15.0% (0.94) | 17.4% (0.95) |
Appendix A.3. Additional Plots



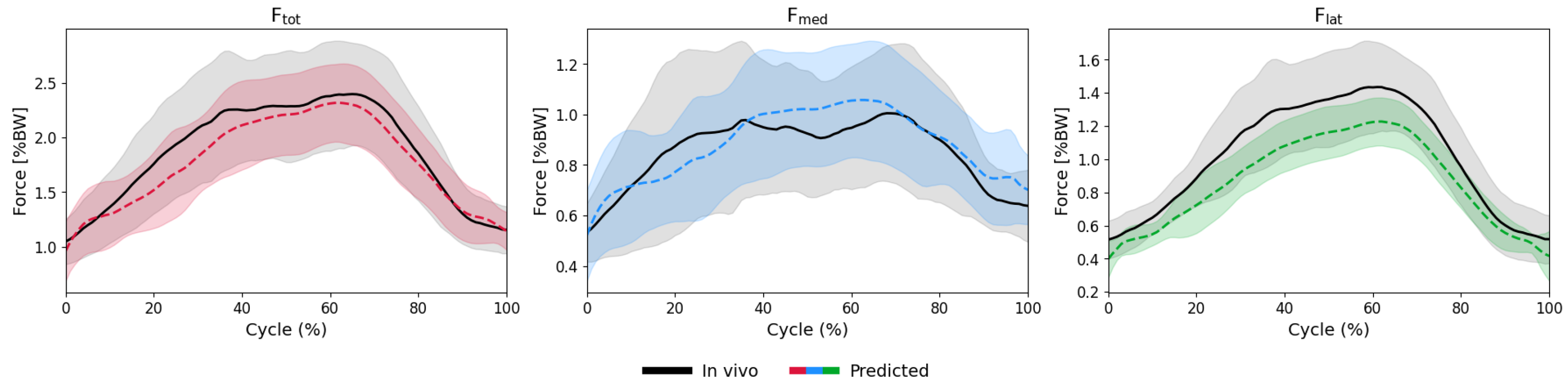



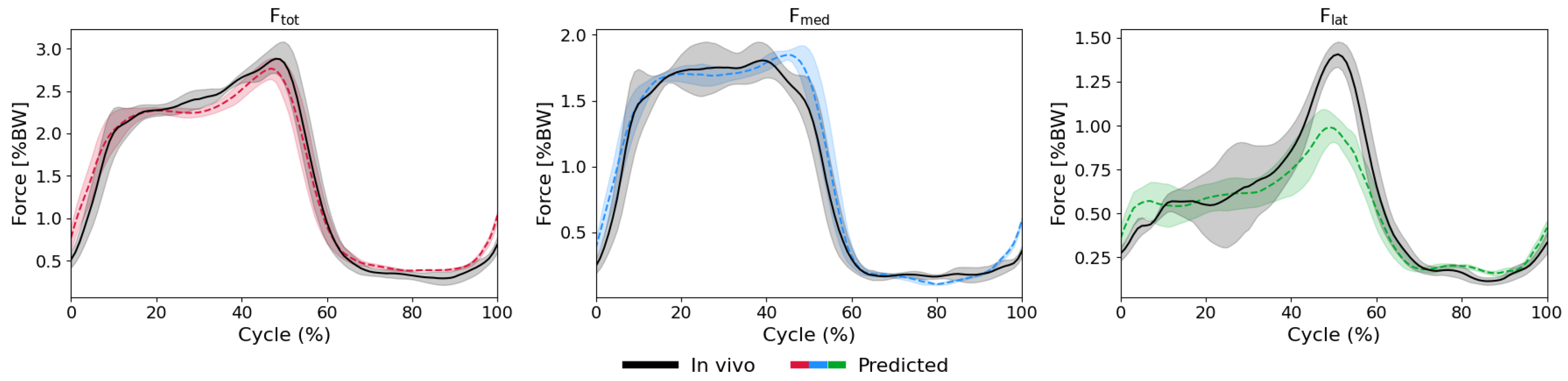



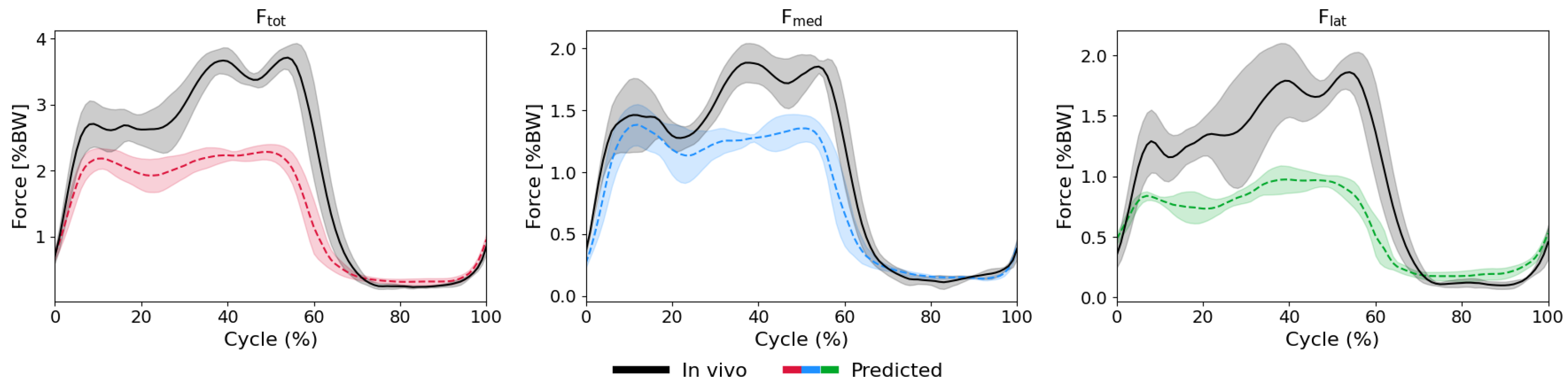






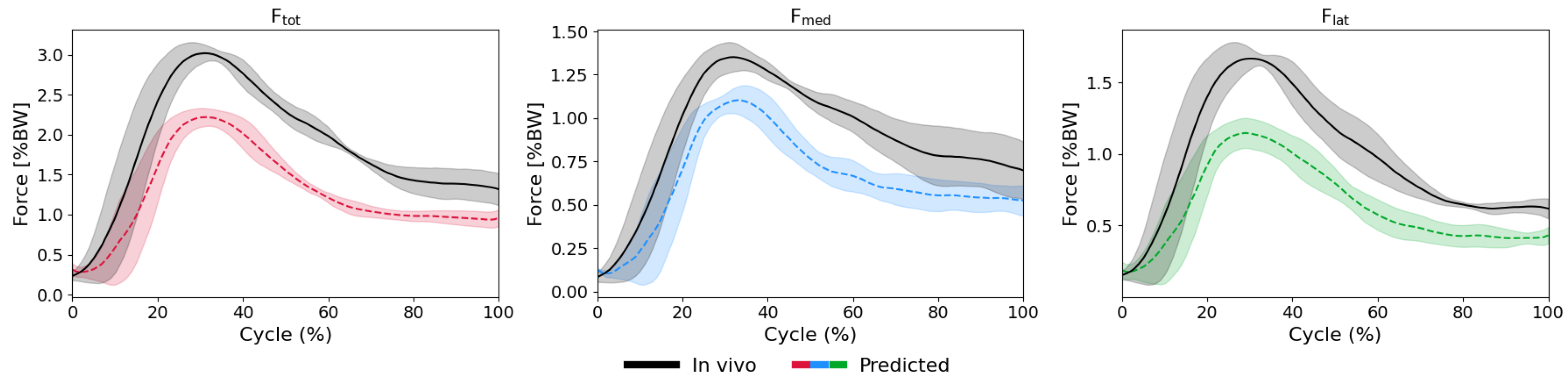
References
- Taylor, W.R.; Schütz, P.; Bergmann, G.; List, R.; Postolka, B.; Hitz, M.; Dymke, J.; Damm, P.; Duda, G.; Gerber, H.; et al. A comprehensive assessment of the musculoskeletal system: The CAMS-Knee data set. J. Biomech. 2017, 65, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Bennett, H.J.; Estler, K.; Valenzuela, K.; Weinhandl, J.T. Predicting Knee Joint Contact Forces During Normal Walking Using Kinematic Inputs With a Long-Short Term Neural Network. J. Biomech. Eng. 2024, 146, 081004. [Google Scholar] [CrossRef] [PubMed]
- Valente, G.; Grenno, G.; Fabbro, G.D.; Zaffagnini, S.; Taddei, F. Medial and lateral knee contact forces during walking, stair ascent and stair descent are more affected by contact locations than tibiofemoral alignment in knee osteoarthritis patients with varus malalignment. Front. Bioeng. Biotechnol. 2023, 11, 1254661. [Google Scholar] [CrossRef] [PubMed]
- Mannisi, M.; Dell’Isola, A.; Andersen, M.; Woodburn, J. Effect of lateral wedged insoles on the knee internal contact forces in medial knee osteoarthritis. Gait Posture 2019, 68, 443–448. [Google Scholar] [CrossRef] [PubMed]
- Heinlein, B.; Graichen, F.; Bender, A.; Rohlmann, A.; Bergmann, G. Design, calibration and pre-clinical testing of an instrumented tibial tray. J. Biomech. 2007, 40, S4–S10. [Google Scholar] [CrossRef]
- Guo, N.; Smith, C.R.; Schütz, P.; Trepczynski, A.; Moewis, P.; Damm, P.; Maas, A.; Grupp, T.M.; Taylor, W.R.; Nasab, S.H.H. Posterior tibial slope influences joint mechanics and soft tissue loading after total knee arthroplasty. Front. Bioeng. Biotechnol. 2024, 12, 1352794. [Google Scholar] [CrossRef] [PubMed]
- Nasab, S.H.H.; Hörmann, S.; Grupp, T.M.; Taylor, W.R.; Maas, A. On the consequences of intra-operative release versus over-tensioning of the posterior cruciate ligament in total knee arthroplasty. J. R. Soc. Interface 2024, 21, 20240588. [Google Scholar] [CrossRef]
