Evaluation of the Lipophilicity of 3,28-Disubstituted Betulin Derivatives with Promising Biological Properties
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Chromatography Procedure
2.3. Methods of Chemometric Analysis
2.3.1. Correlation Analysis
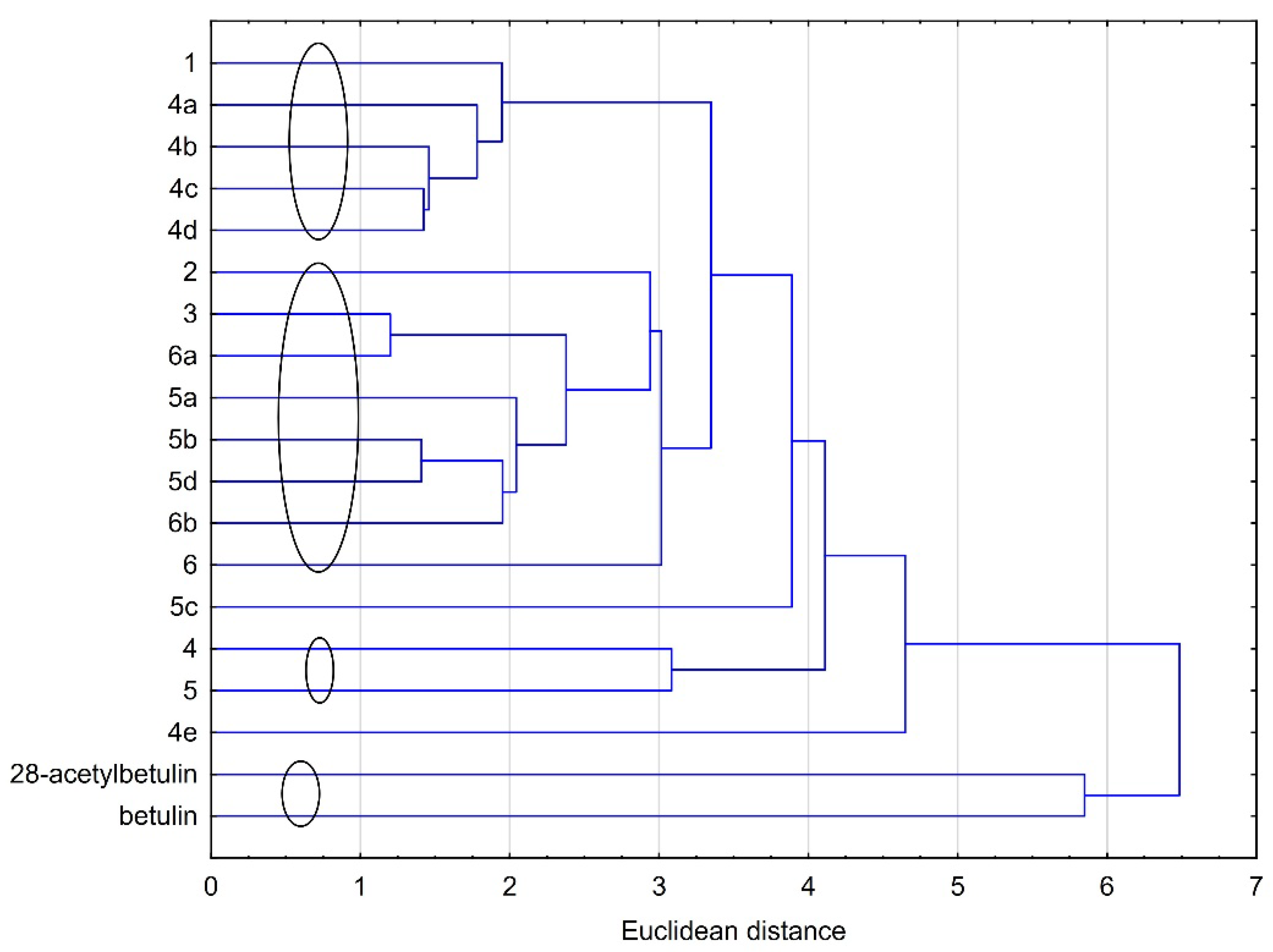
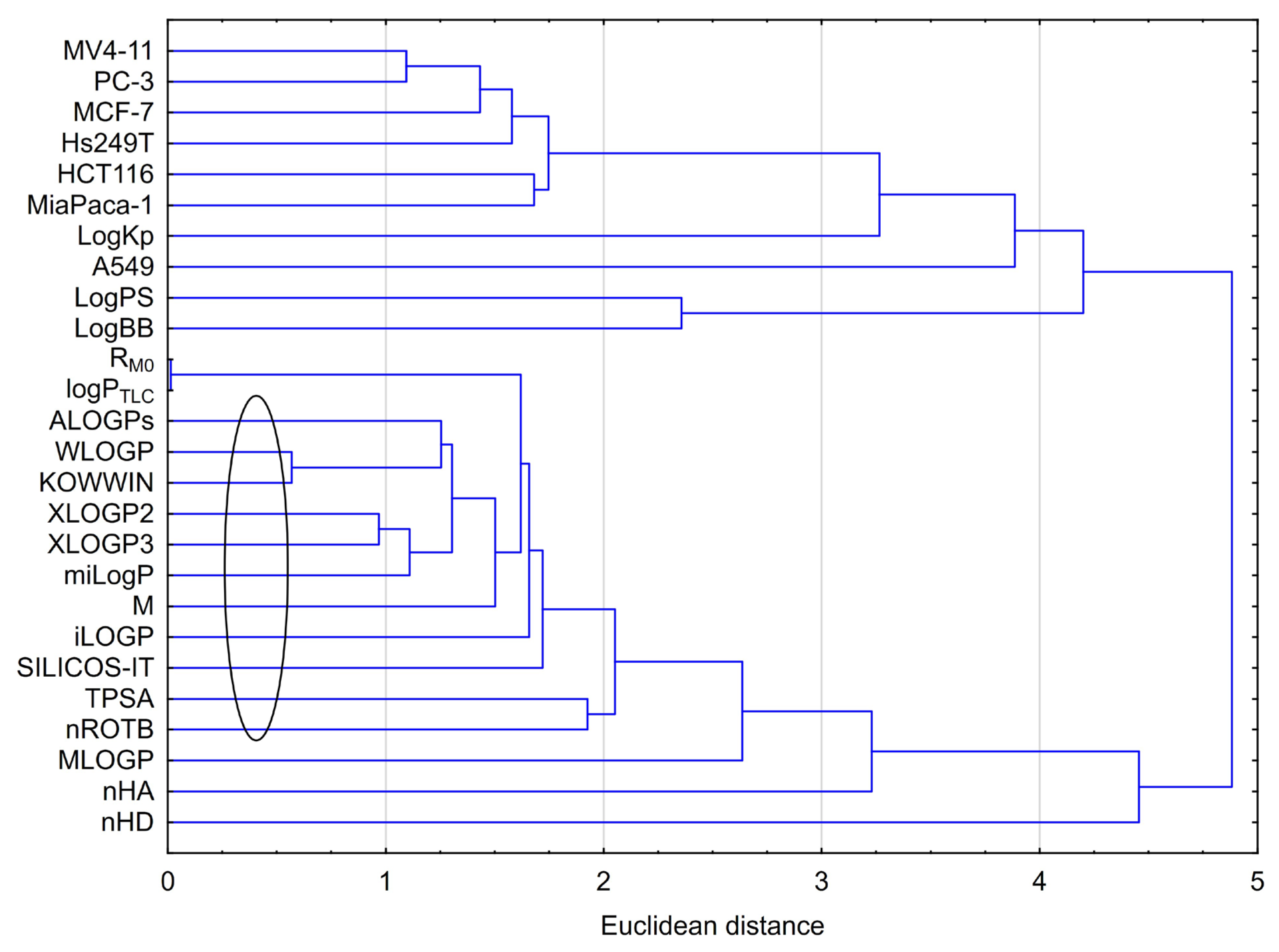
2.3.2. Cluster Analysis (CA)
2.3.3. Principal Component Analysis (PCA)
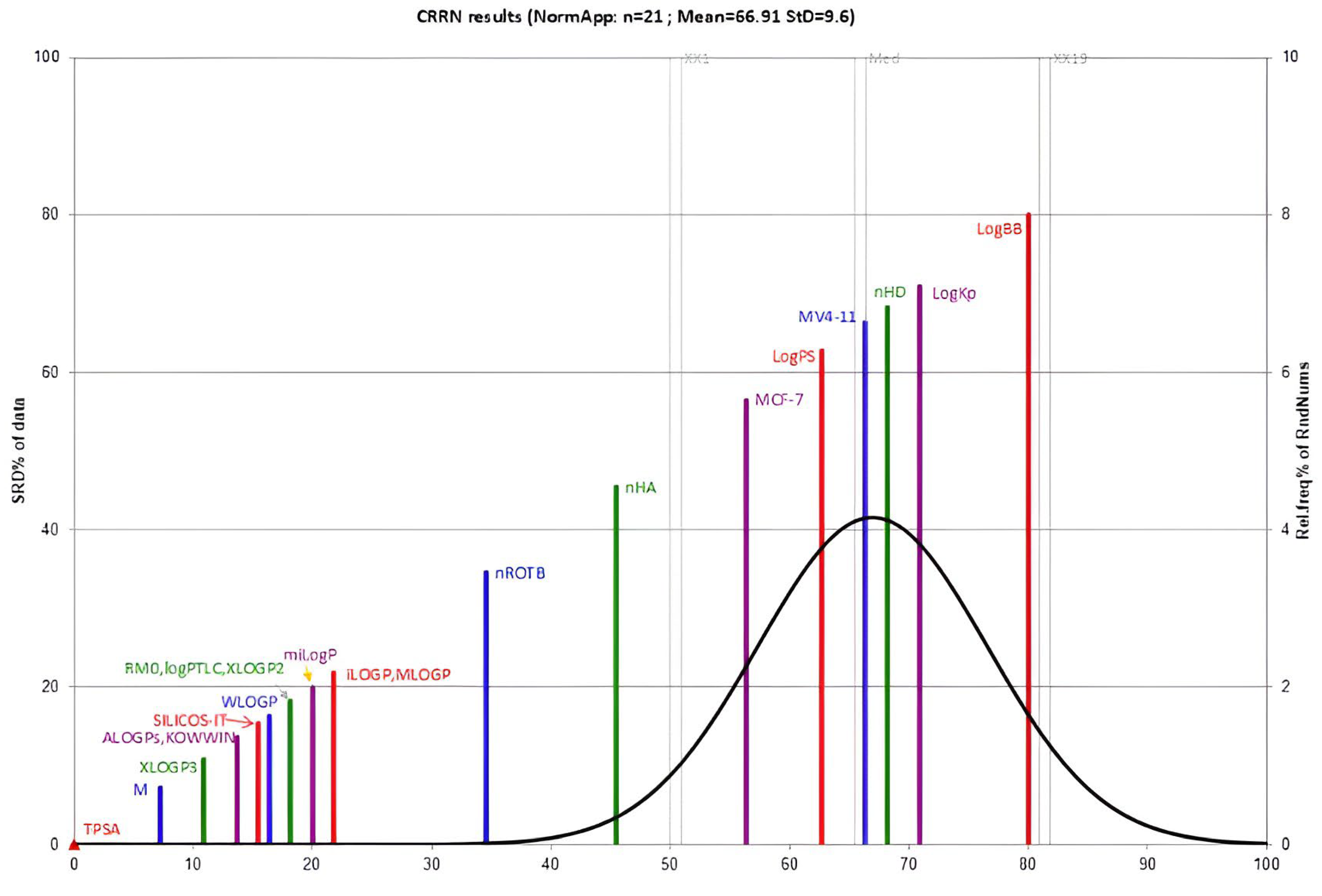
2.3.4. Sum of Ranking Differences (SRD)
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| SAR | structure–activity relationship |
| QSAR | quantitative structure–activity relationship |
| MW | molecular weight |
| RP-TLC | reversed-phase thin-layer chromatography |
| RP-18 | octadecylsilane |
| CA | cluster analysis |
| PCA | principal component analysis |
| SRD | sum of ranking differences |
| DDT | dichlorodiphenyltrichloroethane |
| TPSA | topological surface area |
| nROTB | number of rotatable bonds |
| nHD | number of donors of hydrogen bonds |
| nHA | number of acceptors of hydrogen bonds |
| logKp | skin permeability |
| LogPS | central nervous system permeability |
| LogBB | blood–brain barrier permeability |
References
- Amiri, S.; Dastghaib, S.; Ahmadi, M.; Mehrbod, P.; Khadem, F.; Behrouj, H.; Aghanoori, M.R.; Machaj, F.; Ghamsari, M.; Rosik, J.; et al. Betulin and its derivatives as novel compounds with different pharmacological effects. Biotechnol. Adv. 2020, 38, 107409. [Google Scholar] [CrossRef] [PubMed]
- Patrick, G.L. Part D Tools of the trade. In An Introduction to Medicinal Chemistry, 4th ed.; Oxford University Press: New York, NY, USA, 2009; pp. 377–382. [Google Scholar]
