Specific Avenin Cross-Reactivity with G12 Antibody in a Wide Range of Current Oat Cultivars
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Field Trials
2.2. Chemical Analyses
2.2.1. Determination of Crude Protein Content
2.2.2. Immunoassay Tests
2.2.3. Isolation of Pure Avenins
2.2.4. Avenin Profiling by SDS-PAGE
2.2.5. Pure Avenin Quantification by RP-HPLC
2.2.6. Immunoblotting
2.3. Data Analyses
3. Results and Discussion
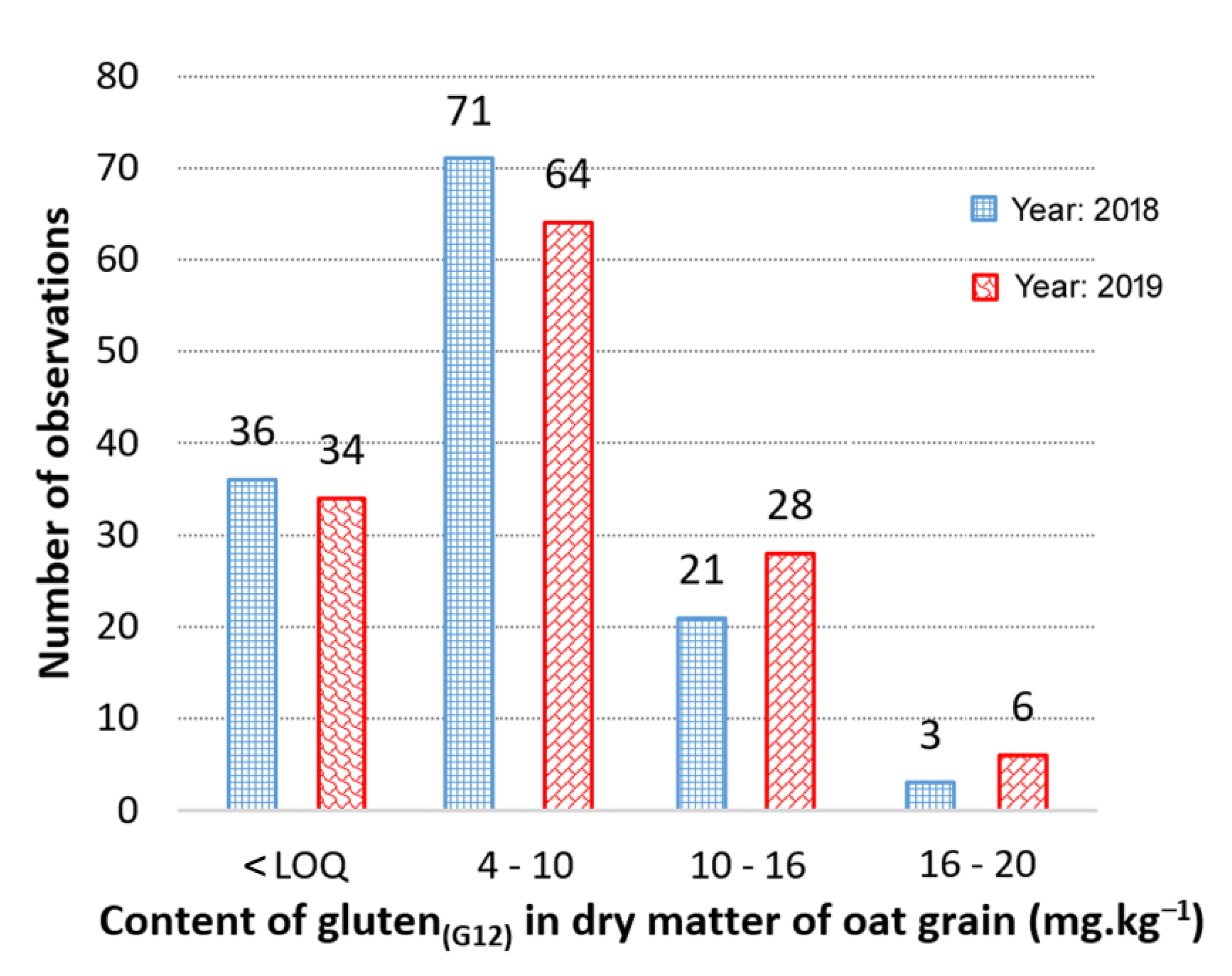
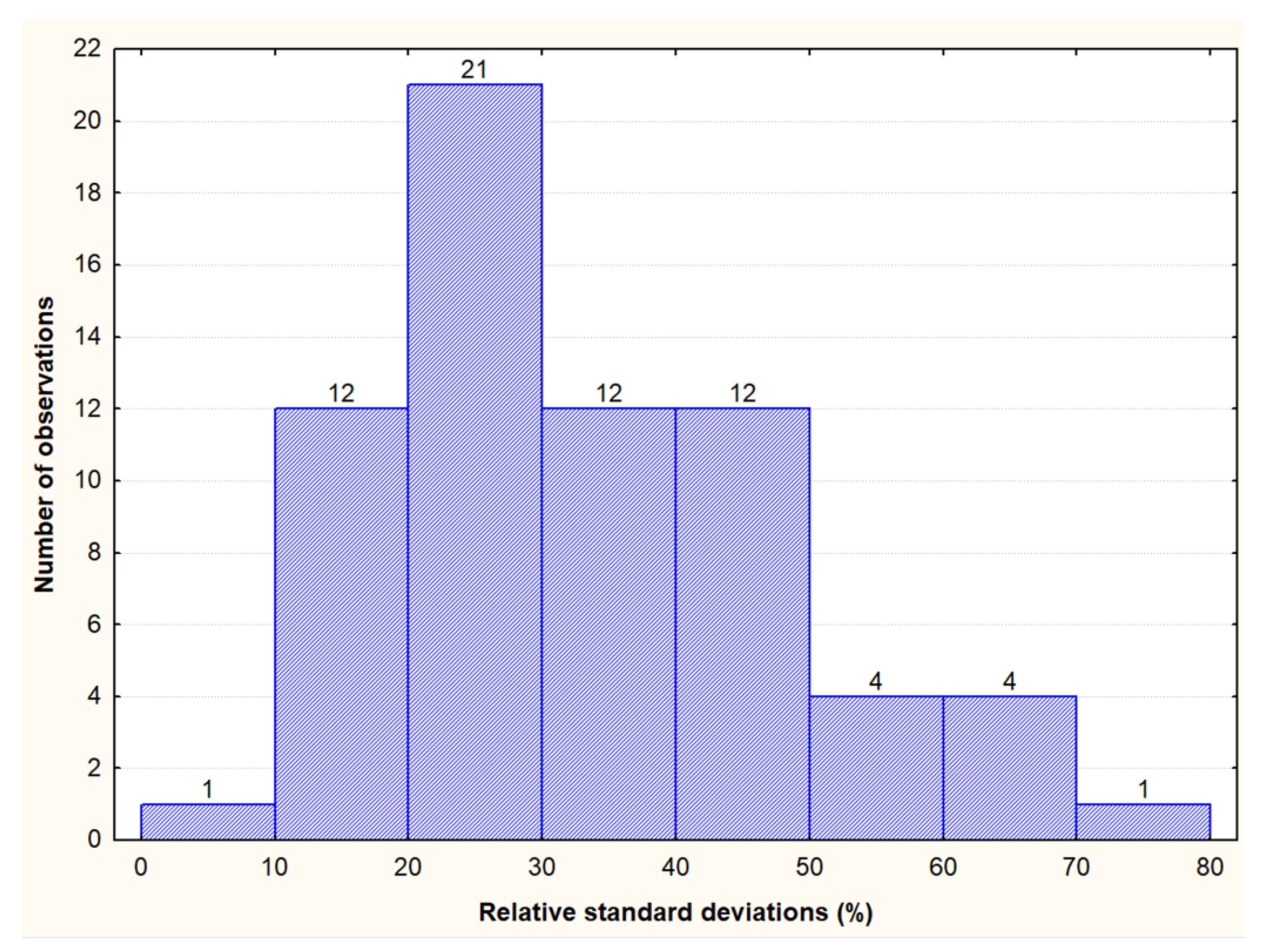
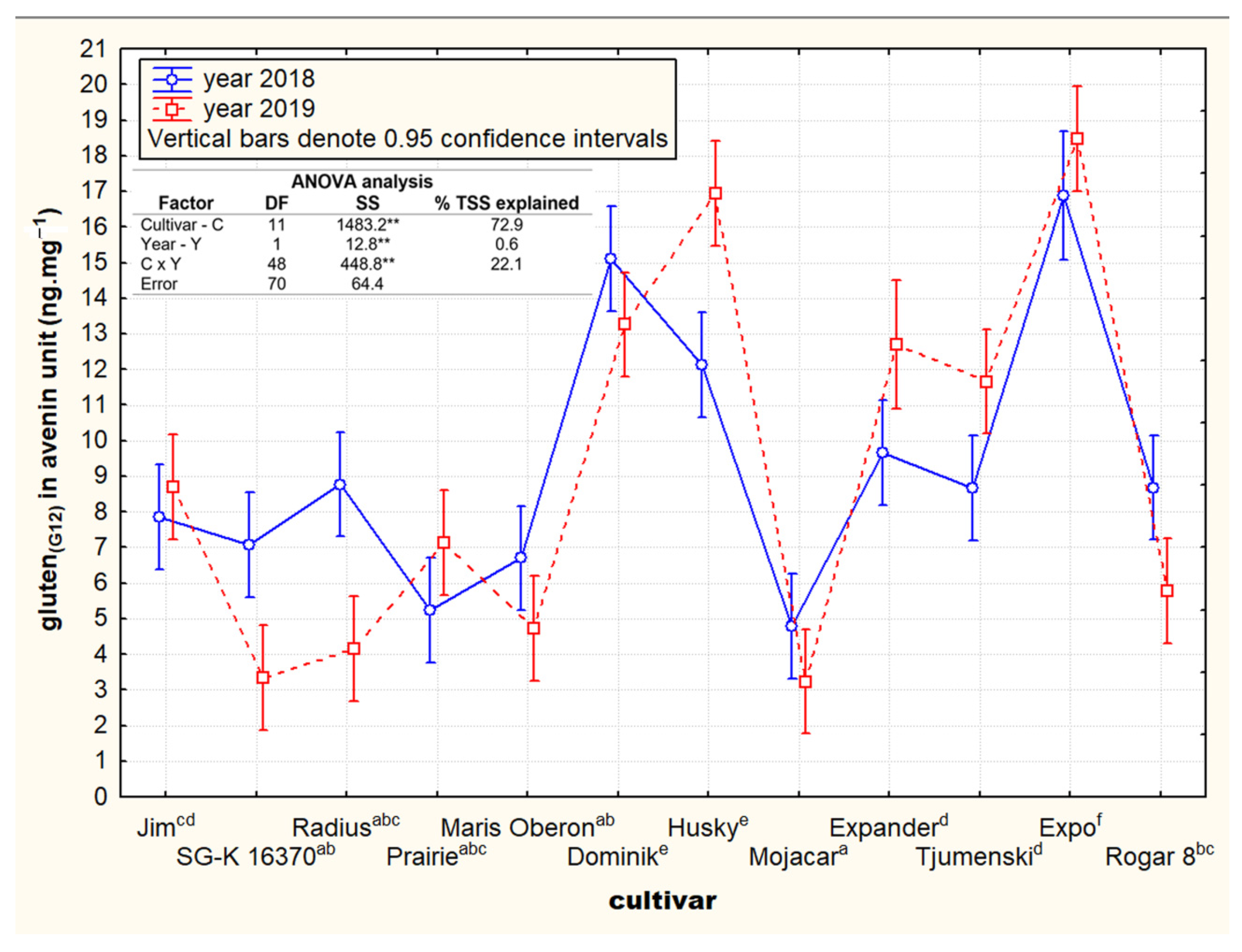
3.1. Contents and Variability of Cross-Reactive Gluten Peptides in the Set of Oat Cultivars
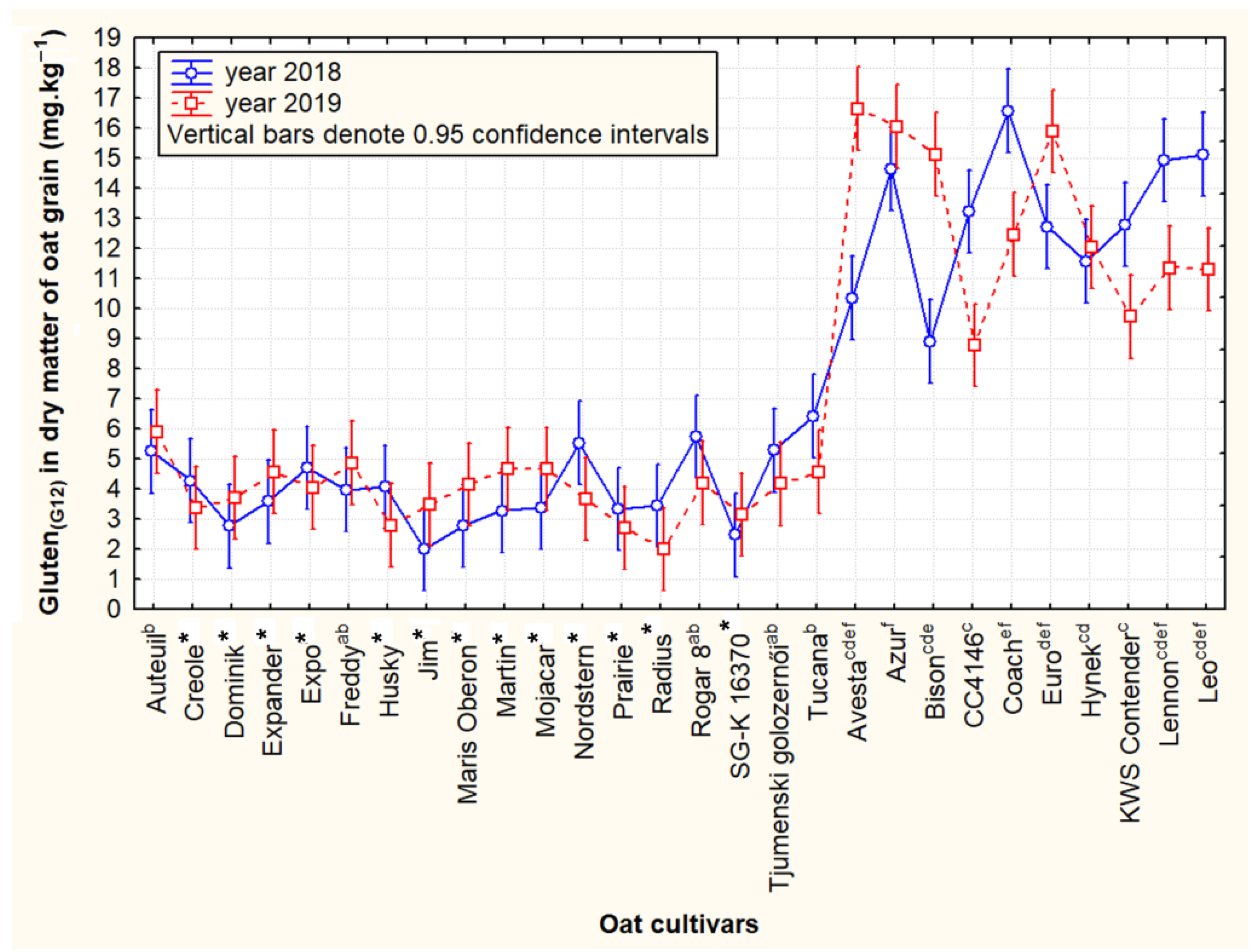
3.2. Contrasting Groups of Oat Cultivars in the Content of Cross-Reactive Avenin Peptides
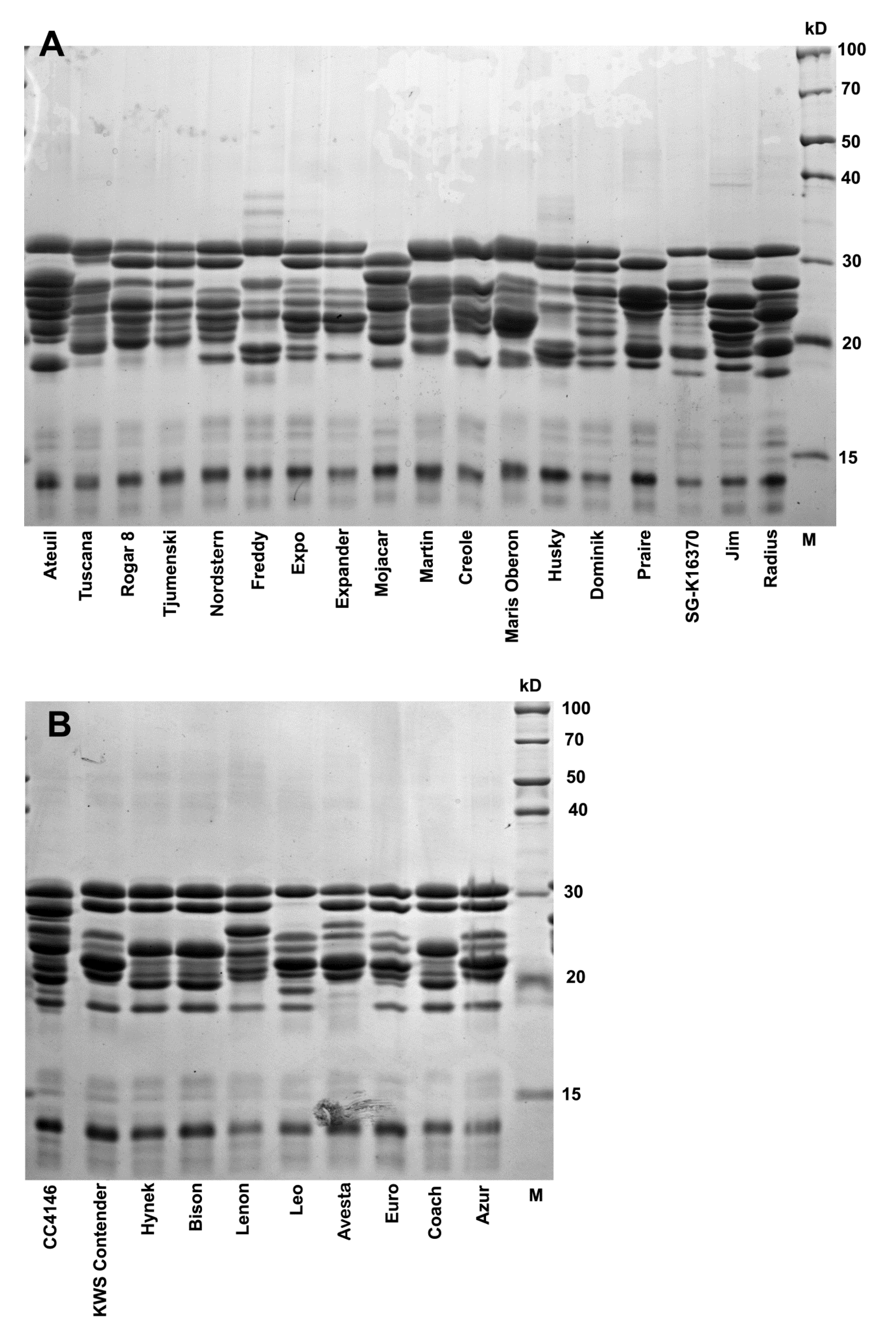
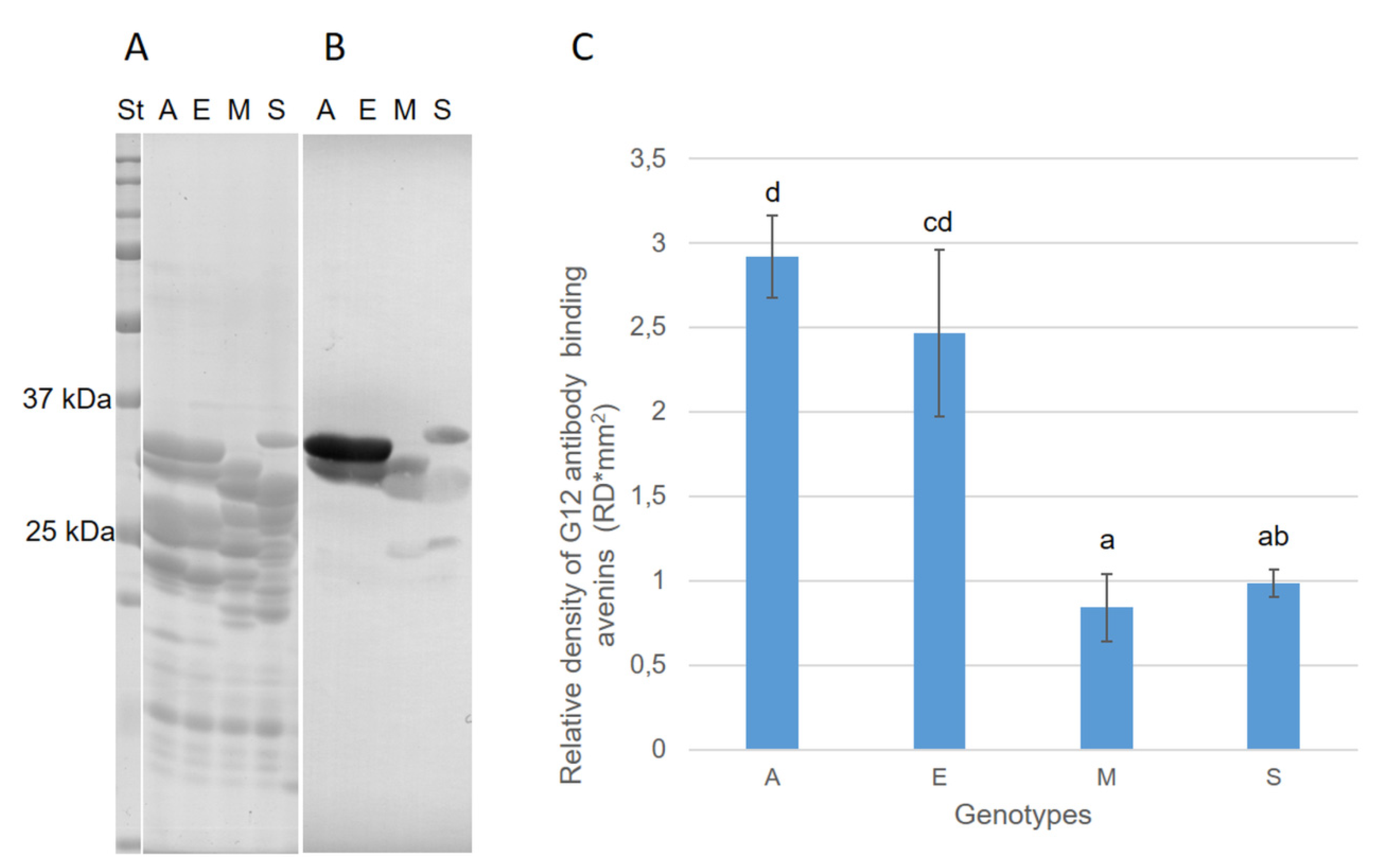
3.3. Avenin Protein Polymorphism and Immunoblot Analysis in Groups of Oat Cultivars with Contrasting Cross-Reactivity
