Alzheimer’s Disease: A Molecular View of β-Amyloid Induced Morbific Events
Abstract
1. Introduction
2. APP Processing
2.1. Post-Translational Modification of APP Alters Aβ Production
2.2. APP-Secretases Processing and Aβ Generation
2.3. Intra-Cellular Aβ
3. Tau Pathology
3.1. Aβ Peptide Promotes Hyperphosphorylation of Tau Protein
3.2. Molecular Mechanism of p-Tau Mediated Neurodegeneration
4. Aβ in Mitochondria Dysfunction
Mitochondrial Dysfunction by Aβ
5. Aβ in ER Stress
ER Dysfunction by Aβ
6. Autophagy/Lysosomal Dysfunction
6.1. Autophagy Physiology
6.2. Autophagy Impairment by Aβ
7. Aβ-Accelerated Golgi Fragmentation
7.1. APP Processing in GC
7.2. Golgi Fragmentation by Aβ
8. Aβ in Gene Regulation
8.1. Aβ as Transcription Factor
8.2. Telomerase, Spliceosome Inhibition and DNA Methylation by Aβ
9. Signalling Mechanism of Aβ Leading to Memory Impairment and Cell Death
9.1. Receptor Mediated Long Term Potentiation Inhibition
9.2. Receptor Mediated Cells Death Induced by Aβ
10. Inflammation a Central Mechanism in AD
10.1. Cellular Mediators Involved in Neuroinflammation
10.1.1. Microglial Cells
10.1.2. Astrocytes
10.1.3. Oligodendrocytes
10.1.4. Neurons
10.2. Inflammatory Mediators in AD
10.2.1. Complement System
10.2.2. Chemokines
10.2.3. Cytokines
10.3. Proinflammatory Mediators
10.3.1. Interleukin 1 (IL-1)
10.3.2. IL-6
10.3.3. Tumor Necrotic Factor Alpha (TNF-α)
10.3.4. NF-κB
10.4. Anti-Inflammatory Mediators
10.4.1. TGF-β (Tumour Growth Factor- β)
10.4.2. IL-10
10.5. Inflammatory Mediators
10.5.1. Cyclooxygenases (COX)
10.5.2. Nitric Oxide (NO)
| Model System | CNS Cells Expressing Receptor | Ligands | Role in AD Pathogenesis | References |
|---|---|---|---|---|
| Complement receptors (CR1, CR3) CD88 | Neurons, microglial cells, astrocytes, and oligodendrocyte | C3b, C4b C3 C5a | Neuroinflammation, uptake and clearance of Aβ | [342,343,344] |
| FPRL1 and FPRL2 | Macrophages, glial cells, astrocytes | Aβ | Proinflammation, Aβ42 internalization, formation of fibrillar aggregates, phagocyte chemotaxis and oxidant stress | [251,298,345,346] |
| Scavenger receptor (SR-A) CD36 (SR-B) LDLR, RAGE, LRP1 | Microglia, human monocytes astrocytes Macrophages neurons | Aβ, β-sheet fibrils, HMGB1 | Aβ clearance, synaptic dysfunction, neuroinflammation, production of chemokines, and neurotoxic ROS, NLRP3 activation | [248,249,250,287,288,302,347,348,349,350,351] |
| Toll-like receptors (TLR2 TLR4, TLR2, TLR9) | Microglia, astrocytes | LPS, Aβ | LTP deficit and neuronal death, neuroinflammation, Aβ uptake and clearance. | [288,289,352,353,354] |
| CX3CR1 | Microglia, neurons, astrocytes | CX3CL | Neuroprotection against AD. AD patients showed reduced level of CX3CR1 which led to enhanced activation of microglial cells with enhanced tau phosphorylation | [355] |
| TREM2 | Microglia and neurons | Aβ | Microglial depolarization, apoptosis activation of Wnt/β-catenin leads to inflammation. | [356,357] |
| CD33 | Microglia | Aβ | Increased expression of CD33 attenuates Aβ uptake leading facilitating plaque formation | [358] |
| NALP3/NLRP3 | Microglia, macrophage cells | Aβ and other mediators | Enhanced caspase-1 activity leading to IL-1β and IL-18 mediated neuroinflammation | [300,301,302] |
11. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tiwari, S.; Atluri, V.; Kaushik, A.; Yndart, A.; Nair, M. Alzheimer’s disease: Pathogenesis, diagnostics, and therapeutics. Int. J. Nanomed. 2019, 14, 5541. [Google Scholar] [CrossRef]
- Nichols, E.; Szoeke, C.E.; Vollset, S.E.; Abbasi, N.; Abd-Allah, F.; Abdela, J.; Aichour, M.T.E.; Akinyemi, R.O.; Alahdab, F.; Asgedom, S.W. Global, regional, and national burden of Alzheimer’s disease and other dementias, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 2019, 18, 88–106. [Google Scholar] [CrossRef]
- Alzheimer’s Association report. Alzheimer’s Disease Facts and Figures. Alzheimers Dement. 2020, 16, 391–460. [Google Scholar]
- Albert, M.S.; DeKosky, S.T.; Dickson, D.; Dubois, B.; Feldman, H.H.; Fox, N.C.; Gamst, A.; Holtzman, D.M.; Jagust, W.J.; Petersen, R.C. The diagnosis of mild cognitive impairment due to Alzheimer’s disease: Recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011, 7, 270–279. [Google Scholar] [CrossRef]
- McKhann, G.M.; Knopman, D.S.; Chertkow, H.; Hyman, B.T.; Jack, C.R., Jr.; Kawas, C.H.; Klunk, W.E.; Koroshetz, W.J.; Manly, J.J.; Mayeux, R. The diagnosis of dementia due to Alzheimer’s disease: Recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011, 7, 263–269. [Google Scholar] [CrossRef] [PubMed]
- Jack, C.R., Jr.; Albert, M.S.; Knopman, D.S.; McKhann, G.M.; Sperling, R.A.; Carrillo, M.C.; Thies, B.; Phelps, C.H. Introduction to the recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011, 7, 257–262. [Google Scholar] [CrossRef] [PubMed]
- Ballard, C.; Khan, Z.; Clack, H.; Corbett, A. Nonpharmacological treatment of Alzheimer disease. Can. J.Psychiatry 2011, 56, 589–595. [Google Scholar] [CrossRef]
- Giri, M.; Zhang, M.; Lü, Y. Genes associated with Alzheimer’s disease: An overview and current status. Clin. Interv. Aging 2016, 11, 665. [Google Scholar] [CrossRef]
- Mahley, R.W. Apolipoprotein E: From cardiovascular disease to neurodegenerative disorders. J. Mol. Med. 2016, 94, 739–746. [Google Scholar] [CrossRef] [PubMed]
- Mayeux, R.; Stern, Y. Epidemiology of Alzheimer disease. Cold Spring Harb. Perspect. Med. 2012, 2, a006239. [Google Scholar] [CrossRef] [PubMed]
- Crews, L.; Masliah, E. Molecular mechanisms of neurodegeneration in Alzheimer’s disease. Hum. Mol. Genet. 2010, 19, R12–R20. [Google Scholar] [CrossRef]
- Thakur, A.; Kamboj, P.; Goswami, K. Pathophysiology and management of Alzheimer’s disease: An overview. J. Anal. Pharm. Res. 2018, 9, 226–235. [Google Scholar] [CrossRef]
- DeTure, M.A.; Dickson, D.W. The neuropathological diagnosis of Alzheimer’s disease. Mol. Neurodegener. 2019, 14, 32. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Singh, A. A review on Alzheimer’s disease pathophysiology and its management: An update. Pharmacol. Rep. 2015, 67, 195–203. [Google Scholar] [CrossRef]
- Folch, J.; Petrov, D.; Ettcheto, M.; Abad, S.; Sánchez-López, E.; García, M.L.; Olloquequi, J.; Beas-Zarate, C.; Auladell, C.; Camins, A. Current research therapeutic strategies for Alzheimer’s disease treatment. Neural Plast. 2016, 2016, 8501693. [Google Scholar] [CrossRef]
- Cao, J.; Hou, J.; Ping, J.; Cai, D. Advances in developing novel therapeutic strategies for Alzheimer’s disease. Mol. Neurodegener. 2018, 13, 64. [Google Scholar] [CrossRef]
- Kurz, A.; Perneczky, R. Novel insights for the treatment of Alzheimer’s disease. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2011, 35, 373–379. [Google Scholar] [CrossRef]
- Wasco, W.; Gurubhagavatula, S.; Paradis, M.D.; Romano, D.M.; Sisodia, S.S.; Hyman, B.T.; Neve, R.L.; Tanzi, R.E. Isolation and characterization of APLP2 encoding a homologue of the Alzheimer’s associated amyloid β protein precursor. Nat. Genet. 1993, 5, 95–100. [Google Scholar] [CrossRef]
- Pearson, P.; Price, D.; Gearhart, J. Introduction and expression of the 400 kilobase amyloid precursor protein gene in transgenic mice (corrected). Nat. Genet. 1993, 5, 22–30. [Google Scholar]
- Izumi, R.; Yamada, T.; Yoshikai, S.-I.; Sasaki, H.; Hattori, M.; Sakaki, Y. Positive and negative regulatory elements for the expression of the Alzheimer’s disease amyloid precursor-encoding gene in mouse. Gene 1992, 112, 189–195. [Google Scholar] [CrossRef]
- Gao, Q.; Dai, Z.; Zhang, S.; Fang, Y.; Yung, K.K.L.; Lo, P.K.; Lai, K.W.C. Interaction of Sp1 and APP promoter elucidates a mechanism for Pb2+ caused neurodegeneration. Arch. Biochem. Biophys. 2020, 681, 108265. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yu, X.; Zack, D.J.; Zhu, H.; Qian, J. TiGER: A database for tissue-specific gene expression and regulation. BMC Bioinform. 2008, 9, 271. [Google Scholar] [CrossRef]
- Sisodia, S.S.; Koo, E.H.; Hoffman, P.N.; Perry, G.; Price, D.L. Identification and transport of full-length amyloid precursor proteins in rat peripheral nervous system. J. Neurosci. 1993, 13, 3136–3142. [Google Scholar] [CrossRef] [PubMed]
- Halim, A.; Brinkmalm, G.; Rüetschi, U.; Westman-Brinkmalm, A.; Portelius, E.; Zetterberg, H.; Blennow, K.; Larson, G.; Nilsson, J. Site-specific characterization of threonine, serine, and tyrosine glycosylations of amyloid precursor protein/amyloid β-peptides in human cerebrospinal fluid. Proc. Natl. Acad. Sci. USA 2011, 108, 11848–11853. [Google Scholar] [CrossRef]
- Lee, M.-S.; Kao, S.-C.; Lemere, C.A.; Xia, W.; Tseng, H.-C.; Zhou, Y.; Neve, R.; Ahlijanian, M.K.; Tsai, L.-H. APP processing is regulated by cytoplasmic phosphorylation. J. Cell Biol. 2003, 163, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyya, R.; Barren, C.; Kovacs, D.M. Palmitoylation of amyloid precursor protein regulates amyloidogenic processing in lipid rafts. J. Neurosci. 2013, 33, 11169–11183. [Google Scholar] [CrossRef] [PubMed]
- Yazaki, M.; Tagawa, K.; Maruyama, K.; Sorimachi, H.; Tsuchiya, T.; Ishiura, S.; Suzuki, K. Mutation of potential N-linked glycosylation sites in the Alzheimer’s disease amyloid precursor protein (APP). Neurosci. Lett. 1996, 221, 57–60. [Google Scholar] [CrossRef]
- Perdivara, I.; Petrovich, R.; Allinquant, B.; Deterding, L.J.; Tomer, K.B.; Przybylski, M. Elucidation of O-glycosylation structures of the β-amyloid precursor protein by liquid chromatography-mass spectrometry using electron transfer dissociation and collision induced dissociation. J. Proteome Res. 2009, 8, 631–642. [Google Scholar] [CrossRef]
- Jacobsen, K.T.; Iverfeldt, K. O-GlcNAcylation increases non-amyloidogenic processing of the amyloid-β precursor protein (APP). Biochem. Biophys. Res. Commun. 2011, 404, 882–886. [Google Scholar] [CrossRef]
- Walter, J.; Capell, A.; Hung, A.Y.; Langen, H.; Schnölzer, M.; Thinakaran, G.; Sisodia, S.S.; Selkoe, D.J.; Haass, C. Ectodomain phosphorylation of β-amyloid precursor protein at two distinct cellular locations. J. Biol. Chem. 1997, 272, 1896–1903. [Google Scholar] [CrossRef]
- Gandy, S.; Czernik, A.J.; Greengard, P. Phosphorylation of Alzheimer disease amyloid precursor peptide by protein kinase C and Ca2+/calmodulin-dependent protein kinase II. Proc. Natl. Acad. Sci. USA 1988, 85, 6218–6221. [Google Scholar] [CrossRef]
- Buxbaum, J.D.; Gandy, S.E.; Cicchetti, P.; Ehrlich, M.E.; Czernik, A.J.; Fracasso, R.P.; Ramabhadran, T.V.; Unterbeck, A.J.; Greengard, P. Processing of Alzheimer beta/A4 amyloid precursor protein: Modulation by agents that regulate protein phosphorylation. Proc. Natl. Acad. Sci. USA 1990, 87, 6003–6006. [Google Scholar] [CrossRef]
- Rebelo, S.; Vieira, S.I.; Esselmann, H.; Wiltfang, J.; da Cruz e Silva, E.F.; da Cruz e Silva, O.A.B. Tyrosine 687 phosphorylated Alzheimer’s amyloid precursor protein is retained intracellularly and exhibits a decreased turnover rate. Neurodegener. Dis. 2007, 4, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-Q.; Sarge, K.D. Sumoylation of amyloid precursor protein negatively regulates Aβ aggregate levels. Biochem. Biophys. Res. Commun. 2008, 374, 673–678. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, M.; Koike, H.; Saito, R.; Kitamura, Y.; Okuma, Y.; Nomura, Y. Loss of HRD1-mediated protein degradation causes amyloid precursor protein accumulation and amyloid-β generation. J. Neurosci. 2010, 30, 3924–3932. [Google Scholar] [CrossRef] [PubMed]
- Morel, E.; Chamoun, Z.; Lasiecka, Z.M.; Chan, R.B.; Williamson, R.L.; Vetanovetz, C.; Dall’Armi, C.; Simoes, S.; Du Jour, K.S.P.; McCabe, B.D. Phosphatidylinositol-3-phosphate regulates sorting and processing of amyloid precursor protein through the endosomal system. Nat. Commun. 2013, 4, 2250. [Google Scholar] [CrossRef] [PubMed]