- Samiulla, D.S.; Vaidyanathan, V.V.; Arun, P.C.; Balan, G.; Blaze, M.; Bondre, S.; Chandrasekhar, G.; Gadakh, A.; Kumar, R.; Kharvi, G.; et al. Rational selection of structurally diverse natural product scaffolds with favorable ADME properties for drug discovery. Mol. Divers. 2005, 9, 131–139. [Google Scholar] [CrossRef] [PubMed]
- Demets, O.V.; Takibayeva, A.T.; Kassenov, R.Z.; Aliyeva, M.R. Methods of Betulin Extraction from Birch Bark. Molecules 2022, 27, 3621. [Google Scholar] [CrossRef] [PubMed]
- Jalmakhanbetova, R.I.; Mukusheva, G.K.; Abdugalimov, A.S.; Zhumagalieva, Z.Z.; Dehaen, W.; Anthonissen, S.; Suleimen, Y.M.; Seidakhmetova, R.B. Synthesis and Investigation of Biological Activity of New Betulonic Acid Derivatives Containing 1,2,3-Triazole Fragments. Molecules 2024, 29, 3149. [Google Scholar] [CrossRef]
- Ali-Seyed, M.; Jantan, I.; Vijayaraghavan, K.; Bukhari, S.N.A. Betulinic acid: Recent advances in chemical modifications, effective delivery, and molecular mechanisms of a promising anticancer therapy. Chem. Biol. Drug Des. 2016, 87, 517–536. [Google Scholar] [CrossRef]
- Pozharitskaya, O.N.; Karlina, M.V.; Shikov, A.N.; Kosman, V.M.; Makarov, V.G.; Casals, E.; Rosenholm, J.M. Pharmacokinetics and tissue disposition of nanosystem entrapped betulin after endotracheal administration to rats. Eur. J. Drug Metab. Pharmacokinet. 2017, 42, 327–332. [Google Scholar] [CrossRef]
- Jaroszewski, B.; Jelonek, K.; Kasperczyk, J. Drug Delivery Systems of Betulin and Its Derivatives: An Overview. Biomedicines 2024, 12, 1168. [Google Scholar] [CrossRef]
- Sousa, J.L.C.; Freire, C.S.R.; Silvestre, A.J.D.; Silva, A.M.S. Recent Developments in the Functionalization of Betulinic Acid and Its Natural Analogues: A Route to New Bioactive Compounds. Molecules 2019, 24, 355. [Google Scholar] [CrossRef]
