5.1. Collapsing Retrospective Provenance
Given a directed graph , where V is the set of vertices (in our graph, a vertex is of type “file” or of type “process”) and E is the set of edges, we denote and as the sets of input and output edges of vertex u. Respectively, , and . The direction of an edge characterizes the dependency of its vertices. For example, a process u spawned by process v is represented by the edge , and a file u read by process v is represented by the edge . The graph G is collapsed based on the following two rules:
Rule 1.
Similarity. Two vertices u and v arecalled similar if and only if they share the same type and have the same input and output connection sets: , and .
The similarity rule groups multiple vertices into a single vertex if the vertices (i) have the same type and (ii) are connected by the same number and type of edges. Additionally, edges of similar vertices will be grouped into a single corresponding edge. Since the provenance graph follows W3C PROV-DM standard, each file is of type entity and each process is of type activity. When applied to our provenance graph, this rule groups different files and processes that are similar each other into the summary groups of files and processes (see
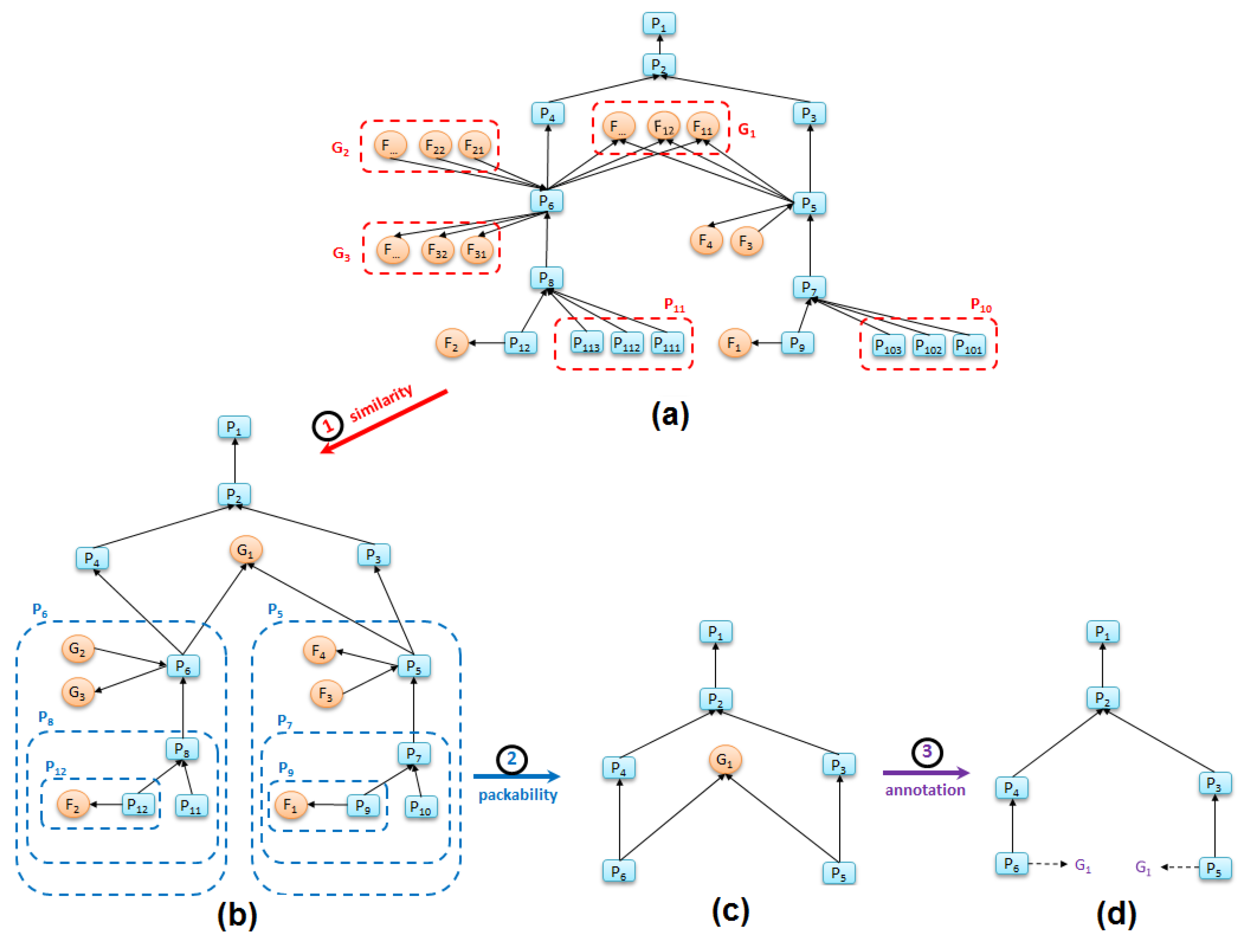
Figure 5a).
