Synthesizing Complex-Valued Multicoil MRI Data from Magnitude-Only Images
Abstract
1. Introduction
2. Methods
2.1. Preliminaries
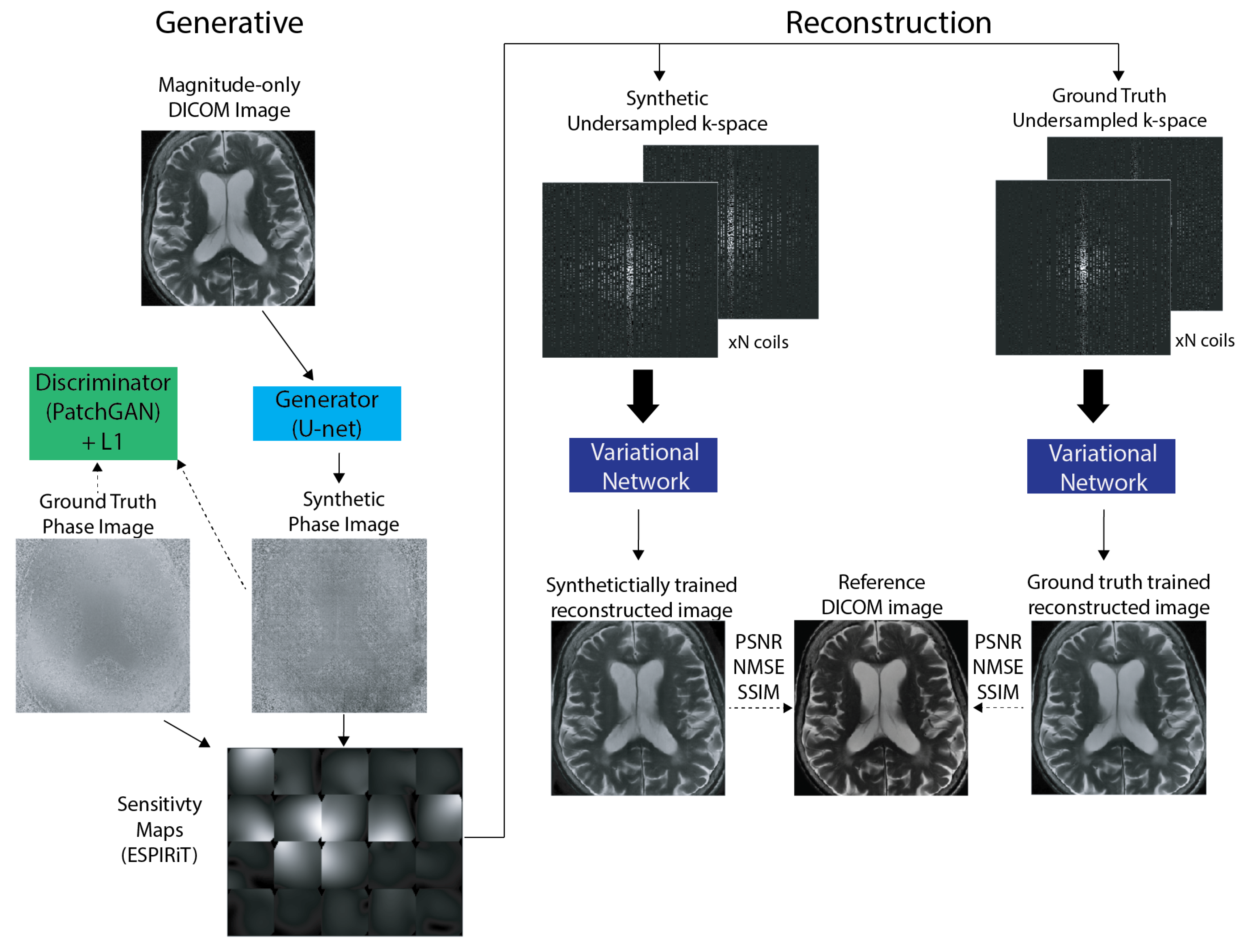
2.2. Generative Modeling
2.2.1. Neural Network Architecture
2.2.2. Synthetic Phase Generation
2.2.3. Multi-Coil Data Generation
2.3. Dataset
