Efficient Methods to Calculate Partial Sphere Surface Areas for a Higher Resolution Finite Volume Method for Diffusion-Reaction Systems in Biological Modeling
Abstract
1. Introduction
2. Methods
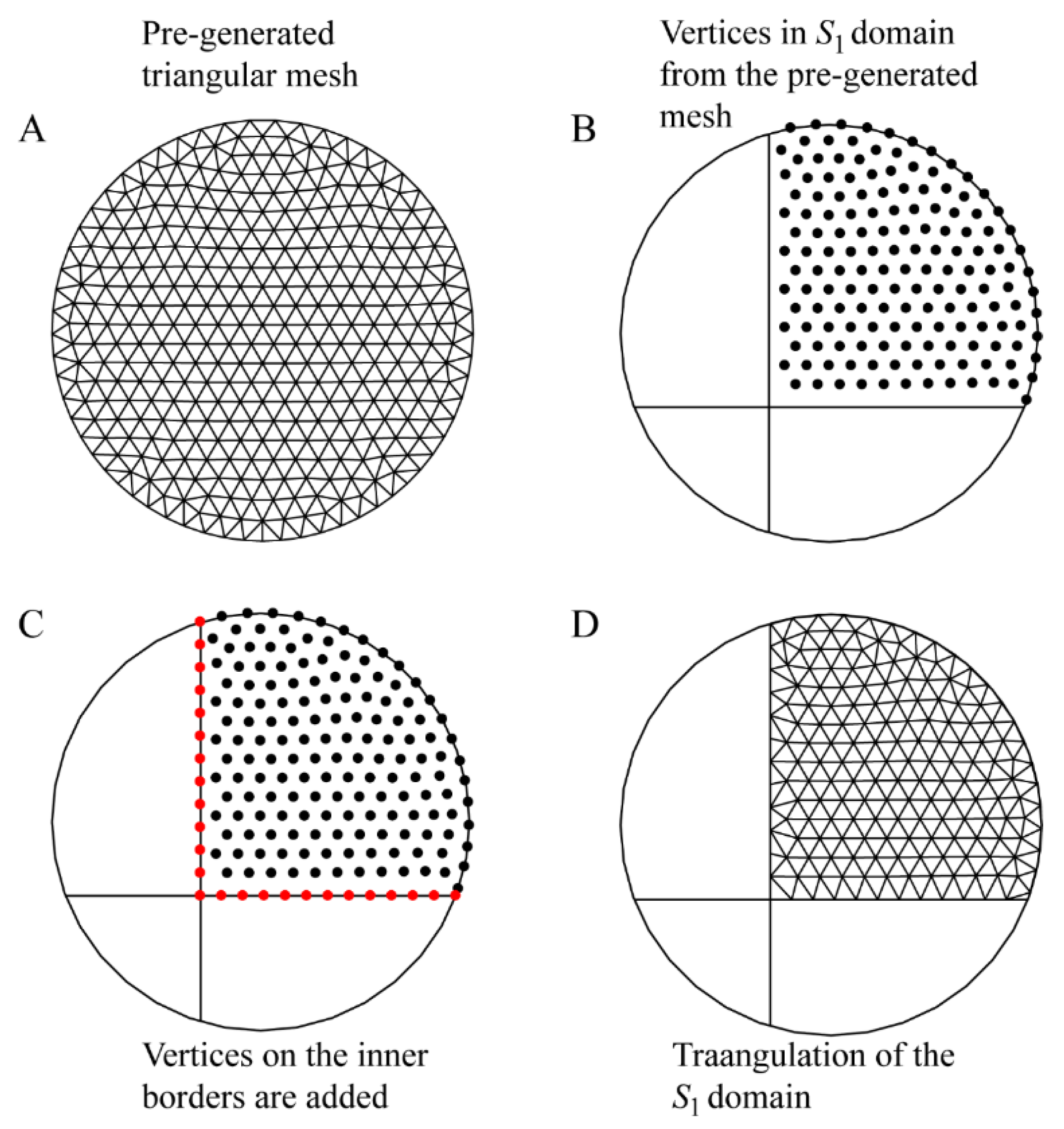
2.1. Surface Area Calculation Using Triangulation
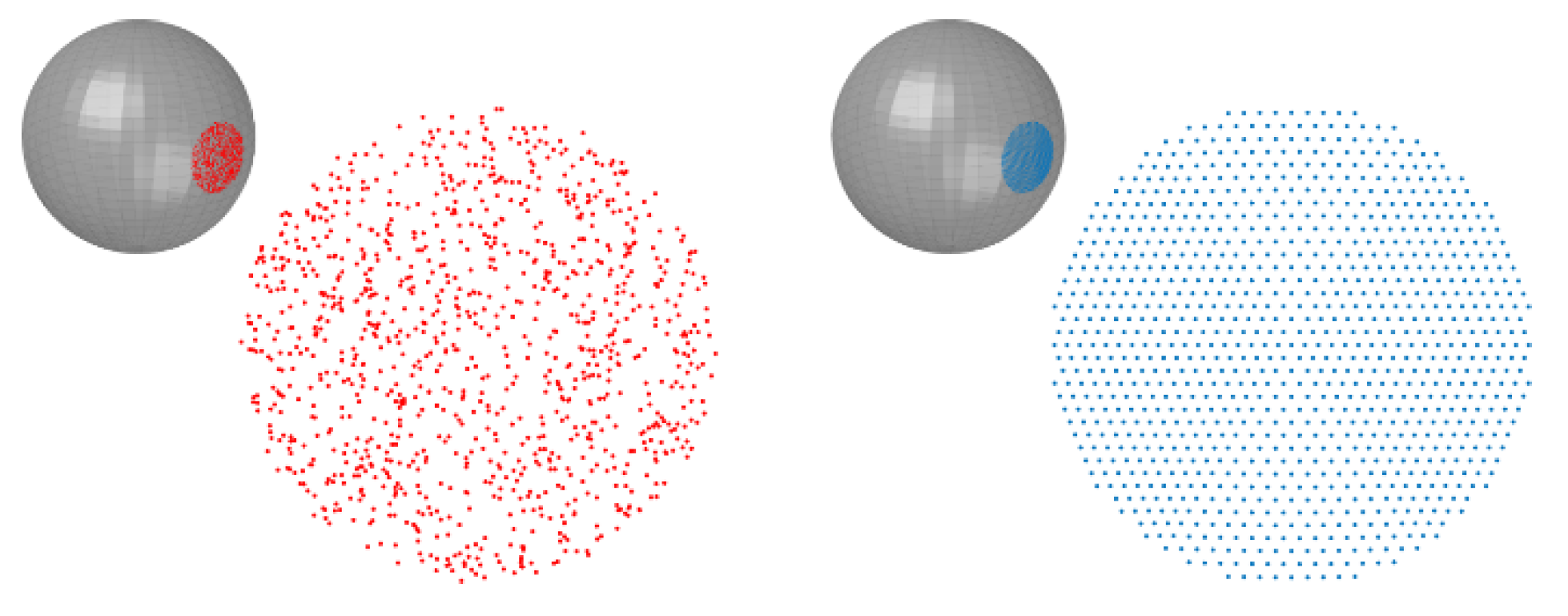
2.2. Monte Carlo Using Sample Points from a Uniform Distribution on Spheres
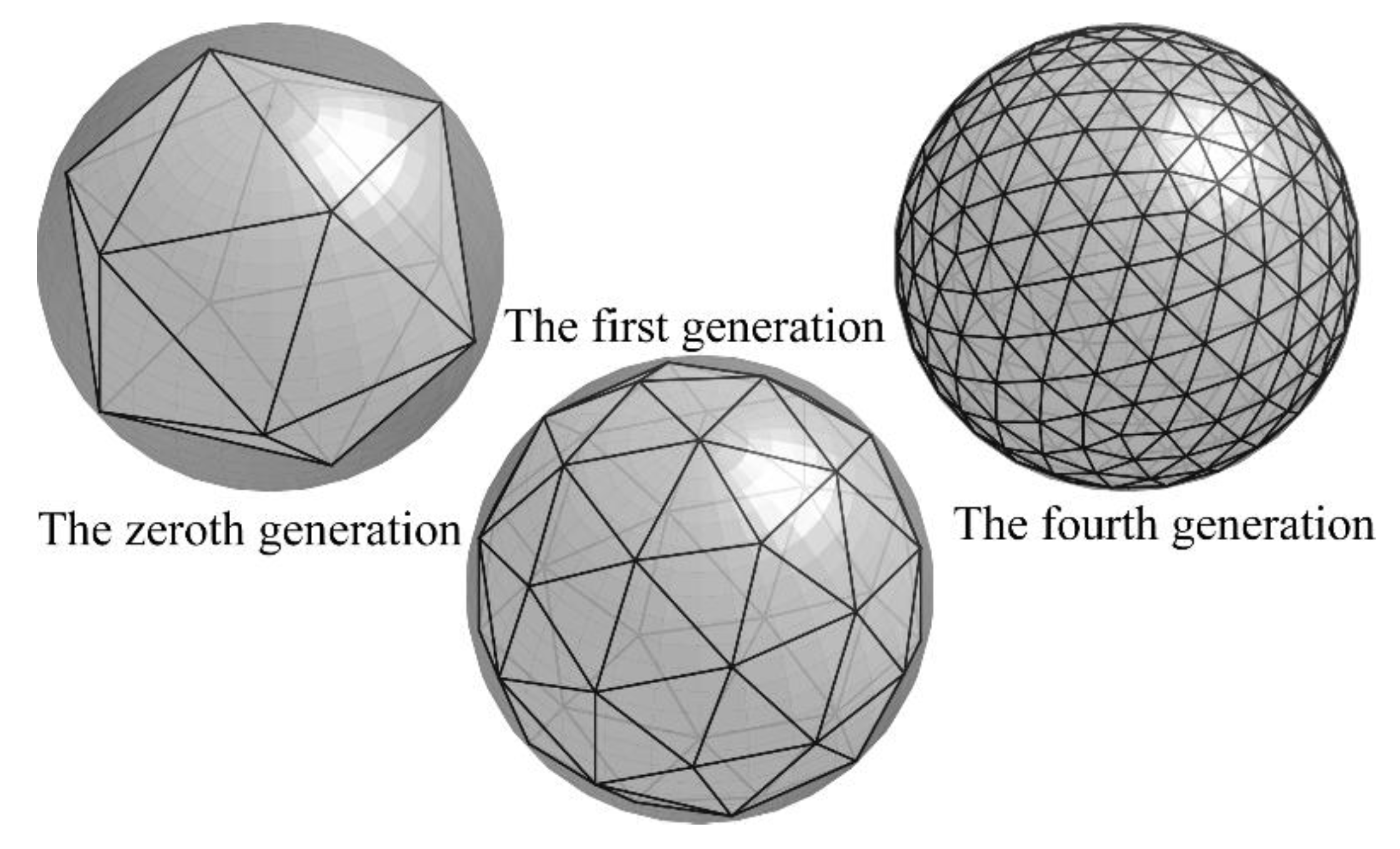
2.3. Monte Carlo Using the Icosahedron-Based Vertices on Spheres
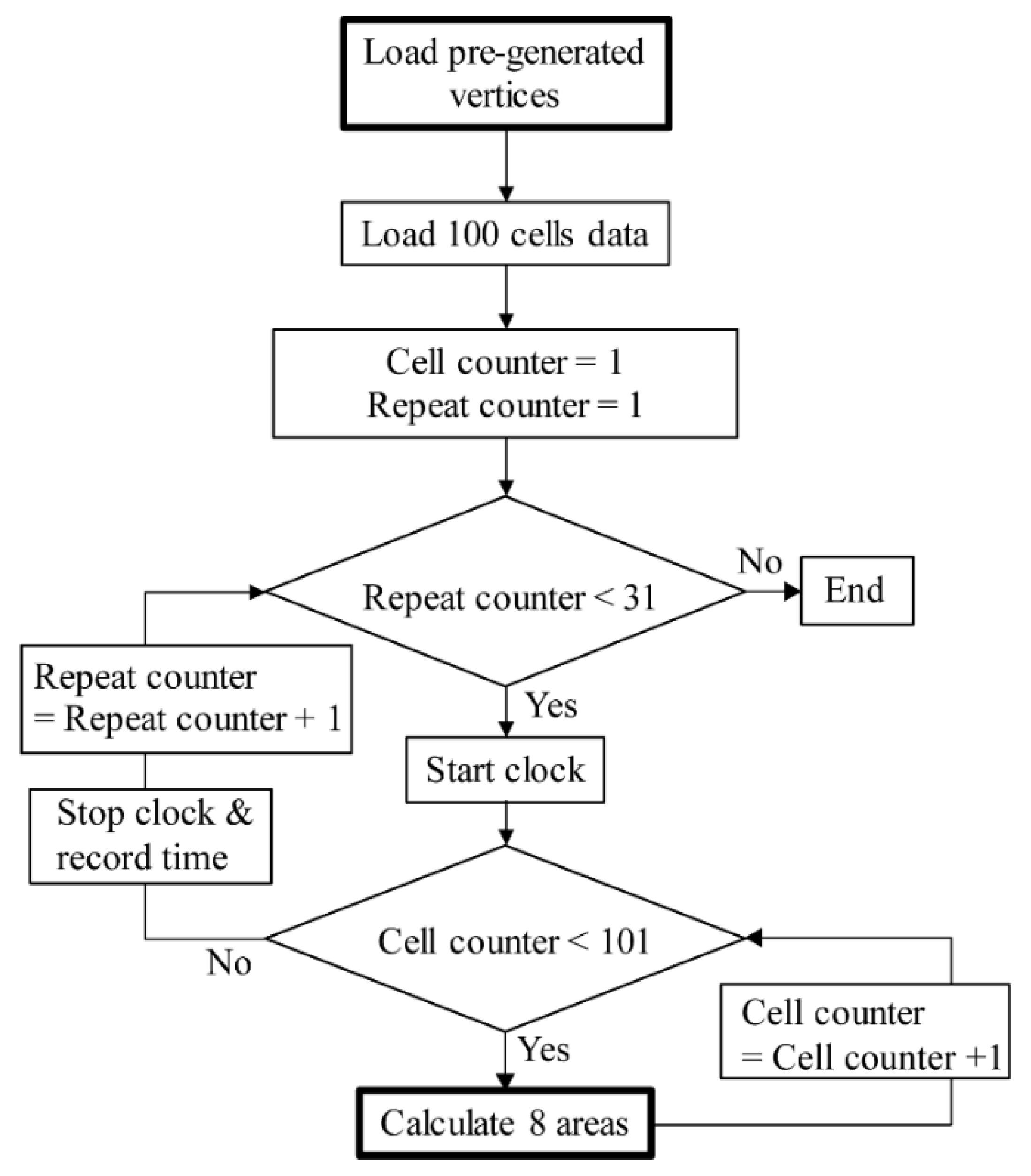
2.4. Implementations for Comparison
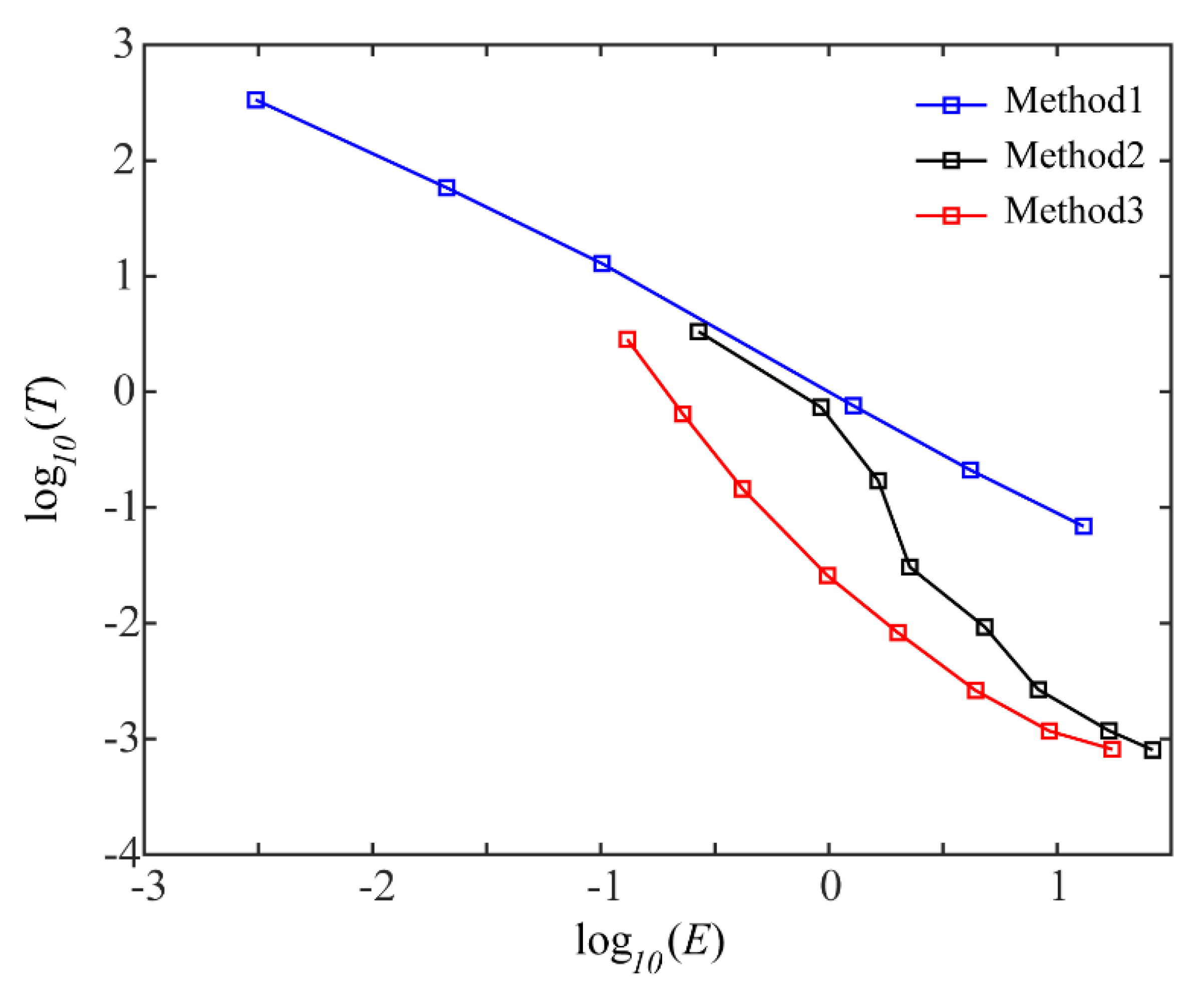
3. Results
4. Discussion
Author Contributions
Acknowledgments
Conflicts of Interest
Data Availability
References
- Rejniak, K.A. Homeostatic Imbalance in Epithelial Ducts and Its Role in Carcinogenesis. Scientifica 2012, 2012, 132978. