Antimicrobial Resistance in Oral Healthcare: A Growing Concern in Dentistry
Abstract
1. Introduction
2. Origins of Antimicrobial Resistance
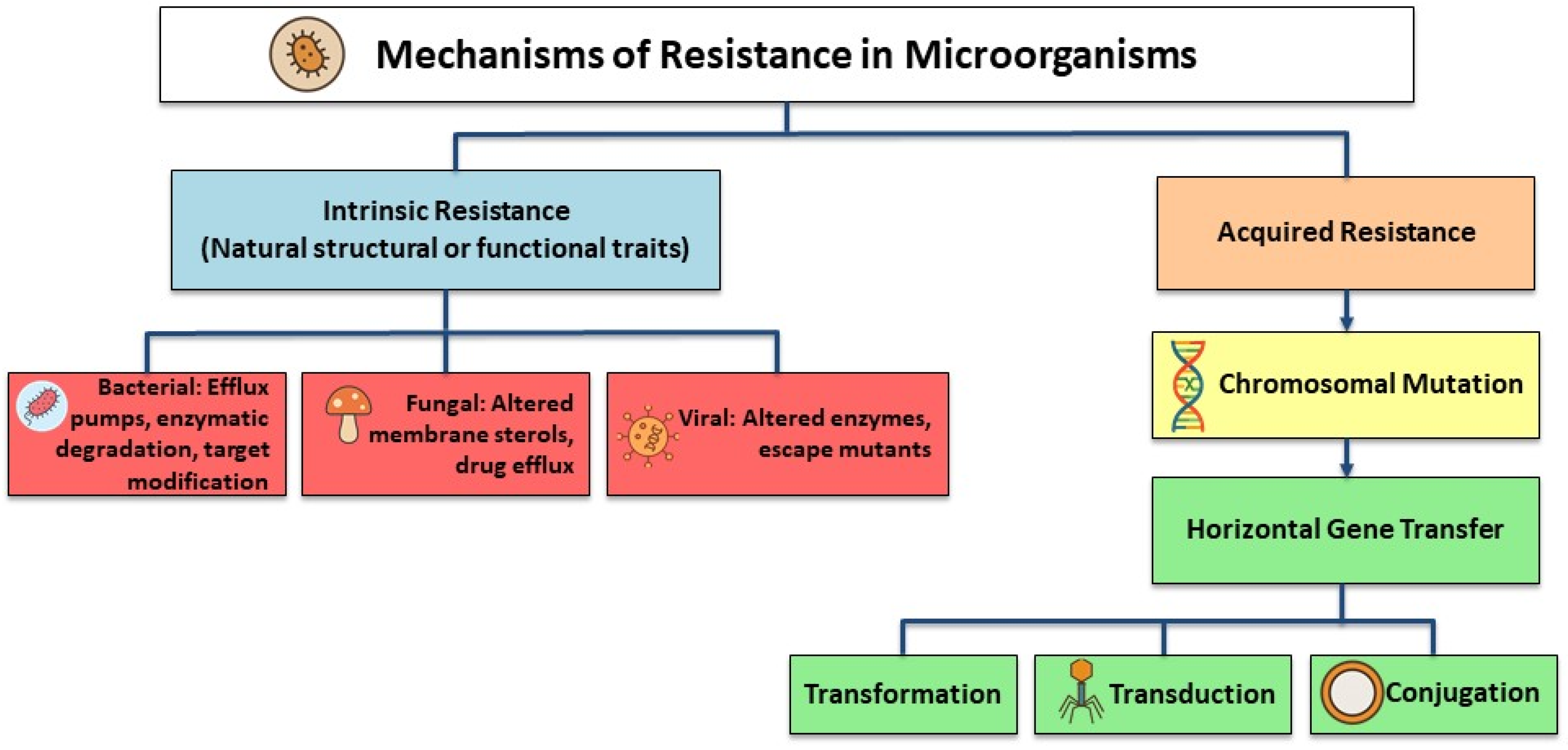
3. Mechanisms of Resistance
- Active drug efflux: Energy-dependent transport proteins, such as those from the ATP-binding cassette (ABC) family or major facilitator superfamily (MFS), pump antimicrobial agents out of the cell before they reach their target concentration [27]. In Gram-negative bacteria, efflux systems like AcrAB-TolC can expel multiple classes of antibiotics, leading to multi-drug resistance [28].
- Drug inactivation: Bacteria produce enzymes that chemically modify or destroy the antimicrobial agent. Classic examples include β-lactamases, which hydrolyse the β-lactam ring in penicillins and cephalosporins, and aminoglycoside-modifying enzymes that acetylate, phosphorylate, or adenylate the drug, making it ineffective [29].
- Limiting drug uptake: Alterations in outer membrane porins or cell wall structure reduce the permeability of the bacterial cell envelope, and as a result, it can prevent antibiotics from entering. This is common in Gram-negative bacteria, where porin loss or modification limits the uptake of β-lactams and fluoroquinolones [30].
- Target modification: Genetic mutations or enzymatic alterations change the binding site of the drug so that it no longer interacts effectively [31]. For instance, mutations in PBPs confer resistance to β-lactams, while alterations in DNA gyrase or topoisomerase II lead to fluoroquinolone resistance [32].
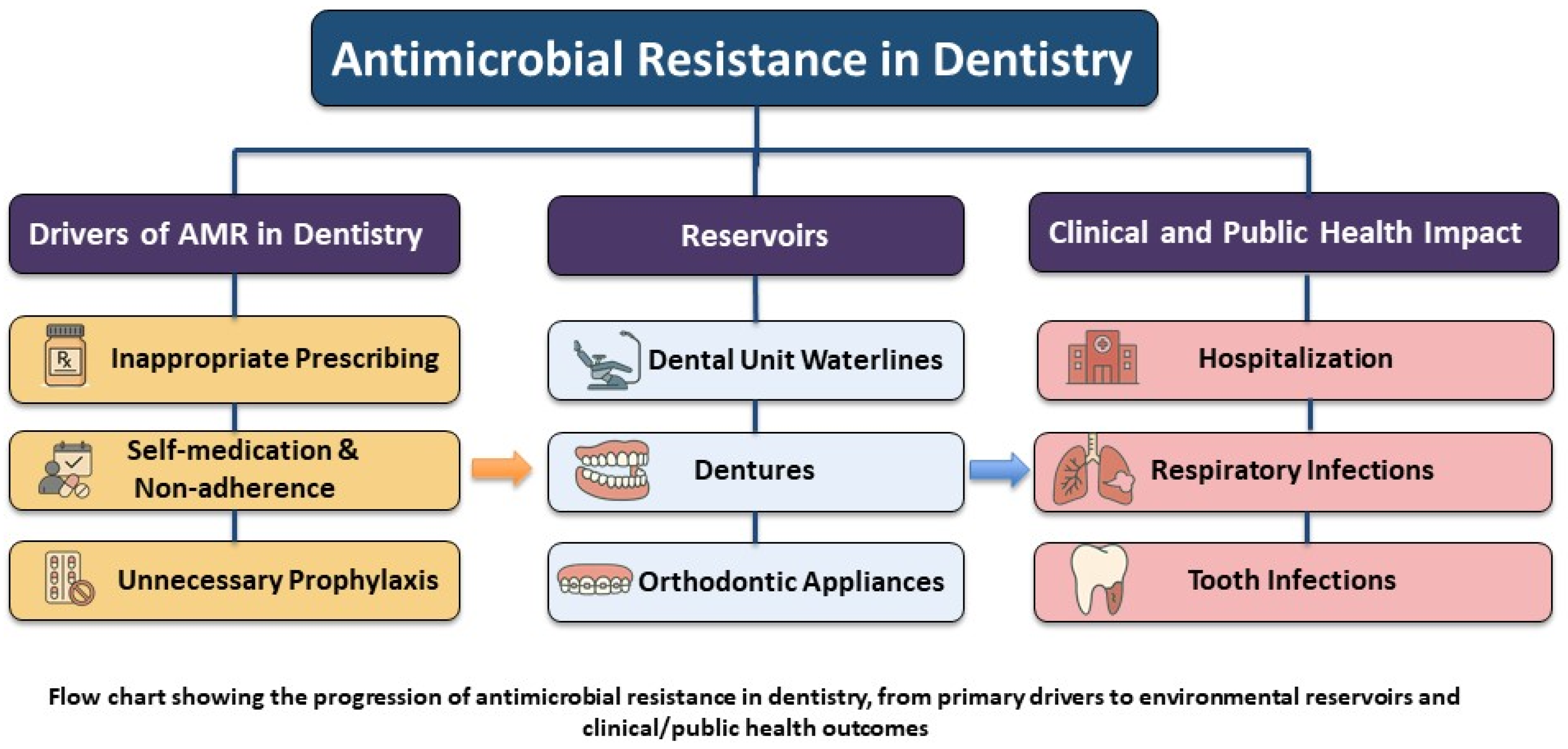
4. Antimicrobial Resistance in Dental Settings
4.1. Antimicrobial Resistance in DUWLs
