1. Introduction
The most widely used method for the detection of SARS-CoV-2 (the infectious agent that causes COVID-19) in nasopharyngeal (NP) swab samples is Reverse Transcription Polymerase Chain Reaction (RT-PCR) [
1,
2,
3]. Inaccurate test results from RT-PCR have been widely reported, with estimated false-negative rates of 10–30% among different implementations of this method [
4,
5,
6,
7]. Such high false-negative rates pose a significant challenge to controlling the spread of infection, and are further exacerbated by poor sample quality or low viral loads that are below the detection limit of standard RT-PCR methods [
8,
9]. This, combined with both the cost and scarcity of reagents [
10,
11,
12], has hampered global scale-up of RT-PCR testing to levels that would be required to adequately monitor communities for COVID-19.
An additional challenge to controlling the spread of COVID-19 is the role of asymptomatic transmission [
13,
14,
15,
16]. Different estimates suggest that 40 to 80% of infected individuals are either pre-symptomatic, asymptomatic, or mildly symptomatic [
17,
18,
19,
20]. Early detection of infection in these individuals is crucial for disease control, which is why many countries and communities have started implementing active screening programs that extend COVID-19 testing to asymptomatic individuals. However, asymptomatic carriers sometimes carry very low viral loads [
21] that may not be detected by a standard RT-PCR test [
22]. Therefore, the development of more sensitive detection methods that can detect low viral loads is crucial.
Most commercial kits for COVID-19 testing utilize either a one-step RT-PCR approach, which combines the RT and qPCR reactions, or a two-step approach in which RT and qPCR are performed sequentially. A target-specific preamplification step has been successfully incorporated in a number of studies to detect and analyze various types of samples with limited amount of genomic materials, including viruses in human samples [
23] or in drinking water [
24], circulating tumor DNA in blood [
25], and even ancient DNA samples [
26]. Therefore, in this study, we implemented a three-step approach involving sequential RT, targeted cDNA preamplification, and qPCR, using a commercially available microfluidics platform. Using this method, we demonstrate reliable ultra-sensitive detection of low SARS-CoV-2 viral loads in both standard positive controls and clinical NP swab samples, including samples previously diagnosed as negative by an accredited diagnostic lab. Overall, this microfluidic RT-PCR assay is a cost-effective strategy with the potential to reduce the false-negative rate of clinical diagnostic tests, and as such, could be a valuable tool in active screening programs aimed at the early detection of SARS-CoV-2.
2. Materials and Methods
We first implemented the three-step SARS-CoV-2 detection method using synthetic SARS-CoV-2 RNA and SARS-CoV-2 plasmids and determined the limit of detection, before validating this method in clinical nasopharyngeal swab samples.
2.1. Ethics Statement
This study was determined as exempt by the NYU Abu Dhabi (NYUAD) Institutional Review Board (HRPP-2020-48) as it involves clinical samples that have already been collected by a diagnostic lab for the primary purpose of SARS-CoV-2 testing, and have been de-identified before being transported to and analyzed at NYUAD.
2.2. Positive Controls
Two types of positive controls were used in this study. The Twist Synthetic SARS-CoV-2 RNA (102024, Twist Biosciences, San Francisco, CA, USA) consists of six non-overlapping ssRNA fragments with a coverage of greater than 99.9% of the viral genome. The SARS-CoV-2 plasmids (10006625, IDT, Leuven, Belgium) contain the complete DNA sequence of the SARS-CoV-2 nucleocapsid gene.
2.3. SARS-CoV-2 Detection (Synthetic RNA and Plasmid)
Two assays (primer/probe sets) were used for SARS-CoV-2 detection, per CDC recommendations: 2019-nCoV_N1 and 2019-nCoV_N2 (2019-nCoV CDC EUA Kit, 10006606, IDT). The human RNase P (RP) assay was used as a control for RNA extraction and RT-qPCR reactions. For both positive controls, 10-fold serial dilutions were prepared, with two replicates at each concentration. Each sample was analyzed using 9 replicates for N1, 9 replicates for N2, and 6 replicates for RP assays.
