Light-Induced Production of An Antibody Fragment and Malaria Vaccine Antigen from Chlamydomonas reinhardtii
Abstract
:1. Introduction
2. Experimental Section
2.1. Gene Constructs for αCD22 scFv and Pfs25
2.2. Cultivation of Recombinant Pfs25 and αCD22 scFv Chlamydomonas reinhardtii Strains
2.3. Effect of Light Duration and Light Intensity on Light-Induced Production of αCD22 scFv
2.4. Protein Extraction
2.5. Protein Analysis
2.6. FLAG Affinity Purification
2.7. Statistical Analysis
3. Results and Discussion
3.1. Algae Cultivation and Accumulation of αCD22scFv and Pfs25
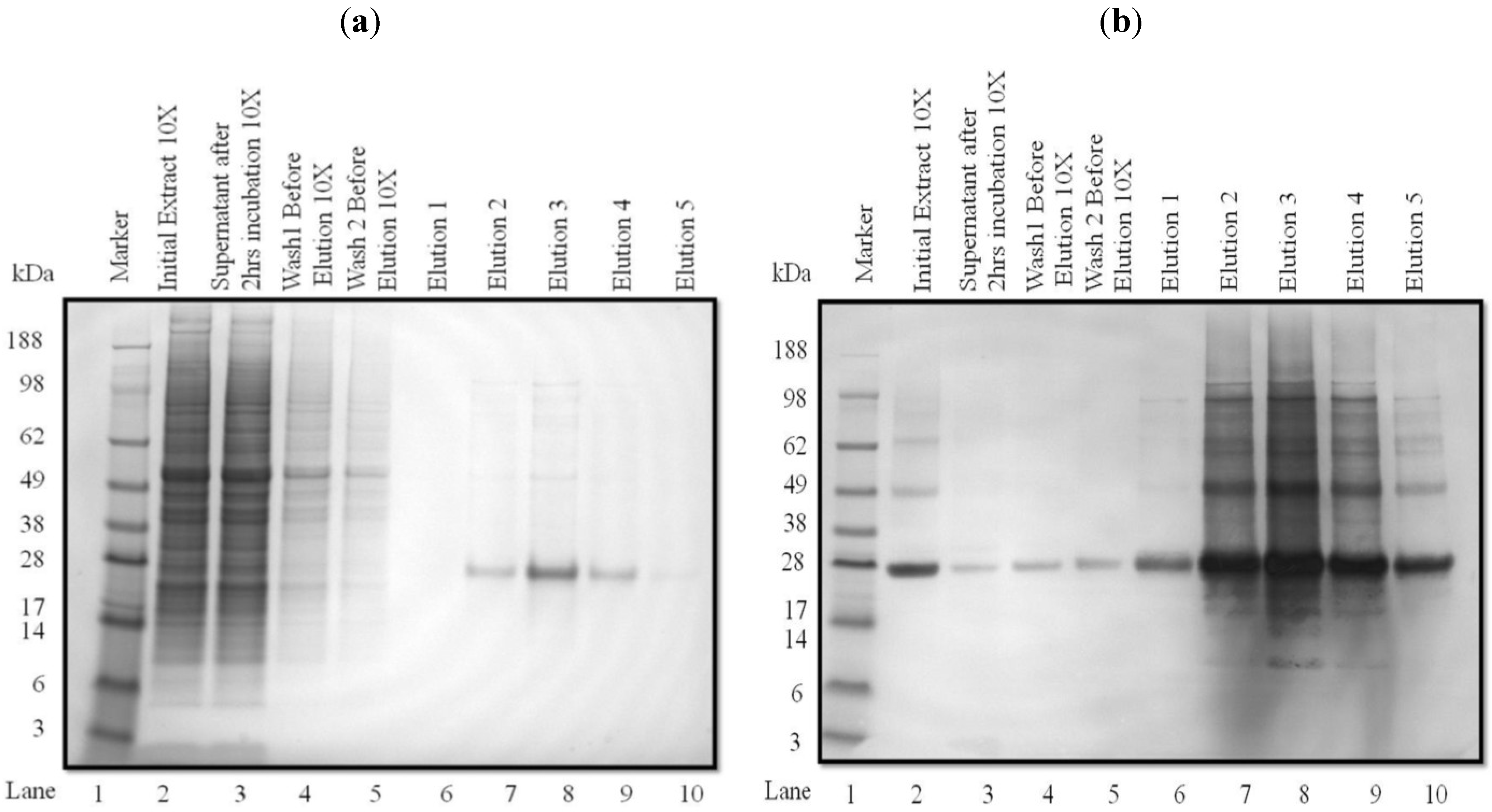
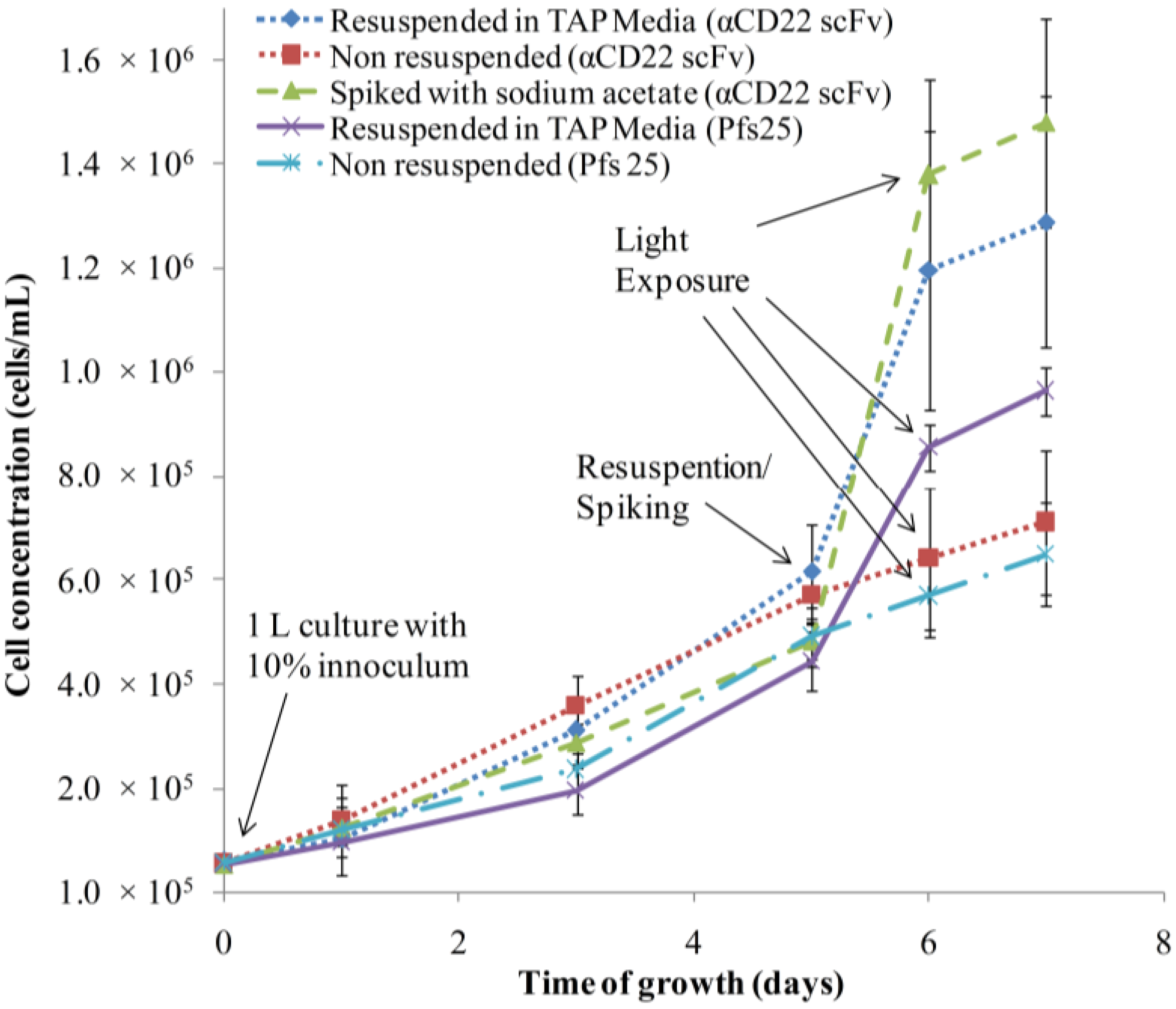
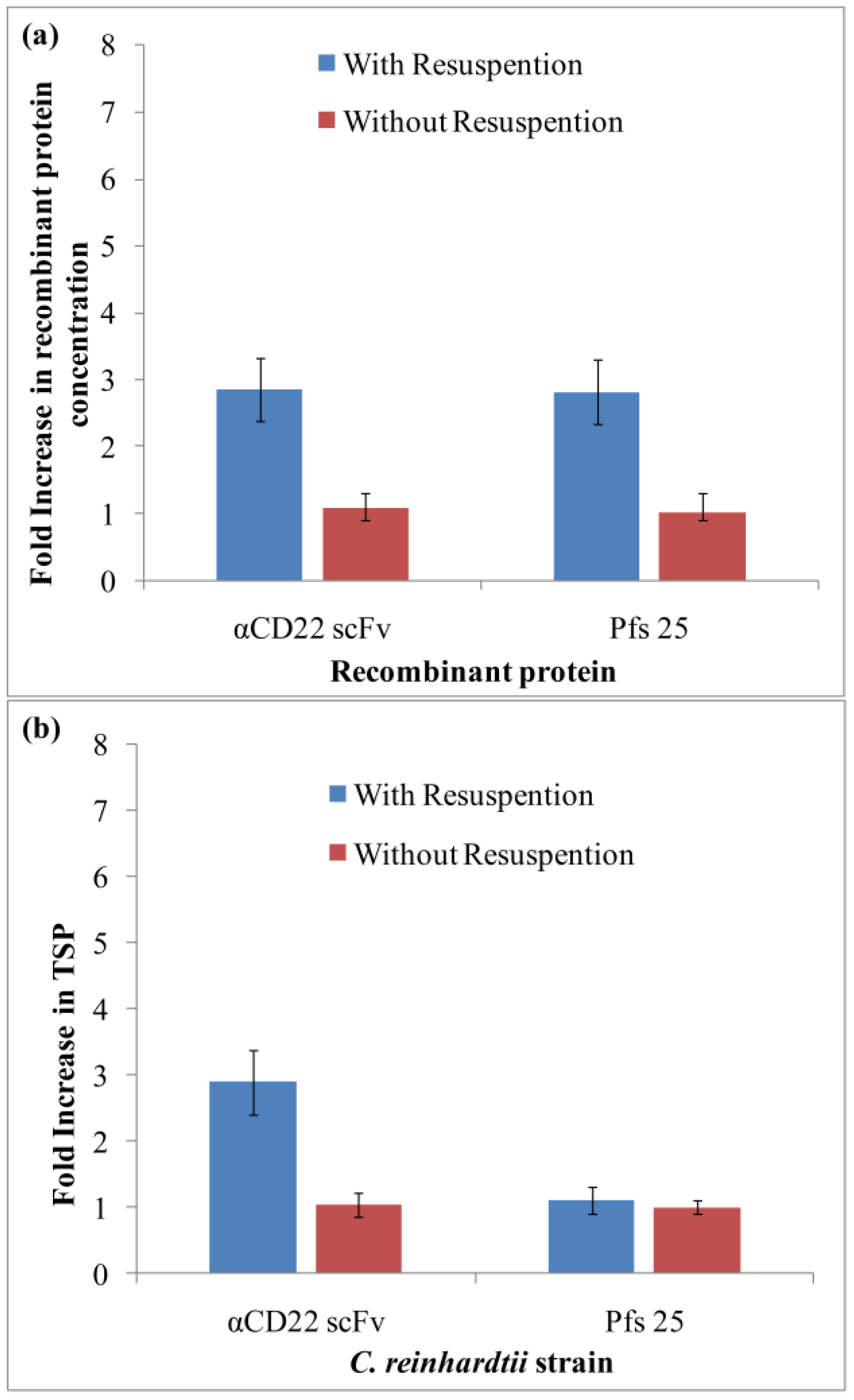

3.2. Purification and Analysis of Recombinant Proteins
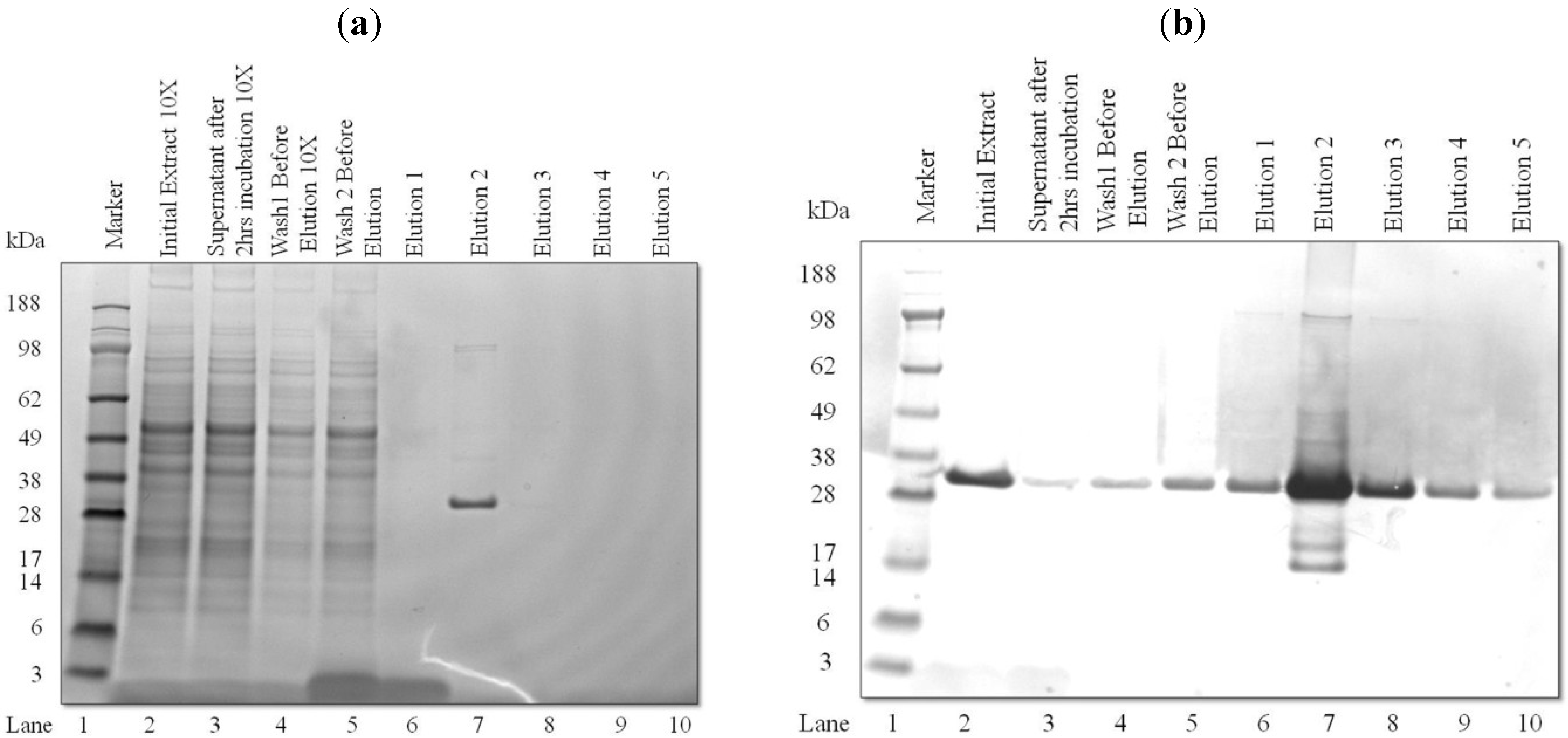

3.3. Light-Induced Accumulation of αCD22scFv
| Photosynthetic Photon Flux (PPF) | Duration | 101 μmol m−2 s−1 | 300 μmol m−2 s−1 |
|---|---|---|---|
| Biomass (g) | 12 h | x 4.0 a ± 0.01 | x 4.9 a ± 1.5 |
| 24 h | x 4.0 a ± 0.01 | x 5.3 a ± 1.0 | |
| TSP in algae extract (μg/mL) | 12 h | x 5939 a ± 257 | x 5485 a ± 802 |
| 24 h | x 6840 a ± 1167 | x 5248 a ± 415 | |
| αCD22 scFv in wet biomass (μg/g) | 12 h | x 21.2 a ± 1.8 | x 26.6 a ± 4.3 |
| 24 h | x 35.1 b ± 5.8 | y 61.5 b ± 14 | |
| αCD22 scFv volumetric conc. (μg/L) | 12 h | x 84.7 a ± 7.1 | x 135 a ± 66 |
| 24 h | x 140 a ± 20 | y 314 b ± 20 | |