- Weidemann, A.; König, G.; Bunke, D.; Fischer, P.; Salbaum, J.M.; Masters, C.L.; Beyreuther, K. Identification, biogenesis, and localization of precursors of Alzheimer’s disease A4 amyloid protein. Cell 1989, 57, 115–126. [Google Scholar] [CrossRef]
- Deolankar, S.C.; Patil, A.H.; Koyangana, S.G.; Subbannayya, Y.; Prasad, T.S.K.; Modi, P.K. Dissecting Alzheimer’s disease molecular substrates by proteomics and discovery of novel post-translational modifications. Omics J. Integr. Biol. 2019, 23, 350–361. [Google Scholar] [CrossRef]
- Osaki, D.; Hiramatsu, H. Itrullination and deamidation affect aggregation properties of amyloid β-proteins. Amyloid 2016, 23, 234–241. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Koo, E.H. Biology and pathophysiology of the amyloid precursor protein. Mol. Neurodegener. 2011, 6, 27. [Google Scholar] [CrossRef]
- Müller, U.C.; Deller, T.; Korte, M. Not just amyloid: Physiological functions of the amyloid precursor protein family. Nat. Rev. Neurosci. 2017, 18, 281–298. [Google Scholar] [CrossRef]
- Yuan, X.-Z.; Sun, S.; Tan, C.-C.; Yu, J.-T.; Tan, L. The role of ADAM10 in Alzheimer’s disease. J. Alzheimers Dis. 2017, 58, 303–322. [Google Scholar] [CrossRef]
- Zhang, X.; Song, W. The role of APP and BACE1 trafficking in APP processing and amyloid-β generation. Alzheimers Res. Ther. 2013, 5, 46. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, Z.; Cai, F.; Zhang, M.; Wu, Y.; Zhang, J.; Song, W. BACE1 cleavage site selection critical for amyloidogenesis and Alzheimer’s pathogenesis. J. Neurosci. 2017, 37, 6915–6925. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; He, G.; Song, W. BACE2, as a novel APP θ-secretase, is not responsible for the pathogenesis of Alzheimer’s disease in Down syndrome. FASEB J. 2006, 20, 1369–1376. [Google Scholar] [CrossRef] [PubMed]
- Azkona, G.; Levannon, D.; Groner, Y.; Dierssen, M. In vivo effects of APP are not exacerbated by BACE2 co-overexpression: Behavioural characterization of a double transgenic mouse model. Amino Acids 2010, 39, 1571–1580. [Google Scholar] [CrossRef]
- Zhang, Z.; Nadeau, P.; Song, W.; Donoviel, D.; Yuan, M.; Bernstein, A.; Yankner, B.A. Presenilins are required for γ-secretase cleavage of β-APP and transmembrane cleavage of Notch-1. Nat. Cell Biol. 2000, 2, 463–465. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Zhang, M.; Cai, F.; Song, W. Biological function of Presenilin and its role in AD pathogenesis. Transl. Neurodegener. 2013, 2, 15. [Google Scholar] [CrossRef]
- He, G.; Luo, W.; Li, P.; Remmers, C.; Netzer, W.J.; Hendrick, J.; Bettayeb, K.; Flajolet, M.; Gorelick, F.; Wennogle, L.P. Gamma-secretase activating protein is a therapeutic target for Alzheimer’s disease. Nature 2010, 467, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.; Liao, G.P.; Chang, J.C.; Xu, P.; Li, Y.-M.; Greengard, P. GSAP modulates γ-secretase specificity by inducing conformational change in PS1. Proc. Natl. Acad. Sci. USA 2019, 116, 6385–6390. [Google Scholar] [CrossRef] [PubMed]
- Angira, D.; Chikhale, R.; Mehta, K.; Bryce, R.A.; Thiruvenkatam, V. Tracing the GSAP–APP C-99 Interaction Site in the β-Amyloid Pathway Leading to Alzheimer’s Disease. ACS Chem. Neurosci. 2019, 10, 3868–3879. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Doms, R.W.; Lee, V.M.-Y. Glu11 site cleavage and N-terminally truncated Aβ production upon BACE overexpression. Biochemistry 2002, 41, 3128–3136. [Google Scholar] [CrossRef] [PubMed]
- Miravalle, L.; Calero, M.; Takao, M.; Roher, A.E.; Ghetti, B.; Vidal, R. Amino-terminally truncated Aβ peptide species are the main component of cotton wool plaques. Biochemistry 2005, 44, 10810–10821. [Google Scholar] [CrossRef]
- Siegel, G.; Gerber, H.; Koch, P.; Bruestle, O.; Fraering, P.C.; Rajendran, L. The Alzheimer’s disease γ-secretase generates higher 42: 40 ratios for β-amyloid than for p3 peptides. Cell Rep. 2017, 19, 1967–1976. [Google Scholar] [CrossRef]
- Willem, M.; Tahirovic, S.; Busche, M.A.; Ovsepian, S.V.; Chafai, M.; Kootar, S.; Hornburg, D.; Evans, L.D.; Moore, S.; Daria, A. η-Secretase processing of APP inhibits neuronal activity in the hippocampus. Nature 2015, 526, 443–447. [Google Scholar] [CrossRef]
- Xu, T.-H.; Yan, Y.; Kang, Y.; Jiang, Y.; Melcher, K.; Xu, H.E. Alzheimer’s disease-associated mutations increase amyloid precursor protein resistance to γ-secretase cleavage and the Aβ42/Aβ40 ratio. Cell Discov. 2016, 2, 16026. [Google Scholar] [CrossRef]
- Sikanyika, N.L.; Parkington, H.C.; Smith, A.I.; Kuruppu, S. Powering amyloid beta degrading enzymes: A possible therapy for Alzheimer’s disease. Neurochem. Res. 2019, 44, 1289–1296. [Google Scholar] [CrossRef]
- Pasternak, S.H.; Bagshaw, R.D.; Guiral, M.; Zhang, S.; Ackerley, C.A.; Pak, B.J.; Callahan, J.W.; Mahuran, D.J. Presenilin-1, nicastrin, amyloid precursor protein, and γ-secretase activity are co-localized in the lysosomal membrane. J. Biol. Chem. 2003, 278, 26687–26694. [Google Scholar] [CrossRef] [PubMed]
- Schreiner, B.; Hedskog, L.; Wiehager, B.; Ankarcrona, M. Amyloid-β peptides are generated in mitochondria-associated endoplasmic reticulum membranes. J. Alzheimers Dis. 2015, 43, 369–374. [Google Scholar] [CrossRef] [PubMed]
- Choy, R.W.Y.; Cheng, Z.; Schekman, R. Amyloid precursor protein (APP) traffics from the cell surface via endosomes for amyloid β (Aβ) production in the trans-Golgi network. Proc. Natl. Acad. Sci. USA 2012, 109, E2077–E2082. [Google Scholar] [CrossRef]
- Fiandaca, M.S.; Kapogiannis, D.; Mapstone, M.; Boxer, A.; Eitan, E.; Schwartz, J.B.; Abner, E.L.; Petersen, R.C.; Federoff, H.J.; Miller, B.L.; et al. Identification of preclinical Alzheimer’s disease by a profile of pathogenic proteins in neurally derived blood exosomes: A case-control study. Alzheimers Dement. 2015, 11, 600–607. [Google Scholar] [CrossRef] [PubMed]
- Perez-Gonzalez, R.; Gauthier, S.A.; Kumar, A.; Levy, E. The exosome secretory pathway transports amyloid precursor protein carboxyl-terminal fragments from the cell into the brain extracellular space. J. Biol. Chem. 2012, 287, 43108–43115. [Google Scholar] [CrossRef] [PubMed]
- Sinha, M.S.; Ansell-Schultz, A.; Civitelli, L.; Hildesjö, C.; Larsson, M.; Lannfelt, L.; Ingelsson, M.; Hallbeck, M. Alzheimer’s disease pathology propagation by exosomes containing toxic amyloid-beta oligomers. Acta Neuropathol. 2018, 136, 41–56. [Google Scholar] [CrossRef]
- Martin, B.L.; Schrader-Fischer, G.; Busciglio, J.; Duke, M.; Paganetti, P.; Yankner, B.A. Intracellular accumulation of β-amyloid in cells expressing the Swedish mutant amyloid precursor protein. J. Biol. Chem. 1995, 270, 26727–26730. [Google Scholar] [CrossRef]
- Rogaeva, E.; Meng, Y.; Lee, J.H.; Gu, Y.; Kawarai, T.; Zou, F.; Katayama, T.; Baldwin, C.T.; Cheng, R.; Hasegawa, H. The neuronal sortilin-related receptor SORL1 is genetically associated with Alzheimer disease. Nat. Genet. 2007, 39, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, A.; Whelan, C.; Smith, C.; Berezovska, O.; Hyman, B. Direct visualization of the gamma secretase-generated carboxyl-terminal domain of the amyloid precursor protein: Association with Fe65 and translocation to the nucleus. J. Neurochem. 2002, 82, 839–847. [Google Scholar] [CrossRef]
- Gyure, K.A.; Durham, R.; Stewart, W.F.; Smialek, J.E.; Troncoso, J.C. Intraneuronal Aβ-amyloid precedes development of amyloid plaques in Down syndrome. Arch. Pathol. Lab. Med. 2001, 125, 489–492. [Google Scholar] [CrossRef]
- Walsh, D.M.; Klyubin, I.; Fadeeva, J.V.; Cullen, W.K.; Anwyl, R.; Wolfe, M.S.; Rowan, M.J.; Selkoe, D.J. Naturally secreted oligomers of amyloid β protein potently inhibit hippocampal long-term potentiation in vivo. Nature 2002, 416, 535–539. [Google Scholar] [CrossRef]
- Cleary, J.P.; Walsh, D.M.; Hofmeister, J.J.; Shankar, G.M.; Kuskowski, M.A.; Selkoe, D.J.; Ashe, K.H. Natural oligomers of the amyloid-β protein specifically disrupt cognitive function. Nat. Neurosci. 2005, 8, 79–84. [Google Scholar] [CrossRef]
- Wilcox, K.C.; Marunde, M.R.; Das, A.; Velasco, P.T.; Kuhns, B.D.; Marty, M.T.; Jiang, H.; Luan, C.-H.; Sligar, S.G.; Klein, W.L. Nanoscale synaptic membrane mimetic allows unbiased high throughput screen that targets binding sites for Alzheimer’s-associated Aβ oligomers. PLoS ONE 2015, 10, e0125263. [Google Scholar] [CrossRef]
- Yang, W.; Ma, K.; Chen, X.; Shi, L.; Bu, G.; Hu, X.; Han, H.; Liu, Y.; Qian, Y. Mitogen-activated protein kinase signaling pathways are involved in regulating α7 nicotinic acetylcholine receptor-mediated amyloid-β uptake in SH-SY5Y cells. Neuroscience 2014, 278, 276–290. [Google Scholar] [CrossRef]
- Bu, G.; Cam, J.; Zerbinatti, C. LRP in Amyloid-β Production and Metabolism. Ann. N. Y. Acad. Sci. 2006, 1086, 35–53. [Google Scholar] [CrossRef]
- Iribarren, P.; Zhou, Y.; Hu, J.; Le, Y.; Wang, J.M. Role of formyl peptide receptor-like 1 (FPRL1/FPR2) in mononuclear phagocyte responses in Alzheimer disease. Immunol. Res. 2005, 31, 165–176. [Google Scholar] [CrossRef]
- Snyder, E.M.; Nong, Y.; Almeida, C.G.; Paul, S.; Moran, T.; Choi, E.Y.; Nairn, A.C.; Salter, M.W.; Lombroso, P.J.; Gouras, G.K. Regulation of NMDA receptor trafficking by amyloid-β. Nat. Neurosci. 2005, 8, 1051–1058. [Google Scholar] [CrossRef]
- Takuma, K.; Fang, F.; Zhang, W.; Yan, S.; Fukuzaki, E.; Du, H.; Sosunov, A.; McKhann, G.; Funatsu, Y.; Nakamichi, N. RAGE-mediated signaling contributes to intraneuronal transport of amyloid-β and neuronal dysfunction. Proc. Natl. Acad. Sci. USA 2009, 106, 20021–20026. [Google Scholar] [CrossRef] [PubMed]
- Waschuk, S.A.; Elton, E.A.; Darabie, A.A.; Fraser, P.E.; McLaurin, J. Cellular membrane composition defines Aβ-lipid interactions. J. Biol. Chem. 2001, 276, 33561–33568. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.I.; Yi, J.S.; Ko, Y.G. Amyloid β oligomerization is induced by brain lipid rafts. J. Cell. Biochem. 2006, 99, 878–889. [Google Scholar] [CrossRef] [PubMed]
- Kawarabayashi, T.; Shoji, M.; Younkin, L.H.; Wen-Lang, L.; Dickson, D.W.; Murakami, T.; Matsubara, E.; Abe, K.; Ashe, K.H.; Younkin, S.G. Dimeric amyloid β protein rapidly accumulates in lipid rafts followed by apolipoprotein E and phosphorylated tau accumulation in the Tg2576 mouse model of Alzheimer’s disease. J. Neurosci. 2004, 24, 3801–3809. [Google Scholar] [CrossRef] [PubMed]
- LaFerla, F.M.; Green, K.N.; Oddo, S. Intracellular amyloid-β in Alzheimer’s disease. Nat. Rev. Neurosci. 2007, 8, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Tackenberg, C.; Nitsch, R.M. The secreted APP ectodomain sAPPα, but not sAPPβ, protects neurons against Aβ oligomer-induced dendritic spine loss and increased tau phosphorylation. Mol. Brain 2019, 12, 27. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Xiong, Z.; Chen, P.; Wei, J.; Chen, S.; Yan, Z. β-Amyloid impairs the regulation of N-methyl-D-aspartate receptors by glycogen synthase kinase 3. Neurobiol. Aging 2014, 35, 449–459. [Google Scholar] [CrossRef]
- Gakhar-Koppole, N.; Hundeshagen, P.; Mandl, C.; Weyer, S.W.; Allinquant, B.; Müller, U.; Ciccolini, F. Activity requires soluble amyloid precursor protein α to promote neurite outgrowth in neural stem cell-derived neurons via activation of the MAPK pathway. Eur. J. Neurosci. 2008, 28, 871–882. [Google Scholar] [CrossRef]
- Fol, R.; Braudeau, J.; Ludewig, S.; Abel, T.; Weyer, S.W.; Roederer, J.-P.; Brod, F.; Audrain, M.; Bemelmans, A.-P.; Buchholz, C.J. Viral gene transfer of APPsα rescues synaptic failure in an Alzheimer’s disease mouse model. Acta Neuropathol. 2016, 131, 247–266. [Google Scholar] [CrossRef]
