Trimeric SARS-CoV-2 Spike Proteins Produced from CHO Cells in Bioreactors Are High-Quality Antigens
Abstract
1. Introduction
2. Material and Methods
2.1. Design of SARS CoV-2 Spike Proteins
2.2. Expression and Purification of SARS-CoV-2 Proteins
2.3. Electron Microscopy Analysis of Trimeric Spikes
2.4. Inhibition of SARS-CoV-2 Infection by SARS-CoV-2 Proteins
2.5. Detection of SARS-CoV-2 Antibodies Using SARS-CoV-2 Proteins
3. Results and Discussion
3.1. Design and Selection of CHO Manufacturable SARS CoV-2 Trimeric Spikes
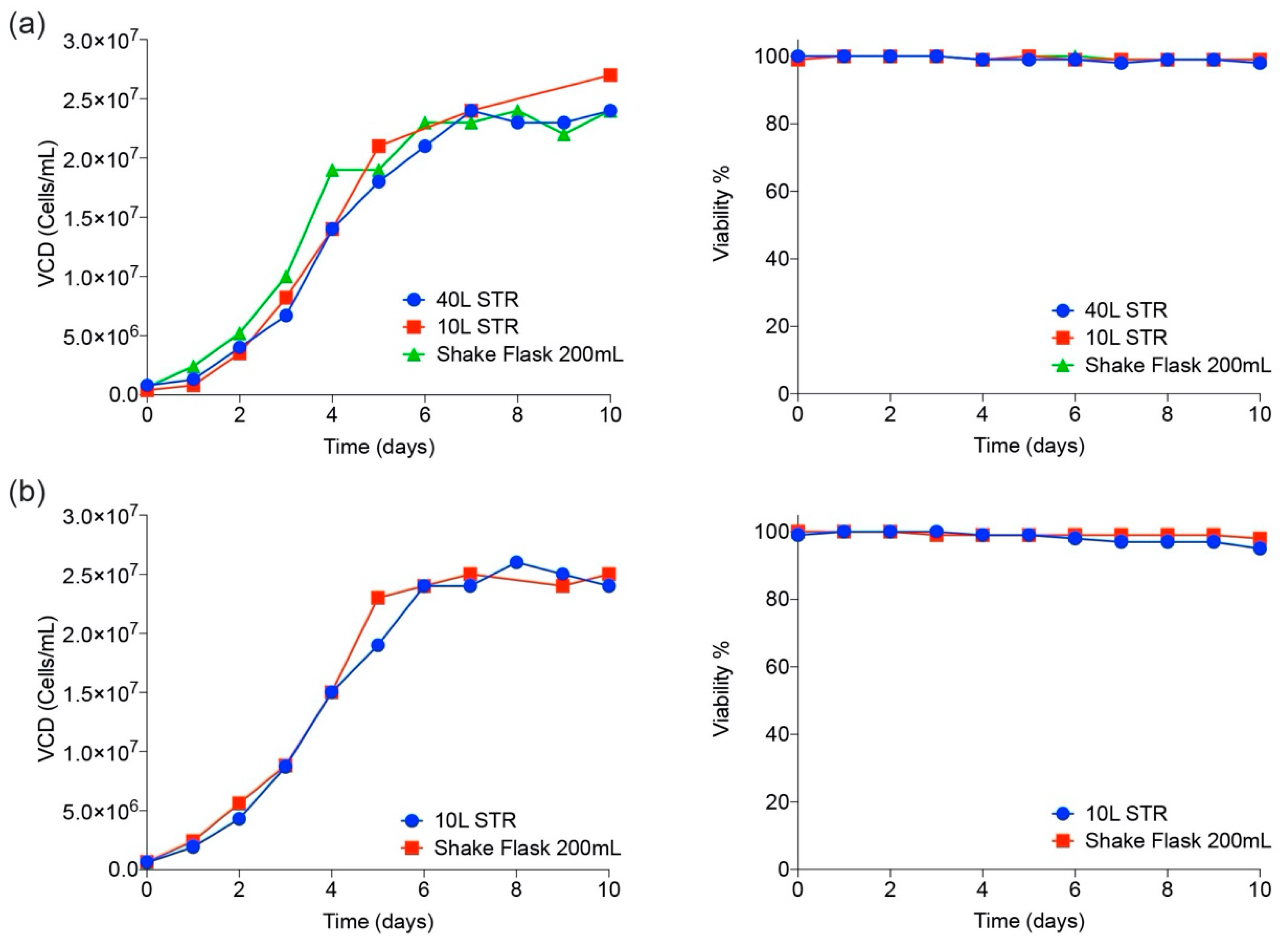
3.2. CHO Expression and Purification of SARS-CoV-2 Proteins
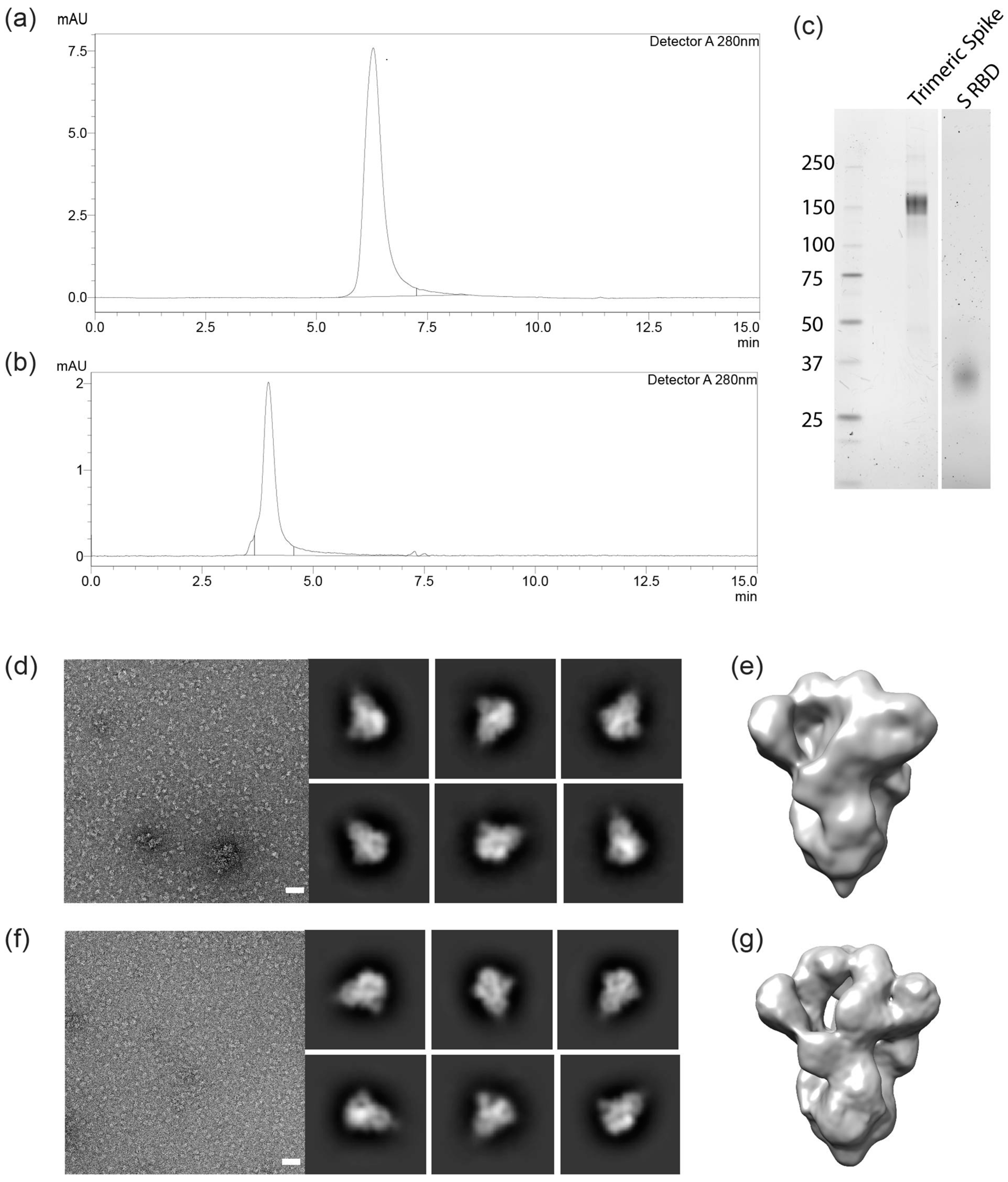
