Application of Machine Learning Methods on Patient Reported Outcome Measurements for Predicting Outcomes: A Literature Review
Abstract
:1. Introduction
2. Methods
2.1. Review Design and Search Strategy
2.2. Article Selection
- Data: The dataset consists of structured questionnaires administered to patients or participants either in-person or via web application before, during and/or after a treatment. Articles that involved objectively measured data or data gathered from online patient forums were excluded from this study.
- Machine Learning: Application of machine learning methods with the intent of data analysis or clustering of patients or assessment of features with prognostic value for one or more target outcomes or building prognostic models for short- or long-term prediction of one or more outcome.
- Full text availability (including institutional access).
- Written in English.
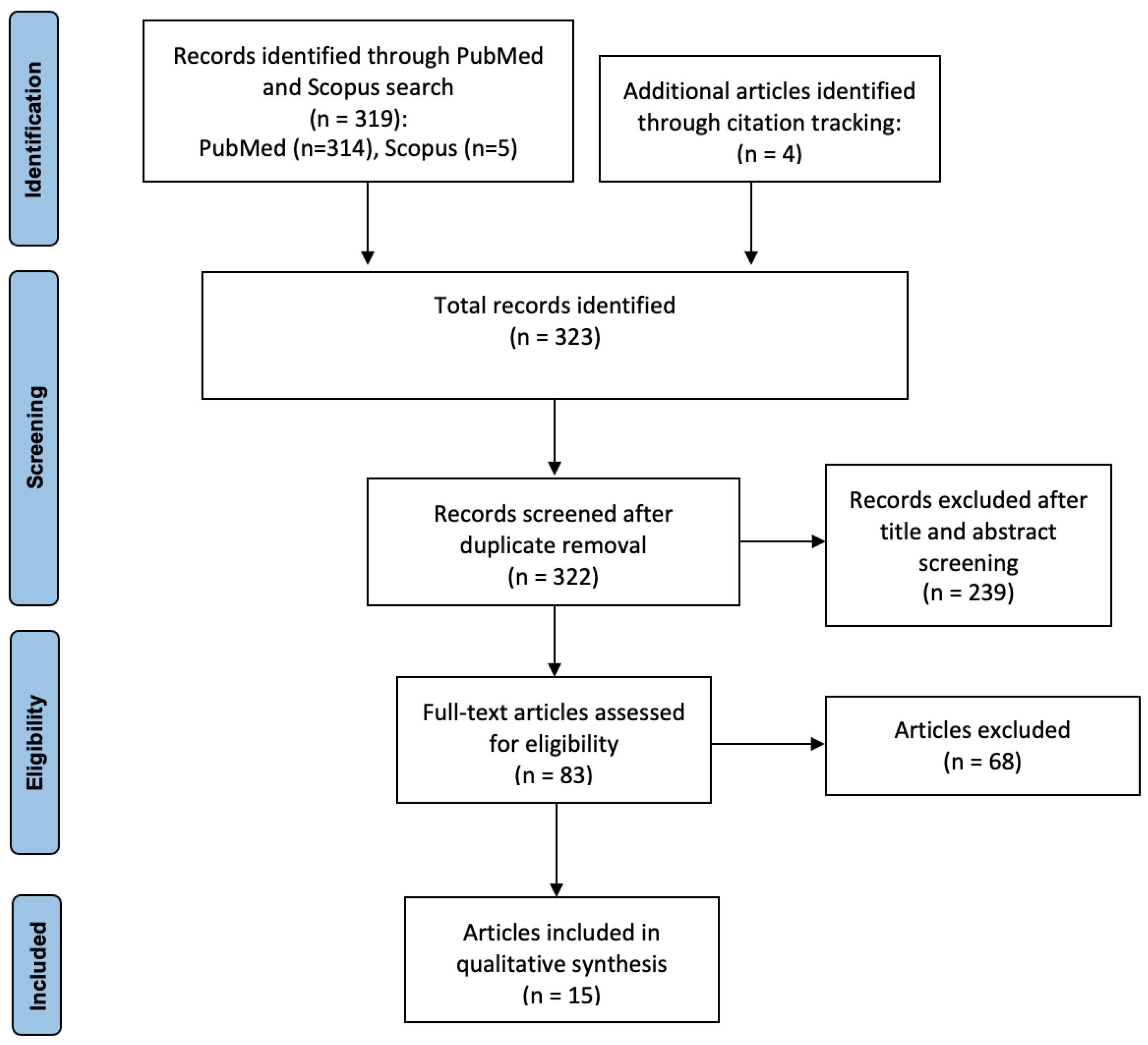
2.3. Search Outcome
2.4. Sources of Evidence
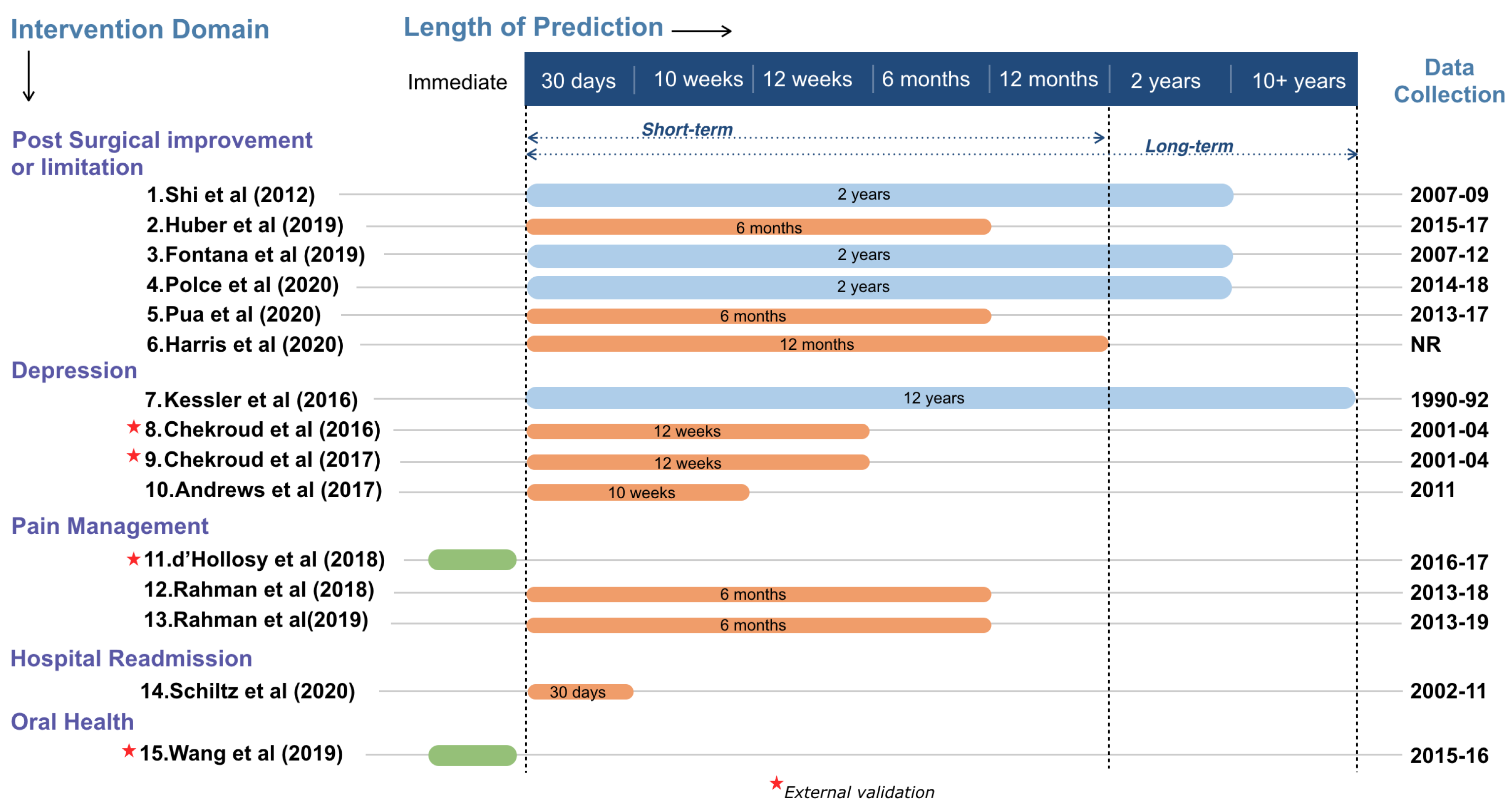
2.5. Intervention Domains and Length of Prediction
2.6. Sources of Data and Availability
2.7. Feature Selection
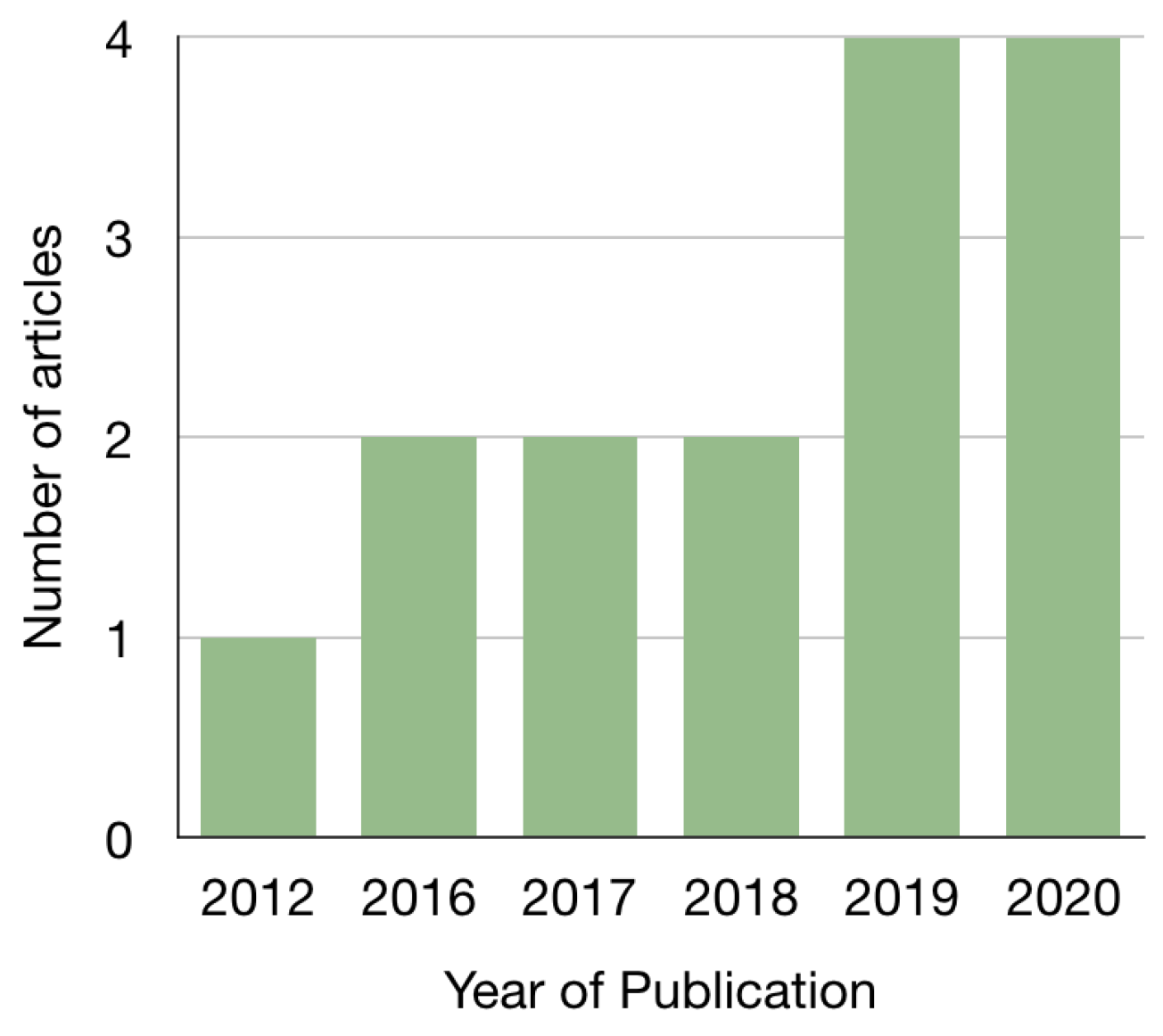
2.8. Trends in the Application of Machine Learning Methods
2.9. Study Design and Model Evaluation
2.10. Model Performance
3. Discussion
3.1. Gaps and Challenges
3.2. Limitations
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| NR | Not Reported |
| PROMs | Patient-Reported Outcome Measures |
| EHR | Electronic Health Records |
| CV | Cross Validation |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| ANOVA | Analysis of Variance |
| RoC | Receiver Operating Characteristic Curve |
| MCID | Minimal Clinically Important Difference |
| NIMH | National Institute of Mental Health |
| NHS | National Health Service |
| HRS | Health and Retirement Study |
References
- Kingsley, C.; Patel, S. Patient-reported outcome measures and patient-reported experience measures. Bja Educ. 2017, 17, 137–144. [Google Scholar] [CrossRef] [Green Version]