- Burton, W.S.; Myers, C.A.; Rullkoetter, P.J. Machine learning for rapid estimation of lower extremity muscle and joint loading during activities of daily living. J. Biomech. 2021, 123, 110439. [Google Scholar] [CrossRef]
- Jung, Y.; Phan, C.B.; Koo, S. Intra-Articular Knee Contact Force Estimation During Walking Using Force-Reaction Elements and Subject-Specific Joint Model. J. Biomech. Eng. 2016, 138. [Google Scholar] [CrossRef]
- Xiang, L.; Gu, Y.; Gao, Z.; Yu, P.; Shim, V.; Wang, A.; Fernandez, J. Integrating an LSTM framework for predicting ankle joint biomechanics during gait using inertial sensors. Comput. Biol. Med. 2024, 170, 108016. [Google Scholar] [CrossRef]
- Zhang, L.; Soselia, D.; Wang, R.; Gutierrez-Farewik, E.M. Lower-Limb Joint Torque Prediction Using LSTM Neural Networks and Transfer Learning. IEEE Trans. Neural Syst. Rehabil. Eng. 2022, 30, 600–609. [Google Scholar] [CrossRef]
- Mundt, M.; Johnson, W.R.; Potthast, W.; Markert, B.; Mian, A.; Alderson, J. A Comparison of Three Neural Network Approaches for Estimating Joint Angles and Moments from Inertial Measurement Units. Sensors 2021, 21, 4535. [Google Scholar] [CrossRef]
- Bai, S.; Kolter, J.Z.; Koltun, V. An Empirical Evaluation of Generic Convolutional and Recurrent Networks for Sequence Modeling. arXiv 2018, arXiv:1803.01271. [Google Scholar] [CrossRef]
- Molinaro, D.D.; Kang, I.; Camargo, J.; Gombolay, M.C.; Young, A.J. Subject-Independent, Biological Hip Moment Estimation During Multimodal Overground Ambulation Using Deep Learning. IEEE Trans. Med. Robot. Bionics 2022, 4, 219–229. [Google Scholar] [CrossRef]
- Nuesslein, C.P.O.; Young, A.J. A Deep Learning Framework for End-to-End Control of Powered Prostheses. IEEE Robot. Autom. Lett. 2024, 9, 3988–3994. [Google Scholar] [CrossRef]
- Febrer-Nafría, M.; Dreyer, M.J.; Maas, A.; Taylor, W.R.; Smith, C.R.; Nasab, S.H.H. Knee kinematics are primarily determined by implant alignment but knee kinetics are mainly influenced by muscle coordination strategy. J. Biomech. 2023, 161, 111851. [Google Scholar] [CrossRef]