- Mandelkow, E.-M.; Mandelkow, E. Biochemistry and cell biology of tau protein in neurofibrillary degeneration. Cold Spring Harb. Perspect. Med. 2012, 2, a006247. [Google Scholar] [CrossRef]
- Woerman, A.L.; Aoyagi, A.; Patel, S.; Kazmi, S.A.; Lobach, I.; Grinberg, L.T.; McKee, A.C.; Seeley, W.W.; Olson, S.H.; Prusiner, S.B. Tau prions from Alzheimer’s disease and chronic traumatic encephalopathy patients propagate in cultured cells. Proc. Natl. Acad. Sci. USA 2016, 113, E8187–E8196. [Google Scholar] [CrossRef]
- Friedrich, M.G.; Skora, A.; Hancock, S.E.; Mitchell, T.W.; Else, P.L.; Truscott, R.J. Tau Is Truncated in Five Regions of the Normal Adult Human Brain. Int. J. Mol. Sci. 2021, 22, 3521. [Google Scholar] [CrossRef]
- Hagestedt, T.; Lichtenberg, B.; Wille, H.; Mandelkow, E.; Mandelkow, E. Tau protein becomes long and stiff upon phosphorylation: Correlation between paracrystalline structure and degree of phosphorylation. J. Cell Biol. 1989, 109, 1643–1651. [Google Scholar] [CrossRef]
- Brion, J.P.; Smith, C.; Couck, A.M.; Gallo, J.M.; Anderton, B.H. Developmental Changes in τ Phosphorylation: Fetal τ Is Transiently Phosphorylated in a Manner Similar to Paired Helical Filament-τ Characteristic of Alzheimer’s Disease. J. Neurochem. 1993, 61, 2071–2080. [Google Scholar] [CrossRef] [PubMed]
- Ebashi, M.; Toru, S.; Nakamura, A.; Kamei, S.; Yokota, T.; Hirokawa, K.; Uchihara, T. Detection of AD-specific four repeat tau with deamidated asparagine residue 279-specific fraction purified from 4R tau polyclonal antibody. Acta Neuropathol. 2019, 138, 163–166. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, C.H.; Garwood, C.J.; Wray, S.; Price, C.; Kellie, S.; Perera, T.; Zvelebil, M.; Yang, A.; Sheppard, P.W.; Varndell, I.M. Phosphorylation regulates tau interactions with Src homology 3 domains of phosphatidylinositol 3-kinase, phospholipase Cγ1, Grb2, and Src family kinases. J. Biol. Chem. 2008, 283, 18177–18186. [Google Scholar] [CrossRef] [PubMed]
- Rossi, G.; Dalprà, L.; Crosti, F.; Lissoni, S.; Sciacca, F.L.; Catania, M.; Mangieri, M.; Giaccone, G.; Croci, D.; Tagliavini, F. A new function of microtubule-associated protein tau: Involvement in chromosome stability. Cell Cycle 2008, 7, 1788–1794. [Google Scholar] [CrossRef]
- Wang, J.Z.; Grundke-Iqbal, I.; Iqbal, K. Kinases and phosphatases and tau sites involved in Alzheimer neurofibrillary degeneration. Eur. J. Neurosci. 2007, 25, 59–68. [Google Scholar] [CrossRef]
- Burns, L.H.; Wang, H.-Y. Altered filamin A enables amyloid beta-induced tau hyperphosphorylation and neuroinflammation in Alzheimer’s disease. Neuroimmunol. Neuroinflamm. 2017, 4, 263–271. [Google Scholar] [CrossRef]
- Patzke, H.; Tsai, L.-H. Calpain-mediated cleavage of the cyclin-dependent kinase-5 activator p39 to p29. J. Biol. Chem. 2002, 277, 8054–8060. [Google Scholar] [CrossRef]
- Plattner, F.; Angelo, M.; Giese, K.P. The roles of cyclin-dependent kinase 5 and glycogen synthase kinase 3 in tau hyperphosphorylation. J. Biol. Chem. 2006, 281, 25457–25465. [Google Scholar] [CrossRef]
- Hernández, F.; Borrell, J.; Guaza, C.; Avila, J.; Lucas, J.J. Spatial learning deficit in transgenic mice that conditionally over-express GSK-3β in the brain but do not form tau filaments. J. Neurochem. 2002, 83, 1529–1533. [Google Scholar] [CrossRef] [PubMed]
- King, M.E.; Kan, H.-M.; Baas, P.W.; Erisir, A.; Glabe, C.G.; Bloom, G.S. Tau-dependent microtubule disassembly initiated by prefibrillar β-amyloid. J. Cell Biol. 2006, 175, 541–546. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Z.; Wu, Q.; Smith, A.; Grundke-Iqbal, I.; Iqbal, K. τ is phosphorylated by GSK-3 at several sites found in Alzheimer disease and its biological activity markedly inhibited only after it is prephosphorylated by A-kinase. FEBS Lett. 1998, 436, 28–34. [Google Scholar] [CrossRef]
- Gu, G.J.; Lund, H.; Wu, D.; Blokzijl, A.; Classon, C.; von Euler, G.; Landegren, U.; Sunnemark, D.; Kamali-Moghaddam, M. Role of individual MARK isoforms in phosphorylation of tau at Ser 262 in Alzheimer’s disease. Neuromol. Med. 2013, 15, 458–469. [Google Scholar] [CrossRef]
- Guise, S.; Braguer, D.; Carles, G.; Delacourte, A.; Briand, C. Hyperphosphorylation of tau is mediated by ERK activation during anticancer drug-induced apoptosis in neuroblastoma cells. J. Neurosci. Res. 2001, 63, 257–267. [Google Scholar] [CrossRef]
- Li, T.; Paudel, H.K. Glycogen synthase kinase 3β phosphorylates Alzheimer’s disease-specific Ser396 of microtubule-associated protein tau by a sequential mechanism. Biochemistry 2006, 45, 3125–3133. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.H.; Johnson, G.V. Primed phosphorylation of tau at Thr231 by glycogen synthase kinase 3β (GSK3β) plays a critical role in regulating tau’s ability to bind and stabilize microtubules. J. Neurochem. 2004, 88, 349–358. [Google Scholar] [CrossRef]
- Phiel, C.J.; Wilson, C.A.; Lee, V.M.-Y.; Klein, P.S. GSK-3α regulates production of Alzheimer’s disease amyloid-β peptides. Nature 2003, 423, 435–439. [Google Scholar] [CrossRef]
- Sotiropoulos, I.; Catania, C.; Pinto, L.G.; Silva, R.; Pollerberg, G.E.; Takashima, A.; Sousa, N.; Almeida, O.F. Stress acts cumulatively to precipitate Alzheimer’s disease-like tau pathology and cognitive deficits. J. Neurosci. 2011, 31, 7840–7847. [Google Scholar] [CrossRef]
- Chow, H.-M.; Guo, D.; Zhou, J.-C.; Zhang, G.-Y.; Li, H.-F.; Herrup, K.; Zhang, J. CDK5 activator protein p25 preferentially binds and activates GSK3β. Proc. Natl. Acad. Sci. USA 2014, 111, E4887–E4895. [Google Scholar] [CrossRef]
- Shen, L.L.; Li, W.W.; Xu, Y.L.; Gao, S.H.; Xu, M.Y.; Bu, X.L.; Liu, Y.H.; Wang, J.; Zhu, J.; Zeng, F. Neurotrophin receptor p75 mediates amyloid β-induced tau pathology. Neurobiol. Dis. 2019, 132, 104567. [Google Scholar] [CrossRef]
- Wang, J.-Z.; Gong, C.-X.; Zaidi, T.; Grundke-Iqbal, I.; Iqbal, K. Dephosphorylation of Alzheimer paired helical filaments by protein phosphatase-2A and− 2B. J. Biol. Chem. 1995, 270, 4854–4860. [Google Scholar] [CrossRef]
- Tanimukai, H.; Grundke-Iqbal, I.; Iqbal, K. Up-regulation of inhibitors of protein phosphatase-2A in Alzheimer’s disease. Am. J. Pathol. 2005, 166, 1761–1771. [Google Scholar] [CrossRef]
- Yu, Y.; Run, X.; Liang, Z.; Li, Y.; Liu, F.; Liu, Y.; Iqbal, K.; Grundke-Iqbal, I.; Gong, C.X. Developmental regulation of tau phosphorylation, tau kinases, and tau phosphatases. J. Neurochem. 2009, 108, 1480–1494. [Google Scholar] [CrossRef] [PubMed]
- Bolmont, T.; Clavaguera, F.; Meyer-Luehmann, M.; Herzig, M.C.; Radde, R.; Staufenbiel, M.; Lewis, J.; Hutton, M.; Tolnay, M.; Jucker, M. Induction of tau pathology by intracerebral infusion of amyloid-β-containing brain extract and by amyloid-β deposition in APP× Tau transgenic mice. Am. J. Pathol. 2007, 171, 2012–2020. [Google Scholar] [CrossRef] [PubMed]
- Nisbet, R.M.; Polanco, J.-C.; Ittner, L.M.; Götz, J. Tau aggregation and its interplay with amyloid-β. Acta Neuropathol. 2015, 129, 207–220. [Google Scholar] [CrossRef]
- Penke, B.; Szűcs, M.; Bogár, F. Oligomerization and Conformational Change Turn Monomeric β-Amyloid and Tau Proteins Toxic: Their Role in Alzheimer’s Pathogenesis. Molecules 2020, 25, 1659. [Google Scholar] [CrossRef]
- Šimić, G.; Babić Leko, M.; Wray, S.; Harrington, C.; Delalle, I.; Jovanov-Milošević, N.; Bažadona, D.; Buée, L.; De Silva, R.; Di Giovanni, G. Tau protein hyperphosphorylation and aggregation in Alzheimer’s disease and other tauopathies, and possible neuroprotective strategies. Biomolecules 2016, 6, 6. [Google Scholar] [CrossRef]
- Alonso, A.d.C.; Grundke-Iqbal, I.; Iqbal, K. Alzheimer’s disease hyperphosphorylated tau sequesters normal tau into tangles of filaments and disassembles microtubules. Nat. Med. 1996, 2, 783–787. [Google Scholar] [CrossRef]
- Elie, A.; Prezel, E.; Guérin, C.; Denarier, E.; Ramirez-Rios, S.; Serre, L.; Andrieux, A.; Fourest-Lieuvin, A.; Blanchoin, L.; Arnal, I. Tau co-organizes dynamic microtubule and actin networks. Sci. Rep. 2015, 5, 9964. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Ramos, A.; Díaz-Hernández, M.; Cuadros, R.; Hernández, F.; Avila, J. Extracellular tau is toxic to neuronal cells. FEBS Lett. 2006, 580, 4842–4850. [Google Scholar] [CrossRef] [PubMed]
- Morozova, V.; Cohen, L.S.; Makki, A.E.-H.; Shur, A.; Pilar, G.; ElIdrissi, A.; Alonso, A.D. Normal & Pathological Tau Uptake Mediated by M1/M3 Muscarinic Receptors Promotes Opposite Neuronal Changes. Front. Cell. Neurosci. 2019, 13, 403. [Google Scholar] [PubMed]
- Alonso, A.D.; Cohen, L.S.; Morozova, V. The tau misfolding pathway to dementia. In Protein Folding Disorders in the Central Nervous System; Ghiso, J., Rostagno, A., Eds.; World Scientific Publishing: New York, NY, USA, 2017; pp. 83–107. [Google Scholar]
- Cripps, D.; Thomas, S.N.; Jeng, Y.; Yang, F.; Davies, P.; Yang, A.J. Alzheimer disease-specific conformation of hyperphosphorylated paired helical filament-Tau is polyubiquitinated through Lys-48, Lys-11, and Lys-6 ubiquitin conjugation. J. Biol. Chem. 2006, 281, 10825–10838. [Google Scholar] [CrossRef] [PubMed]
- Reddy, P.H. Mitochondrial dysfunction in aging and Alzheimer’s disease: Strategies to protect neurons. Antioxid. Redox Signal. 2007, 9, 1647–1658. [Google Scholar] [CrossRef]
- Massaad, C.A.; Pautler, R.G.; Klann, E. Mitochondrial superoxide: A key player in Alzheimer’s disease. Aging 2009, 1, 758. [Google Scholar] [CrossRef][Green Version]
- Kincaid, B.; Bossy-Wetzel, E. Forever young: SIRT3 a shield against mitochondrial meltdown, aging, and neurodegeneration. Front. Aging Neurosci. 2013, 5, 48. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, S.; Li, Y.; Yu, S.; Zhao, Y. SIRT1/PGC-1α signaling promotes mitochondrial functional recovery and reduces apoptosis after intracerebral hemorrhage in rats. Front. Mol. Neurosci. 2018, 10, 443. [Google Scholar] [CrossRef]
- Cheng, Y.; Ren, X.; Gowda, A.S.; Shan, Y.; Zhang, L.; Yuan, Y.; Patel, R.; Wu, H.; Huber-Keener, K.; Yang, J. Interaction of Sirt3 with OGG1 contributes to repair of mitochondrial DNA and protects from apoptotic cell death under oxidative stress. Cell Death Dis. 2013, 4, e731. [Google Scholar] [CrossRef] [PubMed]
- Jęśko, H.; Wencel, P.; Strosznajder, R.P.; Strosznajder, J.B. Sirtuins and their roles in brain aging and neurodegenerative disorders. Neurochem. Res. 2017, 42, 876–890. [Google Scholar] [CrossRef] [PubMed]
- Youle, R.J.; Van Der Bliek, A.M. Mitochondrial fission, fusion, and stress. Science 2012, 337, 1062–1065. [Google Scholar] [CrossRef] [PubMed]
- Swerdlow, R.H. Mitochondria in cybrids containing mtDNA from persons with mitochondriopathies. J. Neurosci. Res. 2007, 85, 3416–3428. [Google Scholar] [CrossRef]
- Hirai, K.; Aliev, G.; Nunomura, A.; Fujioka, H.; Russell, R.L.; Atwood, C.S.; Johnson, A.B.; Kress, Y.; Vinters, H.V.; Tabaton, M. Mitochondrial abnormalities in Alzheimer’s disease. J. Neurosci. 2001, 21, 3017–3023. [Google Scholar] [CrossRef]
- Szablewski, L. Glucose transporters in brain: In health and in Alzheimer’s disease. J. Alzheimers Dis. 2017, 55, 1307–1320. [Google Scholar] [CrossRef]
- Sonntag, K.-C.; Ryu, W.-I.; Amirault, K.M.; Healy, R.A.; Siegel, A.J.; McPhie, D.L.; Forester, B.; Cohen, B.M. Late-onset Alzheimer’s disease is associated with inherent changes in bioenergetics profiles. Sci. Rep. 2017, 7, 14038. [Google Scholar] [CrossRef]
- Manczak, M.; Park, B.S.; Jung, Y.; Reddy, P.H. Differential expression of oxidative phosphorylation genes in patients with Alzheimer’s disease. Neuromol. Med. 2004, 5, 147–162. [Google Scholar] [CrossRef]
- Caspersen, C.; Wang, N.; Yao, J.; Sosunov, A.; Chen, X.; Lustbader, J.W.; Xu, H.W.; Stern, D.; McKhann, G.; Du Yan, S. Mitochondrial Aβ: A potential focal point for neuronal metabolic dysfunction in Alzheimer’s disease. FASEB J. 2005, 19, 2040–2041. [Google Scholar] [CrossRef]