4.1.1. Biofilm Formation in DUWLs and Its Role in Antimicrobial Resistance
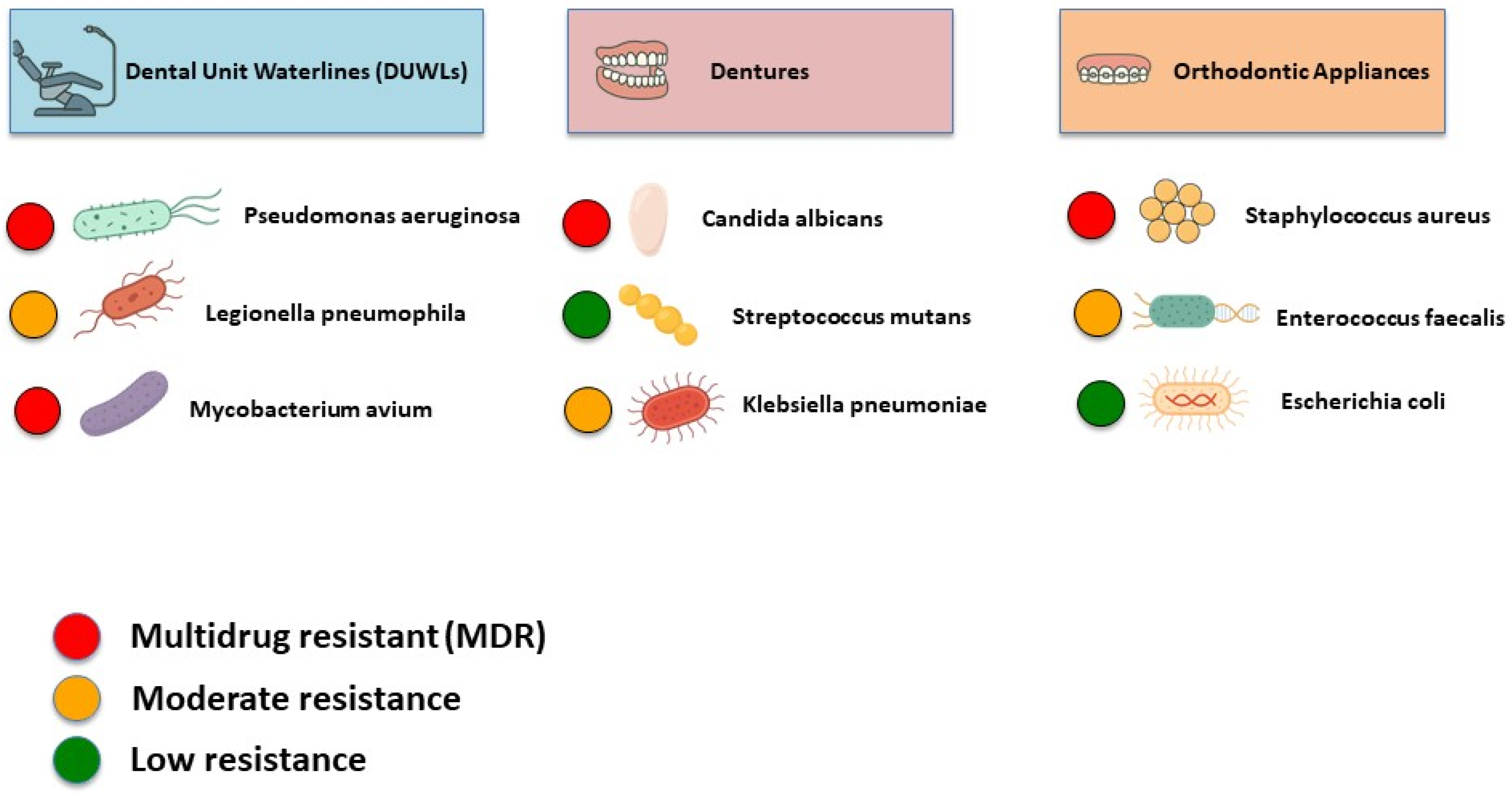
4.1.2. Common Microorganisms in DUWLs
4.1.3. DUWLs and Antimicrobial Resistance
4.1.4. Water Quality and Disinfection Protocols
4.2. Antimicrobial Resistance in Oral Appliances
4.2.1. Microbial Contamination and Biofilm Formation on Dentures
4.2.2. Common Resistant Pathogens in Dentures
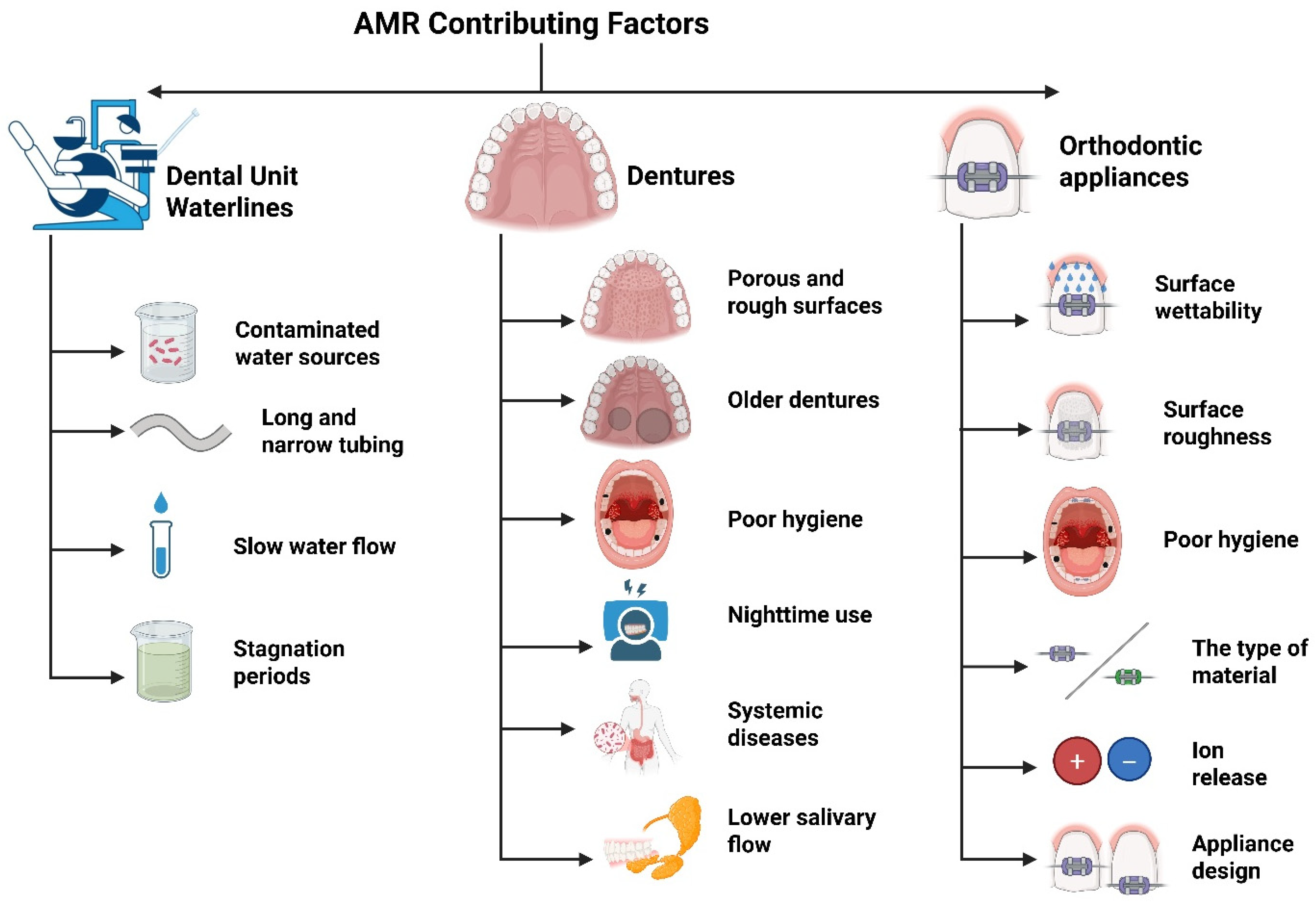
4.2.3. Risk Factors Contributing to Antimicrobial Resistance in Denture Wearers
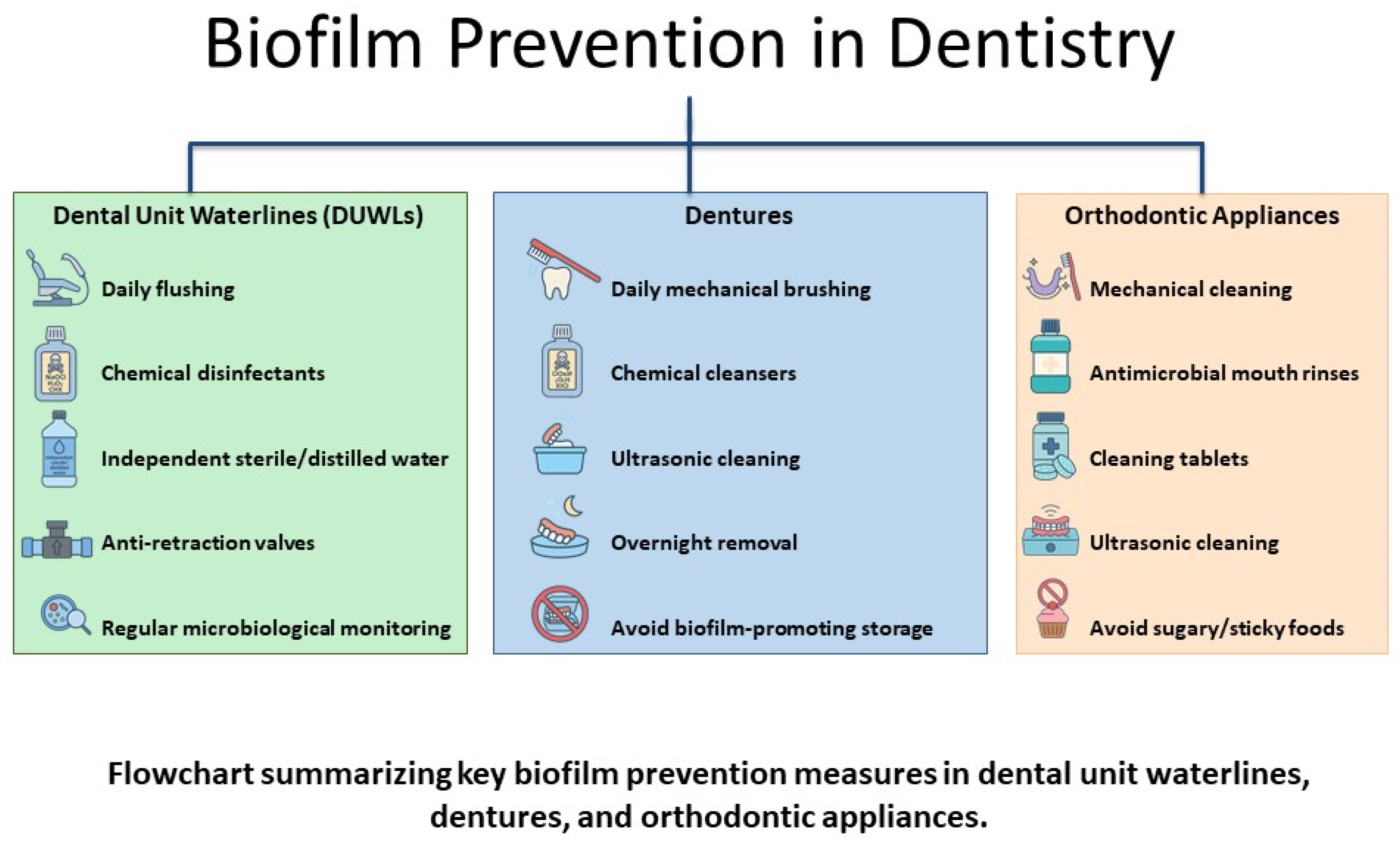
4.2.4. Preventing and Managing Antimicrobial Resistance in Dentures
Pharmacological Strategies
Mechanical and Behavioral Interventions
4.3. Antimicrobial Resistance in Orthodontic Appliances
4.3.1. Biofilm Formation on Orthodontic Devices
4.3.2. Resistant Microorganisms Among Orthodontic Patients
4.3.3. Antimicrobial Resistance Prevention in Orthodontic Device Users
5. Recommendations for AMS Implementation in Dentistry
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial resistance: A global multifaceted phenomenon. Pathog. Glob. Health 2015, 109, 309–318. [Google Scholar] [CrossRef]
- Fernandez, G. Turning the juggernaut. Lancet Planet Health 2022, 6, E75. [Google Scholar]
- Murray, C.J.L.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations; Wellcome Trust: London, UK, 2016. [Google Scholar]
- O’Neill, J. Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations; Review on Antimicrobial Resistance: London, UK, 2014. [Google Scholar]
- Centers for Disease Control and Prevention. Antibiotic Resistance Threats in the United States, 2019; US Department of Health and Human Services, CDC: Atlanta, GA, USA, 2019.