- Kutzner, I.; Küther, S.; Heinlein, B.; Dymke, J.; Bender, A.; Halder, A.M.; Bergmann, G. The effect of valgus braces on medial compartment load of the knee joint—In vivo load measurements in three subjects. J. Biomech. 2011, 44, 1354–1360. [Google Scholar] [CrossRef]
- Mundt, M.; Born, Z.; Goldacre, M.; Alderson, J. Estimating Ground Reaction Forces from Two-Dimensional Pose Data: A Biomechanics-Based Comparison of AlphaPose, BlazePose, and OpenPose. Sensors 2022, 23, 78. [Google Scholar] [CrossRef] [PubMed]
- Nasab, S.H.H.; Smith, C.; Schütz, P.; Postolka, B.; Ferguson, S.; Taylor, W.R.; List, R. Elongation Patterns of the Posterior Cruciate Ligament after Total Knee Arthroplasty. J. Clin. Med. 2020, 9, 2078. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Zhang, X.; Zhang, Y.; Jin, Z. Prediction of medial knee contact force using multisource fusion recurrent neural network and transfer learning. Med. Biol. Eng. Comput. 2024, 62, 1333–1346. [Google Scholar] [CrossRef] [PubMed]
- Liew, B.X.W.; Rügamer, D.; Mei, Q.; Altai, Z.; Zhu, X.; Zhai, X.; Cortes, N. Smooth and accurate predictions of joint contact force time-series in gait using over parameterised deep neural networks. Front. Bioeng. Biotechnol. 2023, 11. [Google Scholar] [CrossRef]
- Hady, D.A.A.; Mabrouk, O.M.; El-Hafeez, T.A. Employing machine learning for enhanced abdominal fat prediction in cavitation post-treatment. Sci. Rep. 2024, 14, 11004. [Google Scholar] [CrossRef]
- Strike.money. Time Series Analysis in Technical Trading. 2024. Available online: https://www.strike.money/technical-analysis/time-series-analysis (accessed on 19 June 2025).
- Li, G.; Shourijeh, M.S.; Ao, D.; Patten, C.; Fregly, B.J. How Well Do Commonly Used Co-contraction Indices Approximate Lower Limb Joint Stiffness Trends During Gait for Individuals Post-stroke? Front. Bioeng. Biotechnol. 2021, 8, 588908. [Google Scholar] [CrossRef]
- Eddo, O.; Lindsey, B.; Caswell, S.V.; Cortes, N. Current Evidence of Gait Modification with Real-time Biofeedback to Alter Kinetic, Temporospatial, and Function-Related Outcomes: A Review. Int. J. Kinesiol. Sport. Sci. 2017, 5, 35. [Google Scholar] [CrossRef]
- van den Noort, J.C.; Steenbrink, F.; Roeles, S.; Harlaar, J. Real-time visual feedback for gait retraining: Toward application in knee osteoarthritis. Med. Biol. Eng. Comput. 2015, 53, 275–286. [Google Scholar] [CrossRef]
