Investigation of DNA Hybridization on Nano-Structured Plasmonic Surfaces for Identifying Nasopharyngeal Viruses
Abstract
:1. Introduction
2. Materials and Methods
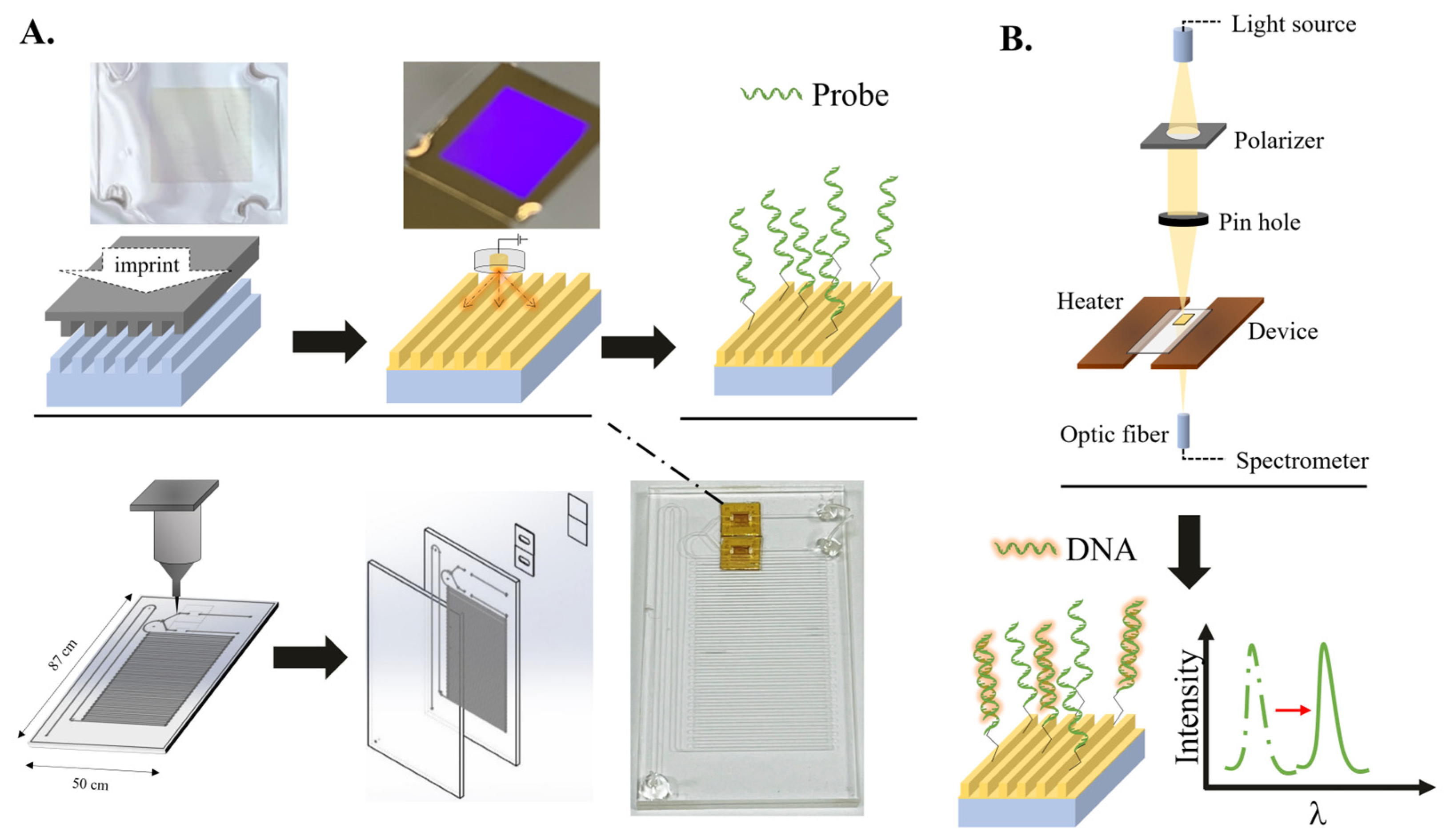
2.1. Integration of the Microfluidic Device
2.2. The Operation of the Device
2.3. PCR Analysis
2.4. Cells
2.5. Plasmids
2.6. Production of the Lentivirus and Cell Infection
2.7. DNA Extraction
2.8. RNA Extraction and Reverse Transcription
3. Results
3.1. Temperature Stability and Sensitivity of the Microfluidic Device
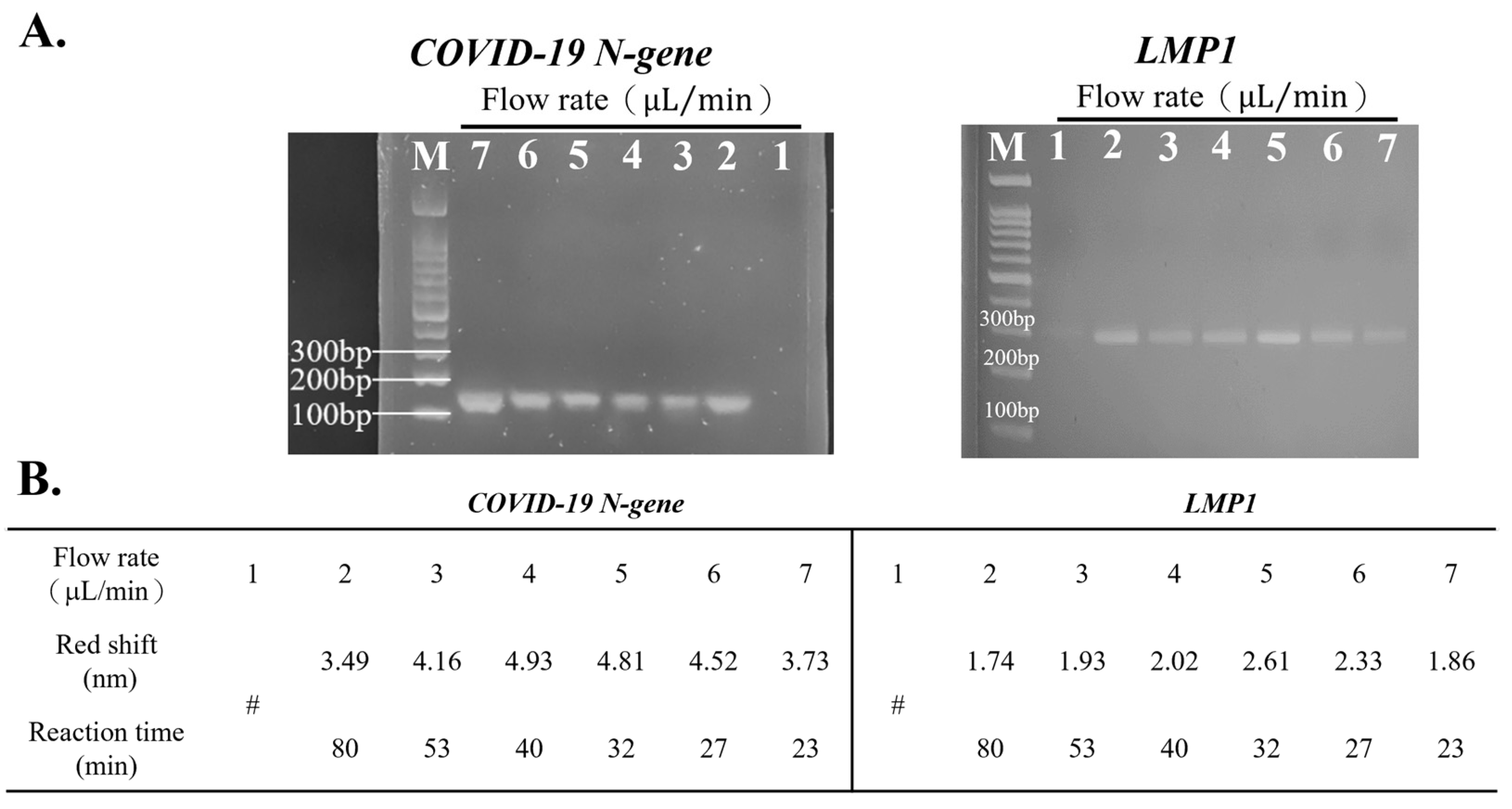
3.2. Optimal Flow Rate for Detection Using the Device
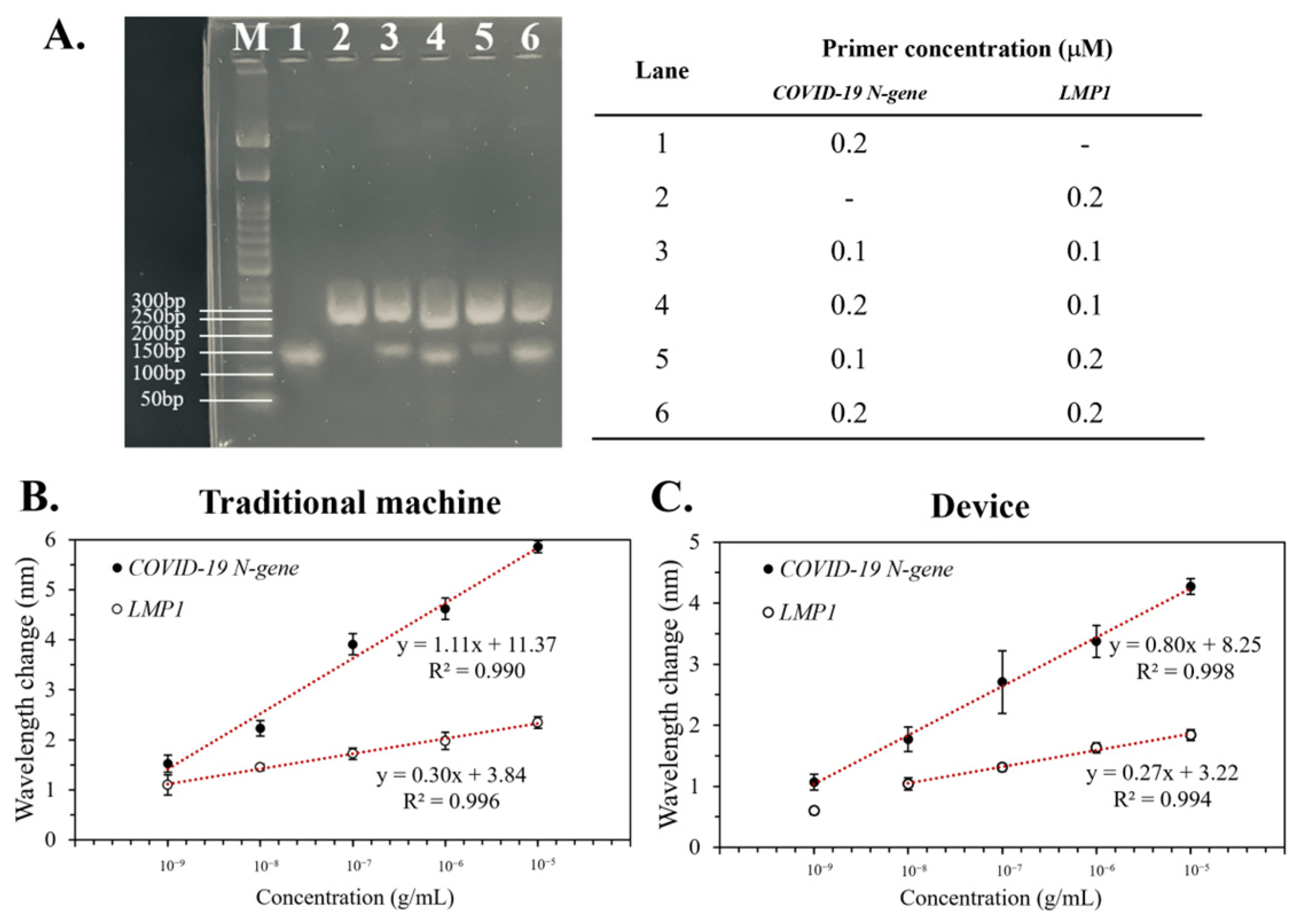
3.3. The Limit of Detection (LOD) of the Device
3.4. Specificity of the Device
3.5. Duplex PCR of the Device
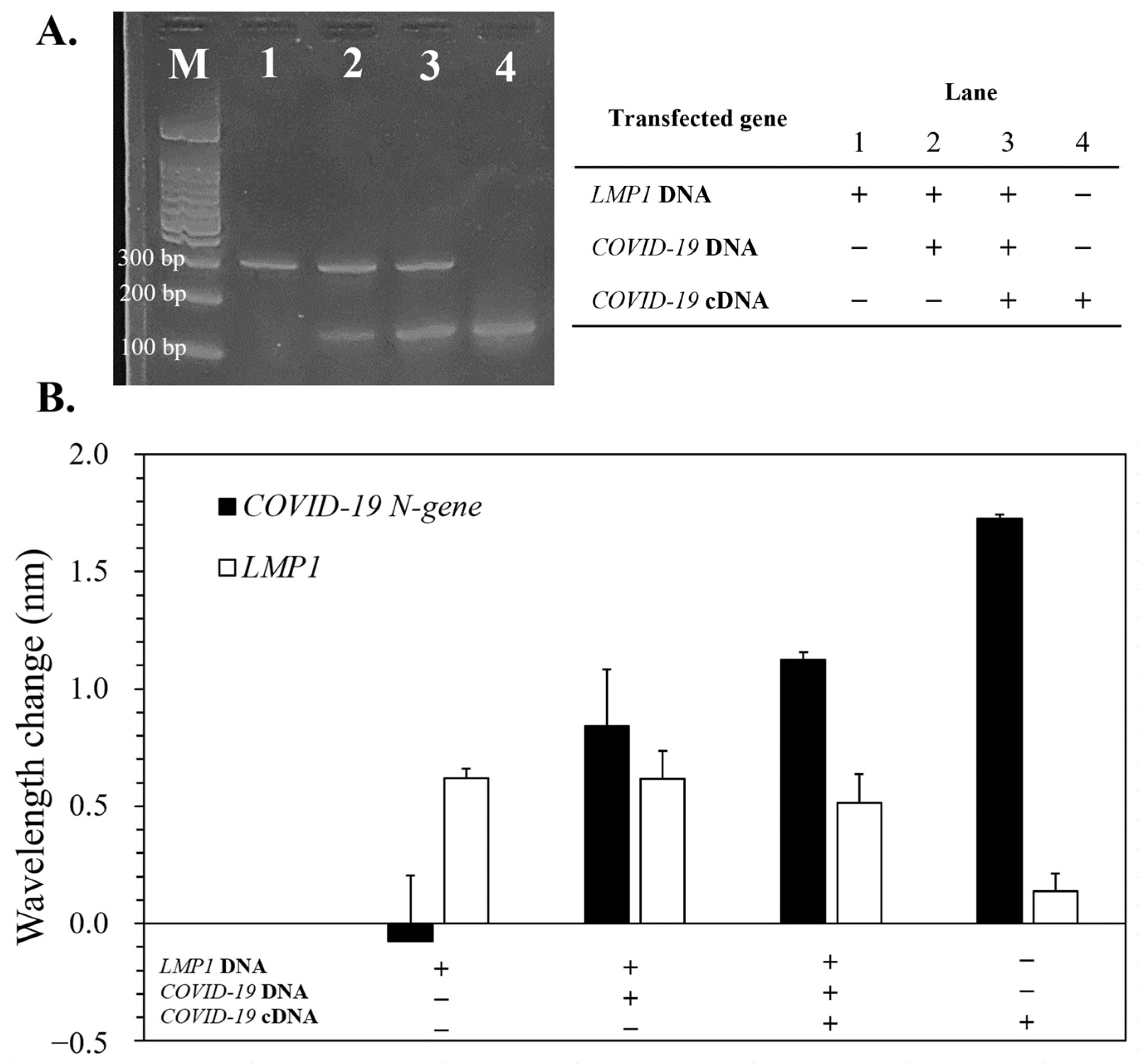
3.6. Detection of LMP1/COVID-19 N-Gene in Artificial HHV-4/SARS-CoV-2-Modified Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chircov, C.; Grumezescu, A.M. Microelectromechanical Systems (MEMS) for Biomedical Applications. Micromachines 2022, 13, 164. [Google Scholar] [CrossRef] [PubMed]
- Ramalingam, N.; Rui, Z.; Liu, H.-B.; Dai, C.-C.; Kaushik, R.; Ratnaharika, B.; Gong, H.-Q. Real-time PCR-based microfluidic array chip for simultaneous detection of multiple waterborne pathogens. Sens. Actuators B Chem. 2010, 145, 543–552. [Google Scholar] [CrossRef]