Manual extraction of synthetic SARS-CoV-2 RNA was performed using the ABIOpureTM Viral DNA/RNA Extraction Kit (M561VT50, Alliance Bio, Bothell, WA, USA) according to the manufacturer’s instructions. The isolated synthetic RNA/SARS-CoV-2 plasmids were then used for reverse transcription (RT) and quantitative PCR (qPCR) using the Fluidigm Real-Time PCR for Viral RNA Detection protocol (FLDM-00103, Fluidigm, San Francisco, CA, USA). The RT was prepared by mixing 5 μL of purified RNA and 1.25 μL RT Master Mix (100-6297, Fluidigm, San Francisco, CA, USA), followed by 3 steps of incubation using the Bio-Rad T100 thermal cycler (Bio-Rad, Hercules, CA, USA). Preamplification of cDNA was performed by first pooling the N1, N2, and RP assays and diluting into dilution reagent (100-8730, Fluidigm, San Francisco, CA, USA) to a final concentration of 100.5 and 25.5 nM for the primers and probes, respectively, and then, mixing it with 2.5 μL Preamp Master Mix (100-5744, Fluidigm, San Francisco, CA, USA) and 0.635 μL PCR water (100-5941, Fluidigm, San Francisco, CA, USA). The final preamplification reaction (12.5 μL) consisted of preamplification pre-mix and RT reaction in a 1:1 ratio. Upon completion of 20 preamplification cycles, reactions were diluted 1:5 in Dilution Reagent (100-8730, Fluidigm, San Francisco, CA, USA), resulting in a total volume of 62.5 μL.
The qPCR mix was prepared using 1.8 μL of diluted cDNA or SARS-CoV-2 plasmid positive controls (with or without preamplification) and 2 μL of 2X TaqMan Fast Advanced Master Mix (4444557, Thermo Fisher Scientific, Waltham, MA, USA) and 0.2 μL 20X GE Sample Loading Reagent (Fluidigm PN 100-7610). Next, 3 μL qPCR mix from each sample was loaded into the sample inlet in the 192.24 integrated fluid circuit (IFC, Fluidigm, see
Figure S1). For each assay, 3 μL of primer/probe mix (13.5X) was mixed with 1 μL of 4X Assay Loading Reagent (Fluidigm, 102-0135), and 3 μL of each assay mixed was loaded in the assay inlet in the 192.24 IFC chip (Fluidigm, 100-626). The chip was then placed in an IFC controller RX machine to pre-load the samples and the assays, and then, loaded onto the BioMark HD instrument (Fluidigm) for RT-qPCR using 35 cycles. In total, 4608 reactions were performed in each 192.24 IFC chip. The raw amplification data were acquired using the Fluidigm data collection software and analyzed using the Fluidigm Real-Time PCR Analysis software 3.0.2. To calculate the Cq (cycle of quantification) of each sample, a global threshold was automatically calculated and applied to all the samples in the chip. The crossing point at which the amplification curve of each sample crossed the threshold line was determined as the Cq. For comparability with other studies, we note that the Cq is numerically equivalent to the Ct value in our system.
2.4. Clinical Samples and SARS-CoV-2 Clinical Diagnostics
A total of 182 de-identified nasopharyngeal (NP) swab samples (91 positive and 91 negatives for SARS-CoV-2) were obtained from an accredited diagnostic lab. The diagnostic lab used the automated NX-48S Viral RNA Kit (NX-48S, Genolution Inc., Seoul, Korea) for RNA extraction, and the U-TOPTM COVID-19 Detection Kit (SS-9930, Seasun Biomaterials Inc., Daejeon, Korea) for SARS-CoV-2 detection, following accredited protocols in the United Arab Emirates that follow guidelines from the US Center for Disease Control and Prevention (CDC). Briefly, three primers were used, two targeting the viral ORF1ab and the N genes, and one acting as an internal control. A sample was classified as positive if at least one of the viral genes was detected with a Ct value ≤38. The reported LoD of the kit was 10 copies/reaction, translating to 1 copy/μL in the RNA sample.
2.5. Three-Step Analysis of SARS-CoV-2 in Clinical Samples
Automated extraction of viral RNA from the clinical samples was performed using the Chemagic 360 automated nucleic acid extraction system (2024-0020, Perkin Elmer, Waltham, MA, USA) and the Chemagic Viral DNA/RNA 300 Kit H96 (CMG-1033S, Perkin Elmer, Waltham, USA) according to the manufacturer’s instructions. For RNA extraction, 300 μL clinical samples were used and eluted in 80 μL elution buffers. Subsequent RT, preamplification, and qPCR were performed as described above. Clinical samples were loaded onto two 96-well plates for RT and preamplification, with each plate containing 10-fold serially diluted SARS-CoV-2 plasmid controls (50–5000 copies/μL in the original experiment; 10–10,000 copies/μL in the replication experiments) used for viral load estimation. Samples were considered valid if the RP gene was reliably detected in at least 4 of the 6 replicates. Samples were classified as positive if at least one of the N assays (N1 or N2) was detected in at least one replicate. Each PCR plate also contained two negative controls: empty transport medium, to control for contamination during RNA extraction (NRX control), and TE (Tris–EDTA) dilution buffer, to control for contamination during pre-amplification and qPCR (NQF control).