3.3. Molecular Characterization and Electron Microscopy Analysis of Trimeric Spikes
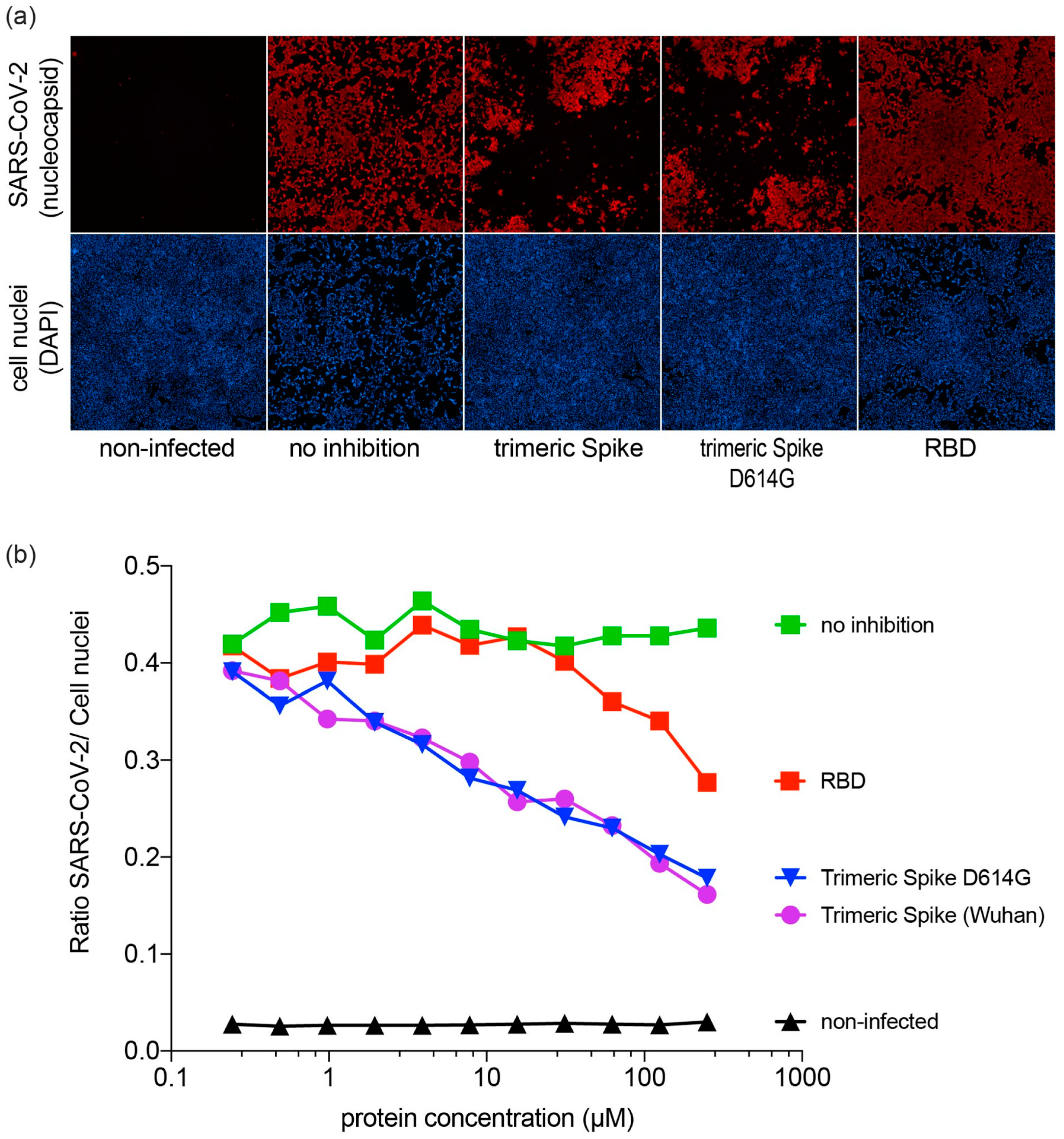
3.4. Inhibition of SARS-CoV-2 Infection by SARS-CoV-2 Proteins
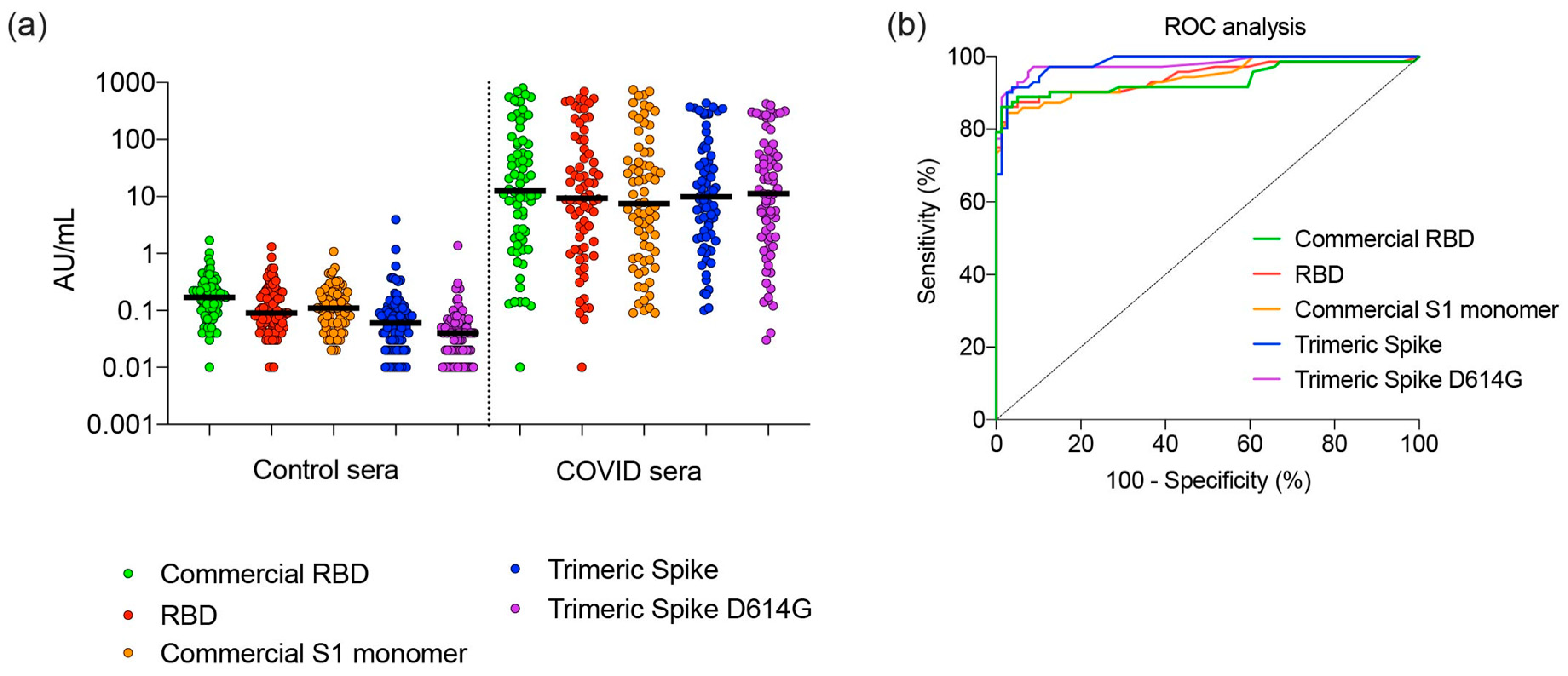
3.5. Sensitivity and Specificity of SARS CoV-2 Spike Proteins for Detection of Antibodies
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- New Pfizer Results: Coronaviurs Vaccine Is Safe and 95% protective. Available online: https://www.nytimes.com/2020/11/18/health/pfizer-covid-vaccine.html (accessed on 18 November 2020).
- Early Data Show Moderna’s Coronavirus Vaccine Is 94.5% Effective. Available online: https://www.nytimes.com/2020/11/16/health/Covid-moderna-vaccine.html (accessed on 22 November 2020).