| αCD22 scFv (%TSP) | 12 h | x 0.07 a ± 0.01 | x 0.1 a ± 0.03 |
| 24 h | x 0.12 a ± 0.03 | y 0.23 b ± 0.05 |
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Mayfield, S.P.; Manuell, A.L.; Chen, S.; Wu, J.; Tran, M.; Siefker, D.; Muto, M.; Marin-Navarro, J. Chlamydomonas reinhardtii chloroplasts as protein factories. Curr Opin Biotechnol. 2007, 18, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Rasala, B.A.; Muto, M.; Lee, P.A.; Jager, M.; Cardoso, R.M.F.; Behnke, C.A.; Kirk, P.; Hokanson, C.A.; Crea, R.; Mendez, M.; et al. Production of therapeutic proteins in algae, analysis of expression of seven human proteins in the chloroplast of Chlamydomonas reinhardtii. Plant Biotechnol. J. 2010, 8, 719–733. [Google Scholar] [CrossRef] [PubMed]
- Tran, M.; Van, C.; Barrera, D.J.; Pettersson, P.L.; Peinado, C.D.; Bui, J.; Mayfield, S.P. Production of unique immunotoxin cancer therapeutics in algal chloroplasts. Proc. Natl. Acad. Sci. USA 2012, 110, 15–22. [Google Scholar] [CrossRef]
- Gregory, J.A.; Li, F.; Tomosada, L.M.; Cox, C.J.; Topol, A.B.; Vinetz, J.M.; Mayfield, S.P. Algae-produced Pfs25 elicits antibodies that inhibit malaria transmission. PLoS One 2012, 7. [Google Scholar] [CrossRef]