2.6. Statistical Analysis
Data analysis was performed with R, associated packages, and GraphPad Prism 8. Data were summarized as mean ± standard deviation. Virus copies were quantified based on a 10-fold dilution series of SARS-CoV-2 plasmids to generate standard curves. The standard curves were used to build log-linear models used to predict viral loads based on Cq values.
3. Results
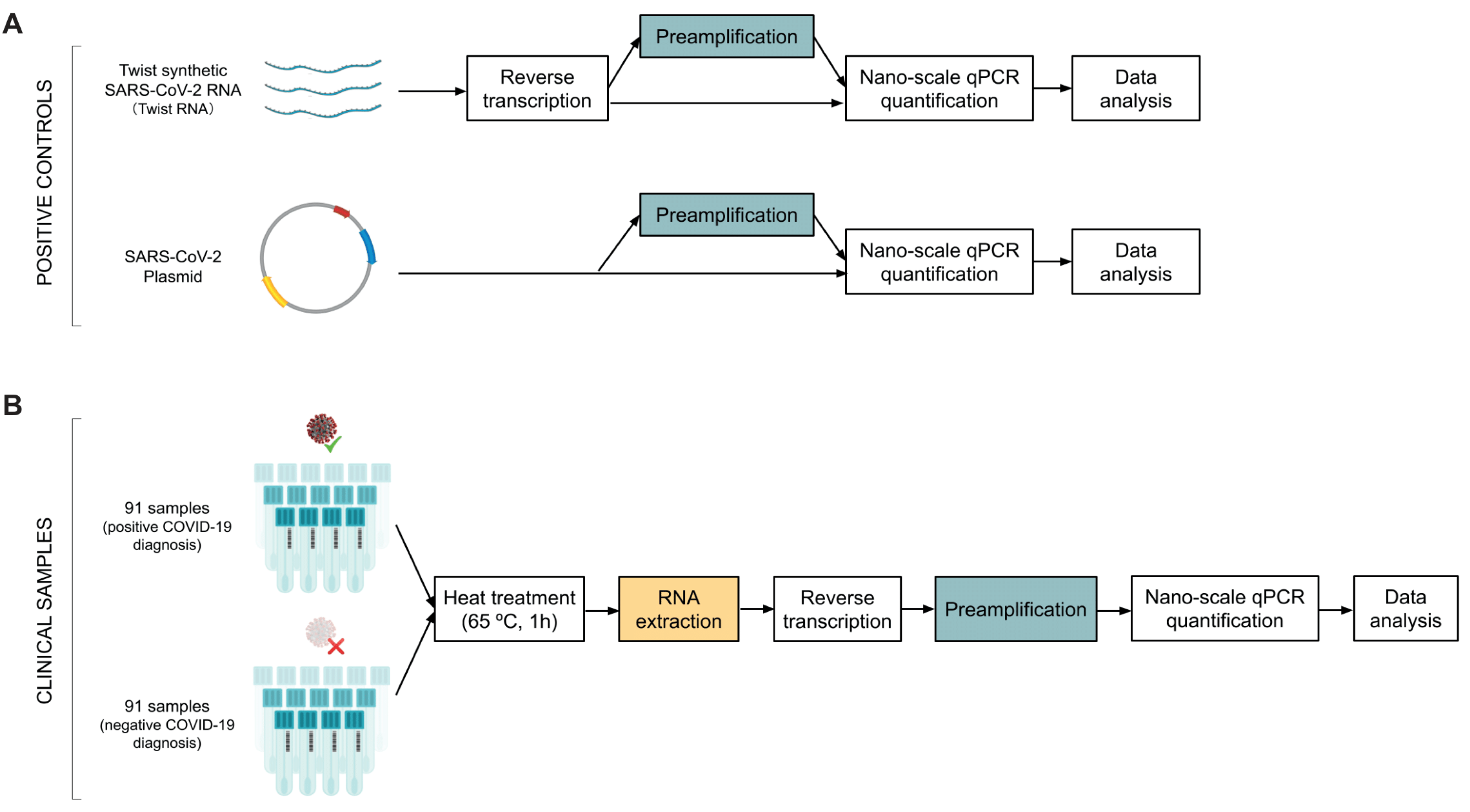
The objective of this study was to evaluate whether a microfluidic nano-scale RT-qPCR system has the potential to enhance the limit of detection (LoD) of SARS-CoV-2 in clinical samples. To do this, we first used synthetic SARS-CoV-2 RNA (Twist RNA) and SARS-CoV-2 plasmids to develop and evaluate our protocols (
Figure 1A), and subsequently applied the method to 182 NP samples that were analyzed in a clinical diagnostic lab using standard RT-PCR protocols (
Figure 1B). For all our experiments, we used a 192.24 microfluidic chip, which allowed 192 samples to be independently analyzed against 24 different qPCR probes (24 assays), totaling 4608 nano-scale qPCR reactions per experiment (
Figure S1A). Following the standards set by the US Center for Disease Control (CDC), we used two probes targeting different regions of the N gene (N1 and N2) for SARS-CoV-2 detection (
Figure S1B), and a probe targeting the human Ribonuclease P (RP) gene as a quality control. To increase the robustness of detection for each assay, we performed nine technical replicates for the N1 and N2 assays, and six replicates for the RP assay.