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.J.; Reed, D.; Rejniak, K.A. The formation of tight tumor clusters affects the efficacy of cell cycle inhibitors: A hybrid model study. J. Theor. Biol. 2014, 352, 31–50. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kim, E.; Rebecca, V.; Fedorenko, I.V.; Messina, J.L.; Mathew, R.; Maria-Engler, S.S.; Basanta, D.; Smalley, K.S.M.; Anderson, A.R.A. Senescent fibroblasts in melanoma initiation and progression: An integrated theoretical, experimental, and clinical approach. Cancer Res. 2013, 73, 6874–6885. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Charypar, D.; Gross, M. Particle-Based Fluid Simulation for Interactive Applications. Available online: https://matthias-research.github.io/pages/publications/sca03.pdf (accessed on 23 December 2019).
- Auer, S. Realtime particle-based fluid simulation. Ph.D. Thesis, Technische Universität München, München, Germany, 2009. [Google Scholar]
- Goedhart, P.W.; Lof, M.E.; Bianchi, F.J.J.A.; Baveco, H.J.M.; van der Werf, W. Modelling mobile agent-based ecosystem services using kernel-weighted predictors. Methods Ecol. Evol. 2018, 9, 1241–1249. [Google Scholar] [CrossRef]
- McLane, A.J.; Semeniuk, C.; McDermid, G.J.; Marceau, D.J. The role of agent-based models in wildlife ecology and management. Ecol. Model. 2011, 222, 1544–1556. [Google Scholar] [CrossRef]
- Kennedy, W.G. Modelling Human Behaviour in Agent-Based Models. In Agent-Based Models of Geographical Systems; Springer: Dordrecht, The Netherlands, 2012; pp. 167–179. [Google Scholar]
- Bonabeau, E. Agent-based modeling: Methods and techniques for simulating human systems. Proc. Natl. Acad. Sci. USA 2002, 99, 7280–7287. [Google Scholar] [CrossRef] [PubMed]
- North, M.J.; Macal, C.M. Managing Business Complexity: Discovering Strategic Solutions with Agent-Based Modeling and Simulation; Oxford University Press: New York, NY, USA, 2007. [Google Scholar]