- Mauk, M.G.; Song, J.; Liu, C.; Bau, H.H. Simple Approaches to Minimally-Instrumented, Microfluidic-Based Point-of-Care Nucleic Acid Amplification Tests. Biosensors 2018, 8, 17. [Google Scholar] [CrossRef] [PubMed]
- Ahrberg, C.D.; Manz, A.; Chung, B.G. Polymerase chain reaction in microfluidic devices. Lab Chip 2016, 16, 3866–3884. [Google Scholar] [CrossRef]
- Shaffaf, T.; Ghafar-Zadeh, E. COVID-19 Diagnostic Strategies. Part I: Nucleic Acid-Based Technologies. Bioengineering 2021, 8, 49. [Google Scholar] [CrossRef]
- Felbel, J.; Reichert, A.; Kielpinski, M.; Urban, M.; Henkel, T.; Häfner, N.; Dürst, M.; Weber, J. Reverse transcription-polymerase chain reaction (RT-PCR) in flow-through micro-reactors: Thermal and fluidic concepts. Chem. Eng. J. 2008, 135, S298–S302. [Google Scholar] [CrossRef]
- Yin, H.; Wu, Z.; Shi, N.; Qi, Y.; Jian, X.; Zhou, L.; Tong, Y.; Cheng, Z.; Zhao, J.; Mao, H. Ultrafast multiplexed detection of SARS-CoV-2 RNA using a rapid droplet digital PCR system. Biosens. Bioelectron. 2021, 188, 113282. [Google Scholar] [CrossRef]
- Syahir, A.; Usui, K.; Tomizaki, K.-y.; Kajikawa, K.; Mihara, H. Label and Label-Free Detection Techniques for Protein Microarrays. Microarrays 2015, 4, 228–244. [Google Scholar] [CrossRef]