- WHO. Antimicrobial Resistance and the United Nations Sustainable Development Cooperation Framework; World Health Organization: Geneva, Switzerland, 2021. [Google Scholar]
- World Health Organization. Global Action Plan on Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015. [Google Scholar]
- Oberoi, S.S.; Dhingra, C.; Sharma, G.; Sardana, D. Antibiotics in dental practice: How justified are we. Int. Dent. J. 2020, 65, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Thompson, W.; Williams, D.; Pulcini, C.; Sanderson, S.; Calfon, P.; Verma, M. Tackling antibiotic resistance: Why dentistry matters. Int. Dent. J. 2021, 71, 450–453. [Google Scholar] [CrossRef] [PubMed]
- Barone, A.; Chatelain, S.; Derchi, G.; Di Spirito, F.; Martuscelli, R.; Porzio, M.; Sbordone, L. Antibiotic’s effectiveness after erupted tooth extractions: A retrospective study. Oral. Dis. 2020, 26, 967–973. [Google Scholar] [CrossRef]
- Contaldo, M.; D’ambrosio, F.; Ferraro, G.A.; Di Stasio, D.; Di Palo, M.P.; Serpico, R.; Simeone, M. Antibiotics in dentistry: A narrative review of the evidence beyond the myth. Int. J. Environ. Res. Public Health 2023, 20, 6025. [Google Scholar] [CrossRef]
- Gill, A.S.; Morrissey, H.; Rahman, A. A systematic review and meta-analysis evaluating antibiotic prophylaxis in dental implants and extraction procedures. Medicina 2018, 54, 95. [Google Scholar] [CrossRef] [PubMed]
- Cuevas-Gonzalez, M.V.; Mungarro-Cornejo, G.A.D.; Espinosa-Cristóbal, L.F.; Donohue-Cornejo, A.; Carrillo, K.L.T.; Acuña, R.A.S.; Calderón, A.G.M.G.; Gastelum, D.A.G.; Zambrano-Galván, G.; Cuevas-Gonzalez, J.C. Antimicrobial resistance in odontogenic infections: A protocol for systematic review. Medicine 2022, 101, e31345. [Google Scholar] [CrossRef] [PubMed]
- Ardila, C.M.; Bedoya-García, J.A. Antimicrobial resistance in patients with odontogenic infections: A systematic scoping review of prospective and experimental studies. J. Clin. Exp. Dent. 2022, 14, e834. [Google Scholar] [CrossRef] [PubMed]
- Abe, F.C.; Kodaira, K.; Motta, C.d.C.B.; Barberato-Filho, S.; Silva, M.T.; Guimarães, C.C.; Martins, C.C.; Lopes, L.C. Antimicrobial resistance of microorganisms present in periodontal diseases: A systematic review and meta-analysis. Front. Microbiol. 2022, 13, 961986. [Google Scholar] [CrossRef] [PubMed]
- Dahlen, G. Biofilms in Dental Unit Water Lines. Monogr. Oral Sci. 2020, 29, 12–18. [Google Scholar]
- O’Donnell, L.E.; Smith, K.; Williams, C.; Nile, C.J.; Lappin, D.F.; Bradshaw, D.; Lambert, M.; Robertson, D.P.; Bagg, J.; Hannah, V.; et al. Dentures are a reservoir for respiratory pathogens. J. Prosthodont. 2015, 25, 99–104. [Google Scholar] [CrossRef]
- Müller, L.K.; Jungbauer, G.; Jungbauer, R.; Wolf, M.; Deschner, J. Biofilm and Orthodontic Therapy. Monogr. Oral Sci. 2021, 29, 201–213. [Google Scholar] [PubMed]
- Teoh, L.; Löffler, C.; Mun, M.; Agnihotry, A.; Kaur, H.; Born, K.; Thompson, W. A Systematic Review of Dental Antibiotic Stewardship Interventions. Community Dent. Oral Epidemiol. 2024, 53, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Cox, G.; Wright, G.D. Intrinsic antibiotic resistance: Mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 2013, 303, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Martinez, J.L. General principles of antibiotic resistance in bacteria. Drug Discov. Today Technol. 2014, 11, 33–39. [Google Scholar] [CrossRef]
- Al-Haroni, M. Bacterial resistance and the dental professionals’ role to halt the problem. J. Dent. 2008, 36, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Munita, J.M.; Arias, C.A. Mechanisms of antibiotic resistance. Microbiol. Spectr. 2016, 4, 481–511. [Google Scholar] [CrossRef] [PubMed]
- Burmeister, A.R. Horizontal gene transfer. Evol. Med. Public Health 2015, 2015, 193–194. [Google Scholar] [CrossRef]
- Michaelis, C.; Grohmann, E. Horizontal gene transfer of antibiotic resistance genes in biofilms. Antibiotics 2023, 12, 328. [Google Scholar] [CrossRef]
- Gaurav, A.; Bakht, P.; Saini, M.; Pandey, S.; Pathania, R. Role of bacterial efflux pumps in antibiotic resistance, virulence, and strategies to discover novel efflux pump inhibitors. Microbiology 2023, 169, 001333. [Google Scholar] [CrossRef] [PubMed]
- Jang, S. AcrAB-TolC, a major efflux pump in Gram negative bacteria: Toward understanding its operation mechanism. BMB Rep. 2023, 56, 326. [Google Scholar] [CrossRef]
- Bush, K.; Bradford, P.A. β-Lactams and β-lactamase inhibitors: An overview. Cold Spring Harb. Perspect. Med. 2016, 6, a025247. [Google Scholar] [CrossRef] [PubMed]
- Delcour, A.H. Outer membrane permeability and antibiotic resistance. Biochim. Biophys. Acta 2009, 1794, 808–816. [Google Scholar] [CrossRef] [PubMed]
- Egorov, A.; Ulyashova, M.; Rubtsova, M. Bacterial enzymes and antibiotic resistance. Acta Nat. 2018, 10, 33–48. [Google Scholar] [CrossRef]
- Pletz, M.W.R.; McGee, L.; Beall, B.; Whitney, C.G.; Klugman, K.P. Interspecies recombination in type II topoisomerase genes is not a major cause of fluoroquinolone resistance in invasive Streptococcus pneumoniae isolates in the United States. Antimicrob. Agents Chemother. 2005, 49, 779–780. [Google Scholar] [CrossRef] [PubMed]
- Reygaert, W.C. An overview of the antimicrobial resistance mechanisms of bacteria. AIMS Microbiol. 2018, 4, 482. [Google Scholar] [CrossRef]
- Hossain, C.M.; Ryan, L.K.; Gera, M.; Choudhuri, S.; Lyle, N.; Ali, K.A.; Diamond, G. Antifungals and drug resistance. Encyclopedia 2022, 2, 1722–1737. [Google Scholar] [CrossRef]
- Vere Hodge, A.; Field, H.J. General mechanisms of antiviral resistance. Genet. Evol. Infect. Dis. 2010, 24, 339–362. [Google Scholar]
- Pallasch, T.J. Global antibiotic resistance and its impact on the dental community. J. Calif. Dent. Assoc. 2000, 28, 215–231. [Google Scholar] [CrossRef]
- Anderson, A.C.; von Ohle, C.; Frese, C.; Boutin, S.; Bridson, C.; Schoilew, K.; Peikert, S.A.; Hellwig, E.; Pelz, K.; Wittmer, A.; et al. The oral microbiota is a reservoir for antimicrobial resistance: Resistome and phenotypic resistance characteristics of oral biofilm in health, caries, and periodontitis. Ann. Clin. Microbiol. Antimicrob. 2023, 22, 37. [Google Scholar] [CrossRef] [PubMed]
- Khalil, D.; Hultin, M.; Rashid, M.; Lund, B. Oral microflora and selection of resistance after a single dose of amoxicillin. Clin. Microbiol. Infect. 2016, 22, 949.e1–949.e4. [Google Scholar] [CrossRef] [PubMed]
- Masuda, K.; Nemoto, H.; Nakano, K.; Naka, S.; Nomura, R.; Ooshima, T. Amoxicillin-resistant oral streptococci identified in dental plaque specimens from healthy Japanese adults. J. Cardiol. 2012, 59, 285–290. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, D.; Kushwah, T.; Kushwah, M.; Singh, V. Antibiotic susceptibility pattern against pathogenic bacteria causing Dental Caries. S. Asian J. Exp. Biol. 2011, 1, 31–35. [Google Scholar] [CrossRef]
- Kuriyama, T.; Williams, D.W.; Yanagisawa, M.; Iwahara, K.; Shimizu, C.; Nakagawa, K.; Yamamoto, E.; Karasawa, T. Antimicrobial susceptibility of 800 anaerobic isolates from patients with dentoalveolar infection to 13 oral antibiotics. Oral Microbiol. Immunol. 2007, 22, 285–288. [Google Scholar] [CrossRef]
- Medina-Palacios, S.E.; Vitales-Noyola, M.; López-González, E.; González-Amaro, A.M.; Méndez-González, V.; Pozos-Guillén, A. Root canal microorganisms and their antibiotic susceptibility in patients with persistent endodontic infections, with and without clinical symptoms. Odontology 2021, 109, 596–604. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.; Coast, J. The true cost of antimicrobial resistance. BMJ 2013, 346, f1493. [Google Scholar] [CrossRef]
- Kang, Y.; Sun, B.; Chen, Y.; Lou, Y.; Zheng, M.; Li, Z.; Fey, P.D. Dental plaque microbial resistomes of periodontal health and disease and their changes after scaling and root planing therapy. mSphere 2021, 6, e0016221. [Google Scholar] [CrossRef] [PubMed]
- Morales-Dorantes, V.; Domínguez-Pérez, R.A.; Pérez-Serrano, R.M.; Solís-Sainz, J.C.; García-Solís, P.; Espinosa-Cristóbal, L.F.; Cabeza-Cabrera, C.V.; Ayala-Herrera, J.L. The distribution of eight antimicrobial resistance genes in Streptococcus oralis, Streptococcus sanguinis, and Streptococcus gordonii strains isolated from dental plaque as oral commensals. Trop. Med. Infect. Dis. 2023, 8, 499. [Google Scholar] [CrossRef]
- Ardila, C.M.; Bedoya-García, J.A. Antimicrobial resistance of Aggregatibacter actinomycetemcomitans, Porphyromonas gingivalis and Tannerella forsythia in periodontitis patients. J. Glob. Antimicrob. Resist. 2020, 22, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Tesauro, M.; Consonni, M.; Grappasonni, I.; Lodi, G.; Mattina, R. Dental unit water content and antibiotic resistance of Pseudomonas aeruginosa and Pseudomonas species: A case study. J. Oral Microbiol. 2022, 14, 2107316. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.T.; Marsh, P.D. Microbial biofilm formation in DUWS and their control using disinfectants. J. Dent. 2007, 35, 721–730. [Google Scholar] [CrossRef] [PubMed]
- Cicciù, M. Water contamination risks at the dental clinic. Biology 2020, 9, 43. [Google Scholar] [CrossRef] [PubMed]
- Spagnolo, A.M.; Sartini, M.; Cristina, M.L. Microbial contamination of dental unit waterlines and potential risk of infection: A narrative review. Pathogens 2020, 9, 651. [Google Scholar] [CrossRef] [PubMed]
- Bayani, M.; Raisolvaezin, K.; Almasi-Hashiani, A.; Mirhoseini, S.H. Bacterial biofilm prevalence in dental unit waterlines: A systematic review and meta-analysis. BMC Oral Health 2023, 23, 158. [Google Scholar] [CrossRef]
- Singh, J.; O’Donnell, K.; Nieves, D.J.; Adler-Shohet, F.C.; Arrieta, A.C.; Ashouri, N.; Ahuja, G.; Cheung, M.; Holmes, W.N.; Huoh, K.; et al. Invasive Mycobacterium abscessus outbreak at a pediatric dental clinic. Open Forum Infect. Dis. 2021, 8, ofab165. [Google Scholar] [CrossRef]