- van Veen, B.C.; Mazza, C.; Viceconti, M. The Uncontrolled Manifold Theory Could Explain Part of the Inter-Trial Variability of Knee Contact Force During Level Walking. IEEE Trans. Neural Syst. Rehabil. Eng. 2020, 28, 1800–1807. [Google Scholar] [CrossRef]
- do Nascimento, T.C.F.; Gervásio, F.M.; Pignolo, A.; Bueno, G.A.S.; do Carmo, A.A.; Ribeiro, D.M.; D’Amelio, M.; dos Santos Mendes, F.A. Assessment of the Kinematic Adaptations in Parkinson’s Disease Using the Gait Profile Score: Influences of Trunk Posture, a Pilot Study. Brain Sci. 2021, 11, 1605. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Yu, H.; Wang, X.; Song, R. Characterizing stroke-induced changes in the variability of lower limb kinematics using multifractal detrended fluctuation analysis. Front. Neurol. 2022, 13, 893999. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.S.; Liao, Y.C.; Li, C.; Lee, B.; Oulasvirta, A. Real-time 3D Target Inference via Biomechanical Simulation. In Proceedings of the CHI Conference on Human Factors in Computing Systems, Honolulu, HI, USA, 11–16 May 2025; ACM: New York, NY, USA, 2024; pp. 1–18. [Google Scholar] [CrossRef]
- Gavrishchaka, V.; Senyukova, O.; Koepke, M. Synergy of physics-based reasoning and machine learning in biomedical applications: Towards unlimited deep learning with limited data. Adv. Phys. X 2019, 4, 1582361. [Google Scholar] [CrossRef]
- Pinheiro, J.M.H.; Oliveira, S.V.B.d.; Silva, T.H.S.; Saraiva, P.A.R.; Souza, E.F.d.; Godoy, R.V.; Ambrosio, L.A.; Becker, M. The Impact of Feature Scaling in Machine Learning: Effects on Regression and Classification Tasks. IEEE Access 2025, 13, 199903–199931. [Google Scholar] [CrossRef]




| Implantation Features | EMG | Skin-Marker Kinematics | Fluoroscopic Kinematics | GRF | Tasks |
|---|---|---|---|---|---|
| Frontal plane limb alignment | Gastrocnemius lateralis | Knee flexion | Knee flexion | Superior force | Walking |
| Posterior tibial slope | Gastrocnemius medialis | Hip flexion | Knee abduction | Anterior force | Squat |