- Fenwick, C.; Croxatto, A.; Coste, A.T.; Pojer, F.; André, C.; Pellaton, C.; Farina, A.; Campos, J.; Hacker, D.; Lau, K.; et al. Changes in SARS-CoV-2 Spike versus nucleoprotein antibody responses impact the estimates of infections in population-based seroprevalence studies. J. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Su, B.; Guo, X.; Sun, W.; Deng, Y.; Bao, L.; Zhu, Q.; Zhang, X.; Zheng, Y.; Geng, C.; et al. Potent neutralizing antibodies against SARS-CoV-2 identified by high-throughput single-cell sequencing of convalescent Patients’ B cells. Cell 2020, 182, 73–84. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wu, N.C.; Yuan, M.; Bangaru, S.; Torres, J.L.; Caniels, T.G.; van Schooten, J.; Zhu, X.; Lee, C.D.; Brouwer, P.J.M.; et al. Cross-neutralization of a SARS-CoV-2 Antibody to a Functionally Conserved Site Is Mediated by Avidity. bioRxiv 2020. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, function, and antigenicity of the SARS-CoV-2 Spike glycoprotein. Cell 2020, 181, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Seydoux, E.; Homad, L.J.; MacCamy, A.J.; Parks, K.R.; Hurlburt, N.K.; Jennewein, M.F.; Akins, N.R.; Stuart, A.B.; Wan, Y.-H.; Feng, J.; et al. Analysis of a SARS-CoV-2-infected individual reveals development of potent neutralizing antibodies with limited somatic mutation. Immunity 2020, 53, 98–105. [Google Scholar] [CrossRef]
- Hsieh, C.-L.; Goldsmith, J.A.; Schaub, J.M.; DiVenere, A.M.; Kuo, H.-C.; Javanmardi, K.; Le, K.C.; Wrapp, D.; Lee, A.G.; Liu, Y.; et al. Structure-based design of prefusion-stabilized SARS-CoV-2 spikes. Science 2020, 369, 1501–1505. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Benton, D.J.; Wrobel, A.G.; Xu, P.; Roustan, C.; Martin, S.R.; Rosenthal, P.B.; Skehel, J.J.; Gamblin, S. Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion. Nat. Cell Biol. 2020, 1–8. [Google Scholar] [CrossRef]
- Johari, Y.B.; Jaffé, S.R.P.; Scarrott, J.M.; Johnson, A.O.; Mozzanino, T.; Pohle, T.H.; Maisuria, S.; Bhayat-Cammack, A.; Lambiase, G.; Brown, A.J.; et al. Production of trimeric SARS-CoV-2 spike protein by CHO cells for serological COVID-19 testing. Biotechnol. Bioeng. 2020. [Google Scholar] [CrossRef] [PubMed]
- Güthe, S.; Kapinos, L.; Möglich, A.; Meier, S.; Grzesiek, S.; Kiefhaber, T. Very fast folding and association of a trimerization domain from bacteriophage T4 fibritin. J. Mol. Biol. 2004, 337, 905–915. [Google Scholar] [CrossRef]