Rule 2. Packability. A vertex u belongs to v’s if and only if vertex u connects to v and satisfies one of the following conditions:
Vertex u is a file that has only one edge to process v: and {}.
Vertex u is a process that has only one output edge to process v: and {}.
Vertex u is a file that has only two edges—an output edge to process v and an input edge from another process x: and {}.
The packability rule identifies hubs in the provenance graph by packing files or processes that are connected by single edges to their parent nodes. It also packs files that are generated and consumed by a single process into their parent processes by producing a process-to-process edge.
When applied in sequence, the similarity and packability rules condense the detail-level of a graph while preserving its core workflow elements.
Figure 5 (all process names and file names are simplified for brevity) illustrates how applying these two rules to a replete graph produces a graph summary that shows the primary processes in a workflow.
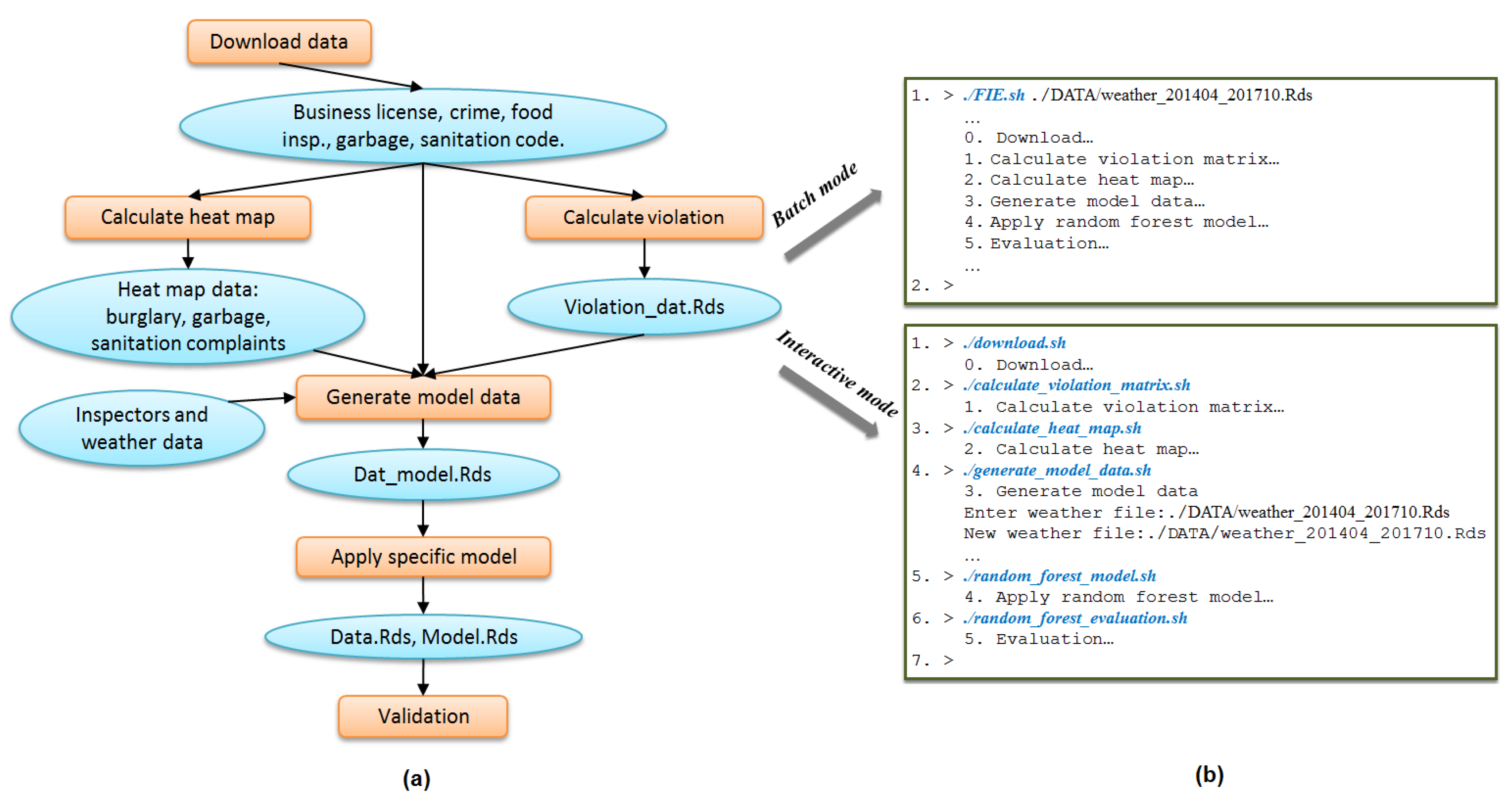
Figure 5a presents the original replete provenance graph of one sub-task of the FIE workflow (the data processing steps “Calculate Violation” and “Calculate Heat Map” of
Figure 1a). Applying the two summarization rules produces the graph in
Figure 5c.
