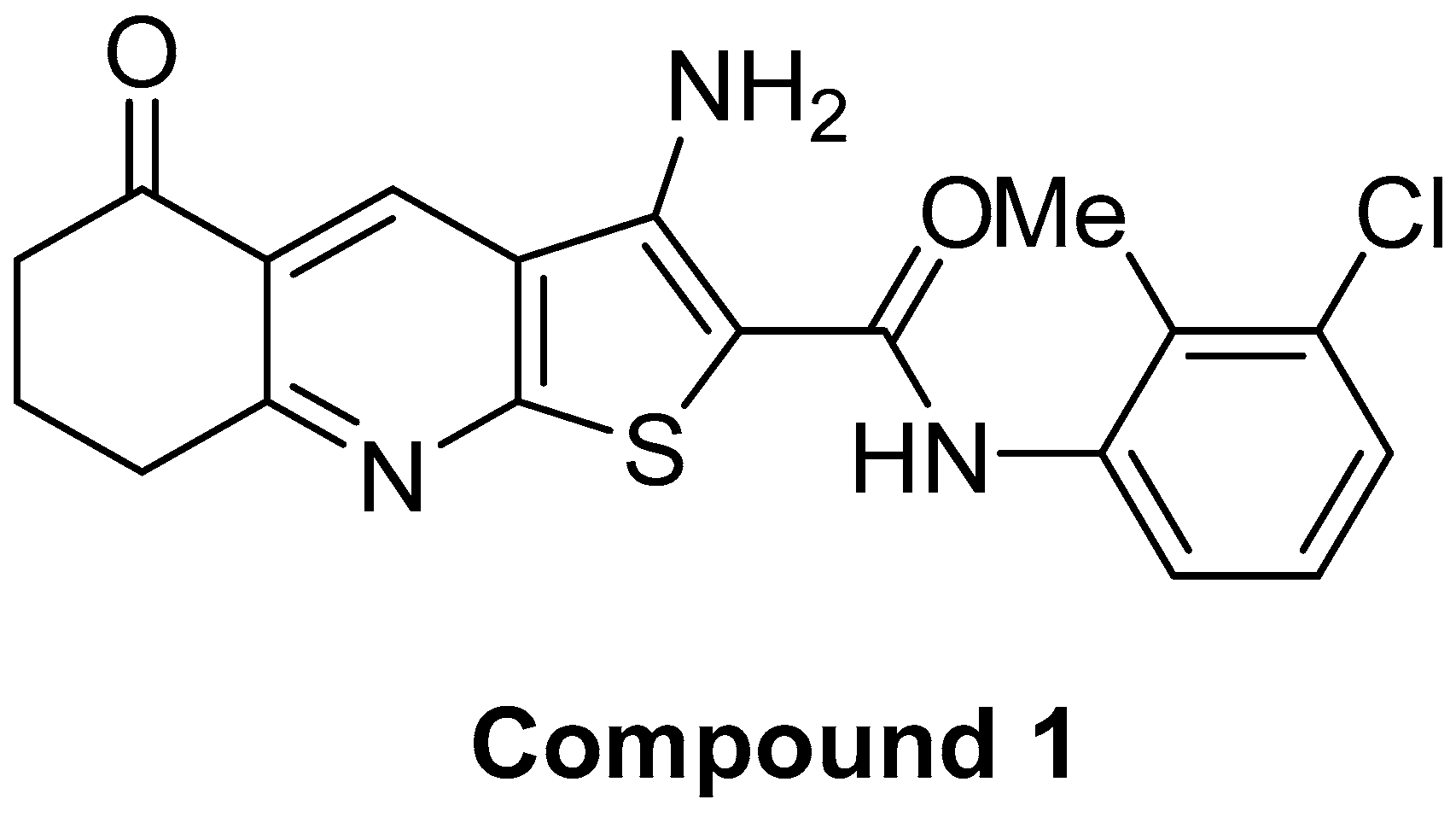
Thieno[2,3-b]Pyridine Derivative Targets Epithelial, Mesenchymal and Hybrid CD15s+ Breast Cancer Cells
Abstract
:1. Introduction
2. Results
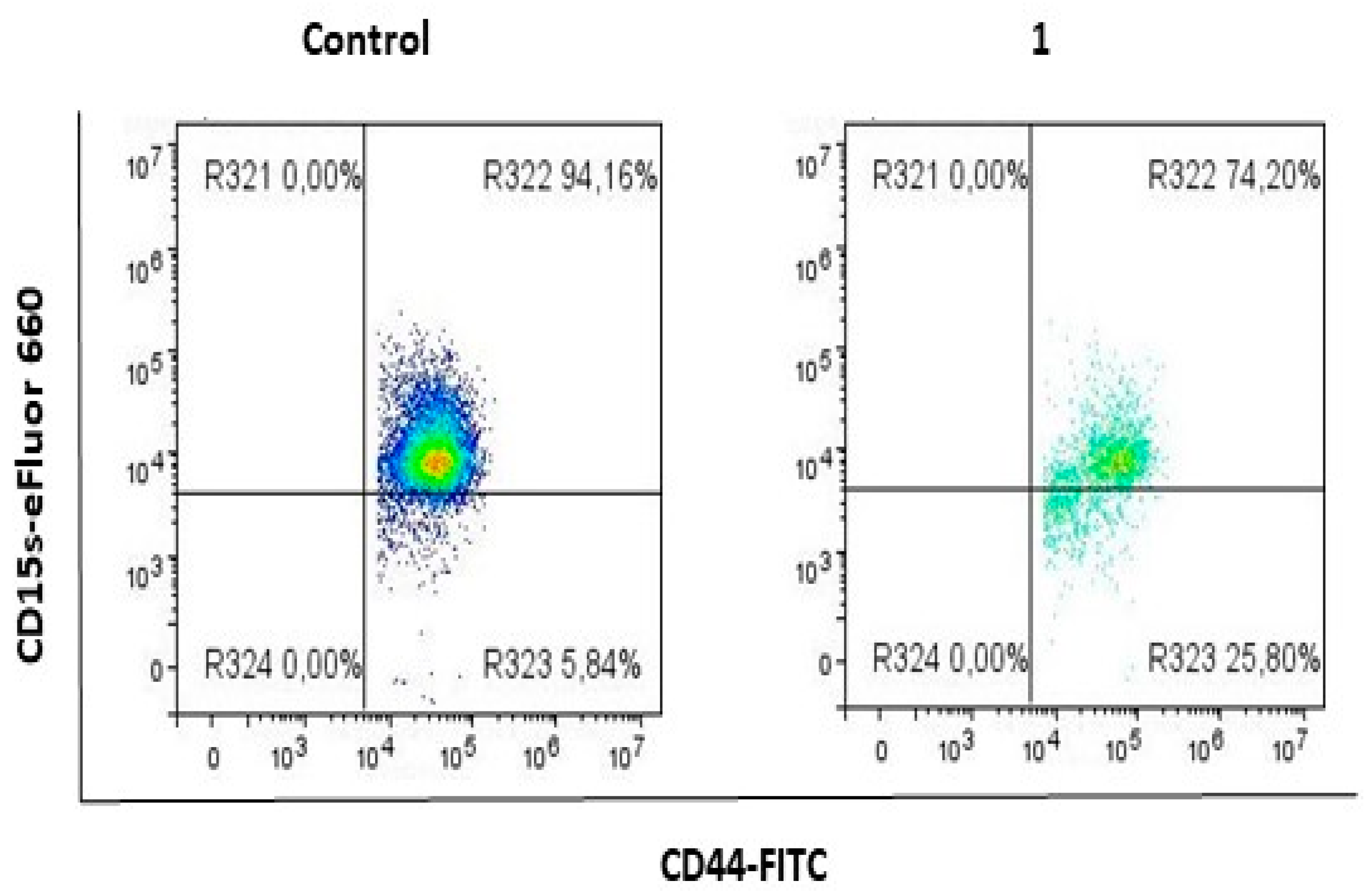
2.1. Compound 1 Decreased the Percentage and Number of Events of CD15s+ CSC
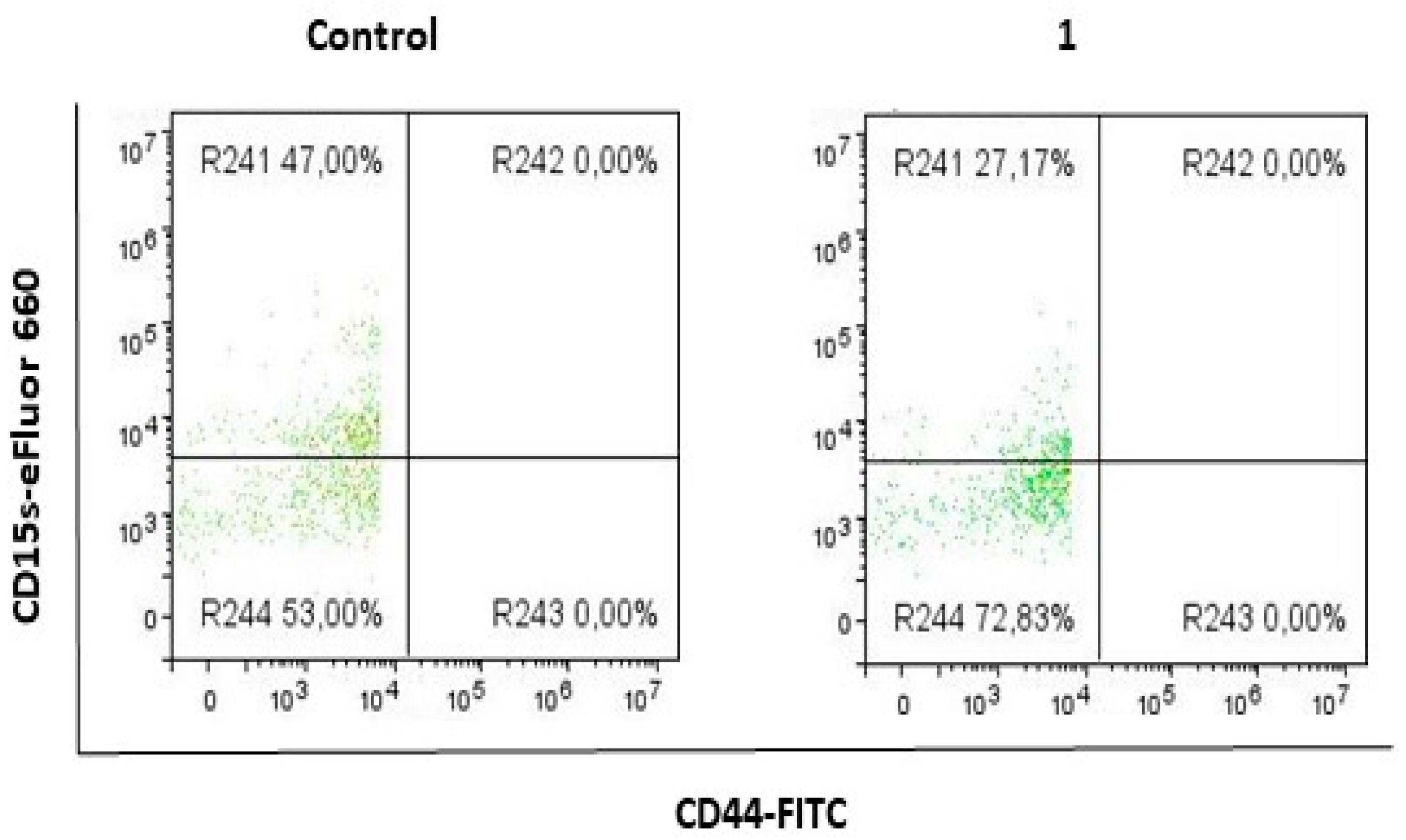
2.2. Compound 1 Decreased the Percentage and Number of Events of CD15s+CD44− Subpopulation
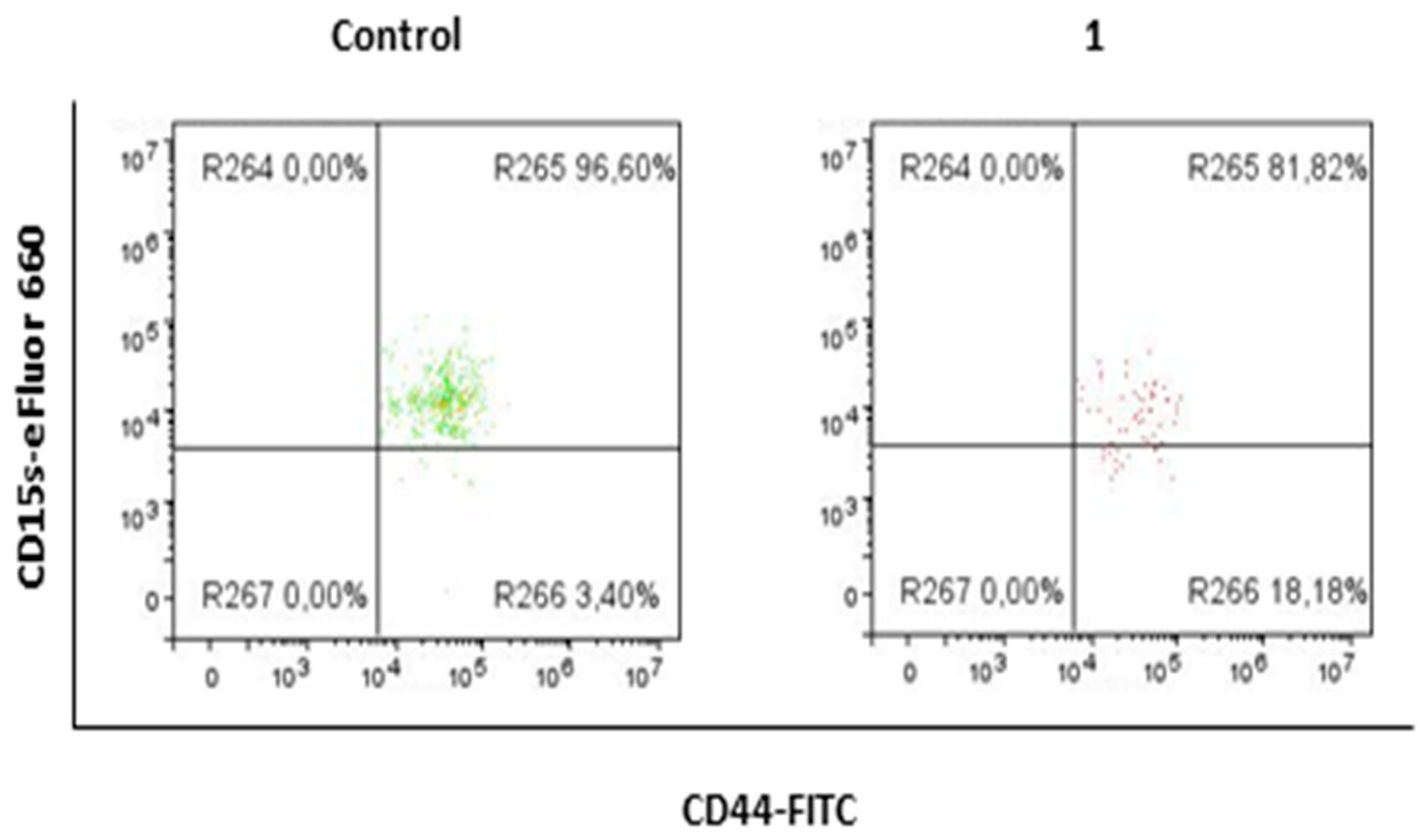
2.3. Compound 1 Decreased the Percentage and Number of Events of CD15s+CD44+CD24+ Subpopulation
2.4. Compound 1 Increased the Percentage of CD15− Cells in CSC, CD44− and CD44+CD24+ Subpopulations
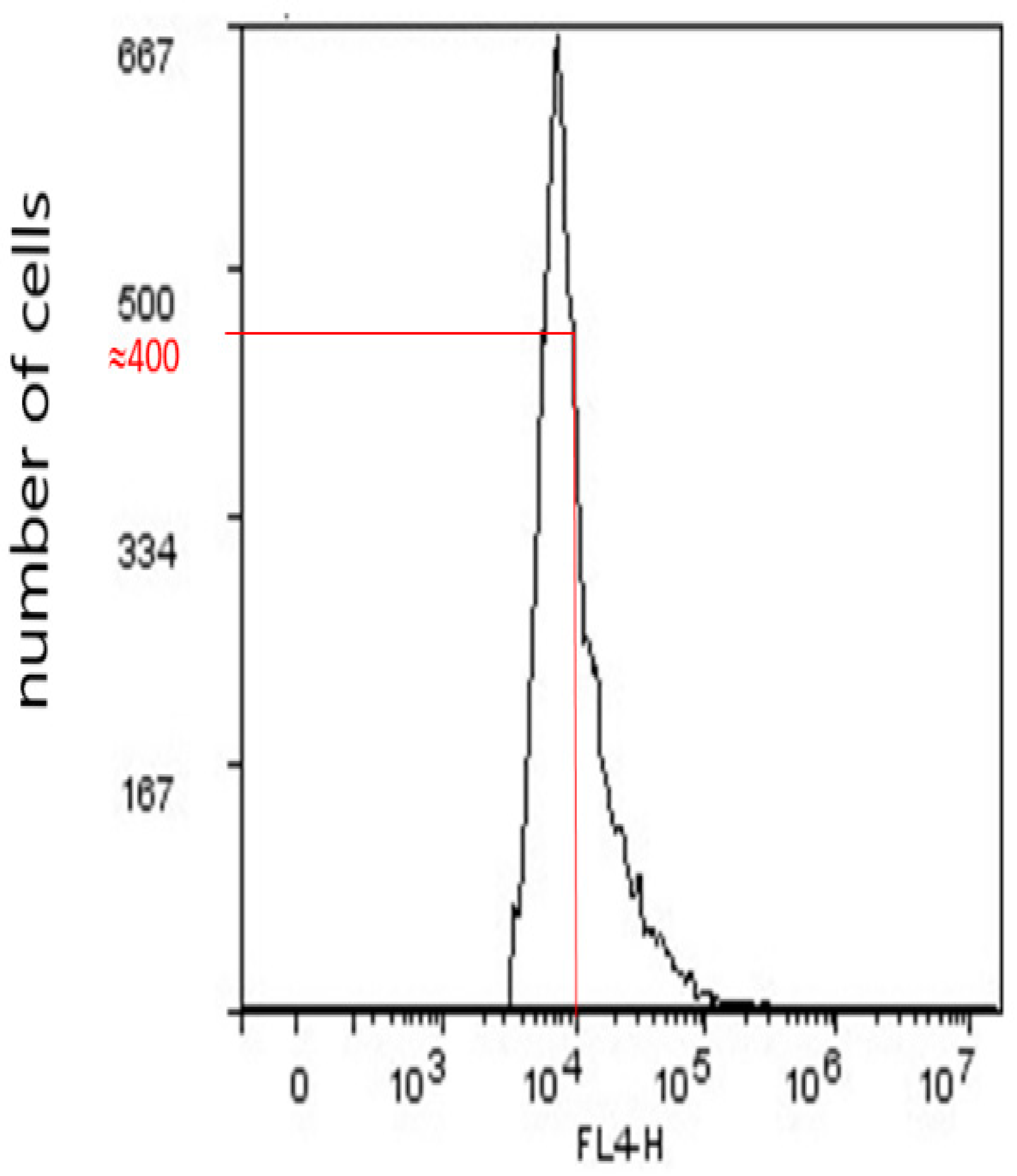
2.5. Geometric Mean Fluorescence Intensity (GMI)
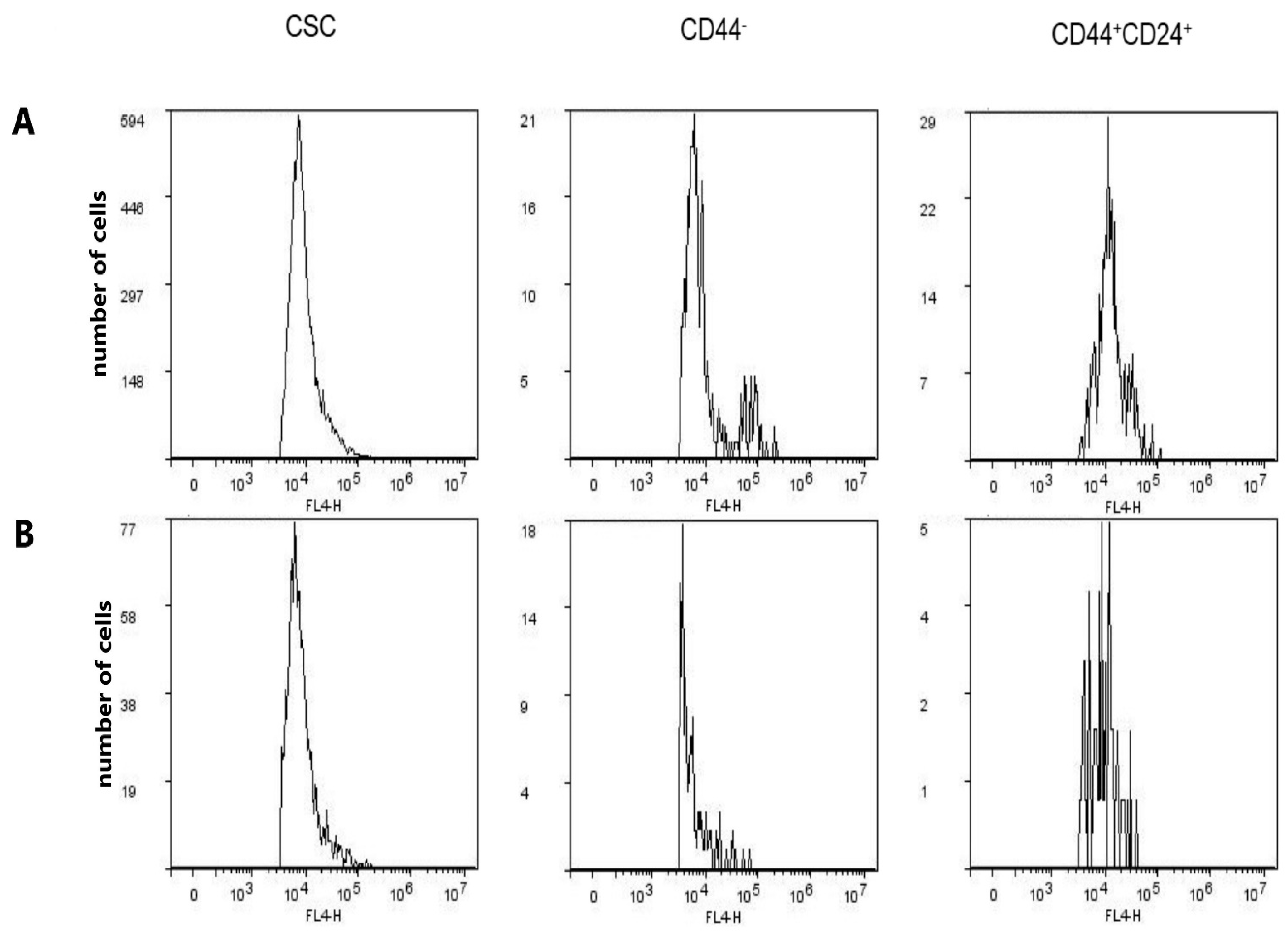
2.6. Expression of CD15s in CSC, CD44−, and CD44+CD24+ Subpopulations after Treatment with Compound 1
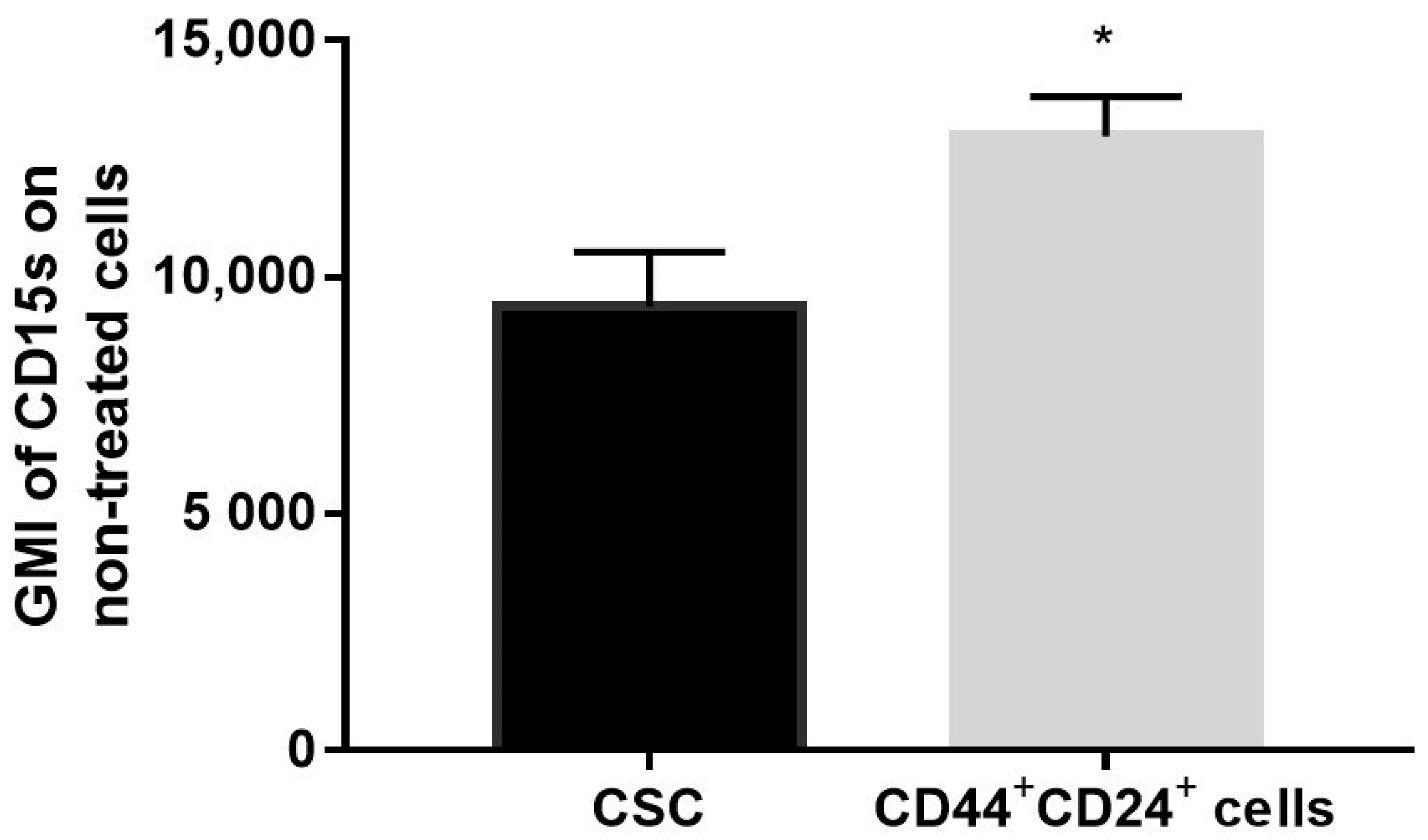
2.7. Expression of CD15s in Untreated CSC and CD44+CD24+ Cells
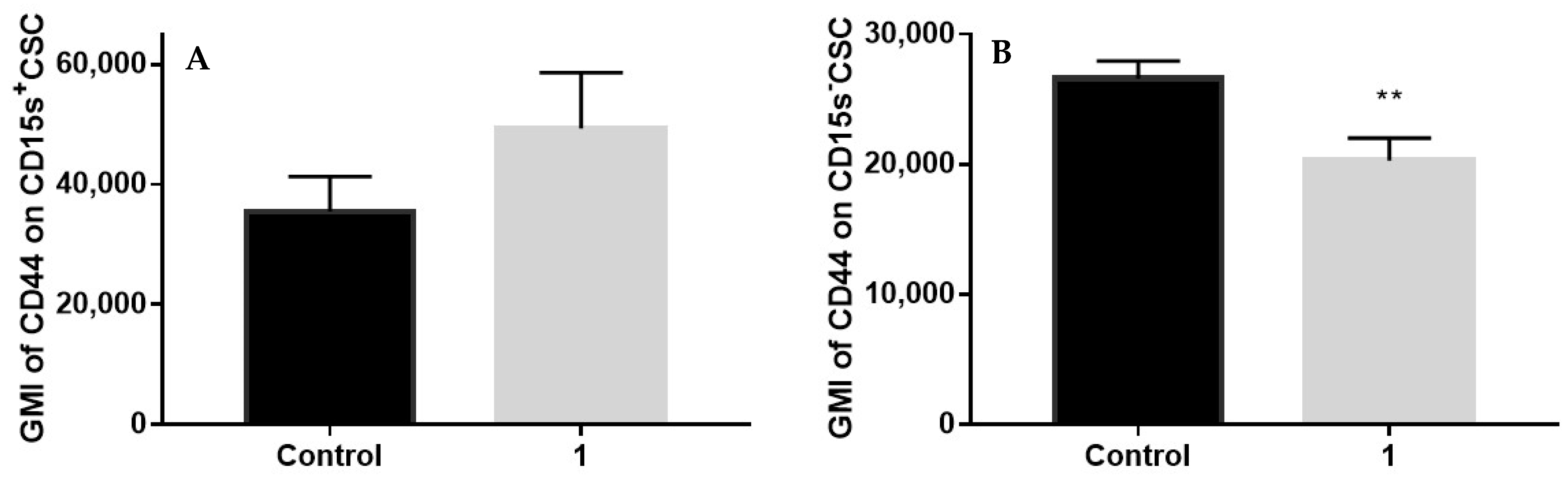
2.8. Expression of CD44 in CSC Positive and Negative for CD15s after Treatment with Compound 1
3. Discussion
4. Materials and Methods
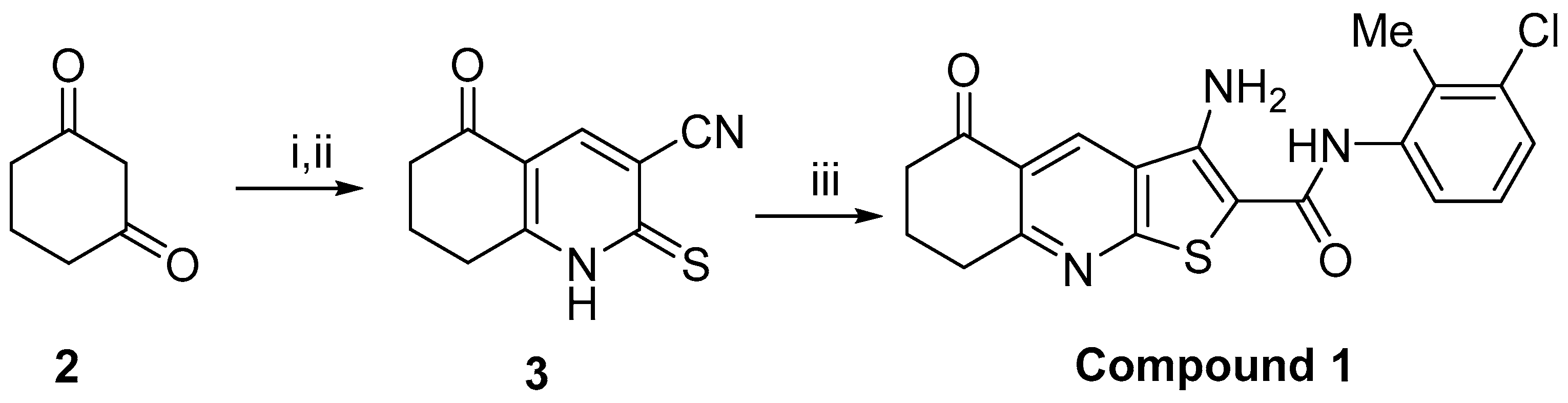
4.1. Chemistry and Cell Line
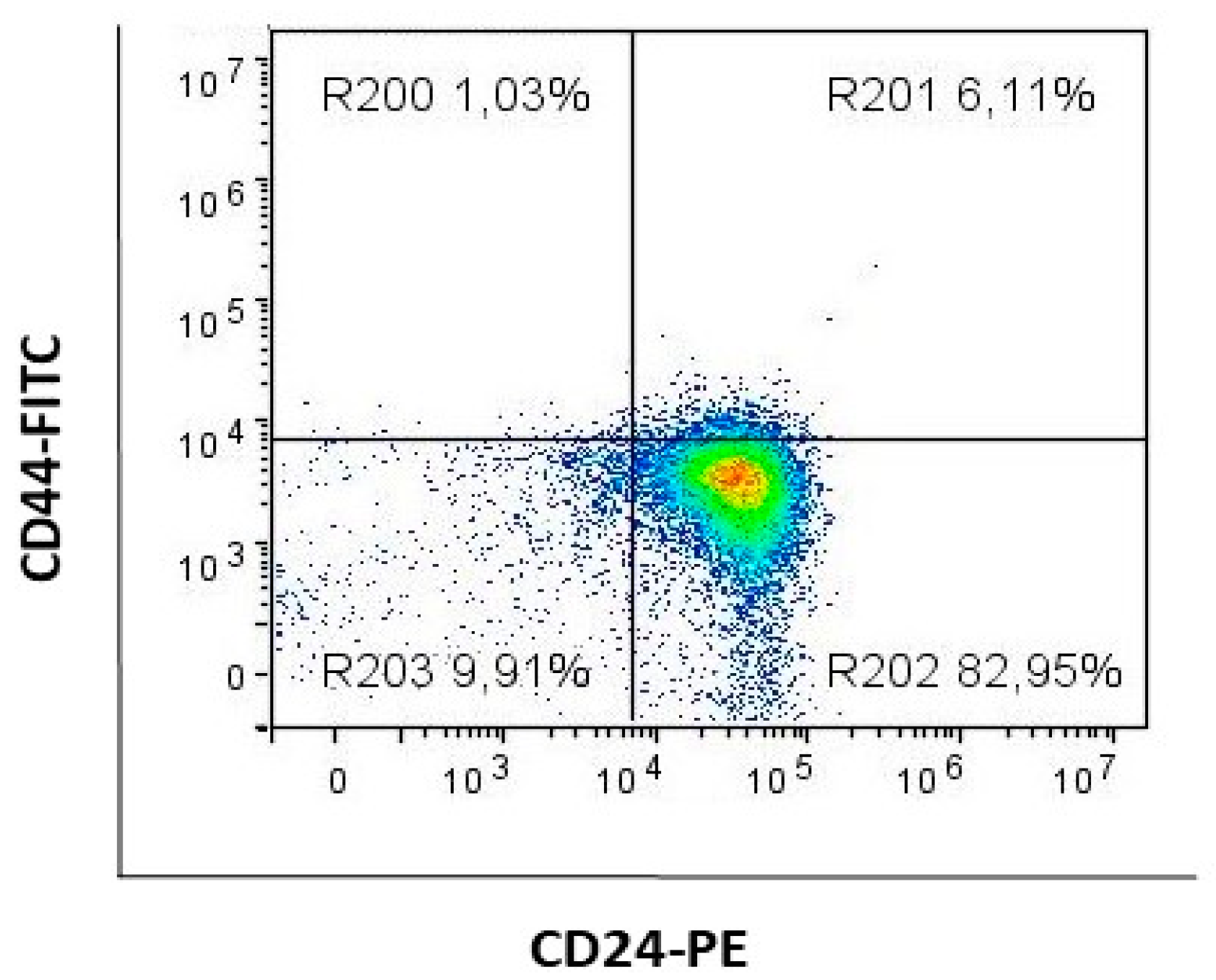
4.2. Flow Cytometric Analysis
4.3. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Konstantopoulos, K.; Thomas, S.N. Cancer cells in transit: The vascular interactions of tumor cells. Annu. Rev. Biomed. Eng. 2009, 11, 177–202. [Google Scholar] [CrossRef]
- Sheridan, C.; Kishimoto, H.; Fuchs, R.K.; Mehrotra, S.; Bhat-Nakshatri, P.; Turner, C.H.; Goulet, R., Jr.; Badve, S.; Nakshatri, H. CD44+/CD24− breast cancer cells exhibit enhanced invasive properties: An early step necessary for metastasis. Breast Cancer Res. BCR 2006, 8, R59. [Google Scholar] [CrossRef] [Green Version]