- Lauersen, K.J.; Berger, H.; Mussgnug, J.H.; Kruse, O. Efficient recombinant protein production and secretion from nuclear transgenes in Chlamydomonas reinhardtii. J. Biotechnol. 2013, 167, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Mayfield, S.P.; Franklin, S.E. Expression of human antibodies in eukaryotic micro-algae. Vaccine 2005, 23, 1828–1832. [Google Scholar] [CrossRef] [PubMed]
- Tran, M.; Zhou, B.; Pettersson, P.L.; Gonzalez, M.J.; Mayfield, S.P. Synthesis and assembly of a full-length human monoclonal antibody in algal chloroplasts. Biotechnol. Bioeng. 2009, 104, 663–673. [Google Scholar] [PubMed]
- Kaslow, D.C.; Shiloach, J. Production, purification and immunogenicity of a malaria transmission-blocking vaccine candidate: TBV25H expressed in yeast and purified using nickel-NTA agarose. Nat. Biotechnol. 1994, 12, 494–499. [Google Scholar] [CrossRef]
- Wilken, L.R.; Nikolov, Z.N. Recovery and purification of plant-made recombinant proteins. Biotechnol. Adv. 2012, 30, 419–433. [Google Scholar] [CrossRef] [PubMed]
- Tran, M.; Henry, R.E.; Siefker, D.; Van, C.; Newkirk, G.; Kim, J.; Bui, J.; Mayfield, S.P. Production of anti-cancer immunotoxins in algae: Ribosome inactivating proteins as fusion partners. Biotechnol. Bioeng. 2013, 110, 2826–2835. [Google Scholar] [CrossRef] [PubMed]
- Manuell, A.L.; Beligni, M.V.; Elder, J.H.; Siefker, D.T.; Tran, M.; Weber, A.; McDonald, T.L.; Mayfield, S.P. Robust expression of a bioactive mammalian protein in Chlamydomonas chloroplast. Plant Biotechnol. J. 2007, 5, 402–412. [Google Scholar] [CrossRef] [PubMed]