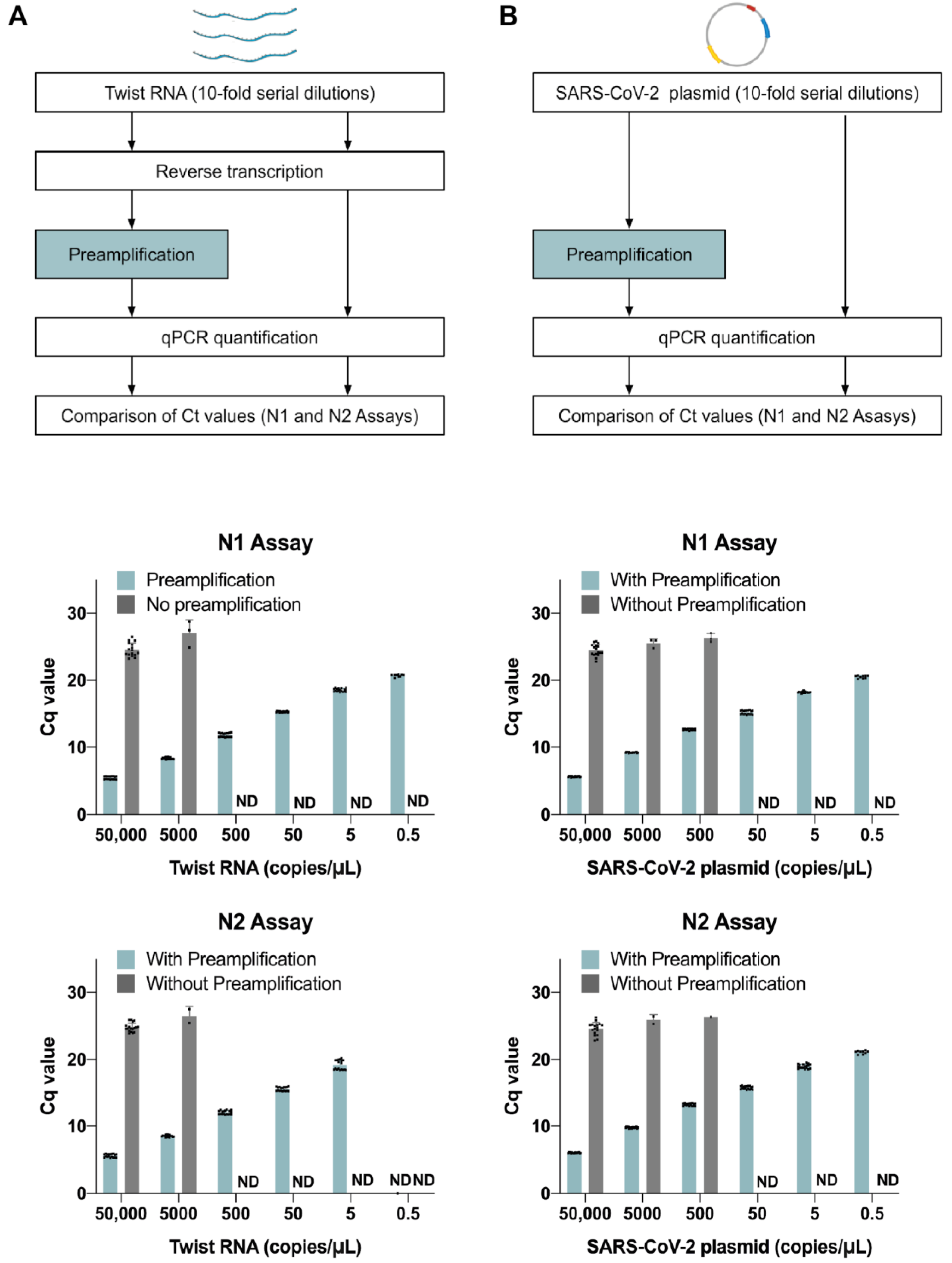
We first used 10-fold dilution series of the synthetic SARS-CoV-2 RNA (Twist RNA) and SARS-CoV-2 plasmid to determine the LoD for the N1 and N2 assays, with and without a preamplification step (
Figure 2). With preamplification, the viral N gene was detectable in Twist RNA at 0.5 copies/µL (N1 assay, 9 out of 18 replicates) and 5 copies/µL (N2 assay, 18 out of 18 replicates), whereas without preamplification, we were only able to detect viral material in a few replicates at 5000 copies/µL (N1 assay, 3 out of 18 replicates; N2 assay, 2 out of 18 replicates). No viral material was detected below 5000 copies/µL (
Figure 2A). We observed similar results using the SARS-CoV-2 plasmid (
Figure 2B), demonstrating that the preamplification step is essential for high detection sensitivity for SARS-CoV-2 in the range of 1 copy/µL. For this reason, we used a preamplification step in all subsequent experiments.