- Weiss, L.; Haydock, K.; Pickren, J.W.; Lane, W.W. Organ vascularity and metastatic frequency. Am. J. Pathol. 1980, 101, 101–113. [Google Scholar]
- Zhang, Z.; Wuhrer, M.; Holst, S. Serum sialylation changes in cancer. Glycoconj. J. 2018, 35, 139–160. [Google Scholar] [CrossRef] [Green Version]
- Shirure, V.S.; Henson, K.A.; Schnaar, R.L.; Nimrichter, L.; Burdick, M.M. Gangliosides expressed on breast cancer cells are E-selectin ligands. Biochem. Biophys. Res. Commun. 2011, 406, 423–429. [Google Scholar] [CrossRef] [PubMed]
- Zen, K.; Cui, L.B.; Zhang, C.Y.; Liu, Y. Critical role of mac-1 sialyl lewis x moieties in regulating neutrophil degranulation and transmigration. J. Mol. Biol. 2007, 374, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Zen, K.; Liu, D.Q.; Guo, Y.L.; Wang, C.; Shan, J.; Fang, M.; Zhang, C.Y.; Liu, Y. CD44v4 is a major E-selectin ligand that mediates breast cancer cell transendothelial migration. PLoS ONE 2008, 3, e1826. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pece, S.; Tosoni, D.; Confalonieri, S.; Mazzarol, G.; Vecchi, M.; Ronzoni, S.; Bernard, L.; Viale, G.; Pelicci, P.G.; Di Fiore, P.P. Biological and molecular heterogeneity of breast cancers correlates with their cancer stem cell content. Cell 2010, 140, 62–73. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Butti, R.; Gunasekaran, V.P.; Kumar, T.V.S.; Banerjee, P.; Kundu, G.C. Breast cancer stem cells: Biology and therapeutic implications. Int. J. Biochem. Cell Biol. 2019, 107, 38–52. [Google Scholar] [CrossRef]