- Bingham III, C.O.; Noonan, V.K.; Auger, C.; Feldman, D.E.; Ahmed, S.; Bartlett, S.J. Montreal Accord on Patient-Reported Outcomes (PROs) use series—Paper 4: Patient-reported outcomes can inform clinical decision making in chronic care. J. Clin. Epidemiol. 2017, 89, 136–141. [Google Scholar] [CrossRef]
- Barry, M.J.; Edgman-Levitan, S. Shared decision making—The pinnacle patient-centered care. N. Engl. J. Med. 2012, 366, 780–781. [Google Scholar] [CrossRef] [Green Version]
- Coronado-Vázquez, V.; Canet-Fajas, C.; Delgado-Marroquín, M.T.; Magallón-Botaya, R.; Romero-Martín, M.; Gómez-Salgado, J. Interventions to facilitate shared decision-making using decision aids with patients in Primary Health Care: A systematic review. Medicine 2020, 99, e21389. [Google Scholar] [CrossRef]
- Sepucha, K.R.; Atlas, S.J.; Chang, Y.; Freiberg, A.; Malchau, H.; Mangla, M.; Rubash, H.; Simmons, L.H.; Cha, T. Informed, Patient-Centered Decisions Associated with Better Health Outcomes in Orthopedics: Prospective Cohort Study. Med. Decis. Mak. 2018, 38, 1018–1026. [Google Scholar] [CrossRef]
- Jayakumar, P.; Di, J.; Fu, J.; Craig, J.; Joughin, V.; Nadarajah, V.; Cope, J.; Bankes, M.; Earnshaw, P.; Shah, Z. A patient-focused technology-enabled program improves outcomes in primary total hip and knee replacement surgery. JBJS Open Access 2017, 2, e0023. [Google Scholar] [CrossRef] [PubMed]
- Giga, A. How health leaders can benefit from predictive analytics. In Healthcare Management Forum; SAGE Publications: Los Angeles, CA, USA, 2017; Volume 30, pp. 274–277. [Google Scholar]
- ShahabiKargar, Z.; Khanna, S.; Good, N.; Sattar, A.; Lind, J.; O’Dwyer, J. Predicting Procedure Duration to Improve Scheduling of Elective Surgery. In PRICAI 2014: Trends in Artificial Intelligence; Pham, D.N., Park, S.B., Eds.; Springer International Publishing: Cham, Switzerland, 2014; pp. 998–1009. [Google Scholar]
- Kargar, Z.S.; Khanna, S.; Sattar, A. Using prediction to improve elective surgery scheduling. Australas. Med. J. 2013, 6, 287. [Google Scholar] [CrossRef] [Green Version]