2.4. Experiments
2.4.1. Generative
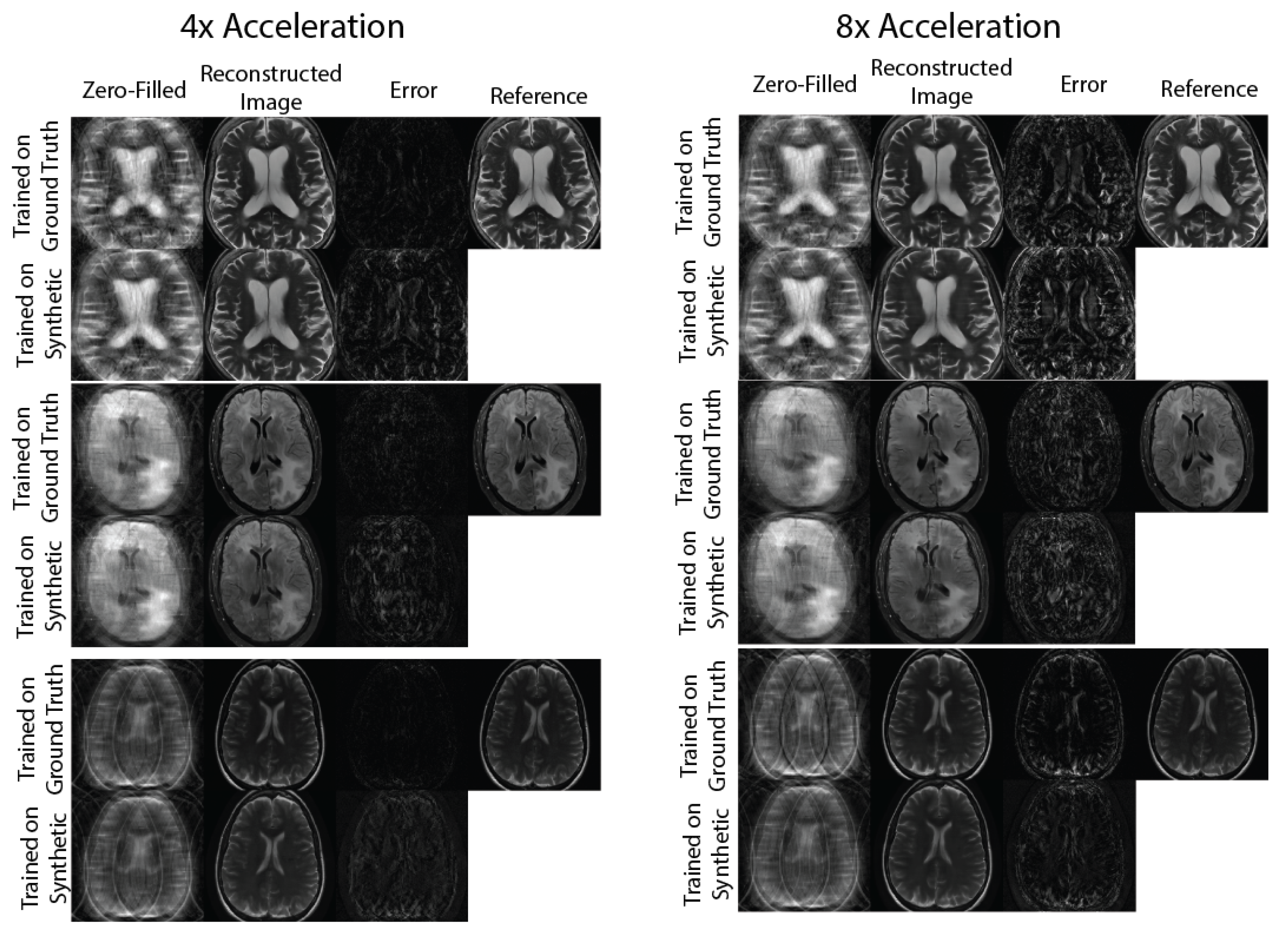
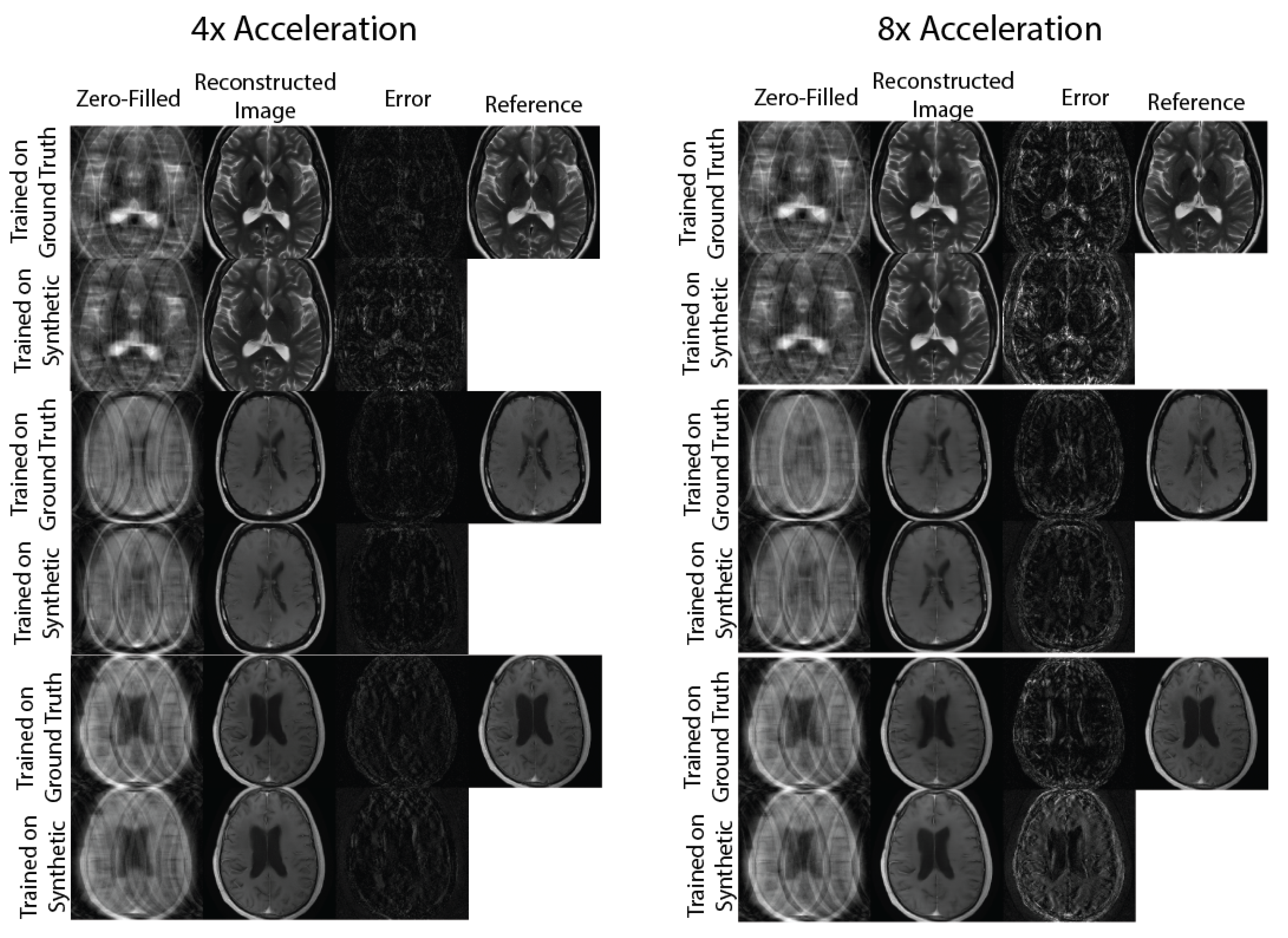
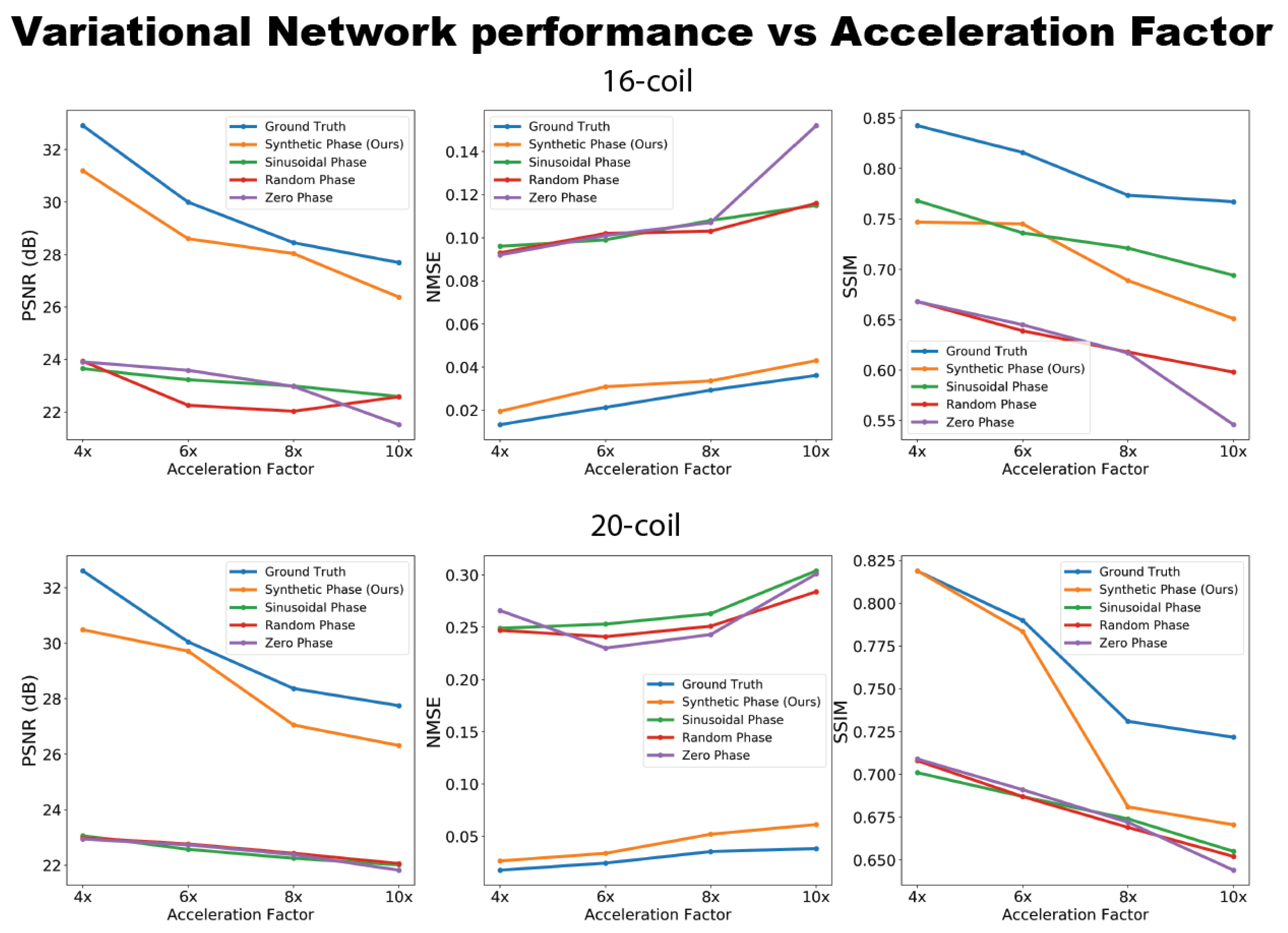
2.4.2. Evaluation: Physics-Based Image Reconstruction
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Knoll, F.; Murrell, T.; Sriram, A.; Yakubova, N.; Zbontar, J.; Rabbat, M.; Defazio, A.; Muckley, M.J.; Sodickson, D.K.; Zitnick, C.L.; et al. Advancing machine learning for MR image reconstruction with an open competition: Overview of the 2019 fastMRI challenge. Magn. Reson. Med. 2020, 84, 3054–3070. [Google Scholar] [CrossRef] [PubMed]
- Ong, F.; Amin, S.; Vasanawala, S.; Lustig, M. Mridata.org: An open archive for sharing MRI raw data. Proc. Intl. Soc. Mag. Reson. Med. 2018, 26, 1. [Google Scholar]
- Calgary Campinas Public Dataset. 2020. Available online: https://sites.google.com/view/calgary-campinas-dataset/mr-reconstruction-challenge?authuser=0 (accessed on 22 March 2021).