- Moreira, M.P.; Brayner, F.A.; Alves, L.C.; Cassali, G.D.; Silva, L.M. Phenotypic, structural, and ultrastructural analysis of triple-negative breast cancer cell lines and breast cancer stem cell subpopulation. Eur. Biophys. J. EBJ 2019, 48, 673–684. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Lee, H.E.; Li, H.; Shipitsin, M.; Gelman, R.; Polyak, K. Heterogeneity for stem cell-related markers according to tumor subtype and histologic stage in breast cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2010, 16, 876–887. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grosse-Wilde, A.; Fouquier d’Herouel, A.; McIntosh, E.; Ertaylan, G.; Skupin, A.; Kuestner, R.E.; del Sol, A.; Walters, K.A.; Huang, S. Stemness of the hybrid Epithelial/Mesenchymal State in Breast Cancer and Its Association with Poor Survival. PLoS ONE 2015, 10, e0126522. [Google Scholar] [CrossRef] [PubMed]
- Sulaiman, A.; McGarry, S.; Han, X.; Liu, S.; Wang, L. CSCs in Breast Cancer-One Size Does Not Fit All: Therapeutic Advances in Targeting Heterogeneous Epithelial and Mesenchymal CSCs. Cancers 2019, 11, 1128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reynisson, J.; Jaiswal, J.K.; Barker, D.; D’mello, S.A.N.; Denny, W.A.; Baguley, B.C.; Leung, E.Y. Evidence that phospholipase C is involved in the antitumour action of NSC768313, a new thieno[2,3-b]pyridine derivative. Cancer Cell Int. 2016, 16, 18–27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zafar, A.; Sari, S.; Leung, E.; Pilkington, L.I.; van Rensburg, M.; Barker, D.; Reynisson, J. GPCR Modulation of Thieno[2,3-b]pyridine Anti-Proliferative Agents. Molecules 2017, 22, 2254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marijan, S.; Markotic, A.; Mastelic, A.; Rezic-Muzinic, N.; Pilkington, L.I.; Reynisson, J.; Culic, V.C. Glycosphingolipid expression at breast cancer stem cells after novel thieno[2,3-b]pyridine anticancer compound treatment. Sci. Rep. 2020, 10, 11876. [Google Scholar] [CrossRef]