- Drag-Zalesińska, M.; Wysocka, T.; Borska, S.; Drąg, M.; Poręba, M.; Choromańska, A.; Kulbacka, J.; Saczko, J. The new esters derivatives of betulin and betulinic acid in epidermoid squamous carcinoma treatment-In vitro studies. Biomed. Pharmacother. 2015, 72, 91–97. [Google Scholar] [CrossRef]
- Cai, S.; Guo, X.; Yang, H.; Zhao, T.; Li, Y.; Deng, N.; Gao, Z.; Meng, O.; Li, X.; Wang, S. Synthesis and antitumor effects of novel betulinic acid derivatives bearing electrophilic moieties. Bioorg. Med. Chem. 2025, 119, 118062. [Google Scholar] [CrossRef]
- Khan, D.A.U. Descriptors and their selection methods in QSAR analysis: Paradigm for drug design. Drug Discov. Today 2016, 21, 1291–1302. [Google Scholar] [CrossRef]
- Andric, F.; Bajusz, D.; Rácz, A.; Segane, S.; Héberger, K. Multivariate assessment of lipophilicity scales—Computational and reversed phase thin-layer chromatographic indices. J. Pharm. Biomed. Anal. 2016, 127, 81–93. [Google Scholar] [CrossRef]
- Šegan, S.; Krunić, M.J.; Andrić, D.B.; Šukalović, V.B.; Penjišević, J.Z.; Jevtić, I.I. Evaluation of lipophilicity and drug-likeness of donepezil-like compounds using reversed-phase thin-layer chromatography. Biomed. Chromatogr. 2024, 38, e5867. [Google Scholar] [CrossRef] [PubMed]
- Soares, J.X.; Santos, Á.; Fernandes, C.; Pinto, M.M.M. Liquid Chromatography on the Different Methods for the Determination of Lipophilicity: An Essential Analytical Tool in Medicinal Chemistry. Chemosensors 2022, 10, 340. [Google Scholar] [CrossRef]
- Chrobak, E.; Świtalska, M.; Wietrzyk, J.; Bębenek, E. New Difunctional Derivatives of Betulin: Preparation, Characterization and Antiproliferative Potential. Molecules 2025, 30, 611. [Google Scholar] [CrossRef] [PubMed]