- Gillardon, F.; Rist, W.; Kussmaul, L.; Vogel, J.; Berg, M.; Danzer, K.; Kraut, N.; Hengerer, B. Proteomic and functional alterations in brain mitochondria from Tg2576 mice occur before amyloid plaque deposition. Proteomics 2007, 7, 605–616. [Google Scholar] [CrossRef] [PubMed]
- Eckert, A.; Hauptmann, S.; Scherping, I.; Rhein, V.; Müller-Spahn, F.; Götz, J.; Müller, W.E. Soluble beta-amyloid leads to mitochondrial defects in amyloid precursor protein and tau transgenic mice. Neurodegener. Dis. 2008, 51, 57–159. [Google Scholar] [CrossRef] [PubMed]
- Eckert, A.; Steiner, B.; Marques, C.; Leutz, S.; Romig, H.; Haass, C.; Müller, W.E. Elevated vulnerability to oxidative stress-induced cell death and activation of caspase-3 by the Swedish amyloid precursor protein mutation. J. Neurosci. Res. 2001, 64, 183–192. [Google Scholar] [CrossRef]
- Marques, C.A.; Keil, U.; Bonert, A.; Steiner, B.; Haass, C.; Müller, W.E.; Eckert, A. Neurotoxic Mechanisms Caused by the Alzheimer’s Disease-linked Swedish Amyloid Precursor Protein Mutation oxidative stress, caspases, and the jnk pathway. J. Biol. Chem. 2003, 278, 28294–28302. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Su, B.; Siedlak, S.L.; Moreira, P.I.; Fujioka, H.; Wang, Y.; Casadesus, G.; Zhu, X. Amyloid-β overproduction causes abnormal mitochondrial dynamics via differential modulation of mitochondrial fission/fusion proteins. Proc. Natl. Acad. Sci. USA 2008, 105, 19318–19323. [Google Scholar] [CrossRef]
- Cieślik, M.; Czapski, G.A.; Wójtowicz, S.; Wieczorek, I.; Wencel, P.L.; Strosznajder, R.P.; Jaber, V.; Lukiw, W.J.; Strosznajder, J.B. Alterations of Transcription of Genes Coding Anti-oxidative and Mitochondria-Related Proteins in Amyloid β Toxicity: Relevance to Alzheimer’s Disease. Mol. Neurobiol. 2020, 57, 1374–1388. [Google Scholar] [CrossRef] [PubMed]
- Singulani, M.P.; Pereira, C.P.M.; Ferreira, A.F.F.; Garcia, P.C.; Ferrari, G.D.; Alberici, L.C.; Britto, L.R. Impairment of PGC-1α-mediated mitochondrial biogenesis precedes mitochondrial dysfunction and Alzheimer’s pathology in the 3xTg mouse model of Alzheimer’s disease. Exp. Gerontol. 2020, 133, 110882. [Google Scholar] [CrossRef]
- Manczak, M.; Reddy, P.H. Abnormal interaction between the mitochondrial fission protein Drp1 and hyperphosphorylated tau in Alzheimer’s disease neurons: Implications for mitochondrial dysfunction and neuronal damage. Hum. Mol. Genet. 2012, 21, 2538–2547. [Google Scholar] [CrossRef]
- Hayashi, T.; Rizzuto, R.; Hajnoczky, G.; Su, T.P. MAM: More than just a housekeeper. Trends Cell Biol. 2009, 19, 81–88. [Google Scholar] [CrossRef]
- Yu, J.; Novgorodov, S.A.; Chudakova, D.; Zhu, H.; Bielawska, A.; Bielawski, J.; Obeid, L.M.; Kindy, M.S.; Gudz, T.I. JNK3 signaling pathway activates ceramide synthase leading to mitochondrial dysfunction. J. Biol. Chem. 2007, 282, 25940–25949. [Google Scholar] [CrossRef] [PubMed]
- Pera, M.; Larrea, D.; Guardia-Laguarta, C.; Montesinos, J.; Velasco, K.R.; Agrawal, R.R.; Xu, Y.; Chan, R.B.; Di Paolo, G.; Mehler, M.F. Increased localization of APP-C99 in mitochondria-associated ER membranes causes mitochondrial dysfunction in Alzheimer disease. EMBO J. 2017, 36, 3356–3371. [Google Scholar] [CrossRef] [PubMed]
- Tillement, L.; Lecanu, L.; Yao, W.; Greeson, J.; Papadopoulos, V. The spirostenol (22R, 25R)-20α-spirost-5-en-3β-yl hexanoate blocks mitochondrial uptake of Aβ in neuronal cells and prevents Aβ-induced impairment of mitochondrial function. Steroids 2006, 71, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Petersen, C.A.H.; Alikhani, N.; Behbahani, H.; Wiehager, B.; Pavlov, P.F.; Alafuzoff, I.; Leinonen, V.; Ito, A.; Winblad, B.; Glaser, E. The amyloid β-peptide is imported into mitochondria via the TOM import machinery and localized to mitochondrial cristae. Proc. Natl. Acad. Sci. USA 2008, 105, 13145–13150. [Google Scholar] [CrossRef] [PubMed]
- Lustbader, J.W.; Cirilli, M.; Lin, C.; Xu, H.W.; Takuma, K.; Wang, N.; Caspersen, C.; Chen, X.; Pollak, S.; Chaney, M. ABAD directly links Aß to mitochondrial toxicity in Alzheimer’s Disease. Science 2004, 304, 448–452. [Google Scholar] [CrossRef]
- Cha, M.-Y.; Cho, H.J.; Kim, C.; Jung, Y.O.; Kang, M.J.; Murray, M.E.; Hong, H.S.; Choi, Y.-J.; Choi, H.; Kim, D.K. Mitochondrial ATP synthase activity is impaired by suppressed O-GlcNAcylation in Alzheimer’s disease. Hum. Mol. Genet. 2015, 24, 6492–6504. [Google Scholar] [CrossRef] [PubMed]
- Halestrap, A. A pore way to die. Nature 2005, 434, 578–579. [Google Scholar] [CrossRef]
- Jacobson, J.; Duchen, M.R. Mitochondrial oxidative stress and cell death in astrocytes requirement for stored Ca2+ and sustained opening of the permeability transition pore. J. Cell Sci. 2002, 115, 1175–1188. [Google Scholar] [CrossRef] [PubMed]
- Du, H.; Guo, L.; Fang, F.; Chen, D.; Sosunov, A.A.; McKhann, G.M.; Yan, Y.; Wang, C.; Zhang, H.; Molkentin, J.D. Cyclophilin D deficiency attenuates mitochondrial and neuronal perturbation and ameliorates learning and memory in Alzheimer’s disease. Nat. Med. 2008, 14, 1097–1105. [Google Scholar] [CrossRef] [PubMed]
- Beck, S.J.; Guo, L.; Phensy, A.; Tian, J.; Wang, L.; Tandon, N.; Gauba, E.; Lu, L.; Pascual, J.M.; Kroener, S. Deregulation of mitochondrial F1FO-ATP synthase via OSCP in Alzheimer’s disease. Nat. Commun. 2016, 7, 11483. [Google Scholar] [CrossRef]
- Sanz-Blasco, S.; Valero, R.A.; Rodríguez-Crespo, I.; Villalobos, C.; Núñez, L. Mitochondrial Ca2+ overload underlies Aβ oligomers neurotoxicity providing an unexpected mechanism of neuroprotection by NSAIDs. PLoS ONE 2008, 3, e2718. [Google Scholar] [CrossRef]
- Alikhani, N.; Ankarcrona, M.; Glaser, E. Mitochondria and Alzheimer’s disease: Amyloid-β peptide uptake and degradation by the presequence protease, hPreP. J.Bioenerg. Biomembr. 2009, 41, 447–451. [Google Scholar] [CrossRef]
- Mossmann, D.; Vögtle, F.-N.; Taskin, A.A.; Teixeira, P.F.; Ring, J.; Burkhart, J.M.; Burger, N.; Pinho, C.M.; Tadic, J.; Loreth, D. Amyloid-β peptide induces mitochondrial dysfunction by inhibition of preprotein maturation. Cell Metab. 2014, 20, 662–669. [Google Scholar] [CrossRef]
- Quintana, D.D.; Garcia, J.A.; Anantula, Y.; Rellick, S.L.; Engler-Chiurazzi, E.B.; Sarkar, S.N.; Brown, C.M.; Simpkins, J.W. Amyloid-β Causes Mitochondrial Dysfunction via a Ca2+-Driven Upregulation of Oxidative Phosphorylation and Superoxide Production in Cerebrovascular Endothelial Cells. J. Alzheimers Dis. 2020, 75, 119–138. [Google Scholar] [CrossRef] [PubMed]
- Rao, R.V.; Bredesen, D.E. Misfolded proteins, endoplasmic reticulum stress and neurodegeneration. Curr. Opin. Cell Biol. 2004, 16, 653–662. [Google Scholar] [CrossRef]
- Toyoshima, C. How Ca2+-ATPase pumps ions across the sarcoplasmic reticulum membrane. Biochim. Biophys. ActaMol. Cell Res. 2009, 1793, 941–946. [Google Scholar] [CrossRef]
- Lodish, H.; Kong, N.; Wikström, L. Calcium is required for folding of newly made subunits of the asialoglycoprotein receptor within the endoplasmic reticulum. J. Biol. Chem. 1992, 267, 12753–12760. [Google Scholar] [CrossRef]
- Kaufman, R.J. Stress signaling from the lumen of the endoplasmic reticulum: Coordination of gene transcriptional and translational controls. Genes Dev. 1999, 13, 1211–1233. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Zhang, K.; Kaufman, R.J. The unfolded protein response—A stress signaling pathway of the endoplasmic reticulum. J. Chem. Neuroanat. 2004, 28, 79–92. [Google Scholar] [CrossRef]
- Hamos, J.E.; Oblas, B.; Pulaski-Salo, D.; Welch, W.; Bole, D.; Drachman, D.A. Expression of heat shock proteins in Alzheimer’s disease. Neurology 1991, 41, 345. [Google Scholar] [CrossRef]
- Görlach, A.; Klappa, P.; Kietzmann, D.T. The endoplasmic reticulum: Folding, calcium homeostasis, signaling, and redox control. Antioxid. Redox Signal. 2006, 8, 1391–1418. [Google Scholar] [CrossRef] [PubMed]
- Leissring, M.A.; Akbari, Y.; Fanger, C.M.; Cahalan, M.D.; Mattson, M.P.; LaFerla, F.M. Capacitative calcium entry deficits and elevated luminal calcium content in mutant presenilin-1 knockin mice. J. Cell Biol. 2000, 49, 793–798. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Cheung, K.-H.; Foskett, J.K. Enhanced ROS generation mediated by Alzheimer’s disease presenilin regulation of InsP3R Ca2+ signaling. Antioxid. Redox Signal. 2011, 14, 1225–1235. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, I.; Bajouco, L.; Mota, S.; Auberson, Y.; Oliveira, C.; Rego, A. Amyloid beta peptide 1–42 disturbs intracellular calcium homeostasis through activation of GluN2B-containing N-methyl-d-aspartate receptors in cortical cultures. Cell Calcium 2012, 51, 95–106. [Google Scholar] [CrossRef]
- Ferreiro, E.; Oliveira, C.R.; Pereira, C. Involvement of endoplasmic reticulum Ca2+ release through ryanodine and inositol 1, 4, 5-triphosphate receptors in the neurotoxic effects induced by the amyloid-β peptide. J. Neurosci. Res. 2004, 76, 872–880. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, T.; Su, T.P. Sigma-1 receptor chaperones at the ER-mitochondrion interface regulate Ca2+ signaling and cell survival. Cell 2007, 131, 596–610. [Google Scholar] [CrossRef] [PubMed]
- Bravo, R.; Vicencio, J.M.; Parra, V.; Troncoso, R.; Munoz, J.P.; Bui, M.; Quiroga, C.; Rodriguez, A.E.; Verdejo, H.E.; Ferreira, J. Increased ER–mitochondrial coupling promotes mitochondrial respiration and bioenergetics during early phases of ER stress. J. Cell Biol. 2011, 124, 2143–2152. [Google Scholar]
- Jakaria, M.; Azam, S.; Haque, M.E.; Jo, S.-H.; Uddin, M.S.; Kim, I.-S.; Choi, D.-K. Taurine and its analogs in neurological disorders: Focus on therapeutic potential and molecular mechanisms. Redox Biol. 2019, 24, 101223. [Google Scholar] [CrossRef]
- Chafekar, S.M.; Hoozemans, J.J.; Zwart, R.; Baas, F.; Scheper, W. A β 1-42 Induces Mild Endoplasmic Reticulum Stress in an Aggregation State—Dependent Manner. Antioxid. Redox Signal. 2007, 9, 2245–2254. [Google Scholar] [CrossRef]
- Barbero-Camps, E.; Fernández, A.; Baulies, A.; Martinez, L.; Fernández-Checa, J.C.; Colell, A. Endoplasmic reticulum stress mediates amyloid β neurotoxicity via mitochondrial cholesterol trafficking. Am. J. Pathol. 2014, 184, 2066–2081. [Google Scholar] [CrossRef]
- Nishitsuji, K.; Tomiyama, T.; Ishibashi, K.; Ito, K.; Teraoka, R.; Lambert, M.P.; Klein, W.L.; Mori, H. The E693Δ mutation in amyloid precursor protein increases intracellular accumulation of amyloid β oligomers and causes endoplasmic reticulum stress-induced apoptosis in cultured cells. Am. J. Pathol. 2009, 174, 957–969. [Google Scholar] [CrossRef]
- Casas-Tinto, S.; Zhang, Y.; Sanchez-Garcia, J.; Gomez-Velazquez, M.; Rincon-Limas, D.E.; Fernandez-Funez, P. The ER stress factor XBP1s prevents amyloid-β neurotoxicity. Hum. Mol. Genet. 2011, 20, 2144–2160. [Google Scholar] [CrossRef]
- Gourmaud, S.; Paquet, C.; Dumurgier, J.; Pace, C.; Bouras, C.; Gray, F.; Laplanche, J.-L.; Meurs, E.F.; Mouton-Liger, F.; Hugon, J. Increased levels of cerebrospinal fluid JNK3 associated with amyloid pathology: Links to cognitive decline. J Psychiatry Neurosci. 2015, 40, 151. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ron, D. Stress-induced phosphorylation and activation of the transcription factor CHOP (GADD153) by p38 MAP kinase. Science 1996, 272, 1347–1349. [Google Scholar] [CrossRef] [PubMed]
- Yoneda, T.; Imaizumi, K.; Oono, K.; Yui, D.; Gomi, F.; Katayama, T.; Tohyama, M. Activation of caspase-12, an endoplastic reticulum (ER) resident caspase, through tumor necrosis factor receptor-associated factor 2-dependent mechanism in response to the ER stress. J. Biol. Chem. 2001, 276, 13935–13940. [Google Scholar] [CrossRef] [PubMed]