- Yang, M.Y.; Wang, C.J.; Chen, N.F.; Ho, W.H.; Lu, F.J.; Tseng, T.H. Luteolin enhances paclitaxel-induced apoptosis in human breast cancer MDA-MB-231 cells by blocking STAT3. Chem. Biol. Interact. 2014, 213, 60–68. [Google Scholar] [CrossRef]
- Hung, J.M.; Arabshahi, H.J.; Leung, E.; Reynisson, J.; Barker, D. Synthesis and cytotoxicity of thieno[2,3-b]pyridine and furo[2,3-b]pyridine derivatives. Eur. J. Med. Chem. 2014, 86, 420–437. [Google Scholar] [CrossRef] [PubMed]
- Leung, E.; Pilkington, L.I.; van Rensburg, M.; Jeon, C.Y.; Song, M.; Arabshahi, H.J.; De Zoysa, G.H.; Sarojini, V.; Denny, W.A.; Reynisson, J.; et al. Synthesis and cytotoxicity of thieno[2,3-b]quinoline-2-carboxamide and cycloalkyl[b]thieno[3,2-e]pyridine-2-carboxamide derivatives. Bioorg. Med. Chem. 2016, 24, 1142–1154. [Google Scholar] [CrossRef]
- Van Rensburg, M.; Leung, E.; Haverkate, N.A.; Eurtivong, C.; Pilkington, L.I.; Reynisson, J.; Barker, D. Synthesis and antiproliferative activity of 2-chlorophenyl carboxamide thienopyridines. Bioorg. Med. Chem. Lett. 2017, 27, 135–138. [Google Scholar] [CrossRef]
- Leung, E.; Hung, J.M.; Barker, D.; Reynisson, J. The effect of a thieno[2,3-b]pyridine PLC-γ inhibitor on the proliferation, morphology, migration and cell cycle of breast cancer cells. MedChemComm 2014, 5, 99–106. [Google Scholar] [CrossRef]
- Dimitroff, C.J.; Lee, J.Y.; Rafii, S.; Fuhlbrigge, R.C.; Sackstein, R. CD44 is a major E-selectin ligand on human hematopoietic progenitor cells. J. Cell Biol. 2001, 153, 1277–1286. [Google Scholar] [CrossRef] [PubMed]
- Hanley, W.D.; Burdick, M.M.; Konstantopoulos, K.; Sackstein, R. CD44 on LS174T colon carcinoma cells possesses E-selectin ligand activity. Cancer Res. 2005, 65, 5812–5817. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takeo, K.; Kawai, T.; Nishida, K.; Masuda, K.; Teshima-Kondo, S.; Tanahashi, T.; Rokutan, K. Oxidative stress-induced alternative splicing of transformer 2beta (SFRS10) and CD44 pre-mRNAs in gastric epithelial cells. Am. J. Physiol. Cell Physiol. 2009, 297, C330–C338. [Google Scholar] [CrossRef]