- Sziklai, B.R.; Héberger, K. Apportionment and districting by Sum of Ranking Differences. PLoS ONE 2020, 15, e0229209. [Google Scholar] [CrossRef]
- Andrić, F.; Héberger, K. Chromatographic and computational assessment of lipophilicity using sum of ranking differences and generalized pair-correlation. J. Chromatogr. A 2015, 1380, 130–138. [Google Scholar] [CrossRef]
- Škrbić, B.; Héberger, K.; Durišić-Mladenović, N. Comparison of multianalyte proficiency test results by sum of ranking differences, principal component analysis, and hierarchical cluster analysis. Anal. Bioanal. Chem. 2013, 405, 8363–8375. [Google Scholar] [CrossRef]
- Héberger, K.; Škrbić, B. Ranking and similarity for quantitative structure–retention relationship models in predicting Lee retention indices of polycyclic aromatic hydrocarbons. Anal. Chim. Acta 2012, 716, 92–100. [Google Scholar] [CrossRef]
- Bodor, N.; Gabanyi, Z.; Wong, C.K. A New Method for the Estimation of Partition Coefficient. J. Am. Chem. Soc. 1989, 111, 3783–3786. [Google Scholar] [CrossRef]
- Berthod, A. Band broadening inside the chromatographic column: The interest of a liquid stationary phase. J. Chromatogr. A 2006, 1126, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Mannhold, R.; Cruciani, G.; Dross, K.; Rekker, R. Multivariate analysis of experimental and computational descriptors of molecular lipophilicity. J. Comput. Aided Mol. Des. 1998, 12, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Kujawski, J.; Popielarska, H.; Myka, A.; Drabińska, B.; Bernard, M.K. The log P Parameter as a Molecular Descriptor in the Computer-aided Drug Design—An Overview. Comput. Methods Sci. Technol. 2012, 18, 81–88. [Google Scholar] [CrossRef]
- Available online: http://swissadme.ch (accessed on 21 May 2025).