- Nishitoh, H.; Matsuzawa, A.; Tobiume, K.; Saegusa, K.; Takeda, K.; Inoue, K.; Hori, S.; Kakizuka, A.; Ichijo, H. ASK1 is essential for endoplasmic reticulum stress-induced neuronal cell death triggered by expanded polyglutamine repeats. Genes Dev. 2002, 16, 1345–1355. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Won, S.M.; Suh, J.; Son, S.J.; Moon, G.J.; Park, U.-J.; Gwag, B.J. Induction of the unfolded protein response and cell death pathway in Alzheimer’s disease, but not in aged Tg2576 mice. Exp. Mol. Med. 2010, 42, 386–394. [Google Scholar] [CrossRef]
- Duran-Aniotz, C.; Cornejo, V.H.; Espinoza, S.; Ardiles, Á.O.; Medinas, D.B.; Salazar, C.; Foley, A.; Gajardo, I.; Thielen, P.; Iwawaki, T. IRE1 signaling exacerbates Alzheimer’s disease pathogenesis. Acta Neuropathol. 2017, 134, 489–506. [Google Scholar] [CrossRef]
- Meusser, B.; Hirsch, C.; Jarosch, E.; Sommer, T. ERAD: The long road to destruction. Nat. Cell Biol. 2005, 7, 766–772. [Google Scholar] [CrossRef]
- Khandelwal, P.J.; Herman, A.M.; Hoe, H.-S.; Rebeck, G.W.; Moussa, C.E.-H. Parkin mediates beclin-dependent autophagic clearance of defective mitochondria and ubiquitinated Aβ in AD models. Hum. Mol. Genet. 2011, 20, 2091–2102. [Google Scholar] [CrossRef]
- Poon, W.W.; Carlos, A.J.; Aguilar, B.L.; Berchtold, N.C.; Kawano, C.K.; Zograbyan, V.; Yaopruke, T.; Shelanski, M.; Cotman, C.W. β-Amyloid (Aβ) oligomers impair brain-derived neurotrophic factor retrograde trafficking by down-regulating ubiquitin C-terminal hydrolase, UCH-L1. J. Biol. Chem. 2013, 288, 16937–16948. [Google Scholar] [CrossRef]
- Saido, T.; Leissring, M.A. Proteolytic degradation of amyloid β-protein. Cold Spring Harb. Perspect. Med. 2012, 2, a006379. [Google Scholar] [CrossRef]
- Salon, M.L.; Pasquini, L.; Moreno, M.B.; Pasquini, J.; Soto, E. Relationship between β-amyloid degradation and the 26S proteasome in neural cells. Exp. Neurol. 2003, 180, 131–143. [Google Scholar] [CrossRef]
- Gallart-Palau, X.; Guo, X.; Serra, A.; Sze, S.K. Alzheimer’s disease progression characterized by alterations in the molecular profiles and biogenesis of brain extracellular vesicles. Alzheimers Res. Ther. 2020, 12, 54. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Yang, J. Amyloid-β peptide is a substrate of the human 20S proteasome. ACS Chem. Neurosci. 2010, 1, 655–660. [Google Scholar] [CrossRef] [PubMed]
- Desikan, R.S.; Fan, C.C.; Wang, Y.; Schork, A.J.; Cabral, H.J.; Cupples, L.A.; Thompson, W.K.; Besser, L.; Kukull, W.A.; Holland, D. Genetic assessment of age-associated Alzheimer disease risk: Development and validation of a polygenic hazard score. PLoS Med. 2017, 14, e1002258. [Google Scholar] [CrossRef]
- Zhu, B.; Jiang, L.; Huang, T.; Zhao, Y.; Liu, T.; Zhong, Y.; Li, X.; Campos, A.; Pomeroy, K.; Masliah, E. ER-associated degradation regulates Alzheimer’s amyloid pathology and memory function by modulating γ-secretase activity. Nat. Commun. 2017, 8, 1472. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, S.; Cuervo, A.M. Chaperone-mediated autophagy: A unique way to enter the lysosome world. Trends Cell Biol. 2012, 22, 407–417. [Google Scholar] [CrossRef]
- Li, W.; Zhu, J.; Dou, J.; She, H.; Tao, K.; Xu, H.; Yang, Q.; Mao, Z. Phosphorylation of LAMP2A by p38 MAPK couples ER stress to chaperone-mediated autophagy. Nat. Commun. 2017, 8, 1763. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N. The ATG conjugation systems in autophagy. Curr. Opin. Cell Biol. 2020, 63, 1–10. [Google Scholar] [CrossRef]
- Yim, W.W.Y.; Mizushima, N. Lysosome biology in autophagy. Cell Discov. 2020, 6, 6. [Google Scholar] [CrossRef]
- Nixon, R.A.; Wegiel, J.; Kumar, A.; Yu, W.H.; Peterhoff, C.; Cataldo, A.; Cuervo, A.M. Extensive involvement of autophagy in Alzheimer disease: An immuno-electron microscopy study. J. Neuropathol. Exp. Neurol. 2005, 64, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.H.; Cuervo, A.M.; Kumar, A.; Peterhoff, C.M.; Schmidt, S.D.; Lee, J.-H.; Mohan, P.S.; Mercken, M.; Farmery, M.R.; Tjernberg, L.O. Macroautophagy—A novel β-amyloid peptide-generating pathway activated in Alzheimer’s disease. J. Cell Biol. 2005, 171, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Fedeli, C.; Filadi, R.; Rossi, A.; Mammucari, C.; Pizzo, P. PSEN2 (presenilin 2) mutants linked to familial Alzheimer disease impair autophagy by altering Ca2+ homeostasis. Autophagy 2019, 15, 2044–2062. [Google Scholar] [CrossRef]
- Pickford, F.; Masliah, E.; Britschgi, M.; Lucin, K.; Narasimhan, R.; Jaeger, P.A.; Small, S.; Spencer, B.; Rockenstein, E.; Levine, B. The autophagy-related protein beclin 1 shows reduced expression in early Alzheimer disease and regulates amyloid β accumulation in mice. J. Clin. Investig. 2008, 118, 2190–2199. [Google Scholar] [CrossRef]
- Swaminathan, G.; Zhu, W.; Plowey, E.D. BECN1/Beclin 1 sorts cell-surface APP/amyloid β precursor protein for lysosomal degradation. Autophagy 2016, 12, 2404–2419. [Google Scholar] [CrossRef]
- Ma, J.F.; Huang, Y.; Chen, S.D.; Halliday, G. Immunohistochemical evidence for macroautophagy in neurones and endothelial cells in Alzheimer’s disease. Neuropathol. Appl. Neurobiol. 2010, 36, 312–319. [Google Scholar] [CrossRef]
- Omata, Y.; Lim, Y.-M.; Akao, Y.; Tsuda, L. Age-induced reduction of autophagy-related gene expression is associated with onset of Alzheimer’s disease. Am. J. Neurodegener. Dis. 2014, 3, 134. [Google Scholar] [PubMed]
- Manczak, M.; Kandimalla, R.; Yin, X.; Reddy, P.H. Hippocampal mutant APP and amyloid beta-induced cognitive decline, dendritic spine loss, defective autophagy, mitophagy and mitochondrial abnormalities in a mouse model of Alzheimer’s disease. Hum. Mol. Genet. 2018, 27, 1332–1342. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.T. Amyloid-β Causes Autophagy Dysfunction by Inhibiting Protein Prenylation; University of Alberta, Canada: Edmonton, AB, Canada, 2016. [Google Scholar]
- Pigino, G.; Morfini, G.; Atagi, Y.; Deshpande, A.; Yu, C.; Jungbauer, L.; LaDu, M.; Busciglio, J.; Brady, S. Disruption of fast axonal transport is a pathogenic mechanism for intraneuronal amyloid beta. Proc. Natl. Acad. Sci. USA 2009, 106, 5907–5912. [Google Scholar] [CrossRef] [PubMed]
- Tammineni, P.; Cai, Q. Defective retrograde transport impairs autophagic clearance in Alzheimer disease neurons. Autophagy 2017, 13, 982–984. [Google Scholar] [CrossRef]
- Aboud, O.; Parcon, P.A.; DeWall, K.M.; Liu, L.; Mrak, R.E.; Griffin, S.T. Aging, Alzheimer’s, and APOE genotype influence the expression and neuronal distribution patterns of microtubule motor protein dynactin-P50. Front. Cell. Neurosci. 2015, 9, 103. [Google Scholar] [CrossRef]
- Xiao, T.; Zhang, L.; Huang, Y.; Shi, Y.; Wang, J.; Ji, Q.; Ye, J.; Lin, Y.; Liu, H. Sestrin2 increases in aortas and plasma from aortic dissection patients and alleviates angiotensin II-induced smooth muscle cell apoptosis via the Nrf2 pathway. Life Sci. 2019, 218, 132–138. [Google Scholar] [CrossRef]
- Rojo, A.I.; Pajares, M.; Rada, P.; Nuñez, A.; Nevado-Holgado, A.J.; Killik, R.; Van Leuven, F.; Ribe, E.; Lovestone, S.; Yamamoto, M. NRF2 deficiency replicates transcriptomic changes in Alzheimer’s patients and worsens APP and TAU pathology. Redox Biol. 2017, 13, 444–451. [Google Scholar] [CrossRef] [PubMed]
- Bahn, G.; Park, J.-S.; Yun, U.J.; Lee, Y.J.; Choi, Y.; Park, J.S.; Baek, S.H.; Choi, B.Y.; Cho, Y.S.; Kim, H.K. NRF2/ARE pathway negatively regulates BACE1 expression and ameliorates cognitive deficits in mouse Alzheimer’s models. Proc. Natl. Acad. Sci. USA 2019, 116, 12516–12523. [Google Scholar] [CrossRef] [PubMed]
- Neely, K.M.; Green, K.N.; LaFerla, F.M. Presenilin is necessary for efficient proteolysis through the autophagy–lysosome system in a γ-secretase-independent manner. J. Neurosci. 2011, 31, 2781–2791. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-H.; Yu, W.H.; Kumar, A.; Lee, S.; Mohan, P.S.; Peterhoff, C.M.; Wolfe, D.M.; Martinez-Vicente, M.; Massey, A.C.; Sovak, G. Lysosomal proteolysis and autophagy require presenilin 1 and are disrupted by Alzheimer-related PS1 mutations. Cell 2010, 141, 1146–1158. [Google Scholar] [CrossRef]
- Coen, K.; Flannagan, R.S.; Baron, S.; Carraro-Lacroix, L.R.; Wang, D.; Vermeire, W.; Michiels, C.; Munck, S.; Baert, V.; Sugita, S. Lysosomal calcium homeostasis defects, not proton pump defects, cause endo-lysosomal dysfunction in PSEN-deficient cells. J. Cell Biol. 2012, 198, 23–35. [Google Scholar] [CrossRef]
- Ditaranto, K.; Tekirian, T.L.; Yang, A.J. Lysosomal membrane damage in soluble Aβ-mediated cell death in Alzheimer’s disease. Neurobiol. Dis. 2001, 8, 19–31. [Google Scholar] [CrossRef]
- Krabbe, G.; Halle, A.; Matyash, V.; Rinnenthal, J.L.; Eom, G.D.; Bernhardt, U.; Miller, K.R.; Prokop, S.; Kettenmann, H.; Heppner, F.L. Functional impairment of microglia coincides with Beta-amyloid deposition in mice with Alzheimer-like pathology. PLoS ONE 2013, 8, e60921. [Google Scholar] [CrossRef]
- Guo, X.; Tang, P.; Chen, L.; Liu, P.; Hou, C.; Zhang, X.; Liu, Y.; Chong, L.; Li, X.; Li, R. Amyloid β-induced redistribution of transcriptional factor EB and lysosomal dysfunction in primary microglial cells. Front. Aging Neurosci. 2017, 9, 228. [Google Scholar] [CrossRef]
- Jiang, Y.; Sato, Y.; Im, E.; Berg, M.; Bordi, M.; Darji, S.; Kumar, A.; Mohan, P.S.; Bandyopadhyay, U.; Diaz, A. Lysosomal dysfunction in Down syndrome is APP-dependent and mediated by APP-βCTF (C99). J. Neurosci. 2019, 39, 5255–5268. [Google Scholar] [CrossRef]
- Soura, V.; Stewart-Parker, M.; Williams, T.L.; Ratnayaka, A.; Atherton, J.; Gorringe, K.; Tuffin, J.; Darwent, E.; Rambaran, R.; Klein, W. Visualization of co-localization in Aβ42-administered neuroblastoma cells reveals lysosome damage and autophagosome accumulation related to cell death. Biochem. J. 2012, 441, 579–590. [Google Scholar] [CrossRef] [PubMed]
- Gowrishankar, S.; Yuan, P.; Wu, Y.; Schrag, M.; Paradise, S.; Grutzendler, J.; De Camilli, P.; Ferguson, S.M. Massive accumulation of luminal protease-deficient axonal lysosomes at Alzheimer’s disease amyloid plaques. Proc. Natl. Acad. Sci. USA 2015, 112, E3699–E3708. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.Z.A.; Fourriere, L.; Wang, J.; Perez, F.; Boncompain, G.; Gleeson, P.A. Distinct anterograde trafficking pathways of BACE1 and amyloid precursor protein from the TGN and the regulation of amyloid-β production. Mol. Biol. Cell. 2020, 31, 27–44. [Google Scholar] [CrossRef]
- Toh, W.H.; Tan, J.Z.A.; Zulkefli, K.L.; Houghton, F.J.; Gleeson, P.A. Amyloid precursor protein traffics from the Golgi directly to early endosomes in an Arl5b-and AP4-dependent pathway. Traffic 2017, 18, 159–175. [Google Scholar] [CrossRef] [PubMed]
- Gosavi, P.; Gleeson, P.A. The function of the Golgi ribbon structure–an enduring mystery unfolds! Bioessays 2017, 39, 1700063. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.-H.; Seemann, J. Golgi ribbon disassembly during mitosis, differentiation and disease progression. Curr. Opin. Cell Biol. 2017, 47, 43–51. [Google Scholar] [CrossRef]
- Kim, M.J.; Je, A.R.; Kim, H.-J.; Huh, Y.H.; Kweon, H.S. Coat protein I depletion-associated Golgi fragmentation in an Alzheimer’s disease model. Anim. Cells Syst. 2015, 19, 8–15. [Google Scholar] [CrossRef]
- Joshi, G.; Chi, Y.; Huang, Z.; Wang, Y. Aβ-induced Golgi fragmentation in Alzheimer’s disease enhances Aβ production. Proc. Natl. Acad. Sci. USA 2014, 111, E1230–E1239. [Google Scholar] [CrossRef]
- Joshi, G.; Bekier, M.I.; Wang, Y. Golgi fragmentation in Alzheimer’s disease. Front. Neurosci. 2015, 9, 340. [Google Scholar] [CrossRef]
- Sun, K.-H.; de Pablo, Y.; Vincent, F.; Johnson, E.O.; Chavers, A.K.; Shah, K. Novel genetic tools reveal Cdk5’s major role in Golgi fragmentation in Alzheimer’s disease. Mol. Biol. Cell. 2008, 19, 3052–3069. [Google Scholar] [CrossRef]