- Pérez-Alfonzo, R.; Brito, L.E.P.; Vergara, M.S.; Damasco, A.R.; Rodríguez, P.L.M.; Quintero, C.E.K.; Martinez, C.C.; Rivera-Oliver, I.A.; Jardin, O.J.D.M.; Rodríguez-Castillo, B.A.; et al. Odontogenic cutaneous sinus tracts due to infection with nontuberculous mycobacteria: A report of three cases. BMC Infect. Dis. 2020, 20, 295. [Google Scholar] [CrossRef] [PubMed]
- Gawish, S.; Abbass, A.; Abaza, A. Occurrence and biofilm forming ability of Pseudomonas aeruginosa in the water output of dental unit waterlines in a dental center in Alexandria, Egypt. Germs 2019, 9, 71–80. [Google Scholar] [CrossRef]
- Mah, T.F. Biofilm-specific antibiotic resistance. Future Microbiol. 2012, 7, 1061–1072. [Google Scholar] [CrossRef]
- Rath, S.; Bal, S.C.B.; Dubey, D. Oral biofilm: Development mechanism, multidrug resistance, and their effective management with novel techniques. Rambam Maimonides Med. J. 2021, 12, e0004. [Google Scholar] [CrossRef] [PubMed]
- Marcinkiewicz, J.; Strus, M.; Pasich, E. Antibiotic resistance: A “dark side” of biofilm-associated chronic infections. Pol. Arch. Med. Wewn. 2013, 123, 309–313. [Google Scholar] [CrossRef]
- Sharma, S.; Mohler, J.; Mahajan, S.D.; Schwartz, S.A.; Bruggemann, L.; Aalinkeel, R. Microbial biofilm: A review on formation, infection, antibiotic resistance, control measures, and innovative treatment. Microorganisms 2023, 11, 1614. [Google Scholar] [CrossRef]
- Soto, S.M. Role of efflux pumps in the antibiotic resistance of bacteria embedded in a biofilm. Virulence 2013, 4, 223–229. [Google Scholar] [CrossRef] [PubMed]
- Van Acker, H.; Van Dijck, P.; Coenye, T. Molecular mechanisms of antimicrobial tolerance and resistance in bacterial and fungal biofilms. Trends Microbiol. 2014, 22, 326–333. [Google Scholar] [CrossRef] [PubMed]
- Barbot, V.; Robert, A.; Rodier, M.-H.; Imbert, C. Update on infectious risks associated with dental unit waterlines. FEMS Immunol. Med. Microbiol. 2012, 65, 196–204. [Google Scholar] [CrossRef]
- Dowdell, K.; Haig, S.-J.; Caverly, L.J.; Shen, Y.; LiPuma, J.J.; Raskin, L. Nontuberculous mycobacteria in drinking water systems—The challenges of characterization and risk mitigation. Curr. Opin. Biotechnol. 2019, 57, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Mtetwa, H.N.; Amoah, I.D.; Kumari, S.; Bux, F.; Reddy, P. The source and fate of Mycobacterium tuberculosis complex in wastewater and possible routes of transmission. BMC Public Health 2022, 22, 145. [Google Scholar] [CrossRef]
- Alkhulaifi, M.M.; Alotaibi, D.H.; Alajlan, H.; Binshoail, T. Assessment of nosocomial bacterial contamination in dental unit waterlines: Impact of flushing. Saudi Dent. J. 2020, 32, 68–73. [Google Scholar] [CrossRef]
- Szymańska, J.; Sitkowska, J.; Dutkiewicz, J. Microbial contamination of dental unit waterlines. Ann. Agric. Environ. Med. 2008, 15, 173–179. [Google Scholar] [PubMed]
- Li, B.; Hu, Y.; Wang, Y.; Zhang, C.; Wang, Z.; Peng, X.; Feng, J. Periodic detection and disinfection maintenance of dental unit waterlines in dental simulation head model laboratories. Sci. Rep. 2025, 15, 5234. [Google Scholar] [CrossRef]
- Atlas, R.M.; Williams, J.F.; Huntington, M.K. Legionella contamination of dental-unit waters. Appl. Environ. Microbiol. 1995, 61, 1208–1213. [Google Scholar] [CrossRef] [PubMed]
- Rudbeck, M.; Viskum, S.; Mølbak, K.; Uldum, S.A. Legionella antibodies in a Danish hospital staff with known occupational exposure. J. Environ. Public Health 2009, 2009, 812829. [Google Scholar] [CrossRef]
- Kevorkyan, A.; Tomova, I.; Raycheva, R.; Stoeva, V.; Stoilova, Y.; Lalabonova, H.; Kondeva, V. Legionella pneumophila antibodies in serum samples from medical and dental personnel: A seroepidemiological survey. Biotechnol. Biotechnol. Equip. 2017, 31, 588–593. [Google Scholar] [CrossRef][Green Version]
- Schönning, C.; Jernberg, C.; Klingenberg, D.; Andersson, S.; Pääjärvi, A.; Alm, E.; Tano, E.; Lytsy, B. Legionellosis acquired through a dental unit: A case study. J. Hosp. Infect. 2017, 96, 89–92. [Google Scholar] [CrossRef]
- Ricci, M.L.; Fontana, S.; Pinci, F.; Fiumana, E.; Pedna, M.F.; Farolfi, P.; Sabattini, M.A.B.; Scaturro, M. Pneumonia associated with a dental unit waterline. Lancet 2012, 379, 684. [Google Scholar] [CrossRef] [PubMed]
- Santajit, S.; Indrawattana, N. Mechanisms of antimicrobial resistance in ESKAPE pathogens. BioMed Res. Int. 2016, 2016, 2475067. [Google Scholar] [CrossRef] [PubMed]
- Desai, A.N.; Hurtado, R.M. Infections and outbreaks of nontuberculous mycobacteria in hospital settings. Curr. Treat. Options Infect. Dis. 2018, 10, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Sachan, R.S.K.; Mistry, V.; Dholaria, M.; Rana, A.; Devgon, I.; Ali, I.; Iqbal, J.; Eldin, S.M.; Al-Tawaha, A.R.M.S.; Bawazeer, S.; et al. Overcoming Mycobacterium tuberculosis drug resistance: Novel medications and repositioning strategies. ACS Omega 2023, 8, 32244–32257. [Google Scholar] [CrossRef]
- Vosooghi, K.; Larypoor, M.; Sakhaee, F.; Jajin, M.G.; Moghaddam, S.; Samieefar, N.; Keramat, R.R.; Sabotki, M.A.; Fateh, A. Distribution of nontuberculous mycobacteria in dental unit waterlines: A potential health hazard in the dental office. Microb. Pathog. 2024, 196, 106963. [Google Scholar] [CrossRef] [PubMed]
- Vincent, A.T.; Freschi, L.; Jeukens, J.; Kukavica-Ibrulj, I.; Emond-Rheault, J.-G.; Leduc, A.; Boyle, B.; Jean-Pierre, F.; Groleau, M.-C.; Déziel, E.; et al. Genomic characterisation of environmental Pseudomonas aeruginosa isolated from dental unit waterlines revealed the insertion sequence IS Pa11 as a chaotropic element. FEMS Microbiol. Ecol. 2017, 93, fix106. [Google Scholar]
- Alsehlawi, Z.S.; Al-Yasiri, I.K.; Fakhriddeen, A.J.; Taher, A.A.Y. Antibiotic susceptibility patterns of legionella pneumophila isolated from water lines of dental settings. Smile Dent. J. 2016, 11, 36–39. [Google Scholar] [CrossRef]
- American Dental Association. ADA statement on dental unit waterlines. J. Am. Dent. Assoc. 1996, 127, 185–186. [Google Scholar] [CrossRef]
- Güngör, N.D.; Kadaifçiler, D.G.; Peker, O.Ö. Investigation of the bacterial load and antibiotic susceptibility of dental units. Environ. Monit. Assess. 2013, 186, 1847–1853. [Google Scholar] [CrossRef] [PubMed]
- Uzel, A.; Cogulu, D.; Oncag, O. Microbiological evaluation and antibiotic susceptibility of dental unit water systems in general dental practice. Int. J. Dent. Hyg. 2008, 6, 43–47. [Google Scholar] [CrossRef] [PubMed]
- Lancellotti, M.; Oliveira, M.P.; Ávila, F.A. Research on Staphylococcus spp. in biofilm formation in water pipes and sensibility to antibiotics. Braz. J. Oral Sci. 2007, 6, 20. [Google Scholar]
- Buitrago, J.M.; Kolbe, R.J.; Siqueira, M.F. Dental unit waterline testing practices: An 11-Year retrospective study. BMC Oral Health 2023, 23, 867. [Google Scholar] [CrossRef]
- Kohn, W.G.; Collins, A.S.; Cleveland, J.L.; Harte, J.A.; Eklund, K.J.; Malvitz, D.M. Guidelines for infection control in dental health-care settings—2003. In MMWR Recommendations and Reports; Centers for Disease Control and Prevention (CDC): Atlanta, GA, USA, 2003; 52, pp. 1–61. [Google Scholar]
- Bowen, C.G.; Greenwood, W.; Guevara, P.; Washington, M.A. Effectiveness of a dental unit waterline treatment protocol with A-Dec ICX and Citrisil Disinfectants. Mil. Med. 2015, 180, 1098–1104. [Google Scholar] [CrossRef]
- O’Donnell, M.J.; Boyle, M.A.; Russell, R.J.; Coleman, D.C. Management of dental unit waterline biofilms in the 21st century. Future Microbiol. 2011, 6, 1209–1226. [Google Scholar] [CrossRef]
- Baudet, A.; Lizon, J.; Martrette, J.M.; Camelot, F.; Florentin, A.; Clément, C. Efficacy of BRS® and Alpron®/Bilpron® disinfectants for dental unit waterlines: A six-year study. Int. J. Environ. Res. Public Health 2020, 17, 2634. [Google Scholar] [CrossRef]
- Noopan, S.; Unchui, P.; Techotinnakorn, S.; Ampornaramveth, R.S. Plasma sterilization effectively reduces bacterial contamination in dental unit waterlines. Int. J. Dent. 2019, 2019, 5720204. [Google Scholar] [CrossRef] [PubMed]
- Okubo, K.; Ito, T.; Shiota, Y.; Kawata, Y.; Yamamoto, T.; Takashiba, S. Effectiveness and safety of low-concentrated ozonized water for the reduction of contamination in dental unit water lines. Heliyon 2019, 5, e02306. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention (CDC). Best Practices for Dental Unit Water Quality. 2024. Available online: https://www.cdc.gov/dental-infection-control/hcp/dental-ipc-faqs/best-practices-dental-unit-water-quality.html (accessed on 15 May 2024).
- College of Dental Surgeons of Saskatchewan. Infection Prevention and Control Standards in the Oral Health Care Facility. Available online: https://saskdentists.com/wp-content/uploads/2023/04/Infection-Prevention-and-Control-Standard.pdf (accessed on 22 June 2025).
- Ji, X.Y.; Fei, C.N.; Zhang, Y.; Zhang, W.; Liu, J.; Dong, J. Evaluation of bacterial contamination of dental unit waterlines and use of a newly designed measurement device to assess retraction of a dental chair unit. Int. Dent. J. 2020, 66, 208–214. [Google Scholar] [CrossRef]
- World Health Organization. Water Safety Plan Manual: Step-by-Step Risk Management for Drinking-Water Suppliers; World Health Organization: Geneva, Switzerland, 2023. [Google Scholar]
- Directive, C. On the quality of water intended for human consumption. In Proceedings of the Commission of The European Communities, Brussels, Belgium, 8 July 1998. [Google Scholar]
- Directive (EU) 2020/2184 of the European Parliament and of the Council of 16 December 2020 on the Quality of Water Intended for Human Consumption; EUR-Lex: Luxembourg, 2020; Volume 435, pp. 1–62.