- Ghaffarizadeh, A.; Friedman, S.H.; MacKlin, P. BioFVM: An efficient, parallelized diffusive transport solver for 3-D biological simulations. Bioinformatics 2015, 32, 1256–1258. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.J.; Baek, S.; Jung, S.H.; Cho, K.H. Dynamical characteristics of bacteria clustering by self-generated attractants. Comput. Biol. Chem. 2007, 31, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.J.; Forbes, N.S. Single-cell analysis demonstrates how nutrient deprivation creates apoptotic and quiescent cell populations in tumor cylindroids. Biotechnol. Bioeng. 2008, 101, 797–810. [Google Scholar] [CrossRef] [PubMed]
- Persson, P.-O.; Strang, G. A Simple Mesh Generator in MATLAB. SIAM Rev. 2004, 46, 329–345. [Google Scholar] [CrossRef]
- Hicks, J.S.; Wheeling, R.F. An efficient method for generating uniformly distributed points on the surface of an n-dimensional sphere. Commun. ACM 1959, 2, 17–19. [Google Scholar] [CrossRef]
- Muller, M.E. A note on a method for generating points uniformly on n-dimensional spheres. Commun. ACM 1959, 2, 19–20. [Google Scholar] [CrossRef]
- Tashiro, Y. On methods for generating uniform random points on the surface of a sphere. Ann. Inst. Stat. Math. 1977, 29, 295–300. [Google Scholar] [CrossRef]
- Sibuya, M. A method for generating uniformly distributed points on N-dimensional spheres. Ann. Inst. Stat. Math. 1962, 14, 81–85. [Google Scholar] [CrossRef]
- Harman, R.; Lacko, V. On decompositional algorithms for uniform sampling from n-spheres and n-balls. J. Multivar. Anal. 2010, 1, 2297–2304. [Google Scholar] [CrossRef]
- Poland, J. Three Different Algorithms for Generating Uniformly Distributed Random Points on the N-Sphere. Available online: https://pdfs.semanticscholar.org/467c/634bc770002ad3d85ccfe05c31e981508669.pdf (accessed on 21 December 2019).