- Jiang, Q.; Wang, L.; Guan, Y.; Xu, H.; Niu, Y.; Han, L.; Wei, Y.-P.; Lin, L.; Chu, J.; Wang, Q. Golgin-84-associated Golgi fragmentation triggers tau hyperphosphorylation by activation of cyclin-dependent kinase-5 and extracellular signal-regulated kinase. Neurobiol. Aging 2014, 35, 1352–1363. [Google Scholar] [CrossRef] [PubMed]
- Liazoghli, D.; Perreault, S.; Micheva, K.D.; Desjardins, M.; Leclerc, N. Fragmentation of the Golgi apparatus induced by the overexpression of wild-type and mutant human tau forms in neurons. Am. J. Pathol. 2005, 166, 1499–1514. [Google Scholar] [CrossRef]
- Cervigni, R.I.; Bonavita, R.; Barretta, M.L.; Spano, D.; Ayala, I.; Nakamura, N.; Corda, D.; Colanzi, A. JNK2 controls fragmentation of the Golgi complex and the G2/M transition through phosphorylation of GRASP65. J. Cell Biol. 2015, 128, 2249–2260. [Google Scholar] [CrossRef] [PubMed]
- Bellouze, S.; Schäfer, M.K.; Buttigieg, D.; Baillat, G.; Rabouille, C.; Haase, G. Golgi fragmentation in pmn mice is due to a defective ARF1/TBCE cross-talk that coordinates COPI vesicle formation and tubulin polymerization. Hum. Mol. Genet. 2014, 23, 5961–5975. [Google Scholar] [CrossRef] [PubMed]
- Gonatas, N.K.; Stieber, A.; Gonatas, J.O. Fragmentation of the Golgi apparatus in neurodegenerative diseases and cell death. J. Neurol. Sci. 2006, 246, 21–30. [Google Scholar] [CrossRef]
- Suga, K.; Saito, A.; Mishima, T.; Akagawa, K. ER and Golgi stresses increase ER–Golgi SNARE Syntaxin5: Implications for organelle stress and βAPP processing. Neurosci. Lett. 2015, 604, 30–35. [Google Scholar] [CrossRef]
- Barucker, C.; Harmeier, A.; Weiske, J.; Fauler, B.; Albring, K.F.; Prokop, S.; Hildebrand, P.; Lurz, R.; Heppner, F.L.; Huber, O. Nuclear translocation uncovers the amyloid peptide Aβ42 as a regulator of gene transcription. J. Biol. Chem. 2014, 289, 20182–20191. [Google Scholar] [CrossRef]
- Maloney, B.; Lahiri, D.K. The Alzheimer’s amyloid β-peptide (Aβ) binds a specific DNA Aβ-interacting domain (AβID) in the APP, BACE1, and APOE promoters in a sequence-specific manner: Characterizing a new regulatory motif. Gene 2011, 488, 1–12. [Google Scholar] [CrossRef]
- Bailey, J.A.; Maloney, B.; Ge, Y.-W.; Lahiri, D.K. Functional activity of the novel Alzheimer’s amyloid β-peptide interacting domain (AβID) in the APP and BACE1 promoter sequences and implications in activating apoptotic genes and in amyloidogenesis. Gene 2011, 488, 13–22. [Google Scholar] [CrossRef][Green Version]
- Gezen-Ak, D.; Atasoy, İ.L.; Candaş, E.; Alaylıoğlu, M.; Dursun, E. The transcriptional regulatory properties of amyloid beta 1–42 may include regulation of genes related to neurodegeneration. Neuromol. Med. 2018, 20, 363–375. [Google Scholar] [CrossRef]
- Barucker, C.; Sommer, A.; Beckmann, G.; Eravci, M.; Harmeier, A.; Schipke, C.G.; Brockschnieder, D.; Dyrks, T.; Althoff, V.; Fraser, P.E. Alzheimer amyloid peptide Aβ 42 regulates gene expression of transcription and growth factors. J. Alzheimers Dis. 2015, 44, 613–624. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Kim, H.S.; Joo, Y.; Choi, Y.; Chang, K.; Park, C.; Shin, K.; Kim, S.; Cheon, Y.; Baik, T. Intracellular domains of amyloid precursor-like protein 2 interact with CP2 transcription factor in the nucleus and induce glycogen synthase kinase-3 β expression. Cell Death Differ. 2007, 14, 79–91. [Google Scholar] [CrossRef]
- Zhu, H.; Fu, W.; Mattson, M.P. The catalytic subunit of telomerase protects neurons against amyloid β-peptide-induced apoptosis. J. Neurochem. 2000, 75, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Flanary, B.E.; Sammons, N.W.; Nguyen, C.; Walker, D.; Streit, W.J. Evidence that aging and amyloid promote microglial cell senescence. Rejuvenation Res. 2007, 10, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Jaskelioff, M.; Muller, F.L.; Paik, J.-H.; Thomas, E.; Jiang, S.; Adams, A.C.; Sahin, E.; Kost-Alimova, M.; Protopopov, A.; Cadinanos, J. Telomerase reactivation reverses tissue degeneration in aged telomerase-deficient mice. Nature 2011, 469, 102–106. [Google Scholar] [CrossRef]
- Wang, J.; Zhao, C.; Zhao, A.; Li, M.; Ren, J.; Qu, X. New insights in amyloid beta interactions with human telomerase. J. Am. Chem. Soc. 2015, 137, 1213–1219. [Google Scholar] [CrossRef]
- Collado, M.; Blasco, M.A.; Serrano, M. Cellular senescence in cancer and aging. Cell 2007, 130, 223–233. [Google Scholar] [CrossRef]
- Boccardi, V.; Pelini, L.; Ercolani, S.; Ruggiero, C.; Mecocci, P. From cellular senescence to Alzheimer’s disease: The role of telomere shortening. Ageing Res. Rev. 2015, 22, 1–8. [Google Scholar] [CrossRef]
- Fuso, A.; Seminara, L.; Cavallaro, R.A.; D’Anselmi, F.; Scarpa, S. S-adenosylmethionine/homocysteine cycle alterations modify DNA methylation status with consequent deregulation of PS1 and BACE and beta-amyloid production. Mol. Cell. Neurosci. 2005, 28, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Taher, N.; McKenzie, C.; Garrett, R.; Baker, M.; Fox, N.; Isaacs, G.D. Amyloid-β alters the DNA methylation status of cell-fate genes in an Alzheimer’s disease model. J. Alzheimers Dis. 2014, 38, 831–844. [Google Scholar] [CrossRef]
- Wang, S.-C.; Oelze, B.; Schumacher, A. Age-specific epigenetic drift in late-onset Alzheimer’s disease. PLoS ONE 2008, 3, e2698. [Google Scholar] [CrossRef]
- Chen, K.-L.; Wang, S.S.-S.; Yang, Y.-Y.; Yuan, R.-Y.; Chen, R.-M.; Hu, C.-J. The epigenetic effects of amyloid-β1–40 on global DNA and neprilysin genes in murine cerebral endothelial cells. Biochem. Biophys. Res. Commun. 2009, 378, 57–61. [Google Scholar] [CrossRef] [PubMed]
- Hales, C.M.; Seyfried, N.T.; Dammer, E.B.; Duong, D.; Yi, H.; Gearing, M.; Troncoso, J.C.; Mufson, E.J.; Thambisetty, M.; Levey, A.I. U1 small nuclear ribonucleoproteins (snRNPs) aggregate in Alzheimer’s disease due to autosomal dominant genetic mutations and trisomy 21. Mol. Neurodegener. 2014, 9, 15. [Google Scholar] [CrossRef] [PubMed]
- Castellano, J.M.; Deane, R.; Gottesdiener, A.J.; Verghese, P.B.; Stewart, F.R.; West, T.; Paoletti, A.C.; Kasper, T.R.; DeMattos, R.B.; Zlokovic, B.V. Low-density lipoprotein receptor overexpression enhances the rate of brain-to-blood Aβ clearance in a mouse model of β-amyloidosis. Proc. Natl. Acad. Sci. USA 2012, 109, 15502–15507. [Google Scholar] [CrossRef]
- Wilkinson, K.; El Khoury, J. Microglial scavenger receptors and their roles in the pathogenesis of Alzheimer’s disease. Int. J. Alzheimers Dis. 2012, 2012, 489456. [Google Scholar] [CrossRef]
- Kanekiyo, T.; Cirrito, J.R.; Liu, C.-C.; Shinohara, M.; Li, J.; Schuler, D.R.; Shinohara, M.; Holtzman, D.M.; Bu, G. Neuronal clearance of amyloid-β by endocytic receptor LRP1. J. Neurosci. 2013, 33, 19276–19283. [Google Scholar] [CrossRef] [PubMed]
- Brandenburg, L.O.; Konrad, M.; Wruck, C.J.; Koch, T.; Lucius, R.; Pufe, T. Functional and physical interactions between formyl-peptide-receptors and scavenger receptor MARCO and their involvement in amyloid beta 1–42-induced signal transduction in glial cells. J. Neurochem. 2010, 113, 749–760. [Google Scholar] [CrossRef]
- Takumi, Y.; Ramírez-León, V.; Laake, P.; Rinvik, E.; Ottersen, O.P. Different modes of expression of AMPA and NMDA receptors in hippocampal synapses. Nat. Neurosci. 1999, 2, 618–624. [Google Scholar] [CrossRef]
- Hayashi, Y.; Shi, S.-H.; Esteban, J.A.; Piccini, A.; Poncer, J.-C.; Malinow, R. Driving AMPA receptors into synapses by LTP and CaMKII: Requirement for GluR1 and PDZ domain interaction. Science 2000, 287, 2262–2267. [Google Scholar] [CrossRef] [PubMed]
- Song, I.; Huganir, R.L. Regulation of AMPA receptors during synaptic plasticity. Trends Neurosci. 2002, 25, 578–588. [Google Scholar] [CrossRef]
- Chan, S.L.; Griffin, W.S.T.; Mattson, M.P. Evidence for caspase-mediated cleavage of AMPA receptor subunits in neuronal apoptosis and Alzheimer’s disease. J. Neurosci. Res. 1999, 57, 315–323. [Google Scholar] [CrossRef]
- Szegedi, V.; Juhász, G.; Budai, D.; Penke, B. Divergent effects of Aβ1–42 on ionotropic glutamate receptor-mediated responses in CA1 neurons in vivo. Brain Res. 2005, 1062, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Malenka, R.C.; Bear, M.F. LTP and LTD: An embarrassment of riches. Neuron 2004, 44, 5–21. [Google Scholar] [CrossRef] [PubMed]
- Nieweg, K.; Andreyeva, A.; Van Stegen, B.; Tanriöver, G.; Gottmann, K. Alzheimer’s disease-related amyloid-β induces synaptotoxicity in human iPS cell-derived neurons. Cell Death Dis. 2015, 6, e1709. [Google Scholar] [CrossRef]
- Popugaeva, E.; Pchitskaya, E.; Bezprozvanny, I. Dysregulation of neuronal calcium homeostasis in Alzheimer’s disease—A therapeutic opportunity? Biochem. Biophys. Res. Commun. 2017, 483, 998–1004. [Google Scholar] [CrossRef]
- Gu, Z.; Liu, W.; Yan, Z. β-Amyloid impairs AMPA receptor trafficking and function by reducing Ca2+/calmodulin-dependent protein kinase II synaptic distribution. J. Biol. Chem. 2009, 284, 10639–10649. [Google Scholar] [CrossRef]
- Zhao, W.-Q.; Santini, F.; Breese, R.; Ross, D.; Zhang, X.D.; Stone, D.J.; Ferrer, M.; Townsend, M.; Wolfe, A.L.; Seager, M.A. Inhibition of calcineurin-mediated endocytosis and α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptors prevents amyloid β oligomer-induced synaptic disruption. J. Biol. Chem. 2010, 285, 7619–7632. [Google Scholar] [CrossRef]
- Kurup, P.; Zhang, Y.; Xu, J.; Venkitaramani, D.V.; Haroutunian, V.; Greengard, P.; Nairn, A.C.; Lombroso, P.J. Aβ-mediated NMDA receptor endocytosis in Alzheimer’s disease involves ubiquitination of the tyrosine phosphatase STEP61. J. Neurosci. 2010, 30, 5948–5957. [Google Scholar] [CrossRef]
- Li, S.; Jin, M.; Koeglsperger, T.; Shepardson, N.E.; Shankar, G.M.; Selkoe, D.J. Soluble Aβ oligomers inhibit long-term potentiation through a mechanism involving excessive activation of extrasynaptic NR2B-containing NMDA receptors. J. Neurosci. 2011, 31, 6627–6638. [Google Scholar] [CrossRef]
- Sheng, M.; Hoogenraad, C.C. The postsynaptic architecture of excitatory synapses: A more quantitative view. Annu. Rev. Biochem. 2007, 76, 823–847. [Google Scholar] [CrossRef] [PubMed]
- Roselli, F.; Tirard, M.; Lu, J.; Hutzler, P.; Lamberti, P.; Livrea, P.; Morabito, M.; Almeida, O. Soluble β-amyloid1-40 induces NMDA-dependent degradation of postsynaptic density-95 at glutamatergic synapses. J. Neurosci. 2005, 25, 11061–11070. [Google Scholar] [CrossRef] [PubMed]
- Lacor, P.N.; Buniel, M.C.; Chang, L.; Fernandez, S.J.; Gong, Y.; Viola, K.L.; Lambert, M.P.; Velasco, P.T.; Bigio, E.H.; Finch, C.E. Synaptic targeting by Alzheimer’s-related amyloid β oligomers. J. Neurosci. 2004, 24, 10191–10200. [Google Scholar] [CrossRef]
- Dohler, F.; Sepulveda-Falla, D.; Krasemann, S.; Altmeppen, H.; Schlüter, H.; Hildebrand, D.; Zerr, I.; Matschke, J.; Glatzel, M. High molecular mass assemblies of amyloid-β oligomers bind prion protein in patients with Alzheimer’s disease. Brain 2014, 137, 873–886. [Google Scholar] [CrossRef] [PubMed]
- Laurén, J.; Gimbel, D.A.; Nygaard, H.B.; Gilbert, J.W.; Strittmatter, S.M. Cellular prion protein mediates impairment of synaptic plasticity by amyloid-β oligomers. Nature 2009, 457, 1128–1132. [Google Scholar] [CrossRef]
- Rushworth, J.V.; Griffiths, H.H.; Watt, N.T.; Hooper, N.M. Prion protein-mediated toxicity of amyloid-β oligomers requires lipid rafts and the transmembrane LRP1. J. Biol. Chem. 2013, 288, 8935–8951. [Google Scholar] [CrossRef]
- Um, J.W.; Kaufman, A.C.; Kostylev, M.; Heiss, J.K.; Stagi, M.; Takahashi, H.; Kerrisk, M.E.; Vortmeyer, A.; Wisniewski, T.; Koleske, A.J. Metabotropic glutamate receptor 5 is a coreceptor for Alzheimer aβ oligomer bound to cellular prion protein. Neuron 2013, 79, 887–902. [Google Scholar] [CrossRef]