- Devi, S.P.; Rao, K.S.; Sangeetha, S.S. Prediction of surgery times and scheduling of operation theaters in optholmology department. J. Med. Syst. 2012, 36, 415–430. [Google Scholar] [CrossRef] [PubMed]
- Wong, D.; Oliver, C.; Moonesinghe, S. Predicting postoperative morbidity in adult elective surgical patients using the Surgical Outcome Risk Tool (SORT). BJA Br. J. Anaesth. 2017, 119, 95–105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moonesinghe, S.R.; Mythen, M.G.; Das, P.; Rowan, K.M.; Grocott, M.P. Risk stratification tools for predicting morbidity and mortality in adult patients undergoing major surgery: Qualitative systematic review. Anesthesiology 2013, 119, 959–981. [Google Scholar] [CrossRef] [Green Version]
- Marufu, T.C.; White, S.; Griffiths, R.; Moonesinghe, S.; Moppett, I.K. Prediction of 30-day mortality after hip fracture surgery by the Nottingham Hip Fracture Score and the Surgical Outcome Risk Tool. Anaesthesia 2016, 71, 515–521. [Google Scholar] [CrossRef]
- Singal, A.G.; Mukherjee, A.; Elmunzer, B.J.; Higgins, P.D.; Lok, A.S.; Zhu, J.; Marrero, J.A.; Waljee, A.K. Machine learning algorithms outperform conventional regression models in predicting development of hepatocellular carcinoma. Am. J. Gastroenterol. 2013, 108, 1723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mansell, G.; Corp, N.; Wynne-Jones, G.; Hill, J.; Stynes, S.; van der Windt, D. Self-reported prognostic factors in adults reporting neck or low back pain: An umbrella review. Eur. J. Pain 2021, 25, 1627–1643. [Google Scholar] [CrossRef] [PubMed]
- Krismer, M.; Van Tulder, M. Low back pain (non-specific). Best Pract. Res. Clin. Rheumatol. 2007, 21, 77–91. [Google Scholar] [CrossRef] [PubMed]
- Waljee, A.K.; Higgins, P.D.; Singal, A.G. A primer on predictive models. Clin. Transl. Gastroenterol. 2014, 5, e44. [Google Scholar] [CrossRef] [PubMed]
- Andrews, J.; Harrison, R.; Brown, L.; MacLean, L.; Hwang, F.; Smith, T.; Williams, E.A.; Timon, C.; Adlam, T.; Khadra, H.; et al. Using the NANA toolkit at home to predict older adults’ future depression. J. Affect. Disord. 2017, 213, 187–190. [Google Scholar] [CrossRef]