We use an annotation method that assigns higher collapsibility to file nodes than process nodes, since an application workflow is typically defined by the primary processes that it runs.
Figure 5d shows how the annotation “G_1”, which is a library dependency used both by “P_5” and “P_6”, is attached to the two process nodes that generated it. Thus, given a file with
n edges (
), we replace this file with
n annotations.
Figure 6 shows the expanded view of node “P_R_27070” (“P_5” in
Figure 5b). In
Figure 6, similarity and packability rules group the nodes within the box into the single node “P_R_27070” (process 27070 runs a subprocess using file “21_calulate_violation_matrix.R” (“F_3” in
Figure 5b) and write data to file “violation_data.Rds” (F_4 in
Figure 5b)). These nodes are application nodes and not system nodes. Here, “Process_G_5” (P_7 in
Figure 5b), another concealing node, correctly hides all the dependencies of the R process calculating the violation matrix.
5.2. Summarizing Retrospective Provenance to Generate Prospective Provenance
The method in the previous section summarizes retrospective provenance by collapsing information. However, such summaries still differ from the conceptual view of the applications. For example,
Figure 1a, which is more familiar to users, is very different in view than
Figure 5d. Another equally important goal of summarization is to summarize retrospective provenance such that it (potentially) matches application workflow. In some situations, this application workflow may be available in the form of prospective provenance [
34]. If available, summary methods can take advantage of this available information. In containers created of ad hoc applications, however, application workflows are rarely available. Therefore, we describe a summarization method that determines the lineage history of nodes and uses this information to summarize retrospective provenance.
Our method to summarize retrospective provenance is based on ideas described in SNAP (Summarization by Grouping Nodes on Attributes and Pairwise Relationships) [
37]. SNAP is a non-statistical method for summarizing undirected and directed graph nodes and edges based on their respective types. In brief, it first groups nodes of the same type. It then recursively sub-divides to form smaller groups of nodes that still have have the same node type but also same relationship type with other groups. SNAP considers direct relationship types amongst group nodes and not relationship types due to ancestry of the nodes. Thus, grouping provided by SNAP can be further improved for provenance graphs by considering ancestral history of nodes while grouping. If ancestral relationships are considered, then, for a node, ancestors or descendants will not be grouped together since, by definition, the ancestral history of an ancestor and its descendent is different. Similarly, nodes in the same group will not share any relationship because then their ancestral history will be different.
To identify nodes with the same derivation history, first nodes of the same type are grouped, defined as
Definition 2 (Node Grouping). Given a provenance graph , is a node-grouping such that
- (1)
or , and ,
- (2)
,
- (3)
and .
In particular, in (1), node grouping is over nodes of two types: activity nodes and entity nodes, in (2), the union of all node grouping is equal to the nodes in G; and, in (3), given a node grouping, groups are not overlapping, but distinct.
We now consider grouping G by ancestry. For this, we identify the ancestors of a node as follows. For a given grouping , the ancestors of a node v is the set . Type of edges is based on types used in W3C PROV standard where . Nodes that do not have an ancestor are assigned the start node as an ancestor, with a start label edge. Now, we define grouping nodes by ancestry.