- Larson, M.; Sherman, M.A.; Amar, F.; Nuvolone, M.; Schneider, J.A.; Bennett, D.A.; Aguzzi, A.; Lesné, S.E. The complex PrPc-Fyn couples human oligomeric Aβ with pathological Tau changes in Alzheimer’s disease. J. Neurosci. 2012, 32, 16857–16871. [Google Scholar] [CrossRef]
- Taylor, D.R.; Whitehouse, I.J.; Hooper, N.M. Glypican-1 mediates both prion protein lipid raft association and disease isoform formation. PLoSPathog. 2009, 5, e1000666. [Google Scholar] [CrossRef]
- Khan, G.M.; Tong, M.; Jhun, M.; Arora, K.; Nichols, R.A. β-Amyloid activates presynaptic α7 nicotinic acetylcholine receptors reconstituted into a model nerve cell system: Involvement of lipid rafts. Eur. J. Neurosci. 2010, 31, 788–796. [Google Scholar] [CrossRef]
- Cissé, M.; Halabisky, B.; Harris, J.; Devidze, N.; Dubal, D.B.; Sun, B.; Orr, A.; Lotz, G.; Kim, D.H.; Hamto, P. Reversing EphB2 depletion rescues cognitive functions in Alzheimer model. Nature 2011, 469, 47–52. [Google Scholar] [CrossRef]
- Vargas, L.M.; Leal, N.; Estrada, L.D.; González, A.; Serrano, F.; Araya, K.; Gysling, K.; Inestrosa, N.C.; Pasquale, E.B.; Alvarez, A.R. EphA4 activation of c-Abl mediates synaptic loss and LTP blockade caused by amyloid-β oligomers. PLoS ONE 2014, 9, e92309. [Google Scholar]
- Fu, A.K.; Hung, K.-W.; Huang, H.; Gu, S.; Shen, Y.; Cheng, E.Y.; Ip, F.C.; Huang, X.; Fu, W.-Y.; Ip, N.Y. Blockade of EphA4 signaling ameliorates hippocampal synaptic dysfunctions in mouse models of Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 2014, 111, 9959–9964. [Google Scholar] [CrossRef]
- Kumar, R.; Herbert, P.; Warrens, A. An introduction to death receptors in apoptosis. Int. J. Surg. 2005, 3, 268–277. [Google Scholar] [CrossRef]
- Ivins, K.J.; Thornton, P.L.; Rohn, T.T.; Cotman, C.W. Neuronal apoptosis induced by β-amyloid is mediated by caspase-8. Neurobiol. Dis. 1999, 6, 440–449. [Google Scholar] [CrossRef]
- Fossati, S.; Ghiso, J.; Rostagno, A. TRAIL death receptors DR4 and DR5 mediate cerebral microvascular endothelial cell apoptosis induced by oligomeric Alzheimer’s A β. Cell Death Dis. 2012, 3, e321. [Google Scholar] [CrossRef]
- Rabizadeh, S.; Bitler, C.M.; Butcher, L.L.; Bredesen, D.E. Expression of the low-affinity nerve growth factor receptor enhances beta-amyloid peptide toxicity. Proc. Natl. Acad. Sci. USA 1994, 91, 10703–10706. [Google Scholar] [CrossRef] [PubMed]
- Yaar, M.; Zhai, S.; Pilch, P.F.; Doyle, S.M.; Eisenhauer, P.B.; Fine, R.E.; Gilchrest, B.A. Binding of beta-amyloid to the p75 neurotrophin receptor induces apoptosis. A possible mechanism for Alzheimer’s disease. J. Clin. Investig. 1997, 100, 2333–2340. [Google Scholar] [CrossRef] [PubMed]
- Perini, G.; Della-Bianca, V.; Politi, V.; Della Valle, G.; Dal-Pra, I.; Rossi, F.; Armato, U. Role of p75 neurotrophin receptor in the neurotoxicity by β-amyloid peptides and synergistic effect of inflammatory cytokines. J. Exp. Med. 2002, 195, 907–918. [Google Scholar] [CrossRef]
- Sotthibundhu, A.; Sykes, A.M.; Fox, B.; Underwood, C.K.; Thangnipon, W.; Coulson, E.J. β-amyloid1–42 induces neuronal death through the p75 neurotrophin receptor. J. Neurosci. 2008, 28, 3941–3946. [Google Scholar] [CrossRef]
- Morishima, Y.; Gotoh, Y.; Zieg, J.; Barrett, T.; Takano, H.; Flavell, R.; Davis, R.J.; Shirasaki, Y.; Greenberg, M.E. β-Amyloid induces neuronal apoptosis via a mechanism that involves the c-Jun N-terminal kinase pathway and the induction of Fas ligand. J. Neurosci. 2001, 21, 7551–7560. [Google Scholar] [CrossRef]
- Knowles, J.K.; Rajadas, J.; Nguyen, T.-V.V.; Yang, T.; LeMieux, M.C.; Vander Griend, L.; Ishikawa, C.; Massa, S.M.; Wyss-Coray, T.; Longo, F.M. The p75 neurotrophin receptor promotes amyloid-β (1-42)-induced neuritic dystrophy in vitro and in vivo. J. Neurosci. 2009, 29, 10627–10637. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Yang, C.; Zhao, H.; Jian, C. Involvement of p75NTR in the effects of Aβ on L-type Ca2+ channel in cultured neuronal networks. Life Sci. 2020, 243, 117293. [Google Scholar] [CrossRef]
- Sasaki, N.; Toki, S.; Chowei, H.; Saito, T.; Nakano, N.; Hayashi, Y.; Takeuchi, M.; Makita, Z. Immunohistochemical distribution of the receptor for advanced glycation end products in neurons and astrocytes in Alzheimer’s disease. Brain Res. 2001, 888, 256–262. [Google Scholar] [CrossRef]
- Paudel, Y.N.; Angelopoulou, E.; Piperi, C.; Othman, I.; Aamir, K.; Shaikh, M. Impact of HMGB1, RAGE, and TLR4 in Alzheimer’s Disease (AD): From Risk Factors to Therapeutic Targeting. Cells 2020, 9, 383. [Google Scholar] [CrossRef]
- Hughes, C.; Choi, M.L.; Yi, J.-H.; Kim, S.-C.; Drews, A.; George-Hyslop, P.S.; Bryant, C.; Gandhi, S.; Cho, K.; Klenerman, D. Beta amyloid aggregates induce sensitised TLR4 signalling causing long-term potentiation deficit and rat neuronal cell death. Commun. Biol. 2020, 3, 79. [Google Scholar] [CrossRef]
- Walters, A.; Phillips, E.; Zheng, R.; Biju, M.; Kuruvilla, T. Evidence for neuroinflammation in Alzheimer’s disease. Prog. Neurol. Psychiatry 2016, 20, 25–31. [Google Scholar] [CrossRef]
- Varnum, M.M.; Ikezu, T. The classification of microglial activation phenotypes on neurodegeneration and regeneration in Alzheimer’s disease brain. Arch. Immunol. Ther. Exp. 2012, 60, 251–266. [Google Scholar] [CrossRef] [PubMed]
- Mietelska-Porowska, A.; Wojda, U. T lymphocytes and inflammatory mediators in the interplay between brain and blood in Alzheimer’s disease: Potential pools of new biomarkers. J. Immunol. Res. 2017, 2017, 4626540. [Google Scholar] [CrossRef]
- Dionisio-Santos, D.A.; Olschowka, J.A.; O’Banion, M.K. Exploiting microglial and peripheral immune cell crosstalk to treat Alzheimer’s disease. J. Neuroinflamm. 2019, 16, 74. [Google Scholar] [CrossRef] [PubMed]
- Mcgeer, P.; Mcgeer, E.; Rogers, J.; Sibley, J. Anti-inflammatory drugs and Alzheimer disease. Lancet 1990, 335, 1037. [Google Scholar] [CrossRef]
- Giometto, B.; Argentiero, V.; Sanson, F.; Ongaro, G.; Tavolato, B. Acute-phase proteins in Alzheimer’s disease. Eur. Neurol. 1988, 28, 30–33. [Google Scholar] [CrossRef] [PubMed]
- Miguel-Álvarez, M.; Santos-Lozano, A.; Sanchis-Gomar, F.; Fiuza-Luces, C.; Pareja-Galeano, H.; Garatachea, N.; Lucia, A. Non-steroidal anti-inflammatory drugs as a treatment for Alzheimer’s disease: A systematic review and meta-analysis of treatment effect. Drugs Aging 2015, 32, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Chan, C. The role of inflammasome in Alzheimer’s disease. Ageing Res. Rev. 2014, 15, 6–15. [Google Scholar] [CrossRef]
- Salter, M.W.; Stevens, B. Microglia emerge as central players in brain disease. Nat. Med. 2017, 23, 1018. [Google Scholar] [CrossRef]
- Yu, Y.; Richard, D.Y. Microglial Aβ receptors in Alzheimer’s disease. Cell. Mol. Neurobiol. 2015, 35, 71–83. [Google Scholar] [CrossRef] [PubMed]
- Halle, A.; Hornung, V.; Petzold, G.C.; Stewart, C.R.; Monks, B.G.; Reinheckel, T.; Fitzgerald, K.A.; Latz, E.; Moore, K.J.; Golenbock, D.T. The NALP3 inflammasome is involved in the innate immune response to amyloid-β. Nat. Immunol. 2008, 9, 857. [Google Scholar] [CrossRef]
- Heneka, M.T.; Kummer, M.P.; Stutz, A.; Delekate, A.; Schwartz, S.; Vieira-Saecker, A.; Griep, A.; Axt, D.; Remus, A.; Tzeng, T.C. NLRP3 is activated in Alzheimer’s disease and contributes to pathology in APP/PS1 mice. Nature 2013, 493, 674–678. [Google Scholar] [CrossRef]
- Sheedy, F.J.; Grebe, A.; Rayner, K.J.; Kalantari, P.; Ramkhelawon, B.; Carpenter, S.B.; Becker, C.E.; Ediriweera, H.N.; Mullick, A.E.; Golenbock, D.T. CD36 coordinates NLRP3 inflammasome activation by facilitating intracellular nucleation of soluble ligands into particulate ligands in sterile inflammation. Nat. Immunol. 2013, 14, 812–820. [Google Scholar] [CrossRef] [PubMed]
- Calsolaro, V.; Edison, P. Neuroinflammation in Alzheimer’s disease: Current evidence and future directions. Alzheimers Dement. 2016, 12, 719–732. [Google Scholar] [CrossRef]
- Sasaki, A. Microglia and brain macrophages: An update. Neuropathology 2017, 37, 452–464. [Google Scholar] [CrossRef] [PubMed]
- Perez, J.; Carrero, I.; Gonzalo, P.; Arevalo-Serrano, J.; Sanz-Anquela, J.; Ortega, J.; Rodriguez, M.; Gonzalo-Ruiz, A. Soluble oligomeric forms of beta-amyloid (Aβ) peptide stimulate Aβ production via astrogliosis in the rat brain. Exp. Neurol. 2010, 223, 410–421. [Google Scholar] [CrossRef]
- Carrero, I.; Gonzalo, M.; Martin, B.; Sanz-Anquela, J.; Arevalo-Serrano, J.; Gonzalo-Ruiz, A. Oligomers of beta-amyloid protein (Aβ1-42) induce the activation of cyclooxygenase-2 in astrocytes via an interaction with interleukin-1beta, tumour necrosis factor-alpha, and a nuclear factor kappa-B mechanism in the rat brain. Exp. Neurol. 2012, 236, 215–227. [Google Scholar] [CrossRef] [PubMed]
- Ringman, J.M.; O’Neill, J.; Geschwind, D.; Medina, L.; Apostolova, L.G.; Rodriguez, Y.; Schaffer, B.; Varpetian, A.; Tseng, B.; Ortiz, F. Diffusion tensor imaging in preclinical and presymptomatic carriers of familial Alzheimer’s disease mutations. Brain 2007, 130, 1767–1776. [Google Scholar] [CrossRef]
- Desai, M.K.; Guercio, B.J.; Narrow, W.C.; Bowers, W.J. An Alzheimer’s disease-relevant presenilin-1 mutation augments amyloid-beta-induced oligodendrocyte dysfunction. Glia 2011, 59, 627–640. [Google Scholar] [CrossRef]
- Vodovotz, Y.; Lucia, M.S.; Flanders, K.C.; Chesler, L.; Xie, Q.-W.; Smith, T.W.; Weidner, J.; Mumford, R.; Webber, R.; Nathan, C. Inducible nitric oxide synthase in tangle-bearing neurons of patients with Alzheimer’s disease. J. Exp. Med. 1996, 184, 1425–1433. [Google Scholar] [CrossRef]
- Limatola, C.; Ransohoff, R.M. Modulating neurotoxicity through CX3CL1/CX3CR1 signaling. Front. Cell. Neurosci. 2014, 8, 229. [Google Scholar] [CrossRef] [PubMed]
- D’Ambrosio, A.L.; Pinsky, D.J.; Connolly, E.S. The role of the complement cascade in ischemia/reperfusion injury: Implications for neuroprotection. Mol. Med. 2001, 7, 367–382. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Yang, L.; Li, R. What does complement do in Alzheimer’s disease? Old molecules with new insights. Transl. Neurodegener. 2013, 2, 21. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Meri, S. Yin and Yang: Complement activation and regulation in Alzheimer’s disease. Prog. Neurobiol. 2003, 70, 463–472. [Google Scholar] [CrossRef]
- Chibnik, L.B.; Shulman, J.M.; Leurgans, S.E.; Schneider, J.A.; Wilson, R.S.; Tran, D.; Aubin, C.; Buchman, A.S.; Heward, C.B.; Myers, A.J. CR1 is associated with amyloid plaque burden and age-related cognitive decline. Ann. Neurol. 2011, 69, 560–569. [Google Scholar] [CrossRef] [PubMed]
- Azizi, G.; Khannazer, N.; Mirshafiey, A. The potential role of chemokines in Alzheimer’s disease pathogenesis. Am. J. Alzheimers Dis. Other Dement. 2014, 29, 415–425. [Google Scholar] [CrossRef] [PubMed]
- Zilka, N.; Kazmerova, Z.; Jadhav, S.; Neradil, P.; Madari, A.; Obetkova, D.; Bugos, O.; Novak, M. Who fans the flames of Alzheimer’s disease brains? Misfolded tau on the crossroad of neurodegenerative and inflammatory pathways. J. Neuroinflamm. 2012, 9, 47. [Google Scholar] [CrossRef]
- Westin, K.; Buchhave, P.; Nielsen, H.; Minthon, L.; Janciauskiene, S.; Hansson, O. CCL2 is associated with a faster rate of cognitive decline during early stages of Alzheimer’s disease. PLoS ONE 2012, 7, e30525. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Hampel, H.; Prvulovic, D.; Wallin, A.; Blennow, K.; Li, R.; Shen, Y. Elevated CSF levels of TACE activity and soluble TNF receptors in subjects with mild cognitive impairment and patients with Alzheimer’s disease. Mol. Neurodegener. 2011, 6, 69. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.-Y.; Tan, M.-S.; Yu, J.-T.; Tan, L. Role of pro-inflammatory cytokines released from microglia in Alzheimer’s disease. Ann. Transl. Med. 2015, 3, 136. [Google Scholar]