- Tuvo, B.; Totaro, M.; Cristina, M.L.; Spagnolo, A.M.; Di Cave, D.; Profeti, S.; Baggiani, A.; Privitera, G.; Casini, B. Prevention and control of Legionella and Pseudomonas spp. colonization in dental units. Pathogens 2020, 9, 305. [Google Scholar] [CrossRef]
- Vaidya, R.Y.; Hegde, V.; Sherigar, P.; Prabhu, N. Exploring microbial interactions with denture resin surface and implications for plant based plaque control strategies: A narrative review. Beni-Suef Univ. J. Basic Appl. Sci. 2024, 13, 110. [Google Scholar] [CrossRef]
- de Foggi, C.C.; Machado, A.L.; Zamperini, C.A.; Fernandes, D.; Wady, A.F.; Vergani, C.E. Effect of surface roughness on the hydrophobicity of a denture-base acrylic resin and Candida albicans colonization. J. Investig. Clin. Dent. 2016, 7, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Suryawanshi, H.; Hande, A.; Dasari, A.K.; Aileni, K.R.; AlZoubi, I.; Patil, S.R. Metal ion release from orthodontic appliances: Concerns regarding potential carcinogenic effects. Oral Oncol. Rep. 2024, 10, 100309. [Google Scholar] [CrossRef]
- Jeon, D.-M.; An, J.-S.; Lim, B.-S.; Ahn, S.-J. Orthodontic bonding procedures significantly influence biofilm composition. Prog. Orthod. 2020, 21, 14. [Google Scholar] [CrossRef] [PubMed]
- Khawwam, S.I.; Al-Groosh, D.H. Effect of Different Cleaning Regimes on Biofilm Formation of Acrylic-Based Removable Orthodontic Appliance: A Randomized Clinical Trial. Sci. World J. 2023, 2023, 9920850. [Google Scholar] [CrossRef] [PubMed]
- Abbas, M.H.; Al-Yasseen, A.K.; Alhamadi, W.W. Prevalence of Staphylococcus aureus among gingivitis in patient with orthodontic wires in Kufa City/Iraq. Biotechnology 2017, 14, 91–96. [Google Scholar]
- McReynolds, D.E.; Moorthy, A.; Moneley, J.O.; Jabra-Rizk, M.A.; Sultan, A.S. Denture stomatitis—An interdisciplinary clinical review. J. Prosthodont. 2023, 32, 560–570. [Google Scholar] [CrossRef] [PubMed]
- Mayer, C.; Moritz, R.; Kirschner, C.; Borchard, W.; Maibaum, R.; Wingender, J.; Flemming, H.-C. The role of intermolecular interactions: Studies on model systems for bacterial biofilms. Int. J. Biol. Macromol. 1999, 26, 3–16. [Google Scholar] [CrossRef]
- Katsikogianni, M.; Missirlis, Y.F. Concise review of mechanisms of bacterial adhesion to biomaterials and of techniques used in estimating bacteria-material interactions. Eur. Cell Mater. 2004, 8, 37–57. [Google Scholar] [CrossRef] [PubMed]
- Sterzenbach, T.; Helbig, R.; Hannig, C.; Hannig, M. Bioadhesion in the oral cavity and approaches for biofilm management by surface modifications. Clin. Oral Investig. 2020, 24, 4237–4260. [Google Scholar] [CrossRef]
- Ten Cate, J.M. Biofilms, a new approach to the microbiology of dental plaque. Odontology 2006, 94, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Morse, D.J.; Wilson, M.J.; Wei, X.; Lewis, M.A.O.; Bradshaw, D.J.; Murdoch, C.; Williams, D.W. Denture-associated biofilm infection in three-dimensional oral mucosal tissue models. J. Med. Microbiol. 2018, 67, 364–375. [Google Scholar] [CrossRef] [PubMed]
- Sultan, A.S.; Rizk, A.M.; Vila, T.; Ji, Y.; Masri, R.; Jabra-Rizk, M.A. Digital design of a universal rat intraoral device for therapeutic evaluation of a topical formulation against Candida-associated denture stomatitis. Infect. Immun. 2019, 87, e00617-19. [Google Scholar] [CrossRef]
- Yoshizaki, T.; Akiba, N.; Inokoshi, M.; Shimada, M.; Minakuchi, S. Hydrophilic nano-silica coating agents with platinum and diamond nanoparticles for denture base materials. Dent. Mater. J. 2017, 36, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Oliveira Junior, N.M.D.; Mendoza Marin, D.O.; Leite, A.R.P.; Pero, A.C.; Klein, M.I.; Compagnoni, M.A. Influence of the use of complete denture adhesives on microbial adhesion and biofilm formation by single-and mixed-species. PLoS ONE 2018, 13, e0203951. [Google Scholar] [CrossRef]
- Buranarom, N.; Komin, O.; Matangkasombut, O. Hyposalivation, oral health, and Candida colonization in independent dentate elders. PLoS ONE 2020, 15, e0242832. [Google Scholar] [CrossRef]
- Gad, M.M.; Al-Thobity, A.M.; Shahin, S.Y.; Alsaqer, B.T.; Ali, A. Inhibitory effect of zirconium oxide nanoparticles on Candida albicans adhesion to repaired polymethyl methacrylate denture bases and interim removable prostheses: A new approach for denture stomatitis prevention. Int. J. Nanomed. 2017, 12, 5409–5419. [Google Scholar] [CrossRef]
- Naji, S.A.; Kashi, T.S.J.; Pourhajibagher, M.; Behroozibakhsh, M.; Masaeli, R.; Bahador, A. Evaluation of antimicrobial properties of conventional poly(methyl methacrylate) denture base resin materials containing hydrothermally synthesised anatase TiO2 nanotubes against cariogenic bacteria and Candida albicans. Iran. J. Pharm. Res. 2018, 17, 161–172. [Google Scholar]
- Majchrzak, K.; Mierzwinska-Nastalska, E.; Chmura, A.; Kwiatkowski, A.; Paczek, L.; Mlynarczyk, G.; Szymanek-Majchrzak, K. Comparison of staphylococcal flora in denture plaque and the surface of the pharyngeal mucous membrane in kidney transplant recipients. Transplant. Proc. 2016, 48, 1590–1597. [Google Scholar] [CrossRef] [PubMed]
- Nett, J.E.; Andes, D.R. Fungal Biofilms: In vivo models for discovery of anti-biofilm drugs. Microb. Biofilms 2015, 3, 33–49. [Google Scholar] [CrossRef] [PubMed]
- Larijani, M.; Zareshahrabadi, Z.; Alhavaz, A.; Hajipour, R.; Ranjbaran, A.; Giti, R.; Soltankarimi, V.; Zomorodian, K. Evaluation of Candida albicans biofilm formation on conventional and computer-aided-design/computer-aided manufacturing (CAD/CAM) denture base materials. Curr. Med. Mycol. 2022, 8, 23–29. [Google Scholar] [CrossRef] [PubMed]
- VV, N.; Karibasappa, G.; Dodamani, A.; Prashanth, V.K. Microbial contamination of removable dental prosthesis at different interval of usage: An: In vitro: Study. J. Indian Prosthodont. Soc. 2016, 16, 346–351. [Google Scholar]
- Arbelaez, M.I.A.; Vergani, C.E.; Barbugli, P.A.; Pavarina, A.C.; Sanita, P.V.; Jorge, J.H. Long-term effect of daily chemical disinfection on surface topography and Candida albicans biofilm formation on denture base and reline acrylic resins. Oral Health Prev. Dent. 2020, 18, a45521. [Google Scholar]
- Rossi, T.; Peltonen, R.; Laine, J.; Eerola, E.; Vuopio-Varkila, J.; Kotilainen, P. Eradication of the long-term carriage of methicillin-resistant Staphylococcus aureus in patients wearing dentures: A follow-up of 10 patients. J. Hosp. Infect. 1996, 34, 311–320. [Google Scholar] [CrossRef]
- Andonissamy, L.; Karthigeyan, S.; Ali, S.A. Detection of Drug Susceptible and Resistant Viridans Streptococci sp. Staphylococcus aureus, Klebsiella pneumoniae and E. coli in Complete Denture Patients and Visualisation using Scanning Electron Microscopy. J. Clin. Diagn. Res. 2020, 14, 15. [Google Scholar]
- Gaetti-Jardim, E.C.; Marqueti, A.C.; Faverani, L.P.; Júnior, E.G.-J. Antimicrobial resistance of aerobes and facultative anaerobes isolated from the oral cavity. J. Appl. Oral Sci. 2010, 18, 551–559. [Google Scholar] [CrossRef]
- Ramos, M.M.B.; Gartti-Jardim, E.C.; Junior, E.G.-J. Resistance to tetracycline and β-lactams and distribution of resistance markers in enteric microorganisms and pseudomonads isolated from the oral cavity. J. Appl. Oral Sci. 2009, 17, 13–18. [Google Scholar] [CrossRef]
- Abu-Elteen, K.H. Candida albicans strain differentiation in complete denture wearers. New Microbiol. 2000, 23, 329–337. [Google Scholar]
- Martin-Mazuelos, E.; Aller, A.I.; Romero, M.J.; Armijo, A.R.; Gutierrez, M.J.; Bernal, S.; Montero, O. Response to fluconazole and itraconazole of Candida spp. in denture stomatitis. Mycoses 1997, 40, 283–289. [Google Scholar] [CrossRef]
- Singh, T.; Bhojaraju, N.; Vinod, V.; Anjali, K.; Shah, S.; Kumar, N. Evaluation of microbial contamination in removable dental prosthesis at different time of usage. J. Oral Maxillofac. Pathol. 2023, 27, 333–339. [Google Scholar] [CrossRef]
- Smith, A.; Brewer, A.; Kirkpatrick, P.; Jackson, M.; Young, J.; Watson, S.; Thakker, B. Staphylococcal species in the oral cavity from patients in a regional burns unit. J. Hosp. Infect. 2003, 55, 184–189. [Google Scholar] [CrossRef]
- Dorocka-Bobkowska, B.; Konopka, K. Susceptibility of Candida isolates from denture-related stomatitis to antifungal agents in vitro. Int. J. Prosthodont. 2007, 20, 504–506. [Google Scholar] [PubMed]
- Ritchie, G.; Fletcher, A.; Main, D.; Prophet, A. The etiology, exfoliative cytology, and treatment of denture stomatitis. J. Prosthet. Dent. 1969, 22, 185–200. [Google Scholar] [CrossRef] [PubMed]
- Manfredi, M.; McCullough, M.J.; Polonelli, L.; Conti, S.; Al-Karaawi, Z.M.; Vescovi, P.; Porter, S.R. In vitro antifungal susceptibility to six antifungal agents of 229 Candida isolates from patients with diabetes mellitus. Oral Microbiol. Immunol. 2006, 21, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Chandra, J.; Mukherjee, P.; Leidich, S.; Faddoul, F.; Hoyer, L.; Douglas, L.; Ghannoum, M. Antifungal resistance of candidal biofilms formed on denture acrylic in vitro. J. Dent. Res. 2001, 80, 903–908. [Google Scholar] [CrossRef] [PubMed]
- Koga-Ito, C.Y.; Lyon, J.P.; Vidotto, V.; de Resende, M.A. Virulence factors and antifungal susceptibility of Candida albicans isolates from oral candidosis patients and control individuals. Mycopathologia 2006, 161, 219–223. [Google Scholar] [CrossRef] [PubMed]
- Lamfon, H.; Porter, S.R.; McCullough, M.; Pratten, J. Susceptibility of Candida albicans biofilms grown in a constant depth film fermentor to chlorhexidine, fluconazole and miconazole: A longitudinal study. J. Antimicrob. Chemother. 2004, 53, 383–385. [Google Scholar] [CrossRef] [PubMed]
- Perić, M.; Miličić, B.; Pfićer, J.K.; Živković, R.; Arsenijević, V.A. A systematic review of denture stomatitis: Predisposing factors, clinical features, etiology, and global Candida spp. distribution. J. Fungi 2024, 10, 328. [Google Scholar] [CrossRef]
- Le Bars, P.; Kouadio, A.A.; Bandiaky, O.N.; Le Guéhennec, L.; de La Cochetière, M.-F. Host’s Immunity and Candida Species Associated with Denture Stomatitis: A Narrative Review. Microorganisms 2022, 10, 1437. [Google Scholar] [CrossRef] [PubMed]
- Sartawi, S.Y.; Abu-Hammad, S.; Salim, N.A.; Al-Omoush, S. Denture stomatitis revisited: A summary of systematic reviews in the past decade and two case reports of papillary hyperplasia of unusual locations. Int. J. Dent. 2021, 2021, 7338143. [Google Scholar] [CrossRef] [PubMed]
- Jewtuchowicz, V.M.; Brusca, M.I.; Mujica, M.T.; Gliosca, L.A.; Finquelievich, J.L.; Lovannitti, C.A.; Rosa, A.C. Subgingival distribution of yeast and their antifungal susceptibility in immunocompetent subjects with and without dental devices. Acta Odontol. Latinoam. 2007, 20, 17–22. [Google Scholar]