Definition 3 (Ancestry grouping). A grouping has the same ancestry if it satisfies the following:
- (i)
Node Grouping Definition 2,
- (ii)
, if , then , .
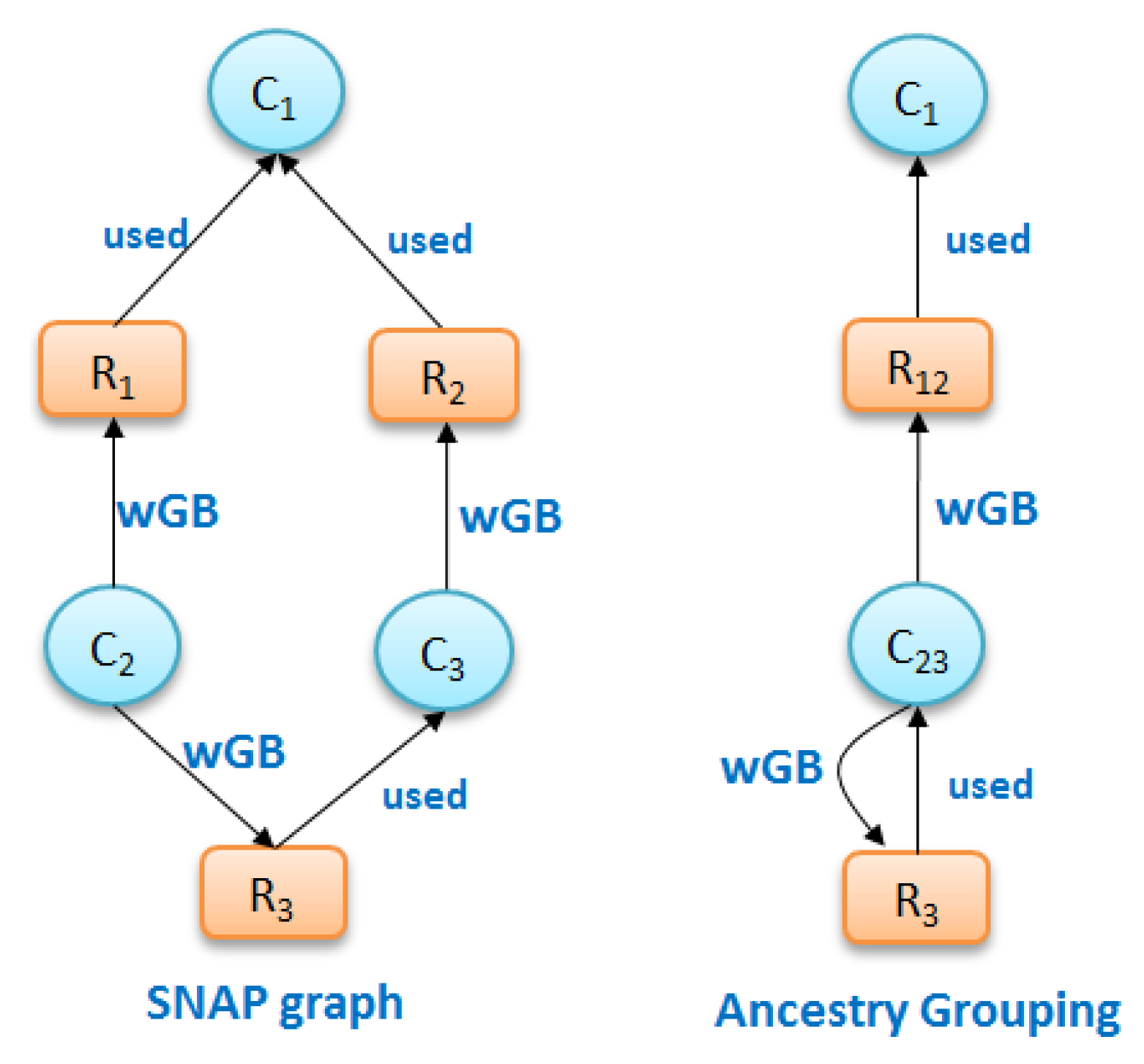
Figure 7a shows the grouping of nodes due to SNAP, in which nodes with the same types and same ancestors are separated into different groups if their descendants are different, and due to ancestry grouping in
Figure 7b, in which nodes of the same type with same ancestors remain grouped.