Since the LoD is dependent on the abundance of input material, we next examined the extent to which RNA extraction affects the quantity of input material. We extracted RNA from the Twist RNA dilution series and eluted it in the same volume as the original samples (
Figure S2). With or without extraction, we were able to detect 5 copies/µL using both the N1 and N2 assays, but the Cq values for the extracted samples increased by 1–3 cycles at different dilutions. Most viral nucleic acid extraction kits recommend using carrier RNA to enhance recovery of viral RNA in samples where the quantity of material is low; however, carrier RNA might compete nonspecifically with the SARS-CoV-2 RNA in reverse transcription reactions. To explore this possibility, we added carrier RNA directly into the Twist RNA serial dilutions and found that Cq values indeed increased by 0.9–2.7 cycles at most concentrations (
Figure S2). This suggests that the presence of carrier RNA, and not RNA extraction itself, can adversely affect the detection of viral material, possibly by interfering with the efficiency of reverse transcription.
To evaluate the sensitivity of our workflow in comparison with standard SARS-CoV-2 detection methods, we next analyzed 182 NP swab samples previously diagnosed by an accredited diagnostic laboratory as SARS-CoV-2 positive or negative (91 samples each). We heated the samples at 65 °C for 1 h to inactivate viral particles and analyzed them using our SARS-CoV-2 detection workflow (
Figure 1B). Any samples with poor qPCR amplification curves and high variation in Cq values among replicates (SEM > 0.5) were flagged as inconsistent, which in our experience is usually due to technical issues such as formation of air bubbles when loading the samples onto the chip. A total of 11 inconsistent samples (4 positive and 7 negative) were removed from subsequent analysis (
Figure 3A). The remaining 171 high quality and valid samples were classified as either positive (either N1 or N2 detected) or negative (neither N1 nor N2 detected). Using this classification, we confirmed 86 positive (Pos_Pos, 94.5%) and found 1 negative sample (Pos_Neg, 1.1%) among the 87 samples with a positive clinical diagnosis (
Figure 3B). The one Pos_Neg sample had a relatively high Cq value (36.4 for the N-gene and 37.2 for the ORF1 gene) in the clinical diagnostic test; therefore, it is likely that RNA degradation during transport or the heat inactivation process may have brought its viral load below the LoD. Exploring how the two methods compared, we found relative consistency between the ranking of Cq values between the microfluidic RT-qPCR test and the clinical diagnostic test, despite the expected difference in Cq values due to the addition of a pre-amplification step (20 cycles) in the microfluidic method (
Figure S3). Among the 84 samples with a negative diagnosis, we detected 17 positives (Neg_Pos, 18.7%): in 9 of these samples, we obtained valid Cq values for all 18 replicates (both N1 and N2), and in the remaining 9, we detected the virus in 9/18 replicates (either all N1 or all N2). All samples with a negative clinical diagnosis showed no detectable signal in the clinical diagnostic qPCR test. These analyses thus show that by performing a large number of replicates, our method can robustly and consistently detect SARS-CoV-2 and identify samples as false-negatives by the clinical diagnostic procedure.
Standard curves are commonly used to estimate viral loads in RT-qPCR reactions, which is why we took that approach in our study. Standard curves based on 100-fold serial dilutions of Twist RNA and SARS-CoV-2 plasmids ranging from 5 to 50,000 copies/µL both showed a nearly-perfect log-linear fit (R
2 > 0.99) with little variation among technical replicates (SEM < 0.2) for both N assays (
Figure S4). Having established technical precision using these curves, we used the SARS-CoV-2 plasmid standards to quantify viral copies in the clinical samples in each of the two PCR plates. Given that the N1 and N2 assays were highly concordant (R
2 = 0.876, Kendall’s Tau correlation), and the difference in Cq values was within one cycle in over 90% of the samples (
Figure S5A,B), we used an average Cq value for the N gene assays. To maximize consistency in viral load estimates among the samples, we then used the standard curve from plate 1 to quantify the viral loads in each clinical sample (
Figure 3C). The Pos_Pos samples showed a wide range of estimated viral loads (0.2–1.17 × 10
6 viral copies/µL) spanning five orders of magnitude. In contrast, the Neg_Pos samples exhibited a much narrower range of viral load estimates (0.2–40.25 viral copies/µL), corresponding to very low amounts of viral material in the NP swab sample (0.05–10.73 viral copies/µL). These results show that the nano-scale qPCR method can detect SARS-CoV-2 across a broad range of viral titers and can confidently detect relatively low viral loads that could otherwise be missed by standard detection methods used in diagnostic labs.