3.4. Interference between Highly Concentrated Avenins of Low-Reactive Cultivars and the G12 Antibody
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gorash, A.; Armonienė, R.; Fetch, J.M.; Liatukas, Ž.; Danytė, V. Aspects in oat breeding: Nutrition quality, nakedness and disease resistance, challenges and perspectives. Ann. Appl. Biol. 2017, 171, 281–302. [Google Scholar] [CrossRef]
- Ahmad, A.; Anjum, F.M.; Zahoor, T.; Nawaz, H.; Ahmed, Z. Extraction and characterization of β-d-glucan from oat for industrial utilization. Int. J. Biol. Macromol. 2010, 46, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Jágr, M.; Dvořáček, V.; Čepková, P.H.; Doležalová, J. Comprehensive analysis of oat avenanthramides using hybrid quadrupole-Orbitrap mass spectrometry: Possible detection of new compounds. Rapid Commun. Mass Spectrom. 2020, 34, e8718. [Google Scholar] [CrossRef] [PubMed]
- Fritz, R.D.; Chen, Y. Oat safety for celiac disease patients: Theoretical analysis correlates adverse symptoms in clinical studies to contaminated study oats. Nutr. Res. 2018, 60, 54–67. [Google Scholar] [CrossRef]
- Pinto-Sánchez, M.I.; Causada-Calo, N.; Bercik, P.; Ford, A.; Murray, J.A.; Armstrong, D.; Semrad, C.; Kupfer, S.S.; Alaedini, A.; Moayyedi, P.; et al. Safety of Adding Oats to a Gluten-Free Diet for Patients with Celiac Disease: Systematic Review and Meta-analysis of Clinical and Observational Studies. Gastroenterology 2017, 153, 395–409.e3. [Google Scholar] [CrossRef] [Green Version]
- Barcchiya, J.; Meena, R.K.; Lal, N. Oat is a multifunctional cereal crop. Innov. Farming 2017, 2, 114–116. [Google Scholar]
- Gilissen, L.J.; Van Der Meer, I.M.; Smulders, M.J.M. Why Oats Are Safe and Healthy for Celiac Disease Patients. Med. Sci. 2016, 4, 21. [Google Scholar] [CrossRef]
- Real, A.; Comino, I.; de Lorenzo, L.; Merchán, F.; Gil-Humanes, J.; Giménez, M.J.; Barro, F. Molecular and immunological characterization of gluten proteins isolated from oat cultivars that differ in toxicity for celiac disease. PLoS ONE 2012, 7, e48365. [Google Scholar] [CrossRef] [Green Version]