- Available online: https://www.molinspiration.com/services/logp.html (accessed on 5 January 2024).
- Tetko, I.V.; Gasteiger, J.; Todeschini, R.; Mauri, A.; Livingstone, D.; Ertl, P.; Palyulin, V.A.; Radchenko, E.V.; Zefirov, N.S.; Makarenko, A.S.; et al. Virtual computational chemistry laboratory-design and description. J. Comput. Aid Mol. Des. 2005, 19, 453–463. [Google Scholar] [CrossRef]
- Available online: https://vcclab.org (accessed on 12 September 2025).
- Estimation Programs Interface Suite™ for Microsoft® Windows, v 4.11; United States Environmental Protection Agency: Washington, DC, USA, 2017.
- Available online: https://biosig.lab.uq.edu.au/pkcsm (accessed on 21 May 2025).
- Sarkar, P.; Alheety, M.A.; Srivastava, V. Molecular Docking and ADMET Study of Spice-Derived Potential Phytochemicals Against Human DNA Topoisomerase III Alpha. Macromol. Symp. 2023, 407, 2200108. [Google Scholar] [CrossRef]
- Stępnik, K.; Kukula-Koch, W.; Plazinski, W.; Rybicka, M.; Gawel, K. Neuroprotective Properties of Oleanolic Acid—Computational-Driven Molecular Research Combined with In Vitro and In Vivo Experiments. Pharmaceuticals 2023, 16, 1234. [Google Scholar] [CrossRef]
- Salaković, B.; Kovačević, S.; Karadžić-Banjac, M.; Jevrić, L.; Podunavac-Kuzmanović, S.; Antonović, D. Comparative chemometric analysis, ranking and selection of lipophilicity parameters of 6-chloro-1,3,5-triazine derivatives with acyclic and cyclic substituents. Acta Period. Technol. 2022, 53, 88–99. [Google Scholar] [CrossRef]
- Sima, I.A.; Kot-Wasik, A.; Wasik, A.; Namieśnik, J.; Sârbu, C. Assessment of Lipophilicity Indices Derived from Retention Behavior of Antioxidant Compounds in RP-HPLC. Molecules 2017, 22, 550. [Google Scholar] [CrossRef]
- Šegan, S.; Božinović, N.; Opsenica, I.; Andrić, F. Consensus-based comparison of chromatographic and computationally estimated lipophilicity of benzothiepino [3,2-c]pyridine derivatives as potential antifungal drugs. J. Sep. Sci. 2017, 40, 2089–2096. [Google Scholar] [CrossRef]



| No. of Compound | Chemical Structure | No. of Compound | Chemical Structure |
|---|---|---|---|
| 1 | 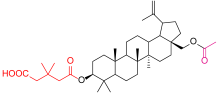 | 5 | 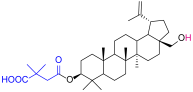 |
| 2 | 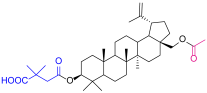 | 5a | 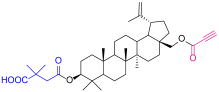 |
| 3 |  | 5b |  |
| 4 | 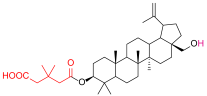 | 5c | 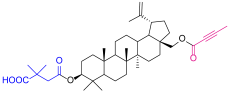 |
| 4a | 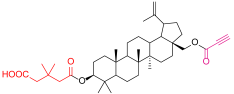 | 5d | 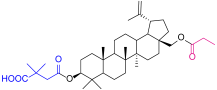 |
| 4b | 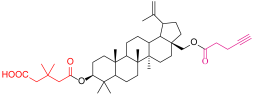 | 6 | 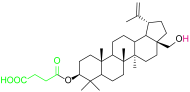 |
| 4c | 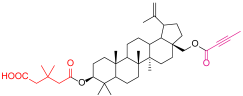 | 6a | 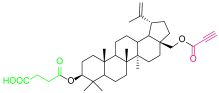 |
| 4d | 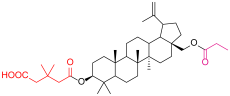 | 6b | 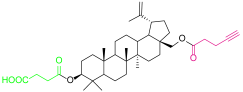 |