- Wang, X.; Gottumukkala, V. Patient Reported Outcomes: Is this the Missing Link in Patient-centered Perioperative Care? Best Pract. Res. Clin. Anaesthesiol. 2020. [Google Scholar] [CrossRef]
- Baumhauer, J. Patient-Reported Outcomes—Are They Living Up to Their Potential? N. Engl. J. Med. 2017, 377, 6–9. [Google Scholar] [CrossRef] [Green Version]
- Shi, H.Y.; Tsai, J.T.; Chen, Y.M.; Culbertson, R.; Chang, H.T.; Hou, M.F. Predicting two-year quality of life after breast cancer surgery using artificial neural network and linear regression models. Breast Cancer Res. Treat. 2012, 135, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Huber, M.; Kurz, C.; Leidl, R. Predicting patient-reported outcomes following hip and knee replacement surgery using supervised machine learning. BMC Med. Inform. Decis. Mak. 2019, 19, 3. [Google Scholar] [CrossRef] [Green Version]
- Pua, Y.H.; Kang, H.; Thumboo, J.; Clark, R.A.; Chew, E.S.X.; Poon, C.L.L.; Chong, H.C.; Yeo, S.J. Machine learning methods are comparable to logistic regression techniques in predicting severe walking limitation following total knee arthroplasty. Knee Surg. Sport. Traumatol. Arthrosc. 2019, 28, 3207–3216. [Google Scholar] [CrossRef]
- Fontana, M.A.; Lyman, S.; Sarker, G.K.; Padgett, D.E.; MacLean, C.H. Can machine learning algorithms predict which patients will achieve minimally clinically important differences from total joint arthroplasty? Clin. Orthop. Relat. Res. 2019, 477, 1267–1279. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Polce, E.M.; Kunze, K.N.; Fu, M.; Garrigues, G.E.; Forsythe, B.; Nicholson, G.P.; Cole, B.J.; Verma, N.N. Development of Supervised Machine Learning Algorithms for Prediction of Satisfaction at Two Years Following Total Shoulder Arthroplasty. J. Shoulder Elb. Surg. 2020, 30, e290–e299. [Google Scholar] [CrossRef] [PubMed]