- Comino Montilla, I.M.; Lorenzo Barrios, L.D.; Cornell, H.; López Casado, M.Á.; Barro, F.; Sousa Martín, C. Diversity in oat potential immunogenicity: Basis for the selection of oat varieties with no toxicity in coeliac disease. Gut 2011, 60, 915–922. [Google Scholar] [CrossRef] [Green Version]
- Ballabio, C.; Uberti, F.; Manferdelli, S.; Vacca, E.; Boggini, G.; Redaelli, R.; Restani, P. Molecular characterisation of 36 oat varieties and in vitro assessment of their suitability for coeliacs’ diet. J. Cereal Sci. 2011, 54, 110–115. [Google Scholar] [CrossRef]
- Londono, D.; Westende, W.V.; Goryunova, S.; Salentijn, E.; Broeck, H.V.D.; van der Meer, I.; Visser, R.; Gilissen, L.; Smulders, M. Avenin diversity analysis of the genus Avena (oat). Relevance for people with celiac disease. J. Cereal Sci. 2013, 58, 170–177. [Google Scholar] [CrossRef]
- Lexhaller, B.; Tompos, C.; Scherf, K.A. Comparative analysis of prolamin and glutelin fractions from wheat, rye, and barley with five sandwich ELISA test kits. Anal. Bioanal. Chem. 2016, 408, 6093–6104. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Kanerva, P.; Salovaara, H.; Stoddard, F.L.; Sontag-Strohm, T. Proposal for C-Hordein as Reference Material in Gluten Quantification. J. Agric. Food Chem. 2017, 65, 2155–2161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanner, G.J.; Blundell, M.J.; Colgrave, M.L.; Howitt, C.A. Quantification of Hordeins by ELISA: The Correct Standard Makes a Magnitude of Difference. PLoS ONE 2013, 8, e56456. [Google Scholar] [CrossRef] [PubMed]
- Wehling, P.; Scherf, K.A. Preparation of validation materials for estimating gluten recovery by ELISA according to SMPR 2017.021. J. AOAC Int. 2020, 103, 210–215. [Google Scholar] [CrossRef]
- Sollid, L.M.; Qiao, S.-W.; Anderson, R.P.; Gianfrani, C.; Koning, F. Nomenclature and listing of celiac disease relevant gluten T-cell epitopes restricted by HLA-DQ molecules. Immunogenetics 2012, 64, 455–460. [Google Scholar] [CrossRef] [Green Version]