- de Melo Costa, D.; Kipnis, A.; de Oliveira Leão-Vasconcelos, L.S.N.; Rocha-Vilefort, L.O.; Telles, S.A.; André, M.C.D.P.B.; Tipple, A.F.V.; Lima, A.B.M.; Ribeiro, N.F.G.; Pereira, M.R. Methicillin-resistant Staphylococcus sp. colonizing health care workers of a cancer hospital. Braz. J. Microbiol. 2014, 45, 799–805. [Google Scholar] [CrossRef]
- Lewis, N.; Parmar, N.; Hussain, Z.; Baker, G.; Green, I.; Howlett, J.; Kearns, A.; Cookson, B.; McDonald, A.; Wilson, M.; et al. Colonisation of dentures by Staphylococcus aureus and MRSA in out-patient and in-patient populations. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 1823–1826. [Google Scholar] [CrossRef] [PubMed]
- Chaulagain, R.; Thakur, S.N.; Khanal, B.; Sapkota, S.M.; Chaudhary, N.K. Microbial Spectrum of Complete Denture Wearer in Old Age People of Chitwan. J. Nepal Health Res. Counc. 2023, 21, 122–128. [Google Scholar] [CrossRef]
- Nishihama, S.; Kawada-Matsuo, M.; Le, M.N.-T.; Fujii, A.; Haruta, A.; Kajihara, T.; Hashimoto, Y.; Yoshikawa, M.; Aikawa, T.; Tsuga, K.; et al. Oral colonization of antimicrobial-resistant bacteria in home health care participants and their association with oral and systemic status. Sci. Rep. 2025, 15, 5776. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Archilla, A.; Garcia-Galan, C. Etiological factors related to denture stomatitis: A meta-analysis. Dent. Med. Res. 2020, 8, 37–42. [Google Scholar] [CrossRef]
- Martori, E.; Ayuso-Montero, R.; Martinez-Gomis, J.; Viñas, M.; Peraire, M. Risk factors for denture-related oral mucosal lesions in a geriatric population. J. Prosthet. Dent. 2014, 111, 273–279. [Google Scholar] [CrossRef]
- RB, O.; Khoder, G.; Fayed, B.; RA, K.; Elkareimi, Y.; Alharbi, N.J.P. Influence of fabrication technique on adhesion and biofilm formation of Candida albicans to conventional, milled, and 3D-printed denture base resin materials: A comparative in vitro study. Polymers 2023, 15, 1836. [Google Scholar] [CrossRef]
- Zheng, S.; Bawazir, M.; Dhall, A.; Kim, H.-E.; He, L.; Heo, J.; Hwang, G. Implication of surface properties, bacterial motility, and hydrodynamic conditions on bacterial surface sensing and their initial adhesion. Front. Bioeng. Biotechnol. 2021, 9, 643722. [Google Scholar] [CrossRef] [PubMed]
- Cankovic, M.; Bokor-Bratic, M.; Marinoski, J.; Stojanovic, D. Prevalence and possible predictors of the occurence of denture stomatitis in patients older than 60 years. Vojn. Pregl. 2017, 74, 311–316. [Google Scholar] [CrossRef]
- Emami, E.; de Grandmont, P.; Rompré, P.; Barbeau, J.; Pan, S.; Feine, J. Favoring Trauma as an Etiological Factor in Denture Stomatitis. J. Dent. Res. 2008, 87, 440–444. [Google Scholar] [CrossRef]
- Duyck, J.; Vandamme, K.; Krausch-Hofmann, S.; Boon, L.; De Keersmaecker, K.; Jalon, E.; Teughels, W.; Bencharit, S. Impact of denture cleaning method and overnight storage condition on denture biofilm mass and composition: A cross-over randomized clinical trial. PLoS ONE 2016, 11, e0145837. [Google Scholar] [CrossRef] [PubMed]
- Ben-Ami, R.; Olshtain-Pops, K.; Krieger, M.; Oren, I.; Bishara, J.; Dan, M.; Wiener-Well, Y.; Weinberger, M.; Zimhony, O.; Chowers, M.; et al. Antibiotic exposure as a risk factor for fluconazole-resistant candida bloodstream infection. Antimicrob. Agents Chemother. 2012, 56, 2518–2523. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Sae-Tia, S.; Fries, B.C. Candidiasis and mechanisms of antifungal resistance. Antibiotics 2020, 9, 312. [Google Scholar] [CrossRef]
- Twigg, J.A.; Smith, A.; Haury, C.; Wilson, M.J.; Lees, J.; Waters, M.; Williams, D.W. Compositional shifts within the denture-associated bacteriome in pneumonia—An analytical cross-sectional study. J. Med. Microbiol. 2023, 72, 001702. [Google Scholar] [CrossRef]
- Mhatre, S.; Srichand, R.; Sethumadhavan, J.; Mishra, P.B.; Patil, S.D.; Chavan, R.S.; Joshi, M.; Shetty, U. Dry mouth dilemma: A comprehensive review of xerostomia in complete denture wearers. Cureus 2024, 16, e58564. [Google Scholar] [CrossRef]
- Ichigaya, N.; Kawanishi, N.; Adachi, T.; Sugimoto, M.; Kimoto, K.; Hoshi, N. Effects of Denture Treatment on Salivary Metabolites: A Pilot Study. Int. J. Mol. Sci. 2023, 24, 13959. [Google Scholar] [CrossRef]
- Devcic, M.K.; Simonic-Kocijan, S.; Prpic, J.; Paskovic, I.; Cabov, T.; Kovac, Z.; Glazar, I. Oral candidal colonization in patients with different prosthetic appliances. J. Fungi 2021, 7, 662. [Google Scholar] [CrossRef] [PubMed]
- Quindos, G.; Gil-Alonso, S.; Marcos-Arias, C.; Sevillano, E.; Mateo, E.; Jauregizar, N.; Eraso, E. Therapeutic tools for oral candidiasis: Current and new antifungal drugs. Med. Oral Patol. Oral Cirugia Bucal 2019, 24, e172. [Google Scholar] [CrossRef]
- Lewis, M.; Williams, D.W. Diagnosis and management of oral candidosis. Br. Dent. J. 2017, 223, 675–681. [Google Scholar] [CrossRef]
- Xiao, Y.; Yuan, P.; Sun, Y.; Xu, Y.; Deng, X.; Wang, X.; Liu, R.; Chen, Q.; Jiang, L. Comparison of topical antifungal agents for oral candidiasis treatment: A systematic review and meta-analysis. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2022, 133, 282–291. [Google Scholar] [CrossRef]
- Abuhajar, E.; Ali, K.; Zulfiqar, G.; Al Ansari, K.; Raja, H.Z.; Bishti, S.; Anweigi, L. Management of chronic atrophic candidiasis (denture stomatitis)—A narrative review. Int. J. Environ. Res. Public Health 2023, 20, 3029. [Google Scholar] [CrossRef]
- Rai, A.; Misra, S.R.; Panda, S.; Sokolowski, G.; Mishra, L.; Das, R.; Lapinska, B. Nystatin effectiveness in Oral candidiasis treatment: A Systematic Review & Meta-Analysis of clinical trials. Life 2022, 12, 1677. [Google Scholar] [CrossRef]
- Matteis, D.V.; Cascione, M.; Toma, C.C.; Albanese, G.; De Giorgi, M.L.; Corsalini, M.; Rinaldi, R. Silver nanoparticles addition in poly(methyl methacrylate) dental matrix: Topographic and antimycotic studies. Int. J. Mol. Sci. 2019, 20, 4691. [Google Scholar] [CrossRef] [PubMed]
- Aoun, G.; Cassia, A. Effectiveness of a Chlorhexidine Digluconate 0.12% and Cetylpyridinium Chloride 0.05% Solution in eliminating Candida albicans Colonizing Dentures: A Randomized Clinical in vivo Study. J. Contemp. Dent. Pract. 2015, 16, 433–436. [Google Scholar] [PubMed]
- Procópio, A.L.F.; Lara, V.S.; Porto, V.C.; Soares, S.; Fernandes, M.H.; Urban, V.M.; Neppelenbroek, K.H. Resilient liner modified by antimicrobials for denture stomatitis treatment: A randomized controlled trial. J. Dent. 2022, 126, 104297. [Google Scholar] [CrossRef] [PubMed]
- Mendes, L.V.; Marques, M.B.e.S.; Teixeira, A.B.V.; de Castro, D.T. Incorporation of antimicrobial materials into denture adhesives: A systematic review of in vitro studies. Int. J. Adhes. Adhes. 2025, 140, 104022. [Google Scholar] [CrossRef]
- Wang, J.; Lu, X.; Wang, C.; Yue, Y.; Wei, B.; Zhang, H.; Wang, H.; Chen, J. Research progress on the combination of quorum-sensing inhibitors and antibiotics against bacterial resistance. Molecules 2024, 29, 1674. [Google Scholar] [CrossRef]
- Zhao, X.; Yu, Z.; Ding, T. Quorum-sensing regulation of antimicrobial resistance in bacteria. Microorganisms 2020, 8, 425. [Google Scholar] [CrossRef]
- Wang, S.; Zhao, Y.; Breslawec, A.P.; Liang, T.; Deng, Z.; Kuperman, L.L.; Yu, Q. Strategy to combat biofilms: A focus on biofilm dispersal enzymes. npj Biofilms Microbiomes 2023, 9, 63. [Google Scholar] [CrossRef]
- Sharma, A.; Gupta, V.K.; Pathania, R. Efflux pump inhibitors for bacterial pathogens: From bench to bedside. Indian J. Med. Res. 2019, 149, 129–145. [Google Scholar] [CrossRef]
- Compagnoni, M.A.; Souza, R.F.; Marra, J.; Pero, A.C.; Barbosa, D.B. Relationship between Candida and nocturnal denture wear: Quantitative study. J. Oral Rehabil. 2007, 34, 600–605. [Google Scholar] [CrossRef] [PubMed]
- Aslam, A.; Gajdács, M.; Zin, C.S.; Ab Rahman, N.S.; Ahmed, S.I.; Zafar, M.Z.; Jamshed, S. Evidence of the practice of self-medication with antibiotics among the lay public in low- and middle-income countries: A scoping review. Antibiotics 2020, 9, 597. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.Y.; Prentice, E.L.; Webber, M.A. Mechanisms of antimicrobial resistance in biofilms. npj Antimicrob. Resist. 2024, 2, 27. [Google Scholar] [CrossRef] [PubMed]
- AOAd, F.; Marquezan, M.; MdCG, N.; DS, A.; Maia, L.C. The influence of orthodontic fixed appliances on the oral microbiota: A systematic review. Dent. Press J. Orthod. 2014, 19, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Mulimani, P.; Popowics, T. Effect of orthodontic appliances on the oral environment and microbiome. Front. Dent. Med. 2022, 3, 924835. [Google Scholar] [CrossRef]
- Lucchese, A.; Bondemark, L.; Marcolina, M.; Manuelli, M. Changes in oral microbiota due to orthodontic appliances: A systematic review. J. Oral Microbiol. 2018, 10, 1476645. [Google Scholar] [CrossRef]
- Hamdoon, S.M.; AlSamak, S.; Ahmed, M.K.; Gasgoos, S. Evaluation of biofilm formation on different clear orthodontic retainer materials. J. Orthod. Sci. 2022, 11, 34. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, L.E.A.G.D.; Pithon, M.M.; dos Santos, R.L.; Freitas, A.O.A.; Alviano, D.S.; Nojima, L.I.; Nojima, M.C.G.; Ruellas, A.C.d.O. Colonization of Streptococcus mutans on esthetic brackets: Self-ligating vs. conventional. Am. J. Orthod. Dentofac. Orthop. 2013, 143, S72–S77. [Google Scholar] [CrossRef]
- Pellissari, B.A.; Sabino, G.S.P.; Lima, R.N.d.S.; Motta, R.H.L.; Suzuki, S.S.; Garcez, A.S.; Basting, R.T.; Barbosa, J.A.; Montalli, V.A.M. Antimicrobial resistance of bacterial strains in patients undergoing orthodontic treatment with and without fixed appliances. Angle Orthod. 2021, 91, 672–679. [Google Scholar] [CrossRef] [PubMed]
- Sabino, G.; Suzuki, S.; Segundo, A.G.; Czezacki, A.; Pugliesi, T.; Saba, G.; Almeida, T.; Caselato, I.; Montalli, V. Prevalence of drug-resistant microorganisms in oral cavity during orthodontic treatment. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2019, 128, e62. [Google Scholar] [CrossRef]
- Lazar, L.; Vlasa, A.; Beresescu, L.; Bud, A.; Lazar, A.P.; Matei, L.; Bud, E. White spot lesions (WSLs)—Post-orthodontic occurrence, management and treatment alternatives: A narrative review. J. Clin. Med. 2023, 12, 1908. [Google Scholar] [CrossRef] [PubMed]
- Papageorgiou, S.N.; Papadelli, A.A.; Eliades, T. Effect of orthodontic treatment on periodontal clinical attachment: A systematic review and meta-analysis. Eur. J. Orthod. 2017, 40, 176–194. [Google Scholar] [CrossRef]
- Verrusio, C.; Iorio-Siciliano, V.; Blasi, A.; Leuci, S.; Adamo, D.; Nicolò, M. The effect of orthodontic treatment on periodontal tissue inflammation: A systematic review. Quintessence Int. 2018, 49, 69–77. [Google Scholar]