- Harris, A.H.; Kuo, A.C.; Weng, Y.; Trickey, A.W.; Bowe, T.; Giori, N.J. Can machine learning methods produce accurate and easy-to-use prediction models of 30-day complications and mortality after knee or hip arthroplasty? Clin. Orthop. Relat. Res. 2019, 477, 452. [Google Scholar] [CrossRef] [PubMed]
- Kessler, R.C.; van Loo, H.M.; Wardenaar, K.J.; Bossarte, R.M.; Brenner, L.A.; Cai, T.; Ebert, D.D.; Hwang, I.; Li, J.; de Jonge, P.; et al. Testing a machine-learning algorithm to predict the persistence and severity of major depressive disorder from baseline self-reports. Mol. Psychiatry 2016, 21, 1366–1371. [Google Scholar] [CrossRef]
- Chekroud, A.M.; Zotti, R.J.; Shehzad, Z.; Gueorguieva, R.; Johnson, M.K.; Trivedi, M.H.; Cannon, T.D.; Krystal, J.H.; Corlett, P.R. Cross-trial prediction of treatment outcome in depression: A machine learning approach. Lancet Psychiatry 2016, 3, 243–250. [Google Scholar] [CrossRef]
- Chekroud, A.M.; Gueorguieva, R.; Krumholz, H.M.; Trivedi, M.H.; Krystal, J.H.; McCarthy, G. Reevaluating the efficacy and predictability of antidepressant treatments: A symptom clustering approach. JAMA Psychiatry 2017, 74, 370–378. [Google Scholar] [CrossRef]
- Rahman, Q.A.; Janmohamed, T.; Pirbaglou, M.; Clarke, H.; Ritvo, P.; Heffernan, J.M.; Katz, J. Defining and predicting pain volatility in users of the Manage My Pain app: Analysis using data mining and machine learning methods. J. Med. Internet Res. 2018, 20, e12001. [Google Scholar] [CrossRef]
- Rahman, Q.A.; Janmohamed, T.; Clarke, H.; Ritvo, P.; Heffernan, J.; Katz, J. Interpretability and class imbalance in prediction models for pain volatility in manage my pain app users: Analysis using feature selection and majority voting methods. JMIR Med. Inform. 2019, 7, e15601. [Google Scholar] [CrossRef]
- Nijeweme-d’Hollosy, W.; van Velsen, L.; Poel, M.; Groothuis-Oudshoorn, C.; Soer, R.; Hermens, H. Evaluation of Three Machine Learning Models for Self-Referral Decision Support on Low Back Pain in Primary Care. Int. J. Med. Inform. 2018, 110, 31–41. [Google Scholar] [CrossRef]