- Desai, A.D.; Schmidt, A.M.; Rubin, E.B.; Sandino, C.M.; Black, M.S.; Mazzoli, V.; Stevens, K.J.; Boutin, R.; Re, C.; Gold, G.E.; et al. SKM-TEA: A Dataset for Accelerated MRI Reconstruction with Dense Image Labels for Quantitative Clinical Evaluation. In Proceedings of the Thirty-Fifth Conference on Neural Information Processing Systems Datasets and Benchmarks Track (Round 2); Stanford University: Stanford, CA, USA, 2021. [Google Scholar]
- Lim, Y.; Toutios, A.; Bliesener, Y.; Tian, Y.; Lingala, S.G.; Vaz, C.; Sorensen, T.; Oh, M.; Harper, S.; Chen, W.; et al. A multispeaker dataset of raw and reconstructed speech production real-time MRI video and 3D volumetric images. arXiv 2021, arXiv:2102.07896. [Google Scholar] [CrossRef] [PubMed]
- IXI Dataset. 2010. Available online: http://brain-development.org/ixi-dataset/ (accessed on 22 March 2021).
- The Human Connectome Project, University of Souterhn California. 2011. Available online: http://www.humanconnectomeproject.org/ (accessed on 22 March 2021).
- AccelMR Dataset. 2020. Available online: https://accelmrorg.wordpress.com/organizers/#jp-carousel-135/ (accessed on 2 May 2021).
- Oasis Dataset. 2007. Available online: https://www.oasis-brains.org/ (accessed on 2 May 2021).
- ADNI Dataset. 2004. Available online: http://adni.loni.usc.edu/data-samples/data-types/ (accessed on 2 May 2021).
- UC Biobank. 2006. Available online: https://www.ukbiobank.ac.uk/ (accessed on 2 May 2021).
- The Cancer Imaging Archive. 2010. Available online: http://www.cancerimagingarchive.net (accessed on 2 May 2021).
- Brain Tumor MRI Dataset. 2020. Available online: https://www.kaggle.com/shlezinger/brain-mri-data/ (accessed on 2 May 2021).
- Shimron, E.; Tamir, J.I.; Wang, K.; Lustig, M. Implicit data crimes: Machine learning bias arising from misuse of public data. Proc. Natl. Acad. Sci. USA 2022, 119, e2117203119. [Google Scholar] [CrossRef]
- Hammernik, K.; Küstner, T.; Yaman, B.; Huang, Z.; Rueckert, D.; Knoll, F.; Akçakaya, M. Physics-Driven Deep Learning for Computational Magnetic Resonance Imaging. arXiv 2022, arXiv:2203.12215. [Google Scholar] [CrossRef]
- Muckley, M.J.; Riemenschneider, B.; Radmanesh, A.; Kim, S.; Jeong, G.; Ko, J.; Jun, Y.; Shin, H.; Hwang, D.; Mostapha, M.; et al. Results of the 2020 fastMRI Challenge for Machine Learning MR Image Reconstruction. IEEE Trans. Med. Imaging 2021, 40, 2306–2317. [Google Scholar] [CrossRef]
- Caliva’, F.; Shah, R.; Bharadwaj, U.U.; Majumdar, S.; Larson, P.; Pedoia, V. Breaking Speed Limits with Simultaneous Ultra-Fast {MRI} Reconstruction and Tissue Segmentation. In Proceedings of the Medical Imaging with Deep Learning; ML Research Press: Maastricht, The Netherlands, 2020. [Google Scholar]
- Eggers, H.; Börnert, P. Chemical shift encoding-based water-fat separation methods. J. Magn. Reson. Imaging 2014, 40, 251–268. [Google Scholar] [CrossRef]
- He, X.; Yablonskiy, D.A. Biophysical mechanisms of phase contrast in gradient echo MRI. Proc. Natl. Acad. Sci. USA 2009, 106, 13558–13563. [Google Scholar] [CrossRef]
- Markl, M.; Chan, F.P.; Alley, M.T.; Wedding, K.L.; Draney, M.T.; Elkins, C.J.; Parker, D.W.; Wicker, R.; Taylor, C.A.; Herfkens, R.J.; et al. Time-resolved three-dimensional phase-contrast MRI. J. Magn. Reson. Imaging 2003, 17, 499–506. [Google Scholar] [CrossRef]
- Zhuang, B.; Sirajuddin, A.; Zhao, S.; Lu, M. The role of 4D flow MRI for clinical applications in cardiovascular disease: Current status and future perspectives. Quant. Imaging Med. Surg. 2021, 11, 4193–4210. [Google Scholar] [CrossRef]
- Duyn, J.H.; van Gelderen, P.; Li, T.Q.; de Zwart, J.A.; Koretsky, A.P.; Fukunaga, M. High-field MRI of brain cortical substructure based on signal phase. Proc. Natl. Acad. Sci. USA 2007, 104, 11796–11801. [Google Scholar] [CrossRef] [PubMed]