| 4e |  | betulin (BET) | 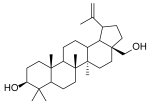 |
| 4f | 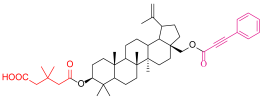 | 28-acetylbetulin (28-BET) | 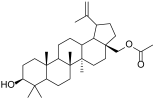 |
| Standard Substance | Chemical Name | RM0 | logPlit |
|---|---|---|---|
| A1 | acetanilide | 0.71 | 1.21 |
| A2 | prednisone | 0.85 | 1.62 |
| A3 | benzophenone | 2.02 | 3.18 |
| A4 | testosterone | 1.72 | 3.32 |
| A5 | anthracene | 3.03 | 4.45 |
| A6 | dibenzyl | 3.64 | 4.79 |
| A7 | 9-phenylanthracene | 3.98 | 6.01 |
| A8 | DDT | 4.67 | 6.38 |
| Compound | RM0 | logPTLC | iLOGP | AlogPs | XLOGP2 | XLOGP3 | WLOGP | MLOGP | SILICOS-IT | KOWWIN | miLogP |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 6.31 | 8.59 | 5.48 | 6.86 | 9.93 | 10.05 | 9.01 | 6.40 | 7.98 | 10.81 | 8.69 |
| 2 | 6.06 | 8.27 | 5.44 | 6.66 | 9.16 | 9.79 | 8.62 | 7.56 | 7.53 | 10.40 | 8.55 |
| 3 | 5.92 | 8.09 | 4.59 | 6.23 | 8.66 | 8.86 | 7.98 | 5.87 | 7.84 | 9.53 | 7.84 |
| 4 | 6.05 | 8.26 | 4.76 | 6.22 | 9.19 | 9.48 | 8.44 | 6.14 | 7.43 | 9.88 | 8.26 |
| 4a | 6.87 | 9.29 | 5.75 | 6.7 | 10.20 | 10.53 | 8.70 | 6.48 | 9.23 | 10.61 | 8.67 |
| 4b | 6.53 | 8.86 | 5.84 | 7.14 | 10.18 | 10.56 | 9.48 | 6.82 | 9.08 | 11.59 | 8.78 |
| 4c | 6.95 | 9.40 | 5.27 | 7.36 | 10.59 | 11.04 | 9.09 | 6.65 | 8.50 | 11.15 | 9.02 |
| 4d | 6.99 | 9.45 | 5.83 | 7.21 | 10.18 | 10.52 | 9.40 | 6.57 | 8.40 | 11.38 | 8.86 |
| 4e | 7.16 | 9.66 | 6.35 | 7.16 | 11.01 | 11.59 | 9.42 | 6.99 | 8.94 | 11.95 | 9.18 |
| 4f | 8.12 | 10.87 | 6.27 | 7.54 | 12.30 | 12.30 | 10.12 | 7.24 | 9.57 | 12.37 | 9.29 |
| 5 | 5.35 | 7.37 | 4.59 | 6.11 | 8.42 | 9.22 | 8.05 | 5.96 | 7.01 | 9.39 | 8.03 |
| 5a | 6.20 | 8.45 | 4.94 | 6.65 | 9.43 | 10.27 | 8.31 | 6.31 | 7.81 | 10.12 | 8.52 |
| 5b | 6.65 | 9.02 | 5.79 | 7.07 | 9.41 | 10.30 | 9.09 | 6.65 | 8.65 | 11.10 | 8.65 |
| 5c | 6.56 | 8.90 | 5.39 | 7.26 | 9.82 | 10.78 | 8.70 | 6.48 | 8.08 | 10.66 | 8.91 |
| 5d | 6.83 | 9.24 | 5.40 | 7.01 | 9.41 | 10.26 | 9.01 | 6.40 | 7.98 | 10.89 | 8.74 |
| 6 | 5.89 | 8.06 | 4.44 | 5.45 | 7.92 | 8.29 | 7.41 | 5.60 | 6.50 | 8.52 | 7.14 |
| 6a | 5.92 | 8.09 | 4.80 | 6.24 | 8.94 | 9.34 | 7.68 | 5.96 | 7.29 | 9.25 | 7.79 |
| 6b | 6.90 | 9.33 | 5.20 | 6.79 | 8.92 | 9.37 | 8.46 | 6.31 | 8.13 | 10.23 | 8.02 |
| BET | 4.51 | 6.31 | 4.47 | 5.34 | 7.81 | 8.28 | 7.00 | 6.00 | 6.21 | 8.18 | 7.16 |
| 28-BET | 5.33 | 7.35 | 4.93 | 5.53 | 8.55 | 8.86 | 7.57 | 6.20 | 6.85 | 8.77 | 7.87 |
| RM0 | logPTLC | iLOGP | ALOGPs | XLOGP2 | XLOGP3 | WLOGP | MLOGP | SILICOS-IT | KOWWIN | miLogP | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| RM0 | 1.000 | ||||||||||
| logPTLC | 0.999 | 1.000 | |||||||||
| iLOGP | 0.819 | 0.818 | 1.000 | ||||||||
| ALOGPs | 0.877 | 0.877 | 0.830 | 1.000 | |||||||