- Schiltz, N.; Dolansky, M.; Warner, D.; Stange, K.; Gravenstein, S.; Koroukian, S. Impact of Instrumental Activities of Daily Living Limitations on Hospital Readmission: An Observational Study Using Machine Learning. J. Gen. Intern. Med. 2020, 35, 2865–2872. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hays, R.D.; Marcus, M.; Maida, C.; Shen, J.; Xiong, D.; Coulter, I.; Lee, S.; Spolsky, V.; Crall, J.; et al. Developing Children’s Oral Health Assessment Toolkits Using Machine Learning Algorithm. JDR Clin. Transl. Res. 2020, 5, 233–243. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Bleeker, S.; Moll, H.; Steyerberg, E.; Donders, A.; Derksen-Lubsen, G.; Grobbee, D.; Moons, K. External validation is necessary in prediction research:: A clinical example. J. Clin. Epidemiol. 2003, 56, 826–832. [Google Scholar] [CrossRef]
- Steyerberg, E.W.; Harrell, F.E., Jr. Prediction models need appropriate internal, internal-external, and external validation. J. Clin. Epidemiol. 2016, 69, 245. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.; Tong, L.; Yao, J.; Guo, Z.; Lui, K.Y.; Hu, X.; Cao, L.; Zhu, Y.; Huang, F.; Guan, X.; et al. Association of sex with clinical outcome in critically ill sepsis patients: A retrospective analysis of the large clinical database MIMIC-III. Shock 2019, 52, 146. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; McDermott, M.B.; Chauhan, G.; Ghassemi, M.; Hughes, M.C.; Naumann, T. Mimic-extract: A data extraction, preprocessing, and representation pipeline for mimic-iii. In Proceedings of the ACM Conference on Health, Inference, and Learning, Toronto, ON, Canada, 2–4 April 2020; pp. 222–235. [Google Scholar]
- Feng, M.; McSparron, J.I.; Kien, D.T.; Stone, D.J.; Roberts, D.H.; Schwartzstein, R.M.; Vieillard-Baron, A.; Celi, L.A. Transthoracic echocardiography and mortality in sepsis: Analysis of the MIMIC-III database. Intensive Care Med. 2018, 44, 884–892. [Google Scholar] [CrossRef] [PubMed]
| Article | Outcome | Dataset Size | Total No. of Features | Features Selected | Feature Selection Method | Hyperparameter Tuning | Model Evaluation | Data Availability | External Validation |