| Hamstring lateralis | Hip adduction | Knee rotation | Medial force | Downhill Walking | |
| Hamstring medialis | Hip rotation | Free torque | Stairs Descent | ||
| Rectus femoris | Ankle flexion | Sit Down | |||
| Tibialis anterior | Stand Up | ||||
| Vastus lateralis | |||||
| Vastus medialis |
| K1L | K2L | K3R | K5R | K7L | K8L | Average | ||
|---|---|---|---|---|---|---|---|---|
| Walking | 12.3% (0.96) | 12.3% (0.99) | 14.4% (0.93) | 11.6% (0.99) | 14.3% (0.97) | 6.7% (0.99) | 11.9% (0.97) | |
| 11.4% (0.97) | 8.7% (0.99) | 17.0% (0.96) | 10.1% (0.98) | 17.9% (0.99) | 6.3% (0.99) | 11.9% (0.98) | ||
| 21.3% (0.74) | 51.4% (0.73) | 30.9% (0.23) | 19.2% (0.85) | 23.8% (0.64) | 20.1% (0.79) | 27.8% (0.66) | ||
| Squat | 14.2% (0.71) | 20.2% (0.89) | 36.3% (0.97) | 25.4% (0.89) | 12.3% (0.67) | 32.2% (0.93) | 23.4% (0.84) | |
| 19.7% (0.00) | 51.8% (0.75) | 60.5% (0.87) | 18.3% (0.89) | 20.4% (0.38) | 38.4% (0.74) | 34.9% (0.61) | ||
| 11.3% (0.92) | 11.6% (0.88) | 15.5% (0.97) | 31.5% (0.85) | 12.5% (0.92) | 24.6% (0.94) | 17.8% (0.92) | ||
| Downhill Walking | 17.0% (0.94) | 7.3% (0.99) | NA | 18.4% (0.98) | 17.4% (0.99) | 5.6% (0.99) | 13.1% (0.98) | |
| 19.0% (0.93) | 7.0% (0.99) | NA | 11.4% (0.97) | 21.7% (0.98) | 7.5% (0.98) | 13.3% (0.97) | ||
| 15.5% (0.88) | 24.3% (0.88) | NA | 27.1% (0.92) | 14.1% (0.88) | 12.6% (0.93) | 18.7% (0.90) | ||
| Stairs Descent | 16.8% (0.92) | 15.4% (0.98) | 16.4% (0.97) | 22.5% (0.95) | 17.1% (0.98) | 10.1% (0.97) | 16.4% (0.96) | |
| 17.6% (0.88) | 12.4% (0.98) | 24.2% (0.94) | 17.7% (0.95) | 18.5% (0.97) | 8.4% (0.98) | 16.5% (0.95) | ||
| 25.0% (0.86) | 27.9% (0.93) | 13.7% (0.87) | 27.7% (0.92) | 16.0% (0.93) | 12.6% (0.93) | 20.5% (0.91) | ||
| Sit Down | 7.3% (0.96) | 23.1% (0.78) | 16.6% (0.96) | 23.0% (0.93) | 11.0% (0.83) | 24.7% (0.94) | 17.6% (0.90) | |
| 15.4% (0.79) | 24.8% (0.79) | 27.7% (0.93) | 22.5% (0.93) | 19.7% (0.62) | 22.6% (0.90) | 22.1% (0.83) | ||
| 8.3% (0.96) | 17.8% (0.79) | 9.9% (0.94) | 22.9% (0.92) | 16.2% (0.93) | 25.4% (0.94) | 16.8% (0.91) | ||
| Stand Up | 11.5% (0.94) | 20.6% (0.96) | 16.6% (0.94) | 20.7% (0.98) | 9.1% (0.96) | 18.6% (0.98) | 16.2% (0.96) | |