- Zhao, Z.; Sokhansanj, B.A.; Malhotra, C.; Zheng, K.; Rosen, G.L. Genetic grouping of SARS-CoV-2 coronavirus sequences using informative subtype markers for pandemic spread visualization. PLoS Comput. Biol. 2020, 16, e1008269. [Google Scholar] [CrossRef] [PubMed]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G increases infectivity of the COVID-19 virus. Cell 2020, 182, 812–827. [Google Scholar] [CrossRef] [PubMed]
- De Jesus, M.J.; Girard, P.; Bourgeois, M.; Baumgartner, G.; Jacko, B.; Amstutz, H.; Wurm, F.M. TubeSpin satellites: A fast track approach for process development with animal cells using shaking technology. Biochem. Eng. J. 2004, 17, 217–223. [Google Scholar] [CrossRef]
- Gomez, N.; Ambhaikar, M.; Zhang, L.; Huang, C.-J.; Barkhordarian, H.; Lull, J.; Gutierrez, C. Analysis of Tubespins as a suitable scale-down model of bioreactors for high cell density CHO cell culture. Biotechnol. Prog. 2017, 33, 490–499. [Google Scholar] [CrossRef]
- Grant, T.; Rohou, A.; Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 2018, 7, e35383. [Google Scholar] [CrossRef]
- Thao, T.T.N.; Labroussaa, F.; Ebert, N.; V’Kovski, P.; Stalder, H.; Portmann, J.; Kelly, J.; Steiner, S.; Holwerda, M.; Kratzel, A.; et al. Rapid reconstruction of SARS-CoV-2 using a synthetic genomics platform. Nat. Cell Biol. 2020, 582, 561–565. [Google Scholar]
- Holwerda, M.; V’kovski, P.; Wider, M.; Thiel, V.; Dijkman, R. Identification of Five Antiviral Compounds from the Pandemic Response Box Targeting SARS-CoV-2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Hartog, G.D.; Schepp, R.M.; Kuijer, M.; GeurtsvanKessel, C.; Van Beek, J.; Rots, N.; Koopmans, M.P.G.; Van Der Klis, F.R.M.; Van Binnendijk, R.S. SARS-CoV-2–specific antibody detection for seroepidemiology: A multiplex analysis approach accounting for accurate seroprevalence. J. Infect. Dis. 2020, 222, 1452–1461. [Google Scholar] [CrossRef]
- Krammer, F. SARS-CoV-2 vaccines in development. Nat. Cell Biol. 2020, 586, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Wurm, F.M. Production of recombinant protein therapeutics in cultivated mammalian cells. Nat. Biotechnol. 2004, 22, 1393–1398. [Google Scholar] [CrossRef]
- Shi, X. Protein N-glycosylation in the baculovirus-insect cell system. Curr. Drug Targets 2007, 8, 1116–1125. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, Y.; Allen, J.D.; Wrapp, D.; McLellan, J.S.; Crispin, M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science 2020, 369, 330–333. [Google Scholar] [CrossRef]