- Ebrahimi, G.; Samadi Pakchin, P.; Shamloo, A.; Mota, A.; de la Guardia, M.; Omidian, H.; Omidi, Y. Label-free electrochemical microfluidic biosensors: Futuristic point-of-care analytical devices for monitoring diseases. Microchim. Acta 2022, 189, 252. [Google Scholar] [CrossRef]
- Prabowo, B.A.; Purwidyantri, A.; Liu, K.-C. Surface Plasmon Resonance Optical Sensor: A Review on Light Source Technology. Biosensors 2018, 8, 80. [Google Scholar] [CrossRef]
- Lo, S.-C.; Li, S.-S.; Yang, W.-F.; Wu, K.-C.; Wei, P.-K.; Sheen, H.-J.; Fan, Y.-J. A Co-Printed Nanoslit Surface Plasmon Resonance Structure in Microfluidic Device for LMP-1 Detection. Biosensors 2022, 12, 653. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Trinh, K.T.L.; Yoon, W.J.; Lee, N.Y.; Ju, H. Integration of a microfluidic polymerase chain reaction device and surface plasmon resonance fiber sensor into an inline all-in-one platform for pathogenic bacteria detection. Sens. Actuators B Chem. 2017, 242, 1–8. [Google Scholar] [CrossRef]
- Zhu, L.; Wang, K.; Cui, J.; Liu, H.; Bu, X.; Ma, H.; Wang, W.; Gong, H.; Lausted, C.; Hood, L.; et al. Label-Free Quantitative Detection of Tumor-Derived Exosomes through Surface Plasmon Resonance Imaging. Anal. Chem. 2014, 86, 8857–8864. [Google Scholar] [CrossRef] [PubMed]