| XLOGP2 | 0.877 | 0.877 | 0.872 | 0.860 | 1.000 | ||||||
| XLOGP3 | 0.868 | 0.868 | 0.882 | 0.910 | 0.968 | 1.000 | |||||
| WLOGP | 0.886 | 0.886 | 0.900 | 0.933 | 0.908 | 0.914 | 1.000 | ||||
| MLOGP | 0.604 | 0.604 | 0.814 | 0.704 | 0.720 | 0.747 | 0.755 | 1.000 | |||
| SILICOS-IT | 0.889 | 0.888 | 0.881 | 0.888 | 0.890 | 0.882 | 0.901 | 0.656 | 1.000 | ||
| KOWWIN | 0.892 | 0.892 | 0.924 | 0.953 | 0.919 | 0.938 | 0.990 | 0.770 | 0.922 | 1.000 | |
| miLogP | 0.814 | 0.814 | 0.866 | 0.936 | 0.903 | 0.952 | 0.935 | 0.773 | 0.858 | 0.948 | 1.000 |
| RM0 | logPTLC | MV4-11 | A549 | MCF-7 | PC-3 | HCT116 | |
|---|---|---|---|---|---|---|---|
| RM0 | 1.000 | ||||||
| logPTLC | 0.999 | 1.000 | |||||
| MV4-11 | −0.737 | −0.737 | 1.000 | ||||
| A549 | −0.027 | −0.028 | 0.260 | 1.000 | |||
| MCF-7 | −0.692 | −0.692 | 0.918 | 0.448 | 1.000 | ||
| PC-3 | −0.728 | −0.727 | 0.942 | 0.241 | 0.884 | 1.000 | |
| HCT116 | −0.634 | −0.634 | 0.858 | 0.522 | 0.847 | 0.814 | 1.000 |
| Compound | MW [g/mol] | TPSA [Å2] | nROTB | nHD | nHA | Log Kp | Log PS | LogBB |
|---|---|---|---|---|---|---|---|---|
| 1 | 626.91 | 89.90 | 8 | 1 | 5 | −2.735 | −2.227 | −0.570 |
| 2 | 612.88 | 89.90 | 9 | 1 | 6 | −2.734 | −2.176 | −0.525 |
| 3 | 584.83 | 89.90 | 9 | 1 | 6 | −2.734 | −2.324 | −0.521 |
| 4 | 584.87 | 83.83 | 7 | 2 | 4 | −2.735 | −1.556 | −0.213 |
| 4a | 639.90 | 89.90 | 8 | 1 | 5 | −2.735 | −2.275 | −0.554 |
| 4b | 664.95 | 89.90 | 10 | 1 | 5 | −2.735 | −2.271 | −0.591 |
| 4c | 650.93 | 89.90 | 8 | 1 | 5 | −2.735 | −2.220 | −0.548 |
| 4d | 640.93 | 89.90 | 9 | 1 | 5 | −2.735 | −2.225 | −0.605 |
| 4e | 676.96 | 89.90 | 8 | 1 | 5 | −2.735 | −2.165 | −0.510 |
| 4f | 713.00 | 89.90 | 8 | 1 | 5 | −2.735 | −2.058 | −0.516 |
| 5 | 570.84 | 83.83 | 7 | 2 | 5 | −2.734 | −1.513 | 0.153 |
| 5a | 622.87 | 89.90 | 9 | 1 | 6 | −2.734 | −2.224 | −0.508 |
| 5b | 650.93 | 89.90 | 11 | 1 | 6 | −2.734 | −2.169 | −0.503 |
| 5c | 636.90 | 89.90 | 9 | 1 | 6 | −2.734 | −1.666 | 0.106 |
| 5d | 626.91 | 89.90 | 10 | 1 | 6 | −2.734 | −2.173 | −0.559 |
| 6 | 542.79 | 83.83 | 7 | 2 | 5 | −2.734 | −2.276 | −0.538 |
| 6a | 594.82 | 89.90 | 9 | 1 | 6 | −2.734 | −2.320 | −0.558 |
| 6b | 622.87 | 89.90 | 11 | 1 | 6 | −2.734 | −2.220 | −0.545 |
| BET | 442.72 | 40.46 | 2 | 2 | 2 | −2.737 | −1.770 | −0.444 |
| 28-BET | 484.75 | 46.53 | 3 | 1 | 3 | −2.614 | −2.244 | −0.326 |
| RM0 | logPTLC | MW | TPSA | nROTB | nHD | nHA | LogKp | LogPS | LogBB | |
|---|---|---|---|---|---|---|---|---|---|---|
| RM0 | 1.000 | |||||||||
| logPTLC | 0.999 | 1.000 | ||||||||
| MW | 0.917 | 0.916 | 1.000 | |||||||
| TPSA | 0.692 | 0.692 | 0.848 | 1.000 | ||||||
| nROTB | 0.636 | 0.635 | 0.778 | 0.897 | 1.000 | |||||
| nHD | −0.196 | −0.197 | −0.081 | 0.269 | 0.440 | 1.000 | ||||