- Kosová, K.; Leišová-Svobodová, L.; Dvořáček, V. Oats as a Safe Alternative to Triticeae Cereals for People Suffering from Celiac Disease? A Review. Plant Foods Hum. Nutr. 2020, 75, 131–141. [Google Scholar] [CrossRef]
- Mujico, J.R.; Mitea, C.; Gilissen, L.J.; de Ru, A.; van Veelen, P.; Smulders, M.J.; Koning, F. Natural variation in avenin epitopes among oat varieties: Implications for celiac disease. J. Cereal Sci. 2011, 54, 8–12. [Google Scholar] [CrossRef]
- Ahola, H.G.; Sontag-Strohm, T.S.; Schulman, A.H.; Tanhuanpää, P.; Viitala, S.; Huang, X. Immunochemical analysis of oat avenins in an oat cultivar and landrace collection. J. Cereal Sci. 2020, 95, 103053. [Google Scholar] [CrossRef]
- Halbmayr-Jech, E.; Hammer, E.; Fielder, R.; Coutts, J.; Rogers, A.; Cornish, M. Characterization of G12 sandwich ELISA, a next-generation immunoassay for gluten toxicity. J. AOAC Int. 2012, 95, 372–376. [Google Scholar] [CrossRef]
- Tanner, G.; Juhasz, A.; Florides, C.G.; Nye-Wood, M.; Békés, F.; Colgrave, M.L.; Russell, A.K.; Hardy, M.; Tye-Din, J.A. Preparation and Characterization of Avenin-Enriched Oat Protein by Chill Precipitation for Feeding Trials in Celiac Disease. Front. Nutr. 2019, 6, 162. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laemmli, U.K. Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Gojković-Cvjetković, V.S.; Grujić, R.D.; Marjanović-Balaban, R.; Stanojević, L.P.; Stanojević, J.S.; Cakić, M.D. Gliadin analysis by reversed-phase high performance liquid chromatography. Adv. Technol. 2019, 8, 30–36. [Google Scholar] [CrossRef] [Green Version]
- NO, E.R. 41/2009. Commission Regulation Concerning the Composition and Labelling of Foodstuffs Suitable for People Intolerant to Gluten. Available online: https://eur-lex.europa.eu/legal-content/EN/ALL/?uri=celex%3A32009R0041 (accessed on 23 October 2021).