- Lim, W.Y.; Lan, B.L.; Ramakrishnan, N. Emerging Biosensors to Detect Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): A Review. Biosensors 2021, 11, 434. [Google Scholar] [CrossRef] [PubMed]
- Cennamo, N.; Pasquardini, L.; Arcadio, F.; Lunelli, L.; Vanzetti, L.; Carafa, V.; Altucci, L.; Zeni, L. SARS-CoV-2 spike protein detection through a plasmonic D-shaped plastic optical fiber aptasensor. Talanta 2021, 233. [Google Scholar] [CrossRef]
- Chen, T.; Song, J.Y.; Liu, H.L.; Zheng, H.M.; Chen, C.Z. Positive Epstein-Barr virus detection in coronavirus disease 2019 (COVID-19) patients. Sci. Rep. 2021, 11, 10902. [Google Scholar] [CrossRef]
- Meng, M.; Zhang, S.; Dong, X.; Sun, W.Q.; Deng, Y.F.; Li, W.Z.; Li, R.R.; Annane, D.; Wu, Z.X.; Chen, D.C. COVID-19 associated EBV reactivation and effects of ganciclovir treatment. Immun. Inflamm. Dis. 2022, 10, e597. [Google Scholar] [CrossRef] [PubMed]
- Roncati, L.; Lusenti, B.; Nasillo, V.; Manenti, A. Fatal SARS-CoV-2 coinfection in course of EBV-associated lymphoproliferative disease. Ann. Hematol. 2020, 99, 1945–1946. [Google Scholar] [CrossRef]
- Vaisocherová, H.; Mrkvová, K.; Piliarik, M.; Jinoch, P.; Šteinbachová, M.; Homola, J. Surface plasmon resonance biosensor for direct detection of antibody against Epstein-Barr virus. Biosens. Bioelectron. 2007, 22, 1020–1026. [Google Scholar] [CrossRef]