| 16.3% (0.93) | 33.5% (0.96) | 25.7% (0.94) | 19.5% (0.97) | 10.3% (0.94) | 16.4% (0.89) | 20.3% (0.94) | ||
| 11.2% (0.93) | 12.6% (0.94) | 12.9% (0.94) | 21.3% (0.98) | 16.6% (0.95) | 22.4% (0.97) | 16.2% (0.95) |
| With All Inputs | With Only Marker-Based Kinematics | With Only Marker-Basedand Fluoroscopic Kinematics | Without Kinematics | Without GRF | Without EMG | ||
|---|---|---|---|---|---|---|---|
| Walking | 11.9% (0.97) | 19.5% (0.90) | 16.5% (0.93) | 13.3% (0.96) | 14.4% (0.92) | 12.1% (0.97) | |
| 11.9% (0.98) | 18.7% (0.91) | 17.8% (0.94) | 15.1% (0.96) | 15.4% (0.93) | 14.5% (0.97) | ||
| 27.8% (0.66) | 23.9% (0.69) | 25.5% (0.74) | 20.7% (0.72) | 24.0% (0.65) | 24.0% (0.78) | ||
| Squat | 23.4% (0.84) | 26.4% (0.72) | 26.9% (0.62) | 31.0% (0.81) | 24.7% (0.85) | 25.1% (0.84) | |
| 34.9% (0.61) | 40.0% (0.54) | 31.7% (0.32) | 37.2% (0.63) | 31.6% (0.59) | 38.5% (0.66) | ||
| 17.8% (0.92) | 23.4% (0.74) | 26.5% (0.72) | 30.5% (0.89) | 24.8% (0.88) | 20.7% (0.88) | ||
| Downhill Walking | 13.1% (0.98) | 19.2% (0.91) | 17.9% (0.92) | 13.0% (0.97) | 15.9% (0.95) | 13.5% (0.96) | |
| 13.3% (0.97) | 20.7% (0.88) | 20.0% (0.91) | 16.6% (0.96) | 17.0% (0.93) | 15.0% (0.96) | ||
| 18.7% (0.90) | 19.2% (0.87) | 20.2% (0.86) | 19.1% (0.82) | 20.2% (0.79) | 17.3% (0.89) | ||
| Stairs Descent | 16.4% (0.96) | 22.5% (0.91) | 20.9% (0.92) | 17.6% (0.96) | 19.1% (0.95) | 17.1% (0.96) | |
| 16.5% (0.95) | 22.0% (0.88) | 21.5% (0.89) | 18.7% (0.94) | 20.0% (0.93) | 18.2% (0.95) | ||
| 20.5% (0.91) | 22.3% (0.86) | 22.8% (0.88) | 21.1% (0.84) | 20.7% (0.82) | 20.1% (0.90) | ||
| Sit Down | 17.6% (0.90) | 26.3% (0.74) | 23.0% (0.75) | 28.2% (0.87) | 19.9% (0.85) | 19.8% (0.92) | |
| 22.1% (0.83) | 29.1% (0.70) | 23.6% (0.66) | 27.0% (0.84) | 22.5% (0.76) | 23.4% (0.87) | ||
| 16.8% (0.91) | 25.7% (0.68) | 26.6% (0.74) | 30.5% (0.87) | 19.9% (0.85) | 21.4% (0.88) | ||
| Stand Up | 16.2% (0.96) | 26.3% (0.79) | 25.0% (0.75) | 20.3% (0.90) | 17.6% (0.92) | 16.2% (0.96) | |
| 20.3% (0.94) | 27.4% (0.78) | 26.8% (0.70) | 22.3% (0.86) | 19.6% (0.86) | 20.4% (0.92) | ||
| 16.2% (0.95) | 26.4% (0.67) | 27.6% (0.61) | 23.3% (0.87) | 20.7% (0.87) | 19.3% (0.94) |