- Grant, O.C.; Montgomery, D.; Ito, K.; Woods, R.J. Analysis of the SARS-CoV-2 spike protein glycan shield reveals implications for immune recognition. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Hanley, J.A.; McNeil, B.J. A method of comparing the areas under receiver operating characteristic curves derived from the same cases. Radiology 1983, 148, 839–843. [Google Scholar] [CrossRef]
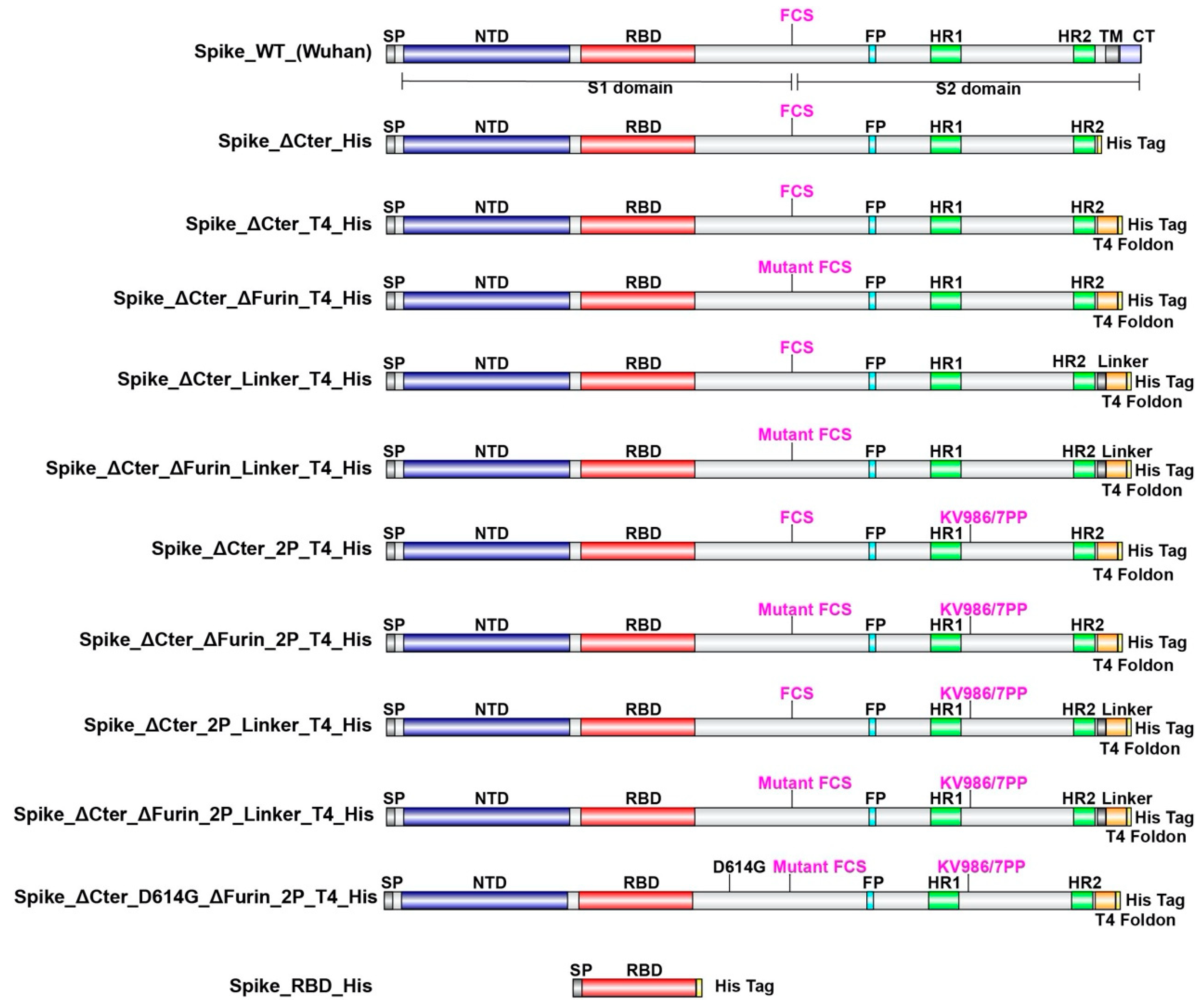

| Construction | CHO Expression | Trimer Formation |
|---|---|---|
| Spike_ΔCter_His | − | n.a. |
| Spike_ΔCter_T4_His | − | + |
| Spike_ΔCter_ΔFurin_T4_His | + | ++ |
| Spike_ΔCter_Linker_T4_His | − | + |
| Spike_ΔCter_ΔFurin_Linker_T4_His | + | ++ |
| Spike_ΔCter_2P_T4_His | − | + |
| Spike_ΔCter_ΔFurin_2P_T4_His | ++ | +++ |
| Spike_ΔCter_2P_Linker_T4_His | − | + |
| Spike_ΔCter_ ΔFurin_2P_Linker_T4_His | + | ++ |
| Spike_RBD_His | +++ | n.a. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pino, P.; Kint, J.; Kiseljak, D.; Agnolon, V.; Corradin, G.; Kajava, A.V.; Rovero, P.; Dijkman, R.; den Hartog, G.; McLellan, J.S.; et al. Trimeric SARS-CoV-2 Spike Proteins Produced from CHO Cells in Bioreactors Are High-Quality Antigens. Processes 2020, 8, 1539. https://doi.org/10.3390/pr8121539
Pino P, Kint J, Kiseljak D, Agnolon V, Corradin G, Kajava AV, Rovero P, Dijkman R, den Hartog G, McLellan JS, et al. Trimeric SARS-CoV-2 Spike Proteins Produced from CHO Cells in Bioreactors Are High-Quality Antigens. Processes. 2020; 8(12):1539. https://doi.org/10.3390/pr8121539
Chicago/Turabian StylePino, Paco, Joeri Kint, Divor Kiseljak, Valentina Agnolon, Giampietro Corradin, Andrey V. Kajava, Paolo Rovero, Ronald Dijkman, Gerco den Hartog, Jason S. McLellan, and et al. 2020. "Trimeric SARS-CoV-2 Spike Proteins Produced from CHO Cells in Bioreactors Are High-Quality Antigens" Processes 8, no. 12: 1539. https://doi.org/10.3390/pr8121539
APA StylePino, P., Kint, J., Kiseljak, D., Agnolon, V., Corradin, G., Kajava, A. V., Rovero, P., Dijkman, R., den Hartog, G., McLellan, J. S., Byrne, P. O., Wurm, M. J., & Wurm, F. M. (2020). Trimeric SARS-CoV-2 Spike Proteins Produced from CHO Cells in Bioreactors Are High-Quality Antigens. Processes, 8(12), 1539. https://doi.org/10.3390/pr8121539