- Hsieh, H.-Y.; Chang, R.; Huang, Y.-Y.; Juan, P.-H.; Tahara, H.; Lee, K.-Y.; Vo, D.N.K.; Tsai, M.-H.; Wei, P.-K.; Sheen, H.-J.; et al. Continuous polymerase chain reaction microfluidics integrated with a gold-capped nanoslit sensing chip for Epstein-Barr virus detection. Biosens. Bioelectron. 2022, 195, 113672. [Google Scholar] [CrossRef]
- Faghih, M.M.; Sharp, M.K. Solvent-based bonding of PMMA–PMMA for microfluidic applications. Microsyst. Technol. 2019, 25, 3547–3558. [Google Scholar] [CrossRef]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed]
- Plesa, C.; Verschueren, D.; Pud, S.; van der Torre, J.; Ruitenberg, J.W.; Witteveen, M.J.; Jonsson, M.P.; Grosberg, A.Y.; Rabin, Y.; Dekker, C. Direct observation of DNA knots using a solid-state nanopore. Nat. Nanotechnol. 2016, 11, 1093–1097. [Google Scholar] [CrossRef] [PubMed]
- Jeong, K.-B.; Luo, K.; Lim, M.-C.; Jung, J.-Y.; Yu, J.-S.; Kim, K.-B.; Kim, Y.-R. Reduction of DNA Folding by Ionic Liquids and Its Effects on the Analysis of DNA–Protein Interaction Using Solid-State Nanopore. Small 2018, 14, 1801375. [Google Scholar] [CrossRef]
- Welch, N.L.; Zhu, M.; Hua, C.; Weller, J.; Mirhashemi, M.E.; Nguyen, T.G.; Mantena, S.; Bauer, M.R.; Shaw, B.M.; Ackerman, C.M.; et al. Multiplexed CRISPR-based microfluidic platform for clinical testing of respiratory viruses and identification of SARS-CoV-2 variants. Nat. Med. 2022, 28, 1083–1094. [Google Scholar] [CrossRef]