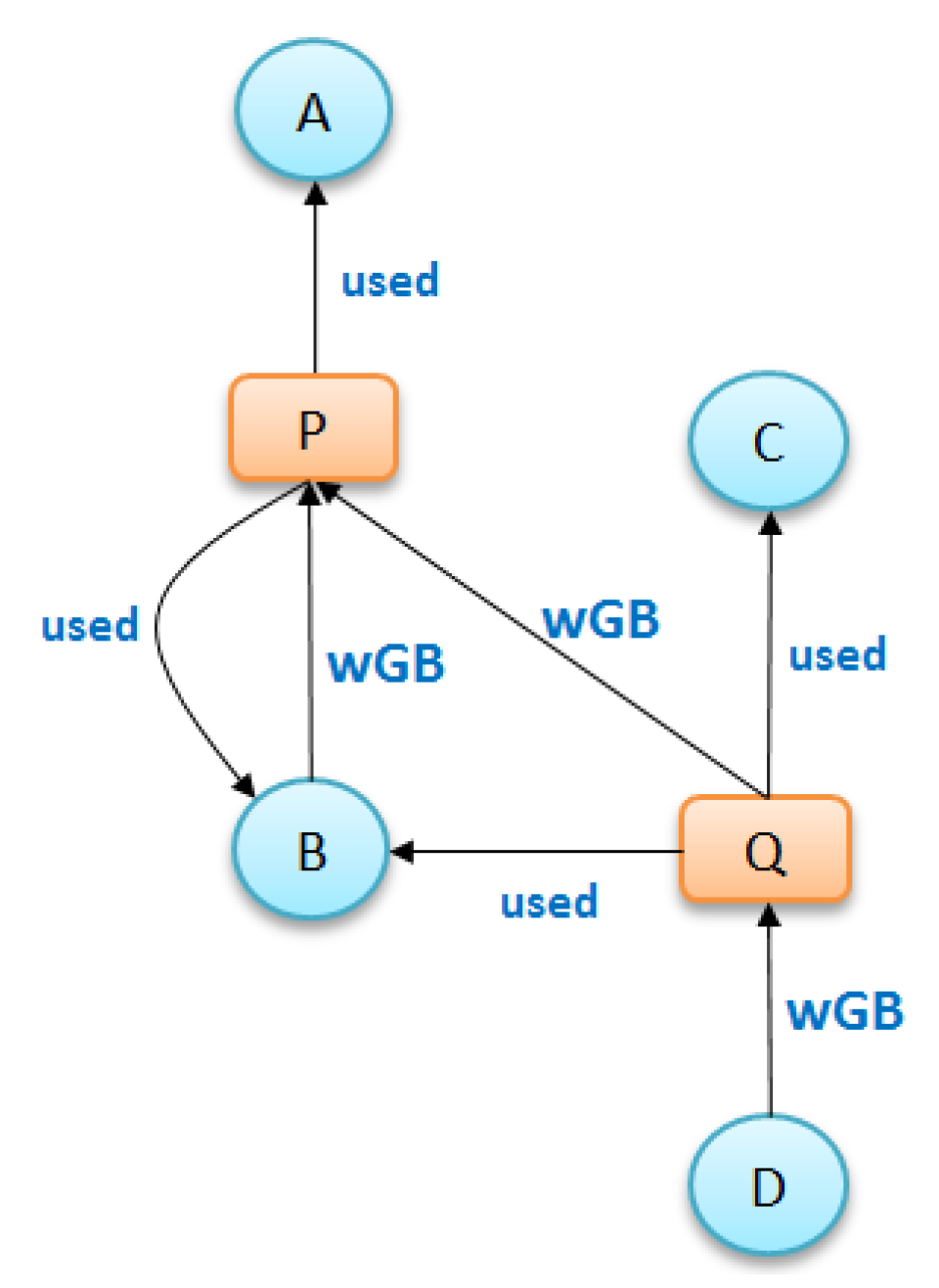
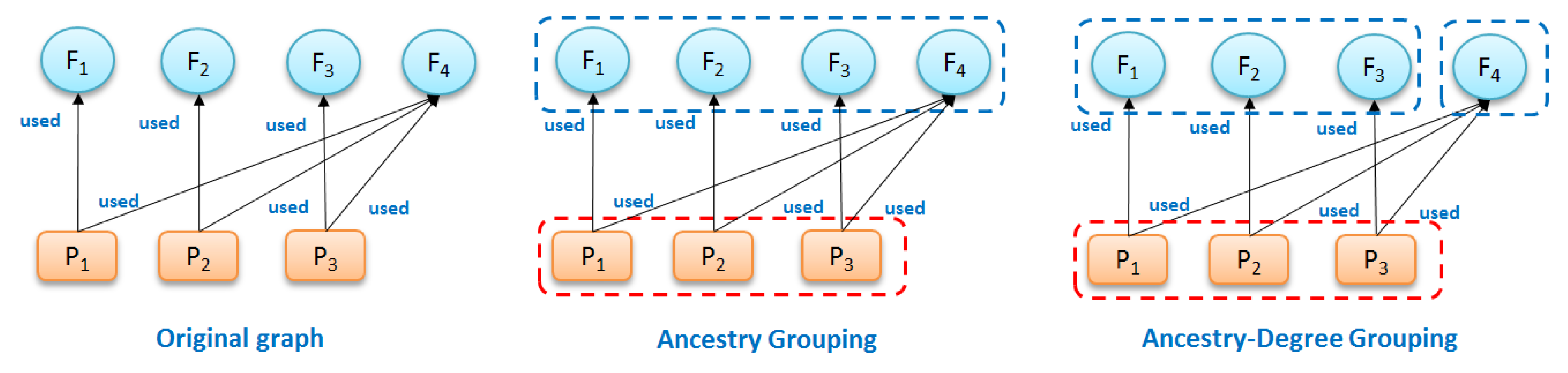
Ancestry grouping, however, may still group system files and dependency information together with application-specific nodes. Consider the example in
Figure 8. Suppose that we have a provenance graph on the left of
Figure 8 that has three nodes of process (
and
), four nodes of file (
and
) and six edges (relationship
). The summary ancestry graph shown in center of
Figure 8 dividing the original graph into two groups
and
, satisfies 3. (All nodes in every group have similar node types and associate with the same ancestry groups). However, from the conceptual point of view, file
, a dependency used by other processes is different from other file nodes in
, since this node is the only node that associates with all processes in group
. In other words,
has
while other nodes in
have
.
To uniquely differentiate , we define ancestry-degree compatible grouping to summarize provenance graphs.
Definition 4 (Ancestry-degree grouping). A grouping has the same ancestry-degree if it satisfies the following:
- (i)
Ancestry grouping Definition 3,
- (ii)
, if , then , and , where and .
Based on this definition, the summary graph shown in
Figure 8 (right) composed of three groups—
,
and
—is ancestry-degree compatible because it is Ancestry-Grouping compatible and all the nodes in every group have the same number of output edges from/to other groups (all nodes in
have one input edge from group
, all nodes in group
have two output edges: one to
and one to
).
We now describe the summary Algorithm 3, which, given a retrospective provenance graph, produces an ancestry-degree grouping. In this algorithm, we first divide the nodes into group with same node type and store them in a stack (Line 2). Next, for each group g in the stack, we re-partition all groups in in stack by calling function divideGroup() with a list of vertices in g (i.e., Lines 5–7). Function divideGroup() is called twice with different direction parameters, since it is applied on two different directions of edges (i.e., input or output). The main purpose of this function is to re-organize all the groups (i.e., ) that are relevant to g by checking all the vertices and edges of vertices in group g of stack (Lines 12 and 26). First, for each types of edges or relationships (in our context, there are three types of edge: ), it calculates the number of edges (or “degree”) from/to vertices in each group of to/from a vertex v in (Lines 16–22). Second, it further divides these vertices in each group of by considering the degree of these vertices (i.e., the number of edges from/to vertices in group g). This means vertices that belong to the same group must have the same degree (Line 22). Third, we add all the new generated groups to if they have never been in (Lines 24–26), before the next consideration of other groups in . Finally, once all the groups g in are considered, we obtain the summary graph by joining all the groups together (Lines 8–10).