| biLSTM-MLP | TCN | ||
|---|---|---|---|
| Walking | 11.9% (0.97) | 17.6% (0.95) | |
| 11.9% (0.98) | 17.2% (0.97) | ||
| 27.8% (0.66) | 25.8% (0.67) | ||
| Squat | 23.4% (0.84) | 28.8% (0.75) | |
| 34.9% (0.61) | 34.4% (0.65) | ||
| 17.8% (0.92) | 25.2% (0.74) | ||
| Downhill Walking | 13.1% (0.98) | 18.4% (0.96) | |
| 13.3% (0.97) | 18.8% (0.96) | ||
| 18.7% (0.90) | 19.8% (0.86) | ||
| Stairs Descent | 16.4% (0.96) | 20.5% (0.96) | |
| 16.5% (0.95) | 20.8% (0.94) | ||
| 20.5% (0.91) | 21.7% (0.86) | ||
| Sit Down | 17.6% (0.90) | 24.0% (0.87) | |
| 22.1% (0.83) | 26.3% (0.79) | ||
| 16.8% (0.91) | 21.4% (0.83) | ||
| Stand Up | 16.2% (0.96) | 20.9% (0.94) | |
| 20.3% (0.94) | 24.0% (0.89) | ||
| 16.2% (0.95) | 23.0% (0.82) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Derungs, Y.N.; Bertsch, M.; Malla, K.; Maas, A.; Grupp, T.M.; Trepczynski, A.; Damm, P.; Nasab, S.H.H. Machine Learning-Based Estimation of Knee Joint Mechanics from Kinematic and Neuromuscular Inputs: A Proof-of-Concept Using the CAMS-Knee Datasets. Bioengineering 2026, 13, 173. https://doi.org/10.3390/bioengineering13020173
Derungs YN, Bertsch M, Malla K, Maas A, Grupp TM, Trepczynski A, Damm P, Nasab SHH. Machine Learning-Based Estimation of Knee Joint Mechanics from Kinematic and Neuromuscular Inputs: A Proof-of-Concept Using the CAMS-Knee Datasets. Bioengineering. 2026; 13(2):173. https://doi.org/10.3390/bioengineering13020173
Chicago/Turabian StyleDerungs, Yara N., Martin Bertsch, Kushal Malla, Allan Maas, Thomas M. Grupp, Adam Trepczynski, Philipp Damm, and Seyyed Hamed Hosseini Nasab. 2026. "Machine Learning-Based Estimation of Knee Joint Mechanics from Kinematic and Neuromuscular Inputs: A Proof-of-Concept Using the CAMS-Knee Datasets" Bioengineering 13, no. 2: 173. https://doi.org/10.3390/bioengineering13020173
APA StyleDerungs, Y. N., Bertsch, M., Malla, K., Maas, A., Grupp, T. M., Trepczynski, A., Damm, P., & Nasab, S. H. H. (2026). Machine Learning-Based Estimation of Knee Joint Mechanics from Kinematic and Neuromuscular Inputs: A Proof-of-Concept Using the CAMS-Knee Datasets. Bioengineering, 13(2), 173. https://doi.org/10.3390/bioengineering13020173