- Sun, F.; Ganguli, A.; Nguyen, J.; Brisbin, R.; Shanmugam, K.; Hirschberg, D.L.; Wheeler, M.B.; Bashir, R.; Nash, D.M.; Cunningham, B.T. Smartphone-based multiplex 30-minute nucleic acid test of live virus from nasal swab extract. Lab Chip 2020, 20, 1621–1627. [Google Scholar] [CrossRef]
- Chu, Y.; Qiu, J.; Wang, Y.; Wang, M.; Zhang, Y.; Han, L. Rapid and High-Throughput SARS-CoV-2 RNA Detection without RNA Extraction and Amplification by Using a Microfluidic Biochip. Chem. Eur. J. 2022, 28, e202104054. [Google Scholar] [CrossRef]
- Masterson, A.N.; Muhoberac, B.B.; Gopinadhan, A.; Wilde, D.J.; Deiss, F.T.; John, C.C.; Sardar, R. Multiplexed and High-Throughput Label-Free Detection of RNA/Spike Protein/IgG/IgM Biomarkers of SARS-CoV-2 Infection Utilizing Nanoplasmonic Biosensors. Anal. Chem. 2021, 93, 8754–8763. [Google Scholar] [CrossRef]
- Funari, R.; Chu, K.-Y.; Shen, A.Q. Detection of antibodies against SARS-CoV-2 spike protein by gold nanospikes in an opto-microfluidic chip. Biosens. Bioelectron. 2020, 169, 112578. [Google Scholar] [CrossRef]