- Surzycki, R.; Greenham, K.; Kitayama, K.; Dibal, F.; Wagner, R.; Rochaix, J.D.; Ajam, T.; Surzycki, S. Factors effecting expression of vaccines in microalgae. Biologicals 2009, 37, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Herman, C.A.; Im, C.S.; Beale, S.I. Light-regulated expression of the gsa gene encoding the chlorophyll biosynthetic enzyme glutamate 1-semialdehyde aminotransferase in carotenoid-deficient Chlamydomonas reinhardtii cells. Plant Mol. Biol. 1999, 39, 289–297. [Google Scholar] [CrossRef] [PubMed]
- Bohne, F.; Linden, H. Regulation of carotenoid biosynthesis genes in response to light in Chlamydomonas reinhardtii. Biochim. Biophys. Acta 2002, 1579, 26–34. [Google Scholar]
- Barneche, F.; Winter, V.; Crevecoeur, M.; Rochaix, J.D. ATAB2 is a novel factor in the signalling pathway of light-controlled synthesis of photosystem proteins. EMBO J. 2006, 25, 5907–5918. [Google Scholar] [CrossRef] [PubMed]
- Franklin, S.; Ngo, B.; Efuet, E.; Mayfield, S.P. Development of a GFP reporter gene for Chlamydomonas reinhardtii chloroplast. Plant J. 2002, 30, 733–744. [Google Scholar] [CrossRef] [PubMed]
- Gregory, J.A.; Topol, A.B.; Doerner, D.Z.; Mayfield, S.P. Alga-Produced Cholera Toxin-Pfs25 Fusion Proteins as Oral Vaccines. Appl. Environ. Microbiol. 2013, 79, 3917–3925. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef] [PubMed]