- Abutayyem, H.; Alshehhi, M.A.; Alameri, M.; Zafar, M.S. Microbial adhesion on different types of orthodontic brackets and wires: An in vitro study. Saudi Dent. J. 2024, 36, 1459–1465. [Google Scholar] [CrossRef]
- Pavlic, A.; Begic, G.; Tota, M.; Abram, M.; Spalj, S.; Gobin, I. Bacterial exposure to nickel: Influence on adhesion and biofilm formation on orthodontic archwires and sensitivity to antimicrobial agents. Materials 2021, 14, 4603. [Google Scholar] [CrossRef] [PubMed]
- Topolska, J.M.; Jagielska, A.; Motyl, S.; Kozub-Budzyń, G.A.; Kępa, L.; Wagner, B.; Wątor, K. Metal leakage from orthodontic appliances chemically alters enamel surface during experimental in vitro simulated treatment. Sci. Rep. 2024, 14, 5412. [Google Scholar] [CrossRef]
- Mei, L.; Chieng, J.; Wong, C.; Benic, G.; Farella, M. Factors affecting dental biofilm in patients wearing fixed orthodontic appliances. Prog. Orthod. 2017, 18, 4. [Google Scholar] [CrossRef] [PubMed]
- Kaluarachchi, H.; Chung, K.C.C.; Zamble, D.B. Microbial nickel proteins. Nat. Prod. Rep. 2010, 27, 681–694. [Google Scholar] [CrossRef]
- Macomber, L.; Hausinger, R.P. Mechanisms of nickel toxicity in microorganisms. Metallomics 2011, 3, 1153–1162. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.-W.; Wang, J.-T.; Li, J.; Shi, X.-Z.; Ma, Y.-B.; Chen, D.; He, J.-Z. Long-term nickel contamination increases the occurrence of antibiotic resistance genes in agricultural soils. Environ. Sci. Technol. 2016, 51, 790–800. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Li, X.; Sun, G.; Zhang, Y.; Su, J.; Ye, J. Heavy metal induced antibiotic resistance in bacterium LSJC7. Int. J. Mol. Sci. 2015, 16, 23390–23404. [Google Scholar] [CrossRef]
- Abdelatey, L.M.; Khalil, W.K.; Ali, T.H.; Mahrous, K.F. Heavy Metal Resistance and Gene Expression Analysis of Metal Resistance Genes in Gram-Positive and Gram-Negative Bacteria Present in Egyptian Soils. J. Appl. Sci. Environ. Sanit. 2011, 6, 201–211. [Google Scholar]
- Dittmer, M.P.; Hellemann, C.F.; Grade, S.; Heuer, W.; Stiesch, M.; Schwestka-Polly, R.; Demling, A.P. Comparative three-dimensional analysis of initial biofilm formation on three orthodontic bracket materials. Head Face Med. 2015, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Asghar, F.; Bano, A.; Waheed, F.; Anjum, A.A.; Ejaz, H.; Javed, N. Association of exogenous factors with molecular epidemiology of Staphylococcus aureus in human oral cavity. Saudi J. Biol. Sci. 2023, 30, 103613. [Google Scholar] [CrossRef]
- Poeta, P.; Igrejas, G.; Gonçalves, A.; Martins, E.; Araújo, C.; Carvalho, C.; Rodrigues, J.; Vinué, L.; López, M.; Torres, C. Influence of oral hygiene in patients with fixed appliances in the oral carriage of antimicrobial-resistant Escherichia coli and Enterococcus isolates. Oral. Surg. Oral Med. Oral Pathol. Oral Radiol. Endodontol. 2009, 108, 557–564. [Google Scholar] [CrossRef]
- Barylyak, A.; Kisała, J.; Fornal, P.; Bobitski, Y.; Dziedzic, A.; Kus-Liśkiewicz, M. Physical, mechanical and antibacterial properties of the orthodontic adhesive doped with S-TiO2. Sci. Rep. 2024, 14, 17862. [Google Scholar] [CrossRef]
- Murphy, A.M.; Patel, U.C.; Wilson, G.M.; Suda, K.J. Prevalence of unnecessary antibiotic prescriptions among dental visits, 2019. Infect. Control Hosp. Epidemiol. 2024, 45, 890–899. [Google Scholar] [CrossRef]
- Okuwaki, S.; Hosomichi, K.; Tajima, A.; Yamaguchi, T. An Observational Study on Changes in the Oral and Gut Microbiota through Professional Mechanical Tooth Cleaning, including Tooth-Brushing Instructions in Patients with Multi-Bracket Appliances. Appl. Sci. 2023, 13, 10843. [Google Scholar] [CrossRef]
- Sycińska-Dziarnowska, M.; Szyszka-Sommerfeld, L.; Ziąbka, M.; Spagnuolo, G.; Woźniak, K. Use of antimicrobial silver coatings on fixed orthodontic appliances, including archwires, brackets, and microimplants: A systematic review. Med. Sci. Monit. 2024, 30, e944255-1. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, O.M.; Sedky, Y.; Shebl, H.R. Comprehensive evaluation of the antibacterial and antibiofilm activities of NiTi orthodontic wires coated with silver nanoparticles and nanocomposites: An in vitro study. BMC Oral Health 2024, 24, 1345. [Google Scholar] [CrossRef] [PubMed]
- Kameda, T.; Sato, H.; Oka, S.; Miyazaki, A.; Ohkuma, K.; Terada, K. Low temperature polytetrafluoroethylene (PTFE) coating improves the appearance of orthodontic wires without changing their mechanical properties. Dent. Mater. J. 2020, 39, 721–734. [Google Scholar] [CrossRef] [PubMed]
- Raji, S.H.; Shojaei, H.; Ghorani, P.S.; Rafiei, E. Bacterial colonization on coated and uncoated orthodontic wires: A prospective clinical trial. Dent. Res. J. 2014, 11, 680–683. [Google Scholar]
- Selvaraj, A.; George, A.M.; Rajeshkumar, S. Efficacy of zirconium oxide nanoparticles coated on stainless steel and nickel titanium wires in orthodontic treatment. Bioinformation 2021, 17, 760. [Google Scholar] [CrossRef]
- Thabit, A.K.; Aljereb, N.M.; Khojah, O.M.; Shanab, H.; Badahdah, A. Towards Wiser Prescribing of Antibiotics in Dental Practice: What Pharmacists Want Dentists to Know. Dent. J. 2024, 12, 345. [Google Scholar] [CrossRef]
- Lockhart, P.B.; Tampi, M.P.; Abt, E.; Aminoshariae, A.; Durkin, M.J.; Fouad, A.F.; Gopal, P.; Hatten, B.W.; Kennedy, E.; Lang, M.S. Evidence-based clinical practice guideline on antibiotic use for the urgent management of pulpal-and periapical-related dental pain and intraoral swelling: A report from the American Dental Association. J. Am. Dent. Assoc. 2019, 150, 906–921.e12. [Google Scholar] [CrossRef] [PubMed]
- Bansal, R.; Jain, A.; Goyal, M.; Singh, T.; Sood, H.; Malviya, H.S. Antibiotic abuse during endodontic treatment: A contributing factor to antibiotic resistance. J. Fam. Med. Prim. Care 2019, 8, 3518–3524. [Google Scholar] [CrossRef] [PubMed]
- Sperotto, F.; France, K.; Gobbo, M.; Bindakhil, M.; Pimolbutr, K.; Holmes, H.; Monteiro, L.; Graham, L.; Hong, C.H.L.; Sollecito, T.P.; et al. Antibiotic prophylaxis and infective endocarditis incidence following invasive dental procedures: A systematic review and meta-analysis. JAMA Cardiol. 2024, 9, 599–610. [Google Scholar] [CrossRef] [PubMed]
- Thornhill, M.H.; Gibson, T.B.; Durkin, M.J.; Dayer, M.J.; Lockhart, P.B.; O’Gara, P.T.; Baddour, L.M. Prescribing of antibiotic prophylaxis to prevent infective endocarditis. J. Am. Dent. Assoc. 2020, 151, 835–845.e31. [Google Scholar] [CrossRef]
- Fluent, M.T.; Jacobsen, P.L.; Hicks, L.A. Considerations for responsible antibiotic use in dentistry. J. Am. Dent. Assoc. 2016, 147, 683–686. [Google Scholar] [CrossRef] [PubMed]
- Cooper, L.; Sneddon, J.; Thompson, W.; Guise, T.; Robertson, D.; Smith, A. Tackling antimicrobial resistance in practice: Dental students’ evaluation of university teaching supplemented by an online course. JAC-Antimicrob. Resist. 2022, 4, dlac039. [Google Scholar] [CrossRef]
- Castro-Sánchez, E.; Drumright, L.N.; Gharbi, M.; Farrell, S.; Holmes, A.H.; Panepinto, J. Mapping antimicrobial stewardship in undergraduate medical, dental, pharmacy, nursing and veterinary education in the United Kingdom. PLoS ONE 2016, 11, e0150056. [Google Scholar] [CrossRef] [PubMed]
- Teoh, L.; Thompson, W.; Suda, K. Antimicrobial stewardship in dental practice. J. Am. Dent. Assoc. 2020, 151, 589–595. [Google Scholar] [CrossRef]
- Martine, C.; Sutherland, S.; Born, K.; Thompson, W.; Teoh, L.; Singhal, S. Dental antimicrobial stewardship: A qualitative study of perspectives among Canadian dentistry sector leaders and experts in antimicrobial stewardship. JAC-Antimicrob. Resist. 2024, 6, dlae082. [Google Scholar] [CrossRef] [PubMed]
- Niemczyk, W.; Żurek, J.; Niemczyk, S.; Kępa, M.; Zięba, N.; Misiołek, M.; Wiench, R. Antibiotic-Loaded Platelet-Rich Fibrin (AL-PRF) as a New Carrier for Antimicrobials: A Systematic Review of In Vitro Studies. Int. J. Mol. Sci. 2025, 26, 2140. [Google Scholar] [CrossRef]
| Feature | Braces | Retainers | References |
|---|---|---|---|
| Dominant Microorganisms | Higher abundance of Streptococcus mutans, Lactobacillus spp., and anaerobes | Candida spp. more prevalent, alongside S. mutans and mixed anaerobes | [101,175,181,190,191] |
| Bacterial Load | Significantly increased compared to non-orthodontic patients | Varies by retainer type; removable retainers may have lower loads than fixed ones | [101,175,181,190,191] |
| Biofilm Characteristics | Thicker and more mature biofilms due to brackets and wires creating retention sites | Biofilms can form on both acrylic and wire components; removable types allow better cleaning | [101,175,181,190,191] |
| Health Implications | Increased risk of enamel demineralization, white spot lesions, and gingivitis | Potential for oral candidiasis, increased caries risk, and periodontal inflammation | [101,175,181,190,191] |
| Pathogen | Location | AMR Profile | Reference |
|---|---|---|---|
| M. chelonae, M. abscessus, M. lentiflavum, M. fortuitum, M. kansasii, M. simiae | DUWLs | High resistance rates were observed to trimethoprim-sulfamethoxazole, doxycycline, imipenem, meropenem, and ciprofloxacin. Moderate resistance to moxifloxacin, cefoxitin, and clarithromycin was observed. In contrast, lower resistance rates were noted for streptomycin, ethambutol, rifampin, and isoniazid. | [75] |
| P. aeruginosa | DUWLs | 30% of strains were resistant. The most common resistance was to colistin (21.4%), followed by piperacillin (5.7%) and ceftazidime (2.9%). Multi-drug resistance included combinations such as levofloxacin-netilmicin-colistin and piperacillin-ceftazidime-colistin. | [47] |
| P. aeruginosa | DUWLs | All isolates were sensitive to all tested antibiotics, including piperacillin, tazobactam, ceftazidime, cefepime, aztreonam, imipenem, meropenem, colistin, polymyxin B, gentamicin, tobramycin, amikacin, and ciprofloxacin. | [54] |
| P. aeruginosa | DUWLs | Isolates lacked several ARGs commonly found in clinical strains, such as PDC-3, PDC-5, PDC-8, and triC. The most basal isolates carried additional resistance-related genes, including golS, mdsA, ceoB, and mdsC. | [76] |
| L. pneumophila | DUWLs | The isolates were 100% resistant to multiple antibiotics, including erythromycin, lincomycin, gentamicin, and several β-lactams. High resistance was also seen to amoxicillin (88.8%), tetracycline (77.7%), and doxycycline and chloramphenicol (55.5%). Amikacin showed the lowest resistance. | [77] |
| P. aeruginosa P. fluorescens P. lutea P. putida Staphylococcus spp. | DUWLs | Pseudomonas spp. were susceptible to cefoperazone, ofloxacin, gentamicin, ciprofloxacin, and piperacillin but resistant to metronidazole and often to rifampicin and erythromycin. Staphylococcus spp. were also susceptible to those antibiotics, but all were resistant to metronidazole. Some strains, such as S. sciuri and S. epidermidis, were methicillin-resistant and multi-drug-resistant. | [79] |