| Algorithm 3: Ancestry-degree grouping |
![Informatics 05 00014 i003]() |
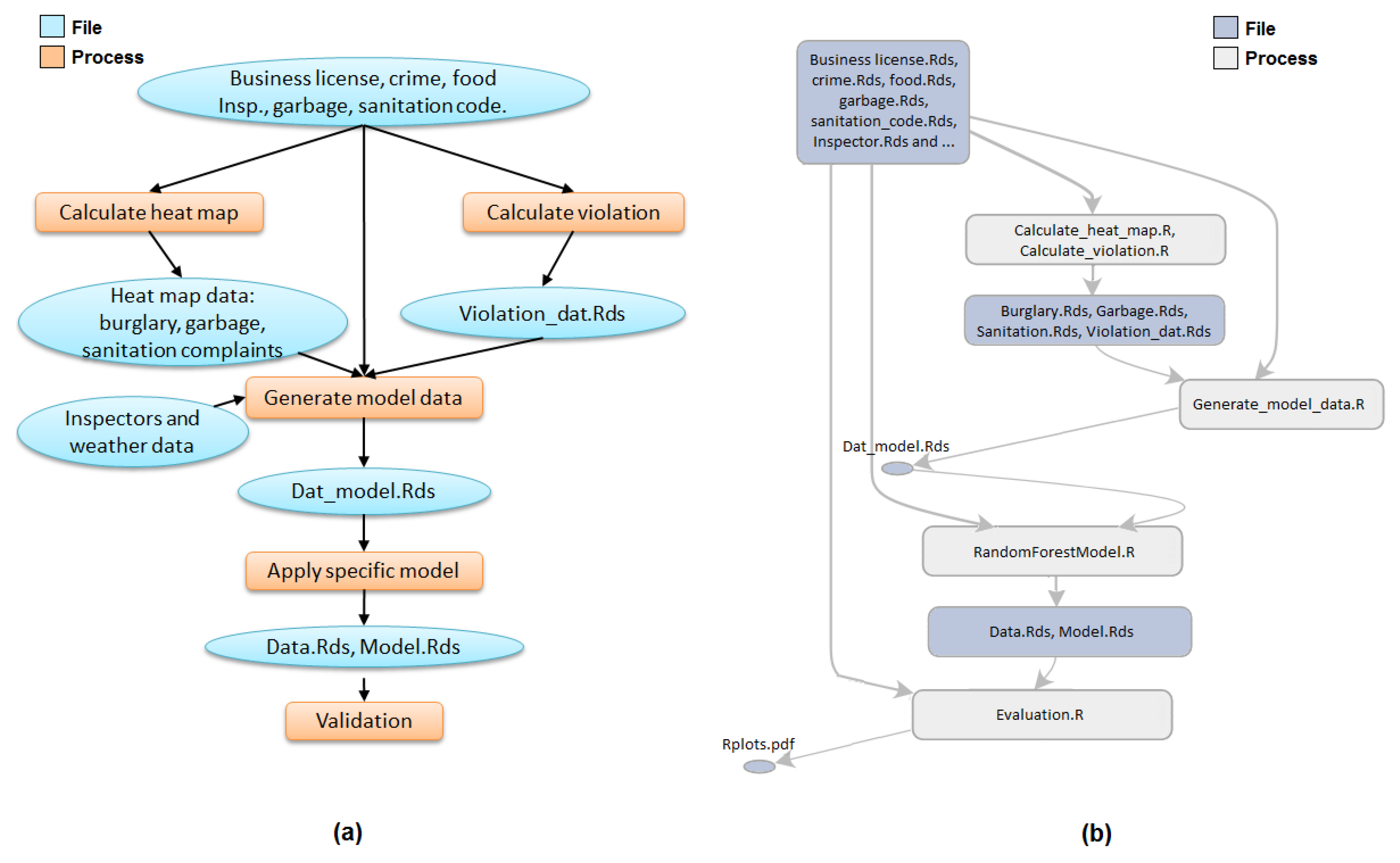
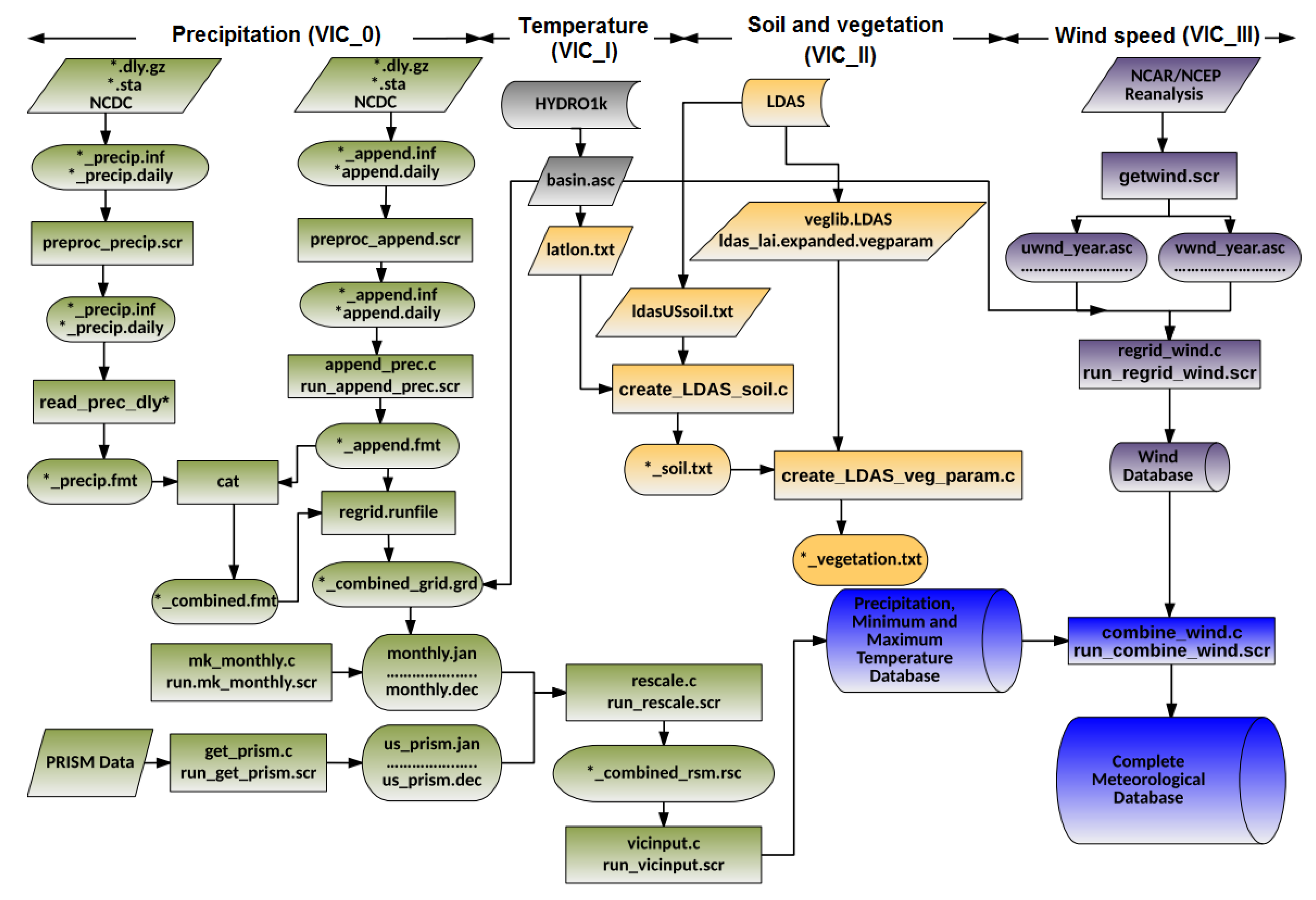
Figure 9 presents an example of graph summarization. The left figure (i.e.,
Figure 9a) shows the work-flow of FIE [
40] application drawn by users that describes the conceptual view of FIE application, while the right figure (i.e.,
Figure 9b) shows the provenance summary graph of FIE application after applying ancestry-degree grouping. As a general observation, these two graphs are fairly close to each other. The summary graph almost captures all the information about the application that users might need at the general view. There are some minor differences between them. For example, the two processes “Calculate heat map” and “Calculate violation” are clearly separate in the left figure while in the right figure they are grouped together. Similarly, the two groups of files “heat map data” and “Violation_dat.Rds” are separate in the left figure and grouped in the right. These groups of files and groups of processes are ancestry-degree compatible, and thus they are grouped in the right figure. While they are separate in the application workflow, ancestry-grouping is helpful in general. For instance, the ancestry-grouping grouped all data files into a single group at the top.