- Boyle, N.; Morgan, J. Flux balance analysis of primary metabolism in Chlamydomonas reinhardtii. BMC Syst. Biol. 2009, 3. [Google Scholar] [CrossRef]
- Malnoë, P.; Mayfield, S.P.; Rochaix, J.D. Comparative analysis of the biogenesis of photosystem II in the wild-type and Y-1 mutant of Chlamydomonas reinhardtii. J. Cell Biol. 1988, 106, 609–616. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, M.J.; Garcin, C.; van Hille, R.P.; Harrison, S.T.L. Interference by pigment in the estimation of microalgal biomass concentration by optical density. J. Microbiol. Methods 2011, 85, 119–123. [Google Scholar] [CrossRef] [PubMed]
- Villaverde, A.; Carrió, M.M. Protein aggregation in recombinant bacteria: Biological role of inclusion bodies. Biotechnol. Lett. 2003, 25, 1385–1395. [Google Scholar] [CrossRef] [PubMed]
- Miles, A.P.; Zhang, Y.; Saul, A.; Stowers, A.W. Large-scale purification and characterization of malariavaccine candidate antigen Pvs25H for use in clinical trials. Protein Expr. Purif. 2002, 25, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Miyata, T.; Harakuni, T.; Tsuboi, T.; Sattabongkot, J.; Kohama, H.; Tachibana, M.; Matsuzaki, G.; Torii, M.; Arakawa, A. Plasmodium vivax ookinete surface protein Pvs25 linked to cholera toxin B subunit induces potent transmission-blocking immunity by intranasal as well as subcutaneous immunization. Infect. Immun. 2010, 78, 3773–3782. [Google Scholar] [CrossRef] [PubMed]