- Knuth, D.E. The Art of Computer Programming. Volume 2: Seminumerical Algorithms; Addison-Wesley: Boston, MA, USA, 1997. [Google Scholar]
- Baumgardner, J.R.; Frederickson, P.O. Icosahedral Discretization of the Two-Sphere. SIAM J. Numer. Anal. 1985, 22, 1107–1115. [Google Scholar] [CrossRef]
- Caflisch, R.E. Monte Carlo and quasi-Monte Carlo methods. Acta Numer. 1998, 7, 1–49. [Google Scholar] [CrossRef]
- Schürer, R. A comparison between (quasi-)Monte Carlo and cubature rule based methods for solving high-dimensional integration problems. Math. Comput. Simul. 2003, 62, 509–517. [Google Scholar] [CrossRef]
- Hislop, J.N.; Von Zastrow, M. Analysis of GPCR localization and trafficking. Methods Mol. Biol. 2011, 746, 425–440. [Google Scholar] [PubMed]
- Steagall, R.J.; Yao, F.; Shaikh, S.R.; Abdel-Rahman, A.A. Estrogen receptor α activation enhances its cell surface localization and improves myocardial redox status in ovariectomized rats. Life Sci. 2017, 182, 41–49. [Google Scholar] [CrossRef] [PubMed]

| Case Number | Triangulation | Uniform Distribution | Icosahedron-Based |
|---|---|---|---|
| 1 | 21 | 42 | 42 |
| 2 | 82 | 162 | 162 |
| 3 | 320 | 642 | 642 |
| 4 | 1243 | 2562 | 2562 |
| 5 | 5120 | 10,242 | 10,242 |
| 6 | 20,504 | 40,962 | 40,962 |
| 7 | N/A | 163,842 | 163,842 |
| 8 | N/A | 655,362 | 655,362 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bowers, A.; Bunn, J.; Kim, M. Efficient Methods to Calculate Partial Sphere Surface Areas for a Higher Resolution Finite Volume Method for Diffusion-Reaction Systems in Biological Modeling. Math. Comput. Appl. 2020, 25, 2. https://doi.org/10.3390/mca25010002
Bowers A, Bunn J, Kim M. Efficient Methods to Calculate Partial Sphere Surface Areas for a Higher Resolution Finite Volume Method for Diffusion-Reaction Systems in Biological Modeling. Mathematical and Computational Applications. 2020; 25(1):2. https://doi.org/10.3390/mca25010002
Chicago/Turabian StyleBowers, Abigail, Jared Bunn, and Myles Kim. 2020. "Efficient Methods to Calculate Partial Sphere Surface Areas for a Higher Resolution Finite Volume Method for Diffusion-Reaction Systems in Biological Modeling" Mathematical and Computational Applications 25, no. 1: 2. https://doi.org/10.3390/mca25010002
APA StyleBowers, A., Bunn, J., & Kim, M. (2020). Efficient Methods to Calculate Partial Sphere Surface Areas for a Higher Resolution Finite Volume Method for Diffusion-Reaction Systems in Biological Modeling. Mathematical and Computational Applications, 25(1), 2. https://doi.org/10.3390/mca25010002