- Shirure, V.S.; Liu, T.; Delgadillo, L.F.; Cuckler, C.M.; Tees, D.F.; Benencia, F.; Goetz, D.J.; Burdick, M.M. CD44 variant isoforms expressed by breast cancer cells are functional E-selectin ligands under flow conditions. Am. J. Physiol. Cell Physiol. 2015, 308, C68–C78. [Google Scholar] [CrossRef]
- Mani, S.A.; Guo, W.; Liao, M.J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Clouthier, S.G.; Wicha, M.S. Role of microRNAs in the regulation of breast cancer stem cells. J. Mammary Gland Biol. Neoplasia 2012, 17, 15–21. [Google Scholar] [CrossRef] [Green Version]
- Kalluri, R.; Weinberg, R.A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 2009, 119, 1420–1428. [Google Scholar] [CrossRef] [Green Version]
- Wu, H.L.; Duan, Z.T.; Jiang, Z.D.; Cao, W.J.; Wang, Z.B.; Hu, K.W.; Gao, X.; Wang, S.K.; He, B.S.; Zhang, Z.Y. Increased endoplasmic reticulum stress response is involved in clopidogrel-induced apoptosis of gastric epithelial cells. PLoS ONE 2013, 8, e74381. [Google Scholar] [CrossRef]
- Nami, B.; Donmez, H.; Kocak, N. Tunicamycin-induced endoplasmic reticulum stress reduces in vitro subpopulation and invasion of CD44+/CD24− phenotype breast cancer stem cells. Exp. Toxicol. Pathol. Off. J Ges. Toxikol. Pathol. 2016, 68, 419–426. [Google Scholar] [CrossRef]
- Binsaleh, N.K.; Wigley, C.A.; Whitehead, K.A.; van Rensburg, M.; Reynisson, J.; Pilkington, L.I.; Barker, D.; Jones, S.; Dempsey-Hibbert, N.C. Thieno[2,3-b]pyridine derivatives are potent anti-platelet drugs, inhibiting platelet activation, aggregation and showing synergy with aspirin. Eur. J. Med. Chem. 2018, 143, 1997–2004. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arabshahi, H.J.; van Rensburg, M.; Pilkington, L.I.; Jeon, C.Y.; Song, M.; Gridel, L.