- Coragliotti, A.; Beligni, M.; Franklin, S.; Mayfield, S.P. Molecular factors affecting the accumulation of recombinant proteins in the Chlamydomonas reinhardtii chloroplast. Mol. Biotechnol. 2011, 48, 60–75. [Google Scholar] [CrossRef] [PubMed]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Munjal, N.; Garzon-Sanabria, A.J.; Quinones, K.W.; Gregory, J.; Nikolov, Z.L. Light-Induced Production of An Antibody Fragment and Malaria Vaccine Antigen from Chlamydomonas reinhardtii. Processes 2014, 2, 625-638. https://doi.org/10.3390/pr2030625
Munjal N, Garzon-Sanabria AJ, Quinones KW, Gregory J, Nikolov ZL. Light-Induced Production of An Antibody Fragment and Malaria Vaccine Antigen from Chlamydomonas reinhardtii. Processes. 2014; 2(3):625-638. https://doi.org/10.3390/pr2030625
Chicago/Turabian StyleMunjal, Neera, Andrea Juliana Garzon-Sanabria, Katelyn Wilson Quinones, James Gregory, and Zivko L. Nikolov. 2014. "Light-Induced Production of An Antibody Fragment and Malaria Vaccine Antigen from Chlamydomonas reinhardtii" Processes 2, no. 3: 625-638. https://doi.org/10.3390/pr2030625
APA StyleMunjal, N., Garzon-Sanabria, A. J., Quinones, K. W., Gregory, J., & Nikolov, Z. L. (2014). Light-Induced Production of An Antibody Fragment and Malaria Vaccine Antigen from Chlamydomonas reinhardtii. Processes, 2(3), 625-638. https://doi.org/10.3390/pr2030625