- Marín-Sanz, M.; Giménez, M.J.; Barro, F.; Savin, R. Prolamin Content and Grain Weight in RNAi Silenced Wheat Lines under Different Conditions of Temperature and Nitrogen Availability. Front. Plant Sci. 2020, 11, 314. [Google Scholar] [CrossRef] [PubMed]
- Shewry, P.R.; Halford, N.G. Cereal seed storage proteins: Structures, properties and role in grain utilization. J. Exp. Bot. 2002, 53, 947–958. [Google Scholar] [CrossRef] [Green Version]
- Gregová, E.; Šliková, S.; Hozlár, P. Seed protein electrophoresis for identification of oat registered cultivars. Potravin. Slovak J. Food Sci. 2015, 9, 411–416. [Google Scholar] [CrossRef] [Green Version]
- Wieser, H. Chemistry of gluten proteins. Food Microbiol. 2007, 24, 115–119. [Google Scholar] [CrossRef]
- Balakireva, A.V.; Zamyatnin, A.A. Properties of Gluten Intolerance: Gluten Structure, Evolution, Pathogenicity and Detoxification Capabilities. Nutrients 2016, 8, 644. [Google Scholar] [CrossRef] [Green Version]
- Comino, I.; Bernardo, D.; Bancel, E.; Moreno, M.D.L.; Sánchez, B.; Barro, F.; Šuligoj, T.; Ciclitira, P.J.; Cebolla, A.; Knight, S.C.; et al. Identification and molecular characterization of oat peptides implicated on coeliac immune response. Food Nutr. Res. 2016, 60, 30324. [Google Scholar] [CrossRef] [Green Version]
- Gimenez, M.J.; Real, A.; García-Molina, M.D.; Sousa, C.; Barro, F. Characterization of celiac disease related oat proteins: Bases for the development of high quality oat varieties suitable for celiac patients. Sci. Rep. 2017, 7, 42588. [Google Scholar] [CrossRef] [Green Version]
- Panda, R.; Zoerb, H.F.; Cho, C.; Jackson, L.S.; Garber, E.A. Detection and Quantification of Gluten during the Brewing and Fermentation of Beer Using Antibody-Based Technologies. J. Food Prot. 2015, 78, 1167–1177. [Google Scholar] [CrossRef] [PubMed]
- Sajic, N.; Oplatowska-Stachowiak, M.; Streppel, L.; Drijfhout, J.-W.; Salden, M.; Koning, F. Development and in-house validation of a competitive ELISA for the quantitative detection of gluten in food. Food Control 2017, 80, 401–410. [Google Scholar] [CrossRef]
- Alves, T.O.; D’Almeida, C.T.; Scherf, K.A.; Ferreira, M.S. Modern approaches in the identification and quanti-fication of immunogenic peptides in cereals by LC-MS/MS. Front. Plant Sci. 2019, 10, 1470. [Google Scholar] [CrossRef] [PubMed]
| Classification | Category | Number of Materials |
|---|---|---|
| Taxon | A. sativa | 100 |
| A. nuda | 21 | |
| A. sativa var. nigra | 11 | |
| State of origin | FIN | 6 |
| CZE | 46 | |
| CAN | 8 | |
| RUS | 5 | |
| GER | 17 | |
| FRA | 10 | |
| GBR | 8 | |
| USA | 7 | |
| BEL | 2 | |
| POL | 2 | |
| AUT | 6 | |
| HUN | 1 | |
| EST | 2 | |
| IRL | 2 | |
| NOR | 1 | |
| SRB | 1 | |
| NLD | 2 | |
| ARG | 2 | |
| ITA | 1 | |
| SWE | 2 | |
| ROU | 1 |
| Parameter | Mean | Min. | Max. | St. Dev. | RSD (%) | |
|---|---|---|---|---|---|---|
| Gluten(G12) (mg·kg−1) | 7.2 * | ≤4.0 | 17.5 | 3.3 | 46.1 | |
| Crude protein (%) | 18.7 | 15.4 | 22.3 | 1.3 | 6.8 | |
| Correlation coef. Gluten(G12) vs. crude protein | −0.13 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dvořáček, V.; Kotrbová-Kozak, A.; Kozová-Doležalová, J.; Jágr, M.; Hlásná Čepková, P.; Vítámvás, P.; Kosová, K. Specific Avenin Cross-Reactivity with G12 Antibody in a Wide Range of Current Oat Cultivars. Foods 2022, 11, 567. https://doi.org/10.3390/foods11040567
Dvořáček V, Kotrbová-Kozak A, Kozová-Doležalová J, Jágr M, Hlásná Čepková P, Vítámvás P, Kosová K. Specific Avenin Cross-Reactivity with G12 Antibody in a Wide Range of Current Oat Cultivars. Foods. 2022; 11(4):567. https://doi.org/10.3390/foods11040567
Chicago/Turabian StyleDvořáček, Václav, Anna Kotrbová-Kozak, Jana Kozová-Doležalová, Michal Jágr, Petra Hlásná Čepková, Pavel Vítámvás, and Klára Kosová. 2022. "Specific Avenin Cross-Reactivity with G12 Antibody in a Wide Range of Current Oat Cultivars" Foods 11, no. 4: 567. https://doi.org/10.3390/foods11040567
APA StyleDvořáček, V., Kotrbová-Kozak, A., Kozová-Doležalová, J., Jágr, M., Hlásná Čepková, P., Vítámvás, P., & Kosová, K. (2022). Specific Avenin Cross-Reactivity with G12 Antibody in a Wide Range of Current Oat Cultivars. Foods, 11(4), 567. https://doi.org/10.3390/foods11040567