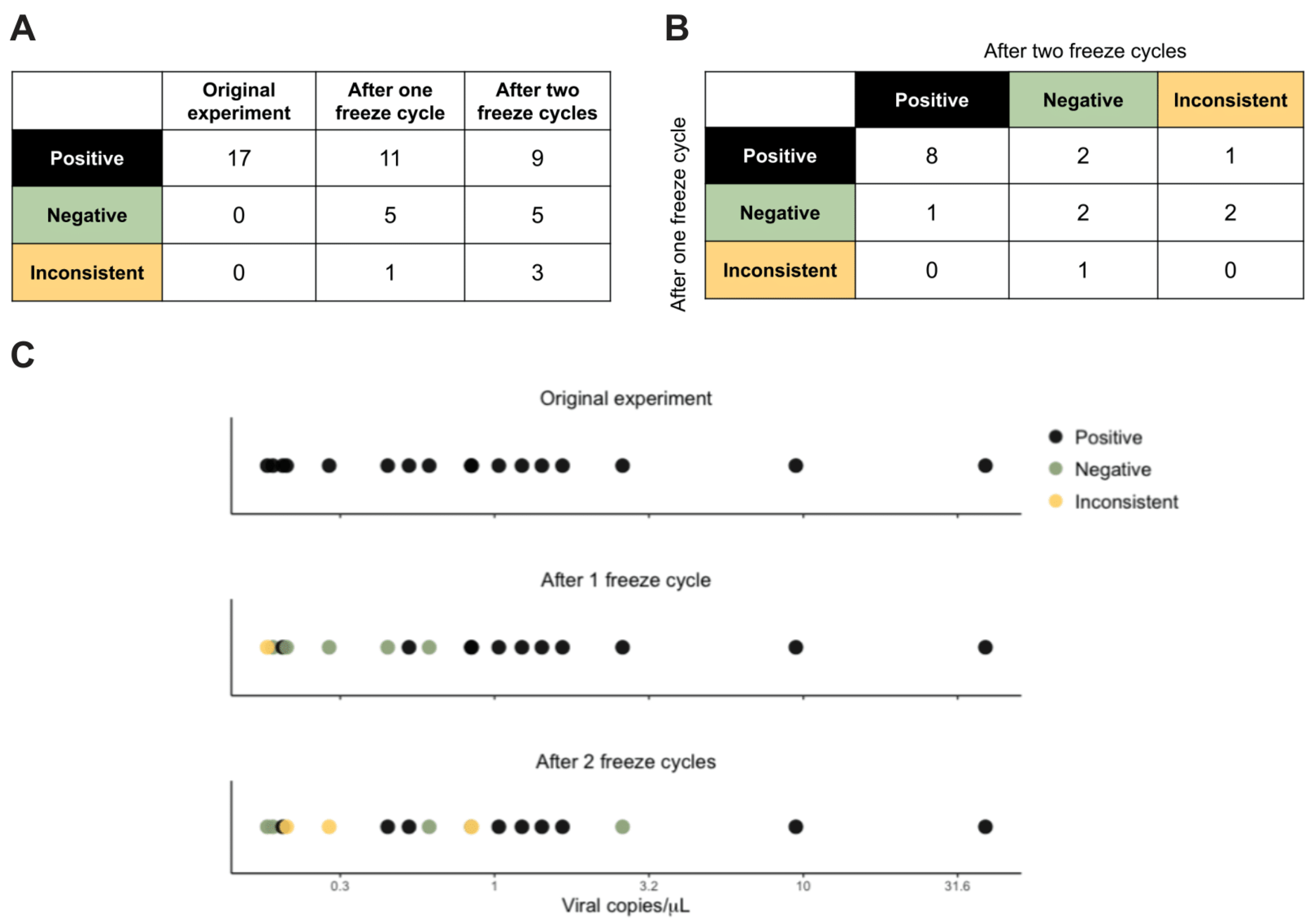
To evaluate the reproducibility of this method in samples with low viral loads, we re-analyzed the 17 Neg_Pos samples after one and after two freeze cycles. After one freeze cycle, 11 samples remained positive, whereas after two freeze cycles, only 9 did (
Figure 4A). This suggests that additional freeze cycles may lead to an increased false-negative rate, likely due to degradation of viral RNA resulting in copy numbers below the LoD. This is consistent with prior observations of increased Cq values following one freeze–thaw cycle [
27]. Nevertheless, 8 of the 17 samples were reproducibly classified as positive in three independent experiments (
Figure 4B). When comparing viral load estimates among the 17 Neg_Pos samples, we found that the 11 samples that retested positive after one freeze cycle had viral loads between 0.21 and 38.89 copies/µL, whereas the 5 samples that retested negative all had viral loads less than 1 copy/µL based on the original experiment (
Figure 4C). This suggests that samples with extremely low viral titers close to the LoD could fail to be consistently detected, likely due to factors such as sample degradation or stochastic variation in the number of viral RNA molecules present in the small reaction volumes used for RT.