|---|---|---|---|---|---|---|---|---|---|
| Shi et al. [21] | Quality of life post surgery | 403 | NR | NR | ANOVA, Fisher exact analysis, Univariate analysis | NR | Holdout (80,20) | NR | no |
| Huber et al. [22] | MCID post surgery | 64,634 | 81 | NR | Algorithm implicit | NR | 5-fold CV | NHS 1 | no |
| Fontana et al. [24] | MCID post surgery | 13,809 | NR | Manual | 5-fold CV | Holdout (80,20) | NR | no | |
| Polce et al. [25] | Satisfaction post surgery | 413 | 16 | 10 | Recursive Feature Elimination, Random Forest | 10-fold CV | Holdout (80,20) | NR | no |
| Pua et al. [23] | Walking limitation post surgery | 4026 | NR | 25 | Manual | 5-fold CV | Holdout (70,30) | NR | no |
| Harris et al. [26] | MCID post surgery | 587 | NR | NR | Manual | NR | 10-fold CV, bootstrapping | NR | no |
| Kessler et al. [27] | Depressive Disorder chronicity, persistence, severity | 1056 | NR | 9–13 | Ensemble and Penalised Regression | NR | 10-fold CV | NR | no |
| Chekroud et al. [28] | Antidepressant treatment | 1949 | 164 | 25 | ElasticNet | RoC maximisation | 10*Repeated 10-fold CV | NIMH 2 | yes |
| Chekroud et al. [29] | Antidepressant treatment | 7221 | 164 | 25 | ElasticNet | NR | 5-fold CV | NIMH 2 | yes |
| Andrews et al. [18] | Depression in older adults | 37 | 6 | 2 | LASSO | Stratified CV | 5-fold CV | NR | no |
| d’Hollosy et al. [32] | Low Back pain self-referral | 1288 | 15 | NR | Algorithm implicit | NR | Holdout (70,30) | On Request | yes |
| Rahman et al. [30] | Pain volatility | 782 | 130 | NR | Algorithm implicit | NR | 5-fold CV | NR | no |
| Rahman et al. [31] | Pain volatility | 879 | 132 | 9 | Gini impurity, Information gain, Class imbalance | NR | 5-fold CV | NR | no |
| Schiltz et al. [33] | Hospital Readmission | 6617 | NR | NR | Random Forest | NR | Holdout (80,20) | HRS | no |
| Wang et al. [34] | Oral Health | 908 | 27 | NA | Manual | Greedy approximation [35] | Holdout (70,30) | NR | yes |