| nHA | 0.422 | 0.423 | 0.359 | 0.0678 | −0.162 | −0.940 | 1.000 | |||
| LogKp | −0.306 | −0.305 | −0.462 | −0.624 | −0.530 | −0.241 | 0.022 | 1.000 | ||
| LogPS | −0.307 | −0.308 | −0.241 | −0.236 | −0.329 | 0.029 | −0.084 | −0.130 | 1.000 | |
| LogBB | −0.330 | −0.331 | −0.232 | −0.164 | −0.247 | 0.196 | −0.230 | 0.129 | 0.846 | 1.000 |
| Dependent Variable | Equation | r | s | F | Number of Equation |
|---|---|---|---|---|---|
| RM0 | 0.8794 (±0.1082) WLOGP − 1.1872 (±0.9320) 0.7684 (±0.0933) SILICOS-IT + 0.7684 (±0.0933) 0.6130 (±0.0733) KOWWIN + 0.0149 (±0.7624) −0.0474 (±0.0105) MV4-11 + 6.6205 (±0.1352) −0.0269 (±0.0062) PC-3 + 6.8227 (±0.1696) 0.2224 (±0.0637) nROTB + 4.5532 (±0.5343) 0.0114 (±0.0012) MW − 0.5814 (±0.7163) 0.0383 (±0.0094) TPSA + 3.1210 (±0.8063) | 0.886 0.889 0.892 −0.737 −0.728 0.636 0.917 0.692 | 0.374 0.370 0.365 0.476 0.484 0.632 0.324 0.583 | 66 68 70 20 19 12 95 17 | (4) (5) (6) (7) (8) (9) (10) (11) |
| logPTLC | 1.1112 (±0.1367) WLOGP − 0.8874 (±1.1774) 0.9703 (±0.1182) SILICOS-IT + 0.9287 (±0.9455) 0.7745 (±0.0926) KOWWIN + 0.6324 (±0.9635) −0.0957 (±0.0133) MV4-11 + 8.9787 (±0.1710) −0.0340 (±0.0078) PC-3 + 9.2341 (±0.2146) 0.2809 (±0.0804) nROTB + 6.3674 (±0.6754) 0.0144 (±0.0015) MW − 0.1194 (±0.9062) 0.0484 (±0.0119) TPSA + 4.5573 (±1.0190) | 0.886 0.888 0.892 −0.737 −0.727 0.635 0.916 0.692 | 0.472 0.468 0.462 0.602 0.612 0.789 0.408 0.737 | 66 67 70 20 19 12 94 17 | (12) (13) (14) (15) (16) (17) (18) (19) |
| Number of Eigenvalue | Eigenvalue | % of Total Variance |
|---|---|---|
| 1 | 15.188 | 58.41 |
| 2 | 4.388 | 75.29 |
| 3 | 1.969 | 82.86 |
| 4 | 1.424 | 88.34 |
| 5 | 1.017 | 92.25 |
| 6 | 0.778 | 95.24 |
| 7 | 0.457 | 97.00 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Chrobak, E.; Bober-Majnusz, K.; Świtalska, M.; Wietrzyk, J.; Bębenek, E. Evaluation of the Lipophilicity of 3,28-Disubstituted Betulin Derivatives with Promising Biological Properties. Processes 2026, 14, 708. https://doi.org/10.3390/pr14040708
Chrobak E, Bober-Majnusz K, Świtalska M, Wietrzyk J, Bębenek E. Evaluation of the Lipophilicity of 3,28-Disubstituted Betulin Derivatives with Promising Biological Properties. Processes. 2026; 14(4):708. https://doi.org/10.3390/pr14040708
Chicago/Turabian StyleChrobak, Elwira, Katarzyna Bober-Majnusz, Marta Świtalska, Joanna Wietrzyk, and Ewa Bębenek. 2026. "Evaluation of the Lipophilicity of 3,28-Disubstituted Betulin Derivatives with Promising Biological Properties" Processes 14, no. 4: 708. https://doi.org/10.3390/pr14040708
APA StyleChrobak, E., Bober-Majnusz, K., Świtalska, M., Wietrzyk, J., & Bębenek, E. (2026). Evaluation of the Lipophilicity of 3,28-Disubstituted Betulin Derivatives with Promising Biological Properties. Processes, 14(4), 708. https://doi.org/10.3390/pr14040708