| Pseudomonas spp. Burkholderia spp. | DUWLs | Pseudomonas spp. were highly susceptible to ceftazidime, meropenem, and ofloxacin but showed strong resistance to ampicillin. P. aeruginosa had only 18.2% susceptibility to ampicillin. Burkholderia spp. showed good susceptibility to ceftazidime and meropenem. However, they were highly resistant to ampicillin, gentamicin, and chloramphenicol. | [80] |
| S. aureus S. epidermidis | DUWLs | S. aureus and S. epidermidis showed high resistance to oxacillin and clindamycin. Both had moderate resistance to ampicillin, amoxicillin, and azithromycin. Ciprofloxacin and vancomycin were effective against all isolates. S. aureus was slightly more susceptible overall than S. epidermidis. | [81] |
| Streptococcus spp. Coagulase-negative Staphylococci, S. aureus E. coli K. pneumoniae P. aeruginosa | Dentures | Amikacin, nalidixic acid, and ciprofloxacin showed the highest sensitivity pattern, while cefixime and amoxicillin/clavulanic acid were the most resistant antibiotics. E. coli exhibited complete sensitivity to all tested antibiotics, while P. aeruginosa showed multi-drug resistance, especially against cotrimoxazole and nitrofurantoin. | [139] |
| Viridans streptococci, S. aureus K. pneumoniae, E. coli | Dentures | Viridans streptococci showed the highest resistance to amoxicillin-clavulanic acid (40%) and methicillin (35%), while S. aureus had 25% resistance to both amoxicillin and methicillin. K. pneumoniae exhibited the highest resistance to amoxicillin-clavulanic acid (40%) and 30% resistance to cefotaxime, gentamicin, and doxycycline; E. coli showed 50% resistance to amoxicillin-clavulanic acid and 30% to multiple other antibiotics. The presence of dense biofilms and extracellular matrices observed via SEM suggested these structural features contributed to the observed multi-drug resistance. | [120] |
| Staphylococcus spp. | Dentures | Isolates showed high resistance to penicillin (91.6%), fosfomycin (87.5%), and cefoxitin (62.5%), indicating widespread methicillin resistance. Moderate resistance was observed to erythromycin (55.5%), tetracycline (43%), and clindamycin (38.8%). Resistance to gentamicin (16.6%) and trimethoprim/sulfamethoxazole (25%) was lower, while all isolates remained susceptible to linezolid and vancomycin. Nearly half (48.6%) of the isolates were multi-drug resistant. | [114] |
| S. aureus | Dentures | MRSA was found on 1% of outpatients’ and 12% of inpatients’ dentures, with most isolates identified as the highly resistant EMRSA-15 strain. All MRSA isolates were resistant to β-lactam antibiotics, including methicillin, oxacillin, and cefoxitin. | [138] |
| Staphylococcus spp. | Dentures | S. aureus and coagulase-negative Staphylococci were isolated. Colonization by MRSE was significantly higher (42.9%) compared to non-denture wearers (16.9%). All MRSE isolates were identified as S. epidermidis, with 94.4% carrying the mecA gene. MRSE strains exhibited greater resistance to antibiotics such as mupirocin while remaining susceptible to dalfopristin/quinupristin and linezolid. | [137] |
| P. aeruginosa, Klebsiella spp. Enterobacter spp. Enterococcus spp. Staphylococcus spp. Streptococcus spp. | Dentures | Resistance was high to ampicillin, amoxicillin, cephalothin, and tetracycline. Carbapenems (imipenem and meropenem) and rifampin were the most effective antibiotics, although some resistance still occurred. | [121] |
| E. faecalis E. faecium E. coli M. morganii P. aeruginosa Klebsiella spp. | Dentures | High resistance was observed to ampicillin (57.4%) and tetracycline (36.5%), with notable β-lactamase production in 41.2% of isolates, particularly blaTEM, blaSHV, and blaCTX-M genes in Gram-negative rods. Tetracycline resistance genes were diverse, with tet(A) and tet(B) common in Gram-negatives and tet(K) and tet(M) dominant in enterococci. | [122] |
| C. albicans and non-albicans Candida isolates | Dentures | All C. albicans isolates were susceptible to amphotericin B, with only 5.6% and 7.0% showing resistance to fluconazole and itraconazole, respectively. Non-albicans species showed higher resistance: 18.4% were resistant to fluconazole and 10.2% to itraconazole, though around 80% remained susceptible to these drugs. Amphotericin B and 5-fluorocytosine were the most effective agents against all Candida isolates. | [127] |
| C. albicans | Dentures | No antifungal resistance was observed in any of the isolates. | [131] |
| C. albicans non-albicans Candida isolates | Dentures | Denture use was associated with a higher oral yeast load and increased resistance to antifungal agents, particularly miconazole and 5-fluorocytosine. Non-albicans species showed higher resistance rates to fluconazole (29%), miconazole (35%), and ketoconazole (23%) compared to C. albicans. | [129] |
| C. albicans | Dentures | C. albicans were highly resistant to fluconazole and miconazole, showing MICs ≥ 256 mg/L, which was over 1000 times higher than for planktonic cells (MIC 0.25 mg/L). Chlorhexidine was more effective, with biofilm inhibition at concentrations 8 times higher than its planktonic MIC (0.3% vs. 0.04%). Young biofilms (2–6 h) were significantly more susceptible to all agents, especially miconazole and chlorhexidine, compared to mature (72 h) biofilms. | [132] |
| Staphylococcus spp. | Dentures | S. epidermidis, S. hominis, and S. xylosus were found on dentures, and methicillin-sensitive S. aureus was recovered from only one denture, and MRSA was found on two partial dentures. No MRSA was recovered from complete dentures. | [126] |
| C. albicans | Dentures | Biofilm-grown C. albicans were significantly more resistant to amphotericin B, nystatin, chlorhexidine, and fluconazole compared to planktonic cells. Biofilm-associated C. albicans required up to 128–256 times higher concentrations to achieve the same metabolic inhibition. | [130] |
| C. albicans non-albicans Candida isolates | Dentures | Fluconazole resistance was observed in 3.2% of isolates, mainly involving C. tropicalis and one strain of C. albicans. Only two of the fluconazole-resistant strains also showed resistance to itraconazole. Clinical cure rates were high with fluconazole (97%), and itraconazole was effective in all fluconazole-resistant cases. | [124] |
| Methicillin-resistant S. aureus | Dentures | All isolates were confirmed to carry the mecA gene and had high oxacillin MICs (96–256 µg/mL). | [119] |
| C. albicans Streptococcus spp. Staphylococcus spp. Lactobacillus spp. | Dentures | Antifungal resistance was noted in Candida strains, with 3 resistant to amphotericin B, 4 to nystatin, and 1 resistant to both. Nystatin was the most effective treatment, though some cases required combination therapies due to drug resistance. | [128] |
| S. aureus | Orthodontic Appliances | 100% of isolates were resistant to oxacillin and cefoxitin, classifying them as MRSA. 61% were resistant to vancomycin. Genetic characterization revealed nuc gene (100%), mecA gene (55.6%), pvl gene (50%), agr gene (33.3%), and coa gene (55.6%). mecA/coa genes showed significant associations with male gender and smoking, as well as with antibiotic use and dietary patterns. The pvl gene showed association with mouth rinse use and buccal cavity inflammation. | [190] |
| S. aureus S. hominis Enterobacter cloacae complex, Klebsiella oxytoca P. aeruginosa | Orthodontic Appliances | 74% of isolated bacteria showed resistance. The most common resistance was to beta-lactam antibiotics (ampicillin, cefoxitin, and cefuroxime), followed by erythromycin, clindamycin, and tetracycline. | [175] |
| S. aureus | Orthodontic Appliances | Bacteria adapted to 250 µg/mL Ni2+ showed the highest adhesion, biofilm biomass, and resistance to some antibiotics. Bacteria adapted to 62.5–250 µg/mL Ni2+ showed increased resistance to gentamicin, benzylpenicillin, rifampicin, trimethoprim-sulfamethoxazole, moxifloxacin, cefoxitin, and linezolid. Strains adapted to higher Ni2+ concentrations (500–1000 µg/mL) were more susceptible. Increased resistance was associated with lower nickel adaptation, while higher adaptation reduced resistance and biofilm adhesion. | [181] |
| S. aureus | Orthodontic Appliances | Increased resistance was observed in isolates treated with NiTi and SS wires. Resistance in both original and mutated isolates to ceftazidime. Some mutated isolates showed resistance to bacitracin, erythromycin, ogmentin, and amikacin, especially after prolonged incubation (48–96 h). Most isolates remained sensitive to cefotaxime. Mutation induced by orthodontic wire exposure increased resistance over time. | [101] |
| E. faecalis E. faecium and E. coli | Orthodontic Appliances | Among the Enterococcus isolates, resistance was detected as follows: 100% to erythromycin (associated with the erm(B) gene), 75% to kanamycin (aph(3′)-IIIa0, 50% to tetracycline (tet(L) with or without tet(M)), 37% to streptomycin (ant(6)-Ia), and 12% to chloramphenicol (catA). One E. coli isolate exhibited a multi-drug resistant phenotype, carrying five resistance genes along with both class 1 and class 2 integrons. All Enterococcus strains produced gelatinase, and four harbored genes for enterocins L50A/B and P. The esp virulence gene was identified in one multi-drug resistant E. faecalis isolate. | [191] |
| C. albicans and non-albicans Candida isolates | Orthodontic Appliances and Dentures | All C. albicans, C. parapsilosis, C. tropicalis, and C. sake isolates were fully susceptible to both fluconazole and voriconazole. Among the non-albicans species, one C. dubliniensis isolate (17%) was resistant to both antifungals, while all C. guilliermondii were fully susceptible to voriconazole. | [136] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farzinnia, G.; Anvari, Y.; Siqueira, M.F. Antimicrobial Resistance in Oral Healthcare: A Growing Concern in Dentistry. Dent. J. 2025, 13, 391. https://doi.org/10.3390/dj13090391
Farzinnia G, Anvari Y, Siqueira MF. Antimicrobial Resistance in Oral Healthcare: A Growing Concern in Dentistry. Dentistry Journal. 2025; 13(9):391. https://doi.org/10.3390/dj13090391
Chicago/Turabian StyleFarzinnia, Golnoush, Yalda Anvari, and Michelle F. Siqueira. 2025. "Antimicrobial Resistance in Oral Healthcare: A Growing Concern in Dentistry" Dentistry Journal 13, no. 9: 391. https://doi.org/10.3390/dj13090391
APA StyleFarzinnia, G., Anvari, Y., & Siqueira, M. F. (2025). Antimicrobial Resistance in Oral Healthcare: A Growing Concern in Dentistry. Dentistry Journal, 13(9), 391. https://doi.org/10.3390/dj13090391