| Article | Supervised | Unsupervised | Machine Learning Task | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ensemble Methods | Linear Methods | DT | SVM | NN | NB | k-NN | QDA | k-Means | Aggl. | ||
| Shi et al. [21] | ✓ | ✓ | Regression | ||||||||
| Huber et al. [22] | ✓ | ✓ | ✓ | ✓ | ✓ | Classification | |||||
| Fontana et al. [24] | ✓ | ✓ | ✓ | Classification | |||||||
| Polce et al. [25] | ✓ | ✓ | ✓ | ✓ | Classification | ||||||
| Pua et al. [23] | ✓ | ✓ | Classification | ||||||||
| Harris et al. [26] | ✓ | ✓ | ✓ | Classification | |||||||
| Kessler et al. [27] | ✓ | ✓ | Classification | ||||||||
| Chekroud et al. [28] | ✓ | Classification | |||||||||
| Chekroud et al. [29] | ✓ | ✓ | Regression | ||||||||
| Andrews et al. [18] | ✓ | Classification | |||||||||
| d’Hollosy et al. [32] | ✓ | ✓ | Classification | ||||||||
| Rahman et al. [30] | ✓ | ✓ | ✓ | ✓ | Classification | ||||||
| Rahman et al. [31] | ✓ | ✓ | ✓ | ✓ | Classification | ||||||
| Schiltz et al. [33] | ✓ | ✓ | ✓ | Classification | |||||||
| Wang et al. [34] | ✓ | ✓ | Regression, Classification | ||||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Verma, D.; Bach, K.; Mork, P.J. Application of Machine Learning Methods on Patient Reported Outcome Measurements for Predicting Outcomes: A Literature Review. Informatics 2021, 8, 56. https://doi.org/10.3390/informatics8030056
Verma D, Bach K, Mork PJ. Application of Machine Learning Methods on Patient Reported Outcome Measurements for Predicting Outcomes: A Literature Review. Informatics. 2021; 8(3):56. https://doi.org/10.3390/informatics8030056
Chicago/Turabian StyleVerma, Deepika, Kerstin Bach, and Paul Jarle Mork. 2021. "Application of Machine Learning Methods on Patient Reported Outcome Measurements for Predicting Outcomes: A Literature Review" Informatics 8, no. 3: 56. https://doi.org/10.3390/informatics8030056
APA StyleVerma, D., Bach, K., & Mork, P. J. (2021). Application of Machine Learning Methods on Patient Reported Outcome Measurements for Predicting Outcomes: A Literature Review. Informatics, 8(3), 56. https://doi.org/10.3390/informatics8030056