- Xu, W.; Liu, J.; Song, D.; Li, C.; Zhu, A.; Long, F. Rapid, label-free, and sensitive point-of-care testing of anti-SARS-CoV-2 IgM/IgG using all-fiber Fresnel reflection microfluidic biosensor. Microchim. Acta 2021, 188, 261. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.-S.; Lu, Y.-J.; Chang, R.; Tsai, M.-H.; Hung, J.-N.; Chen, W.-H.; Fan, Y.-J.; Wei, P.-K.; Sheen, H.-J. Investigation of DNA Hybridization on Nano-Structured Plasmonic Surfaces for Identifying Nasopharyngeal Viruses. Bioengineering 2023, 10, 1189. https://doi.org/10.3390/bioengineering10101189
Li S-S, Lu Y-J, Chang R, Tsai M-H, Hung J-N, Chen W-H, Fan Y-J, Wei P-K, Sheen H-J. Investigation of DNA Hybridization on Nano-Structured Plasmonic Surfaces for Identifying Nasopharyngeal Viruses. Bioengineering. 2023; 10(10):1189. https://doi.org/10.3390/bioengineering10101189
Chicago/Turabian StyleLi, Shao-Sian, Yi-Jung Lu, Ray Chang, Ming-Han Tsai, Jo-Ning Hung, Wei-Hung Chen, Yu-Jui Fan, Pei-Kuen Wei, and Horn-Jiunn Sheen. 2023. "Investigation of DNA Hybridization on Nano-Structured Plasmonic Surfaces for Identifying Nasopharyngeal Viruses" Bioengineering 10, no. 10: 1189. https://doi.org/10.3390/bioengineering10101189
APA StyleLi, S.-S., Lu, Y.-J., Chang, R., Tsai, M.-H., Hung, J.-N., Chen, W.-H., Fan, Y.-J., Wei, P.-K., & Sheen, H.-J. (2023). Investigation of DNA Hybridization on Nano-Structured Plasmonic Surfaces for Identifying Nasopharyngeal Viruses. Bioengineering, 10(10), 1189. https://doi.org/10.3390/bioengineering10101189