-M.; Leung, E.; Barker, D.; Vuica-Ross, M.; Volcho, K.P. A synthesis, In Silico, In Vitro and In Vivo study of thieno[2,3-b]pyridine anticancer analogues. MedChemComm 2015, 6, 1987–1997. [Google Scholar] [CrossRef]
- Eurtivong, C.; Semenov, V.; Semenova, M.; Konyushkin, L.; Atamanenko, O.; Reynisson, J.; Kiselyov, A. 3-Amino-thieno[2,3-b]pyridines as microtubule-destabilising agents: Molecular modelling and biological evaluation in the sea urchin embryo and human cancer cells. Bioorg. Med. Chem. 2017, 25, 658–664. [Google Scholar] [CrossRef] [PubMed]
- Naguib, B.H.; El-Nassan, H.B. Synthesis of new thieno[2,3-b]pyridine derivatives as pim-1 inhibitors. J. Enzym. Inhib. Med. Chem. 2016, 31, 1718–1725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lockman, J.W.; Reeder, M.D.; Suzuki, K.; Ostanin, K.; Hoff, R.; Bhoite, L.; Austin, H.; Baichwal, V.; Adam Willardsen, J. Inhibition of eEF2-K by thieno[2,3-b]pyridine analogues. Bioorg. Med. Chem. Lett. 2010, 20, 2283–2286. [Google Scholar] [CrossRef] [PubMed]
- Sanad, S.M.H.; Mekky, A.E.M. Novel Nicotinonitriles and Thieno[2,3-b]pyridines as Potent Biofilm and COX-2 Inhibitors: Synthesis, In Vitro and In Silico Studies. ChemistrySelect 2020, 5, 8494–8503. [Google Scholar] [CrossRef]
- Dallas, M.R.; Liu, G.; Chen, W.C.; Thomas, S.N.; Wirtz, D.; Huso, D.L.; Konstantopoulos, K. Divergent roles of CD44 and carcinoembryonic antigen in colon cancer metastasis. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2012, 26, 2648–2656. [Google Scholar] [CrossRef] [Green Version]
- Brown, R.L.; Reinke, L.M.; Damerow, M.S.; Perez, D.; Chodosh, L.A.; Yang, J.; Cheng, C. CD44 splice isoform switching in human and mouse epithelium is essential for epithelial-mesenchymal transition and breast cancer progression. J. Clin. Investig. 2011, 121, 1064–1074. [Google Scholar] [CrossRef] [Green Version]