- Shimron, E.; Grissom, W.; Azhari, H. Temporal differences (TED) compressed sensing: A method for fast MRgHIFU temperature imaging. NMR Biomed. 2020, 33, e4352. [Google Scholar] [CrossRef]
- Cole, E.; Cheng, J.; Pauly, J.; Vasanawala, S. Analysis of deep complex-valued convolutional neural networks for MRI reconstruction and phase-focused applications. Magn. Reson. Med. 2021, 86, 1093–1109. [Google Scholar] [CrossRef] [PubMed]
- Bassey, J.; Qian, L.; Li, X. A Survey of Complex-Valued Neural Networks. arXiv 2021, arXiv:2101.12249. [Google Scholar] [CrossRef]
- Ji, S.; Yang, D.; Lee, J.; Choi, S.H.; Kim, H.; Kang, K.M. Synthetic MRI: Technologies and Applications in Neuroradiology. J. Magn. Reson. Imaging 2020, 55, 1013–1025. [Google Scholar] [CrossRef]
- Gulani, V.; Schmitt, P.; Griswold, M.A.; Webb, A.G.; Jakob, P.M. Towards a Single-Sequence Neurologic Magnetic Resonance Imaging Examination: Multiple-Contrast Images From an IR TrueFISP Experiment. Investig. Radiol. 2004, 39, 767–774. [Google Scholar] [CrossRef]
- Warntjes, J.; Leinhard, O.D.; West, J.; Lundberg, P. Rapid magnetic resonance quantification on the brain: Optimization for clinical usage. Magn. Reson. Med. 2008, 60, 320–329. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, S.; Wang, Y.; Kang, Y.; Haacke, E.M. STrategically Acquired Gradient Echo (STAGE) imaging, part I: Creating enhanced T1 contrast and standardized susceptibility weighted imaging and quantitative susceptibility mapping. Magn. Reson. Imaging 2018, 46, 130–139. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, Y.; Wu, D.; Wang, Y.; Sethi, S.K.; Yang, G.; Xie, H.; Xia, S.; Haacke, E.M. STrategically Acquired Gradient Echo (STAGE) imaging, part II: Correcting for RF inhomogeneities in estimating T1 and proton density. Magn. Reson. Imaging 2018, 46, 140–150. [Google Scholar] [CrossRef]
- Haacke, E.M.; Chen, Y.; Utriainen, D.; Wu, B.; Wang, Y.; Xia, S.; He, N.; Zhang, C.; Wang, X.; Lagana, M.M.; et al. STrategically Acquired Gradient Echo (STAGE) imaging, part III: Technical advances and clinical applications of a rapid multi-contrast multi-parametric brain imaging method. Magn. Reson. Imaging 2020, 65, 15–26. [Google Scholar] [CrossRef]
- Callaghan, M.F.; Mohammadi, S.; Weiskopf, N. Synthetic quantitative MRI through relaxometry modelling. NMR Biomed. 2016, 29, 1729–1738. [Google Scholar] [CrossRef]
- Blystad, I.; Warntjes, J.; Smedby, O.; Landtblom, A.M.; Lundberg, P.; Larsson, E.M. Synthetic Mri of the Brain in a Clinical Setting. Acta Radiol. 2012, 53, 1158–1163. [Google Scholar] [CrossRef] [PubMed]
- Loecher, M.; Perotti, L.E.; Ennis, D.B. Using synthetic data generation to train a cardiac motion tag tracking neural network. Med. Image Anal. 2021, 74, 102223. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Liu, J.Z.; Cauley, S.F.; Rosen, B.R.; Rosen, M.S. Image reconstruction by domain-transform manifold learning. Nature 2018, 555, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Knoll, F.; Hammernik, K.; Kobler, E.; Pock, T.; Recht, M.P.; Sodickson, D.K. Assessment of the generalization of learned image reconstruction and the potential for transfer learning. Magn. Reson. Med. 2018, 81, 116–128. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Nets. In Proceedings of the Advances in Neural Information Processing Systems; Ghahramani, Z., Welling, M., Cortes, C., Lawrence, N., Weinberger, K., Eds.; Curran Associates, Inc.: Nice, France, 2014; Volume 27. [Google Scholar]
- Isola, P.; Zhu, J.Y.; Zhou, T.; Efros, A.A. Image-to-Image Translation with Conditional Adversarial Networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017. [Google Scholar] [CrossRef]