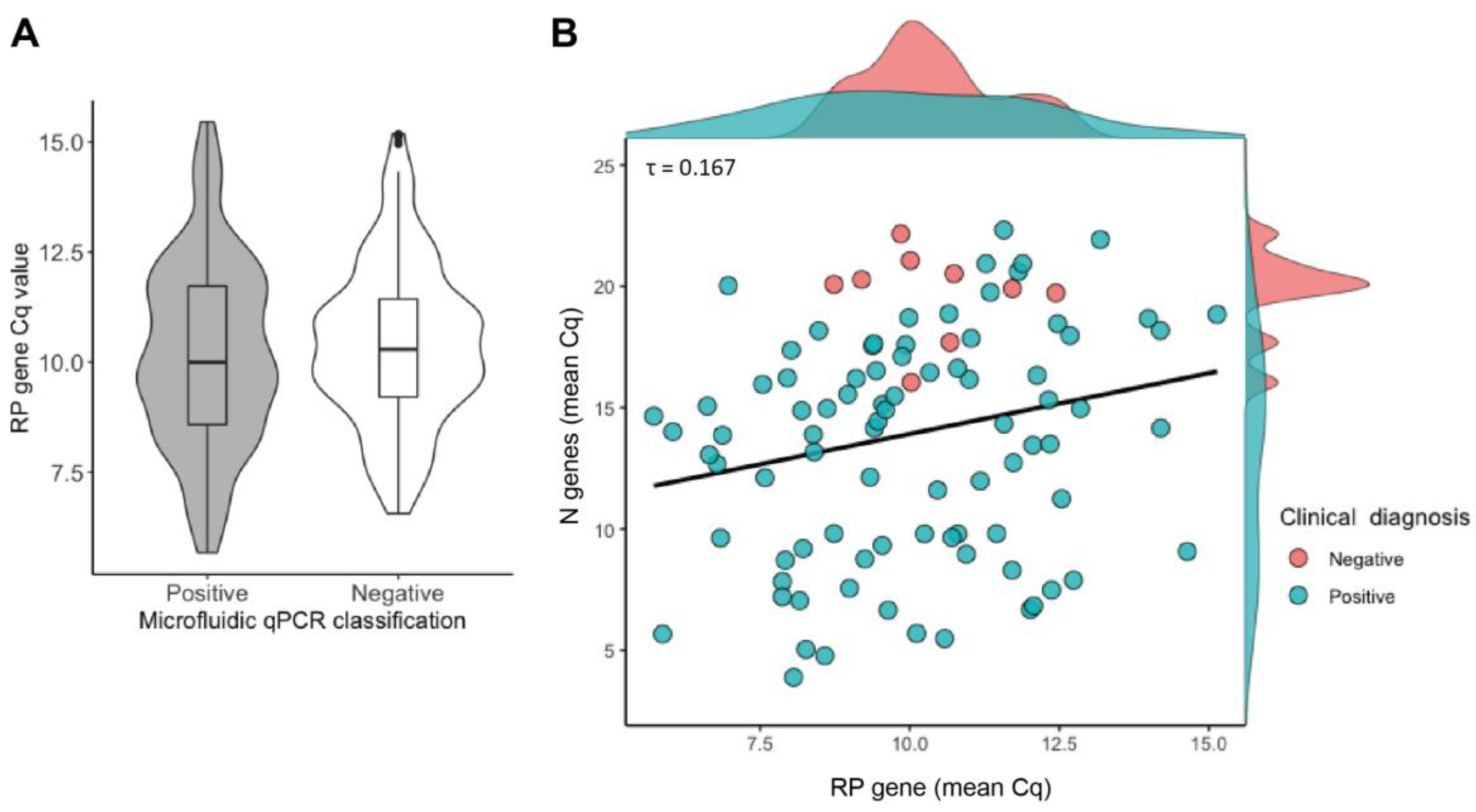
Lastly, we examined the relationship between the distributions of mean Cq values for the host RP gene assay and the N gene assay. If negative samples tended to show much higher Cq values for the RP assay than the positive samples, this would indicate that the negative result was likely due to inadequate sampling of human tissues in the swab. Instead, we observed no significant differences in the Cq values of RP assay between positive and negative samples (
t-test,
p-value = 0.08) (
Figure 5A) and found minimal correlation between the mean Cq values of the RP and the N gene assays, with only 2.8% of the variation in the N gene assays attributable to detection of the RP gene (R
2 = 0.028, Kendall’s Tau correlation) (
Figure 5B). We thus conclude that the high Cq values for the N assays in the Neg_Pos samples were not due to inadequate sampling, but instead, accurately reflected low viral loads in these samples.