- Vikram, R.; Chou, W.C.; Hung, S.C.; Shen, C.Y. Tumorigenic and Metastatic Role of CD44(-/low)/CD24(-/low) Cells in Luminal Breast Cancer. Cancers 2020, 12, 1239. [Google Scholar] [CrossRef]
- Bocci, F.; Gearhart-Serna, L.; Boareto, M.; Ribeiro, M.; Ben-Jacob, E.; Devi, G.R.; Levine, H.; Onuchic, J.N.; Jolly, M.K. Toward understanding cancer stem cell heterogeneity in the tumor microenvironment. Proc. Natl. Acad. Sci. USA 2019, 116, 148–157. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Cong, Y.; Wang, D.; Sun, Y.; Deng, L.; Liu, Y.; Martin-Trevino, R.; Shang, L.; McDermott, S.P.; Landis, M.D.; et al. Breast cancer stem cells transition between epithelial and mesenchymal states reflective of their normal counterparts. Stem Cell Rep. 2014, 2, 78–91. [Google Scholar] [CrossRef] [PubMed]
- Kamal El-Dean, A.M.; Elkhawaga, A.M.; Radwan, S.M.; Ahmed, M.M. Synthesis of Some Pyridothienopyrazolopyrimidopyrimidine and Mercaptomethylpyrazolopyrimidine Derivatives. Phosphorus Sulfur Silicon Relat. Elem. 2009, 184, 2034–2048. [Google Scholar] [CrossRef]
| CSC | |||
|---|---|---|---|
| Control | 1 | p-Value | |
| % CD15s+ | 92.83 ± 3.21 | 74.15 ± 2.21 | 0.0012 |
| Events CD15s+ | 11,585 ± 977 | 1263 ± 107 | 0.00005 |
| CD44− | |||
|---|---|---|---|
| Control | 1 | p-Value | |
| % CD15s+ | 49.03 ± 5.74 | 27.06 ± 5.75 | 0.0094 |
| Events CD15s+ | 429 ± 70 | 153 ± 20 | 0.0028 |
| CD44+CD24+ | |||
|---|---|---|---|
| Control | 1 | p-Value | |
| % CD15s+ | 93.15 ± 3.64 | 82.77 ± 0.82 | 0.0052 |
| Events CD15s+ | 370 ± 176 | 69 ± 8 | 0.0943 |
| CSC | CD44− | CD44+CD24+ | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Control | 1 | p-Value | Control | 1 | p-Value | Control | 1 | p-Value | |