- Sriram, A.; Zbontar, J.; Murrell, T.; Defazio, A.; Zitnick, C.L.; Yakubova, N.; Knoll, F.; Johnson, P. End-to-End Variational Networks for Accelerated MRI Reconstruction. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2020; Springer International Publishing: Berlin/Heidelberg, Germany, 2020; pp. 64–73. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Lecture Notes in Computer Science; Springer International Publishing: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar] [CrossRef]
- Uecker, M.; Lai, P.; Murphy, M.J.; Virtue, P.; Elad, M.; Pauly, J.M.; Vasanawala, S.S.; Lustig, M. ESPIRiT-an eigenvalue approach to autocalibrating parallel MRI: Where SENSE meets GRAPPA. Magn. Reson. Med. 2013, 71, 990–1001. [Google Scholar] [CrossRef]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar] [CrossRef]
- Saharia, C.; Chan, W.; Chang, H.; Lee, C.; Ho, J.; Salimans, T.; Fleet, D.; Norouzi, M. Palette: Image-to-Image Diffusion Models. In Proceedings of the Special Interest Group on Computer Graphics and Interactive Techniques Conference Proceedings; ACM: New York, NY, USA, 2022. [Google Scholar] [CrossRef]
- Angelopoulos, A.N.; Kohli, A.P.; Bates, S.; Jordan, M.; Malik, J.; Alshaabi, T.; Upadhyayula, S.; Romano, Y. Image-to-Image Regression with Distribution-Free Uncertainty Quantification and Applications in Imaging. In Proceedings of the 39th International Conference on Machine Learning, Baltimore, MD, USA, 17–23 July 2022; Volume 162, pp. 717–730. [Google Scholar]
- Borji, A. Pros and Cons of GAN Evaluation Measures. arXiv 2018, arXiv:1802.03446. [Google Scholar] [CrossRef]
- Salimans, T.; Goodfellow, I.; Zaremba, W.; Cheung, V.; Radford, A.; Chen, X.; Chen, X. Improved Techniques for Training GANs. In Proceedings of the Advances in Neural Information Processing Systems; Lee, D., Sugiyama, M., Luxburg, U., Guyon, I., Garnett, R., Eds.; Curran Associates, Inc.: Nice, France, 2016; Volume 29. [Google Scholar]
- Heusel, M.; Ramsauer, H.; Unterthiner, T.; Nessler, B.; Hochreiter, S. GANs Trained by a Two Time-Scale Update Rule Converge to a Local Nash Equilibrium. In Proceedings of the Advances in Neural Information Processing Systems; Guyon, I., Luxburg, U.V., Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S., Garnett, R., Eds.; Curran Associates, Inc.: Nice, France, 2017; Volume 30. [Google Scholar]
- Mei, X.; Liu, Z.; Robson, P.M.; Marinelli, B.; Huang, M.; Doshi, A.; Jacobi, A.; Cao, C.; Link, K.E.; Yang, T.; et al. RadImageNet: An Open Radiologic Deep Learning Research Dataset for Effective Transfer Learning. Radiol. Artif. Intell. 2022, 4. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Deveshwar, N.; Rajagopal, A.; Sahin, S.; Shimron, E.; Larson, P.E.Z. Synthesizing Complex-Valued Multicoil MRI Data from Magnitude-Only Images. Bioengineering 2023, 10, 358. https://doi.org/10.3390/bioengineering10030358
Deveshwar N, Rajagopal A, Sahin S, Shimron E, Larson PEZ. Synthesizing Complex-Valued Multicoil MRI Data from Magnitude-Only Images. Bioengineering. 2023; 10(3):358. https://doi.org/10.3390/bioengineering10030358
Chicago/Turabian StyleDeveshwar, Nikhil, Abhejit Rajagopal, Sule Sahin, Efrat Shimron, and Peder E. Z. Larson. 2023. "Synthesizing Complex-Valued Multicoil MRI Data from Magnitude-Only Images" Bioengineering 10, no. 3: 358. https://doi.org/10.3390/bioengineering10030358
APA StyleDeveshwar, N., Rajagopal, A., Sahin, S., Shimron, E., & Larson, P. E. Z. (2023). Synthesizing Complex-Valued Multicoil MRI Data from Magnitude-Only Images. Bioengineering, 10(3), 358. https://doi.org/10.3390/bioengineering10030358