4. Discussion
In this study, we implement and validate a three-step approach for SARS-CoV-2 detection utilizing RT of SARS-CoV-2 viral RNA, cDNA preamplification, and nano-scale qPCR. Using serial dilutions of positive controls, we demonstrate a 1000-fold improvement in detection sensitivity of the microfluidic qPCR system when adding the preamplification step, consistent with a previous study in which a target preamplification step was found to enhance the LoD by 100-fold in detecting SARS-CoV [
23]. We also show that nano-scale qPCR can be used to quantify viral copies in clinical samples with high confidence. Our data suggest that the LoD of this method (with preamplification) is less than 1 copy/µL: we obtained a LoD of 0.5 copies/µL for Twist RNA and SARS-CoV-2 plasmid, and detected the virus down to 0.2 copies/µL in experiments using clinical NP swab samples. Based on this analysis, our method seems to be more sensitive than standard RT-PCRs, which have reported a LoD ranging between 5.6 and 100 copies/µL [
28,
29,
30], and that its performance is comparable to other highly sensitive methods such as the CDC 2019-nCoV RT-PCR Diagnostic Panel with QIAGEN QIAmp DSP Viral RNA Mini Kit and the QIAGEN EZ1 DSP (1 copy/µL) [
3], ddPCR (0.1 copies/µL) [
8], and RT-LAMP (1 copy/µL) [
31].
Crucially, we demonstrate the power of this method to reduce the false-negative rate of SARS-CoV-2 clinical diagnostic tests: we detected SARS-CoV-2 in 17 samples diagnosed as negative by an accredited diagnostic lab with high confidence, based on a large number of technical replicates (9 for N1, 9 for N2, 6 for RP) and a conservative threshold for Cq value consistency among sample replicates (SEM < 0.5). This is especially important considering that the viral loads of these Neg_Pos samples (0.2–40.25 viral copies/µL) were close to the LoD of standard SARS-CoV-2 tests and are thus, more likely to return false-negative results using standard RT-PCR methods. This three-step microfluidics RT-qPCR method, which includes a preamplification step, nano-scale reactions, and a large number of replicates, can reliably detect samples with low viral load and reduce the false-negative rate compared to standard RT-PCR assays.
Beyond its ultra-sensitive detection of SARS-CoV-2, this microfluidic platform is a cost-effective strategy with several advantages: a nanoliter volume per reaction (lower reagent consumption per assay), a parallelized assay system (increased throughput), amenability to automation (increased precision), capacity to run a large number of replicates per sample (increased confidence in test results), and the capacity to simultaneously test for multiple pathogens (broader diagnostic utility). Based on our experience, and considering the fact that reagents cost more in the UAE than in Europe and the US, the cost of SARS-CoV-2 testing per sample using the 192.24 microfluidic chip and the design used in this study (9 replicates for N1 and N2, 6 replicates for RP) is approximately USD 17, including reagents and the chip, but excluding labor costs. This is approximately four times cheaper than the current cost of standard SARS-CoV-2 testing in the UAE. Lowering the number to three technical replicates for each of the N1, N2, and RP assays would allow for four additional assays, which can be utilized to diagnose other viral or bacterial infections using three technical replicates (or six additional assays with two technical replicates) at no extra cost, bringing down the cost per sample per assay even further. Since each sample is loaded only once and then analyzed against 24 assays simultaneously, no extra time is required for performing multiple assays of replicates. Because of this, the method has great potential for economies of scale, including assay multiplexing (to detect additional pathogens) and sample pooling (to increase throughput), which would further reduce per test costs. These advantages warrant serious consideration of this three-step nano-scale assay system, especially for active screening programs, which aim at the early detection of SARS-CoV-2 in asymptomatic, pre-asymptomatic, or mildly symptomatic individuals who likely carry low viral loads.