| % CD15s− | 7.17 ± 3.21 | 25.85 ± 2.21 | 0.0019 | 50.97 ± 5.74 | 72.94 ± 5.75 | 0.0094 | 4.36 ± 2.66 | 16.4 ± 1.56 | 0.0052 |
| Events CD15s− | 668 ± 33.94 | 443 ± 75.44 | 0.1758 | 444.67 ± 50.29 | 426.33 ± 105.67 | 0.8045 | 14 ± 2 | 13.67 ± 2.51 | 0.8666 |
| CSC | CD44− | CD44+CD24+ | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Control | 1 | p-Value | Control | 1 | p-Value | Control | 1 | p-Value | |
| GMI of CD15s | 9396 ± 1138 | 9212 ± 1006 | 0.844 | 9220 ± 899 | 6090 ± 191 | 0.0041 | 12985 ± 835 | 9469 ± 592 | 0.004 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marijan, S.; Mastelić, A.; Markotić, A.; Režić-Mužinić, N.; Vučenović, N.; Barker, D.; Pilkington, L.I.; Reynisson, J.; Čulić, V.Č. Thieno[2,3-b]Pyridine Derivative Targets Epithelial, Mesenchymal and Hybrid CD15s+ Breast Cancer Cells. Medicines 2021, 8, 32. https://doi.org/10.3390/medicines8070032
Marijan S, Mastelić A, Markotić A, Režić-Mužinić N, Vučenović N, Barker D, Pilkington LI, Reynisson J, Čulić VČ. Thieno[2,3-b]Pyridine Derivative Targets Epithelial, Mesenchymal and Hybrid CD15s+ Breast Cancer Cells. Medicines. 2021; 8(7):32. https://doi.org/10.3390/medicines8070032
Chicago/Turabian StyleMarijan, Sandra, Angela Mastelić, Anita Markotić, Nikolina Režić-Mužinić, Nikolina Vučenović, David Barker, Lisa I. Pilkington, Jóhannes Reynisson, and Vedrana Čikeš Čulić. 2021. "Thieno[2,3-b]Pyridine Derivative Targets Epithelial, Mesenchymal and Hybrid CD15s+ Breast Cancer Cells" Medicines 8, no. 7: 32. https://doi.org/10.3390/medicines8070032
APA StyleMarijan, S., Mastelić, A., Markotić, A., Režić-Mužinić, N., Vučenović, N., Barker, D., Pilkington, L. I., Reynisson, J., & Čulić, V. Č. (2021). Thieno[2,3-b]Pyridine Derivative Targets Epithelial, Mesenchymal and Hybrid CD15s+ Breast Cancer Cells. Medicines, 8(7), 32. https://doi.org/10.3390/medicines8070032