- Goldgaber, D.; Harris, H.W.; Hla, T.; Maciag, T.; Donnelly, R.J.; Jacobsen, J.S.; Vitek, M.P.; Gajdusek, D.C. Interleukin 1 regulates synthesis of amyloid beta-protein precursor mRNA in human endothelial cells. Proc. Natl. Acad. Sci. USA 1989, 86, 7606–7610. [Google Scholar] [CrossRef]
- Griffin, W.S.T.; Sheng, J.G.; Roberts, G.W.; Mrak, R.E. Interleukin-1 expression in different plaque types in Alzheimer’s disease: Significance in plaque evolution. J. Neuropathol. Exp. Neurol. 1995, 54, 276–281. [Google Scholar] [CrossRef]
- Park, S.Y.; Kang, M.J.; Han, J.S. Interleukin-1 beta promotes neuronal differentiation through the Wnt5a/RhoA/JNK pathway in cortical neural precursor cells. Mol. Brain 2018, 11, 39. [Google Scholar] [CrossRef]
- Farrar, W.; Kilian, P.; Ruff, M.; Hill, J.; Pert, C. Visualization and characterization of interleukin 1 receptors in brain. J. Immunol. 1987, 139, 459–463. [Google Scholar]
- Buxbaum, J.D.; Oishi, M.; Chen, H.I.; Pinkas-Kramarski, R.; Jaffe, E.A.; Gandy, S.E.; Greengard, P. Cholinergic agonists and interleukin 1 regulate processing and secretion of the Alzheimer beta/A4 amyloid protein precursor. Proc. Natl. Acad. Sci. USA 1992, 89, 10075–10078. [Google Scholar] [CrossRef] [PubMed]
- Quintanilla, R.A.; Orellana, D.I.; González-Billault, C.; Maccioni, R.B. Interleukin-6 induces Alzheimer-type phosphorylation of tau protein by deregulating the cdk5/p35 pathway. Exp. Cell Res. 2004, 295, 245–257. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, K.; Smale, S.; Premchand, P.; Maiden, L.; Sherwood, R.; Thjodleifsson, B.; Bjornsson, E.; Bjarnason, I. Prevalence and mechanism of nonsteroidal anti-inflammatory drug–induced clinical relapse in patients with inflammatory bowel disease. Clin. Gastroenterol. Hepatol. 2006, 4, 196–202. [Google Scholar] [CrossRef]
- Perry, R.T.; Collins, J.S.; Wiener, H.; Acton, R.; Go, R.C. The role of TNF and its receptors in Alzheimer’s disease. Neurobiol. Aging 2001, 22, 873–883. [Google Scholar] [CrossRef]
- Yamamoto, M.; Kiyota, T.; Horiba, M.; Buescher, J.L.; Walsh, S.M.; Gendelman, H.E.; Ikezu, T. Interferon-γ and tumor necrosis factor-α regulate amyloid-β plaque deposition and β-secretase expression in Swedish mutant APP transgenic mice. Am. J. Pathol. 2007, 170, 680–692. [Google Scholar] [CrossRef] [PubMed]
- Chang, R.; Yee, K.-L.; Sumbria, R.K. Tumor necrosis factor α inhibition for Alzheimer’s disease. J. Cent. Nerv. Syst. Dis. 2017, 9, 1179573517709278. [Google Scholar] [CrossRef]
- Grammas, P.; Ovase, R. Cerebrovascular transforming growth factor-β contributes to inflammation in the Alzheimer’s disease brain. Am. J. Pathol. 2002, 160, 1583–1587. [Google Scholar] [CrossRef]
- Cao, B.-B.; Zhang, X.-X.; Du, C.-Y.; Liu, Z.; Qiu, Y.-H.; Peng, Y.-P. TGF-β1 Provides Neuroprotection via Inhibition of Microglial Activation in 3-Acetylpyridine-Induced Cerebellar Ataxia Model Rats. Front. Neurosci. 2020, 14, 187. [Google Scholar] [CrossRef]
- Caraci, F.; Spampinato, S.; Sortino, M.A.; Bosco, P.; Battaglia, G.; Bruno, V.; Drago, F.; Nicoletti, F.; Copani, A. Dysfunction of TGF-β1 signaling in Alzheimer’s disease: Perspectives for neuroprotection. Cell Tissue Res. 2012, 347, 291–301. [Google Scholar] [CrossRef]
- Lio, D.; Licastro, F.; Scola, L.; Chiappelli, M.; Grimaldi, L.; Crivello, A.; Colonna-Romano, G.; Candore, G.; Franceschi, C.; Caruso, C. Interleukin-10 promoter polymorphism in sporadic Alzheimer’s disease. Genes Immun. 2003, 4, 234–238. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, J.; Tian, C.; Xiao, Y.; Li, X.; He, C.; Huang, J.; Fan, H. The 1082G/A polymorphism in IL-10 gene is associated with risk of Alzheimer’s disease: A meta-analysis. J. Neurol. Sci. 2011, 303, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Ho, L.; Purohit, D.; Haroutunian, V.; Luterman, J.D.; Willis, F.; Naslund, J.; Buxbaum, J.D.; Mohs, R.C.; Aisen, P.S.; Pasinetti, G.M. Neuronal cyclooxygenase 2 expression in the hippocampal formation as a function of the clinical progression of Alzheimer disease. Arch. Neurol. 2001, 58, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Z.; Ho, L.; Yemul, S.; Zhao, Z.; Pompl, P.; Kelley, K.; Dang, A.; Qing, W.; Teplow, D.; Pasinetti, G.M. Cyclooxygenase-2 promotes amyloid plaque deposition in a mouse model of Alzheimer’s disease neuropathology. Gene Expr. J. Liver Res. 2002, 10, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Z.; Ho, L.; Valdellon, J.; Borchelt, D.; Kelley, K.; Spielman, L.; Aisen, P.S.; Pasinetti, G.M. Cyclooxygenase (COX)-2 and cell cycle activity in a transgenic mouse model of Alzheimer’s disease neuropathology. Neurobiol. Aging 2002, 23, 327–334. [Google Scholar] [CrossRef]
- Hoozemans, J.; Veerhuis, R.; Janssen, I.; Rozemuller, A.J.; Eikelenboom, P. Interleukin-1β induced cyclooxygenase. Exp. Gerontol. 2001, 2, 559–570. [Google Scholar] [CrossRef]
- Cinelli, M.A.; Do, H.T.; Miley, G.P.; Silverman, R.B. Inducible nitric oxide synthase: Regulation, structure, and inhibition. Med. Res. Rev. 2020, 40, 158–189. [Google Scholar] [CrossRef]
- Asiimwe, N.; Yeo, S.G.; Kim, M.-S.; Jung, J.; Jeong, N.Y. Nitric oxide: Exploring the contextual link with alzheimer’s disease. Oxid. Med. Cell. Longev. 2016, 2016, 7205747. [Google Scholar] [CrossRef]
- Saez, T.E.; Pehar, M.; Vargas, M.; Barbeito, L.; Maccioni, R.B. Astrocytic nitric oxide triggers tau hyperphosphorylation in hippocampal neurons. In Vivo 2004, 18, 275–280. [Google Scholar]
- Zhang, Q.; Yu, J.-T.; Zhu, Q.-X.; Zhang, W.; Wu, Z.-C.; Miao, D.; Tan, L. Complement receptor 1 polymorphisms and risk of late-onset Alzheimer’s disease. Brain Res. 2010, 1348, 216–221. [Google Scholar] [CrossRef]
- Brubaker, W.D.; Crane, A.; Johansson, J.U.; Yen, K.; Garfinkel, K.; Mastroeni, D.; Asok, P.; Bradt, B.; Sabbagh, M.; Wallace, T.L. Peripheral complement interactions with amyloid β peptide: Erythrocyte clearance mechanisms. Alzheimers Dement. 2017, 13, 1397–1409. [Google Scholar] [CrossRef] [PubMed]
- Ager, R.R.; Fonseca, M.I.; Chu, S.H.; Sanderson, S.D.; Taylor, S.M.; Woodruff, T.M.; Tenner, A.J. Microglial C5aR (CD88) expression correlates with amyloid-β deposition in murine models of Alzheimer’s disease. J. Neurochem. 2010, 113, 389–401. [Google Scholar] [CrossRef] [PubMed]
- Yazawa, H.; Yu, Z.X.; Takeda, K.; Le, Y.; Gong, W.; Ferrans, V.J.; Oppenheim, J.J.; Li, C.C.H.; Wang, J.M. β Amyloid peptide (Aβ42) is internalized via the G-protein-coupled receptor FPRL1 and forms fibrillar aggregates in macrophages1. FASEB J. 2001, 15, 2454–2462. [Google Scholar] [CrossRef] [PubMed]
- Tiffany, H.L.; Lavigne, M.C.; Cui, Y.-H.; Wang, J.-M.; Leto, T.L.; Gao, J.-L.; Murphy, P.M. Amyloid-β induces chemotaxis and oxidant stress by acting at formylpeptide receptor 2, a G protein-coupled receptor expressed in phagocytes and brain. J. Biol. Chem. 2001, 276, 23645–23652. [Google Scholar] [CrossRef] [PubMed]
- Paresce, D.M.; Ghosh, R.N.; Maxfield, F.R. Microglial cells internalize aggregates of the Alzheimer’s disease amyloid β-protein via a scavenger receptor. Neuron 1996, 17, 553–565. [Google Scholar] [CrossRef]
- Coraci, I.S.; Husemann, J.; Berman, J.W.; Hulette, C.; Dufour, J.H.; Campanella, G.K.; Luster, A.D.; Silverstein, S.C.; El Khoury, J.B. CD36, a class B scavenger receptor, is expressed on microglia in Alzheimer’s disease brains and can mediate production of reactive oxygen species in response to β-amyloid fibrils. Am. J. Pathol. 2002, 160, 101–112. [Google Scholar] [CrossRef]
- Stewart, C.R.; Stuart, L.M.; Wilkinson, K.; Van Gils, J.M.; Deng, J.; Halle, A.; Rayner, K.J.; Boyer, L.; Zhong, R.; Frazier, W.A. CD36 ligands promote sterile inflammation through assembly of a Toll-like receptor 4 and 6 heterodimer. Nat. Immunol. 2010, 11, 155. [Google Scholar] [CrossRef]
- Origlia, N.; Bonadonna, C.; Rosellini, A.; Leznik, E.; Arancio, O.; Yan, S.S.; Domenici, L. Microglial receptor for advanced glycation end product-dependent signal pathway drives β-amyloid-induced synaptic depression and long-term depression impairment in entorhinal cortex. J. Neurosci. 2010, 30, 11414–11425. [Google Scholar] [CrossRef]
- Slowik, A.; Merres, J.; Elfgen, A.; Jansen, S.; Mohr, F.; Wruck, C.J.; Pufe, T.; Brandenburg, L.O. Involvement of formyl peptide receptors in receptor for advanced glycation end products (RAGE)-and amyloid beta 1-42-induced signal transduction in glial cells. Mol. Neurodegener. 2012, 7, 55. [Google Scholar] [CrossRef]
- Michaud, J.P.; Hallé, M.; Lampron, A.; Thériault, P.; Préfontaine, P.; Filali, M.; Tribout-Jover, P.; Lanteigne, A.-M.; Jodoin, R.; Cluff, C. Toll-like receptor 4 stimulation with the detoxified ligand monophosphoryl lipid A improves Alzheimer’s disease-related pathology. Proc. Natl. Acad. Sci. USA 2013, 10, 1941–1946. [Google Scholar] [CrossRef]
- Reed-Geaghan, E.G.; Savage, J.C.; Hise, A.G.; Landreth, G.E. CD14 and toll-like receptors 2 and 4 are required for fibrillar Aβ-stimulated microglial activation. J. Neurosci. 2009, 29, 11982–11992. [Google Scholar] [CrossRef]
- Doi, Y.; Mizuno, T.; Maki, Y.; Jin, S.; Mizoguchi, H.; Ikeyama, M.; Doi, M.; Michikawa, M.; Takeuchi, H.; Suzumura, A. Microglia activated with the toll-like receptor 9 ligand CpG attenuate oligomeric amyloid β neurotoxicity in in vitro and in vivo models of Alzheimer’s disease. Am. J. Pathol. 2009, 175, 2121–2132. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Varvel, N.H.; Konerth, M.E.; Xu, G.; Cardona, A.E.; Ransohoff, R.M.; Lamb, B.T. CX3CR1 deficiency alters microglial activation and reduces beta-amyloid deposition in two Alzheimer’s disease mouse models. Am. J. Pathol. 2010, 177, 2549–2562. [Google Scholar] [CrossRef] [PubMed]
- Leyns, C.E.; Ulrich, J.D.; Finn, M.B.; Stewart, F.R.; Koscal, L.J.; Serrano, J.R.; Robinson, G.O.; Anderson, E.; Colonna, M.; Holtzman, D.M. TREM2 deficiency attenuates neuroinflammation and protects against neurodegeneration in a mouse model of tauopathy. Proc. Natl. Acad. Sci. USA 2017, 114, 11524–11529. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Wu, X.; Li, X.; Jiang, L.-L.; Gui, X.; Liu, Y.; Sun, Y.; Zhu, B.; Piña-Crespo, J.C.; Zhang, M. TREM2 is a receptor for β-amyloid that mediates microglial function. Neuron 2018, 97, 1023–1031. [Google Scholar] [CrossRef] [PubMed]
- Griciuc, A.; Serrano-Pozo, A.; Parrado, A.R.; Lesinski, A.N.; Asselin, C.N.; Mullin, K.; Hooli, B.; Choi, S.H.; Hyman, B.T.; Tanzi, R.E. Alzheimer’s disease risk gene CD33 inhibits microglial uptake of amyloid beta. Neuron 2013, 78, 631–643. [Google Scholar] [CrossRef]
- Chen, H.; Liu, S.; Ji, L.; Wu, T.; Ji, Y.; Zhou, Y.; Zheng, M.; Zhang, M.; Xu, W.; Huang, G. Folic acid supplementation mitigates Alzheimer’s disease by reducing inflammation: A randomized controlled trial. Mediat. Inflamm. 2016, 2016, 5912146. [Google Scholar] [CrossRef]
- Rafiee, S.; Asadollahi, K.; Riazi, G.; Ahmadian, S.; Saboury, A.A. Vitamin B12 inhibits tau fibrillization via binding to cysteine residues of tau. ACS Chem. Neurosci. 2017, 8, 2676–2682. [Google Scholar] [CrossRef]
- Jia, L.; Wang, Y.; Wei, W.; Zhao, W.; Lu, F.; Liu, F. Vitamin B12 inhibits α-synuclein fibrillogenesis and protects against amyloid-induced cytotoxicity. Food Funct. 2019, 10, 2861–2870. [Google Scholar] [CrossRef]
- Huang, L.K.; Chao, S.P.; Hu, C.J. Clinical trials of new drugs for Alzheimer disease. J. Biomed. Sci. 2020, 27, 18. [Google Scholar] [CrossRef]
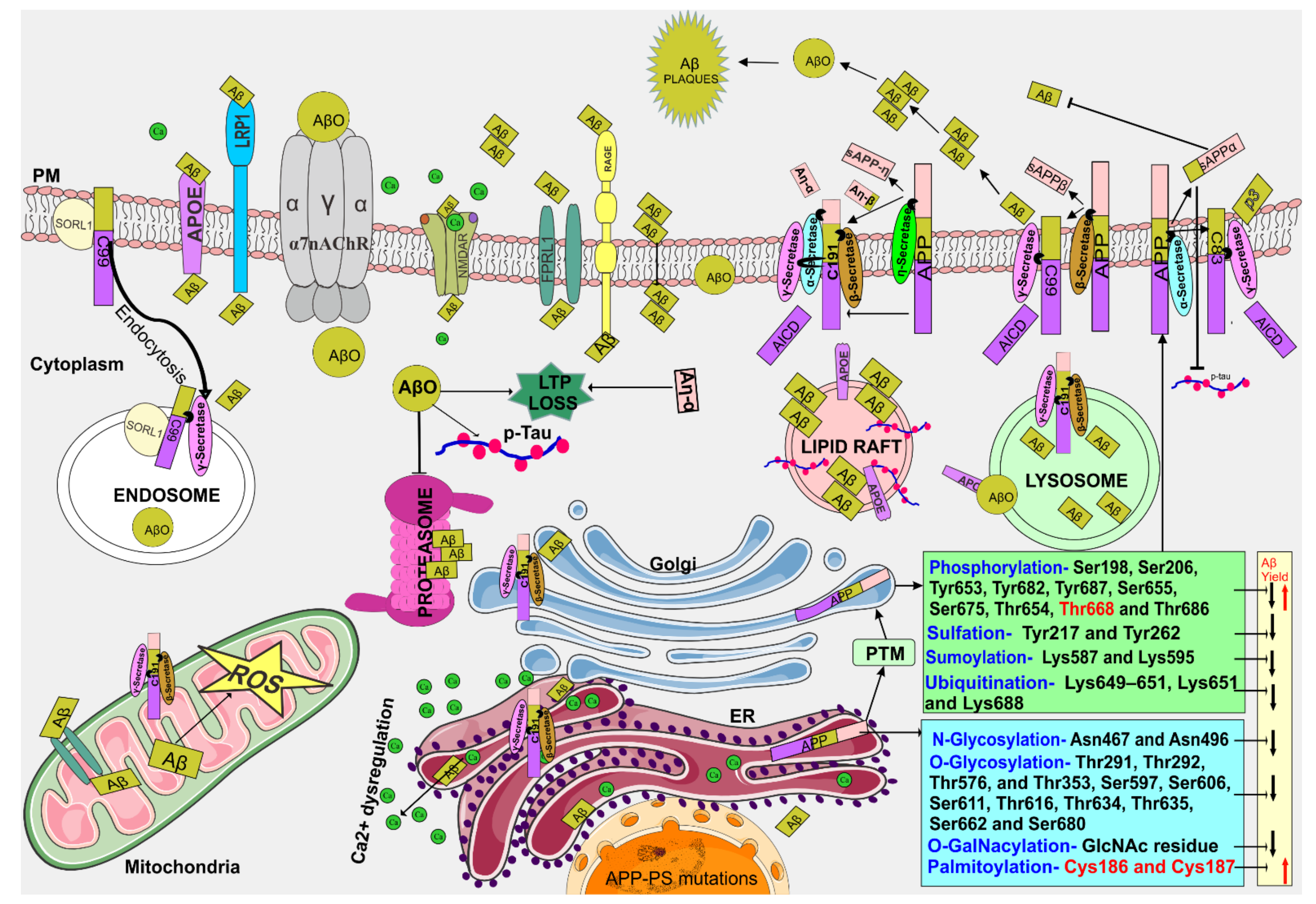
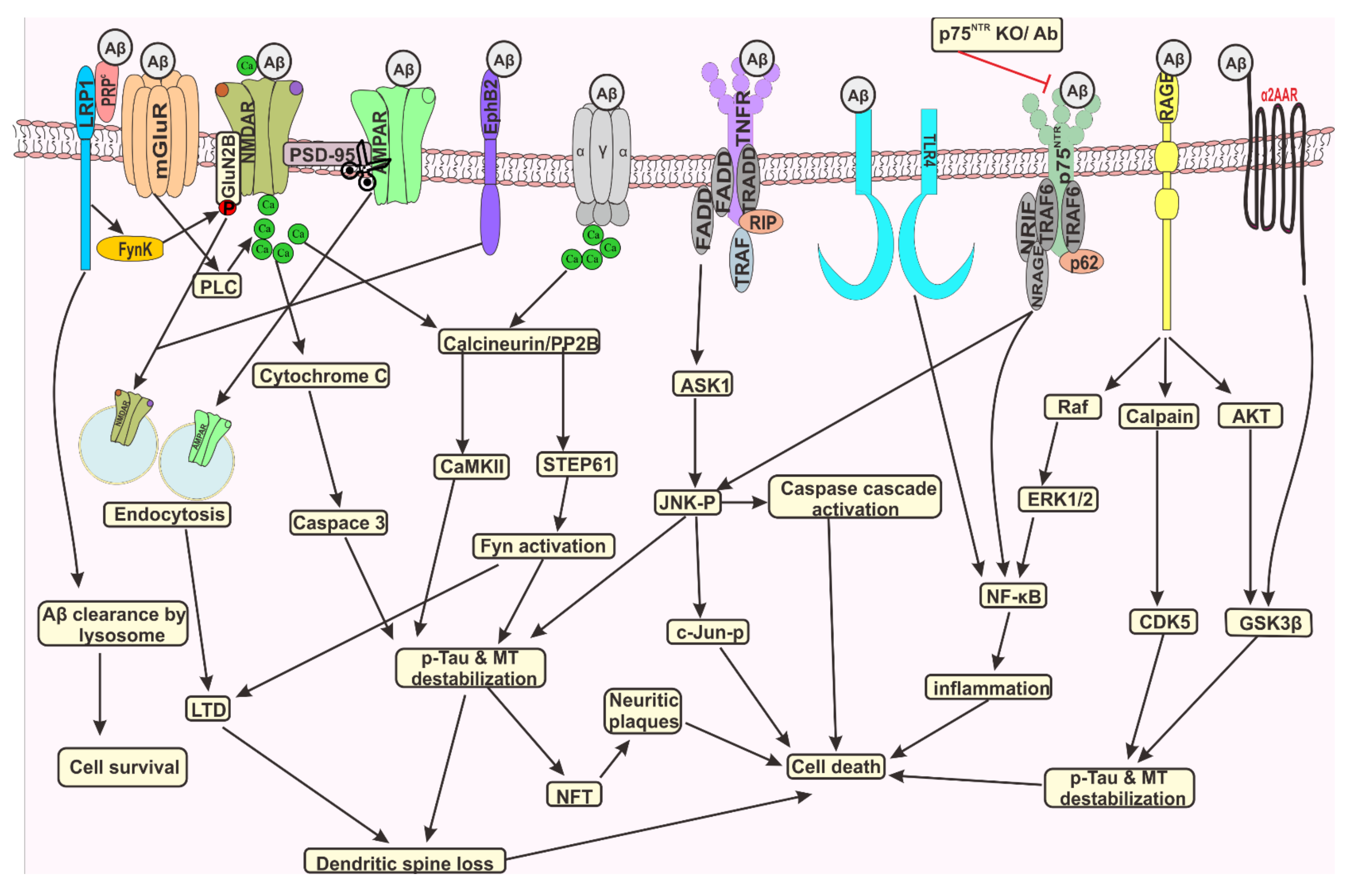
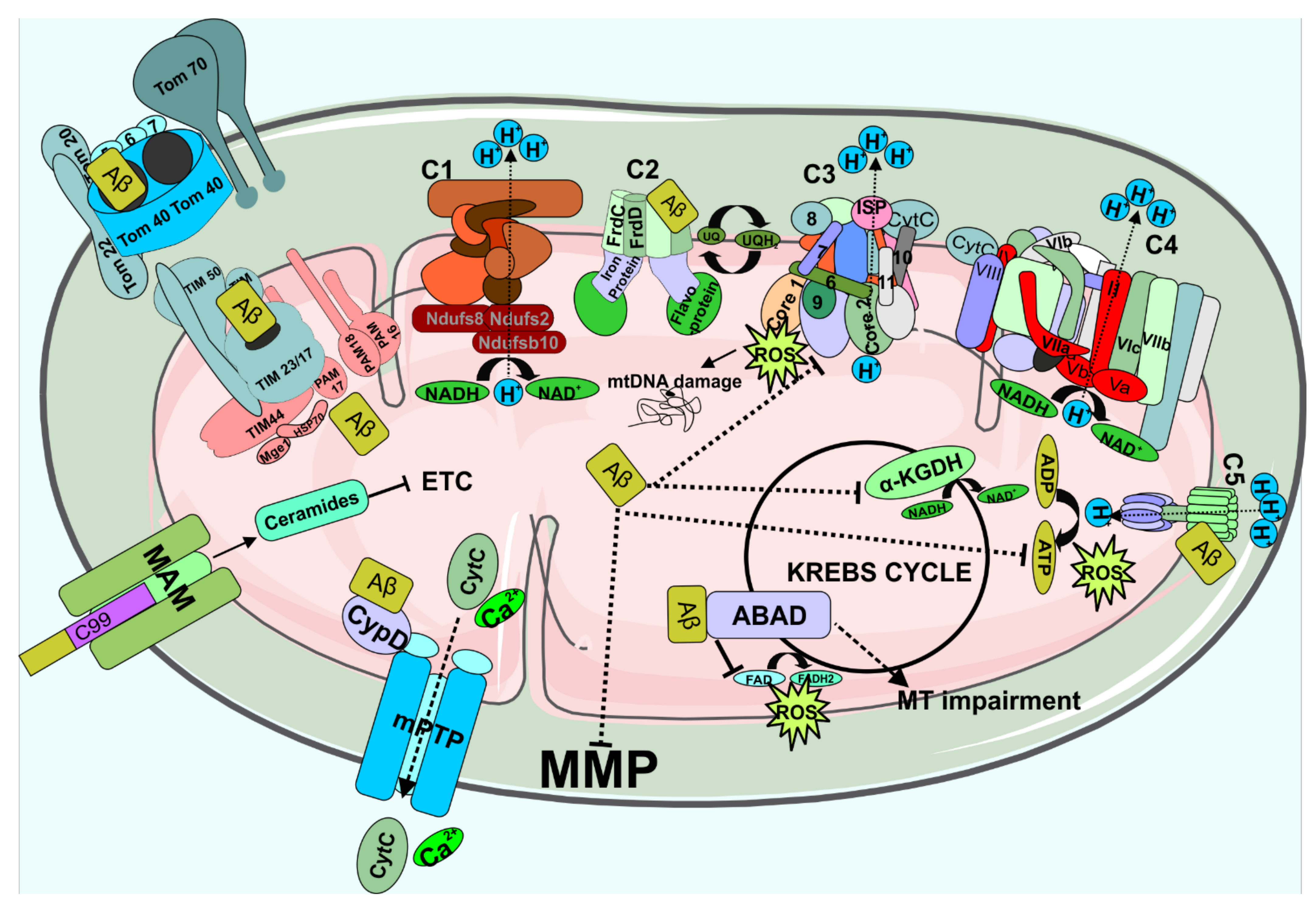
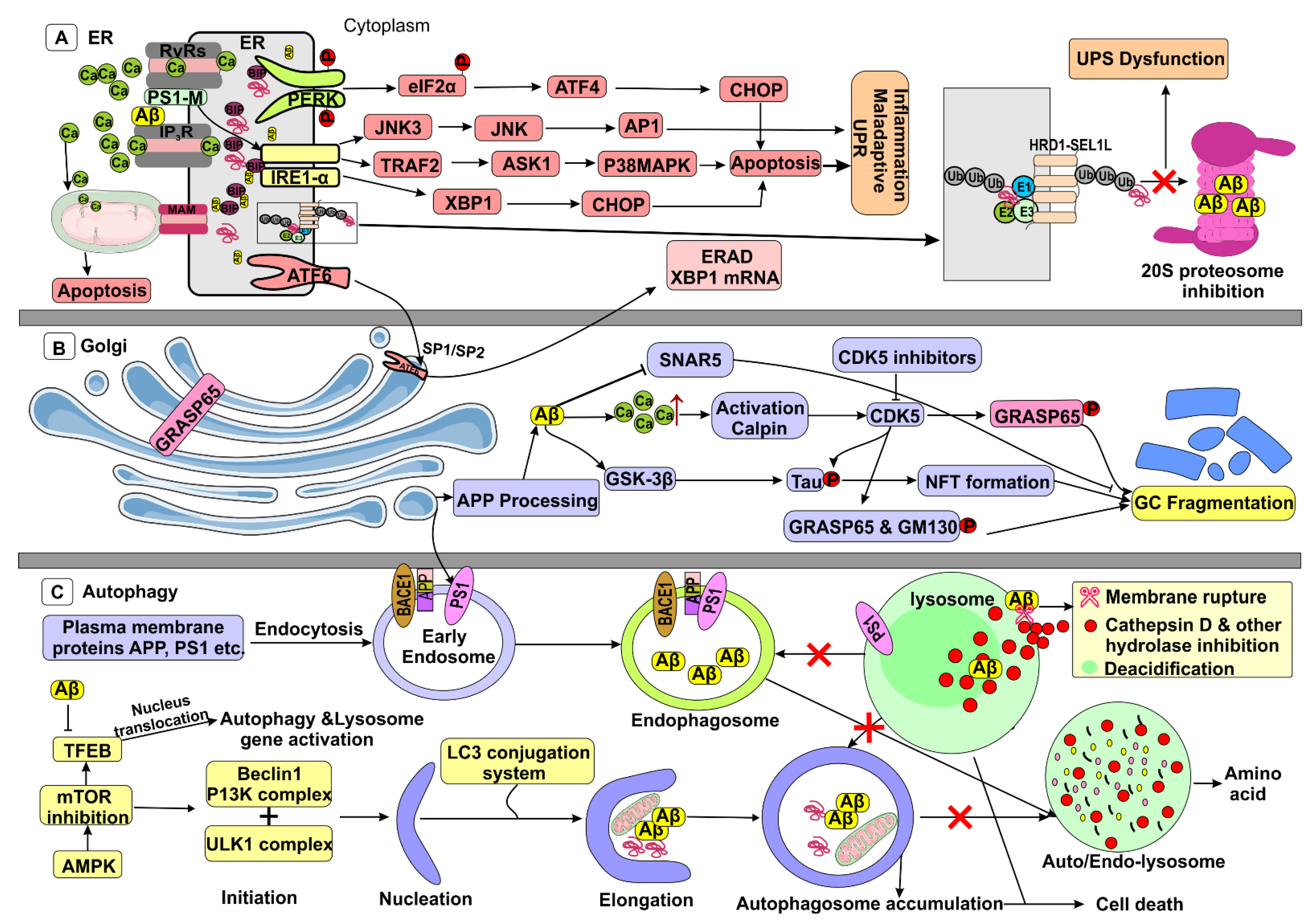
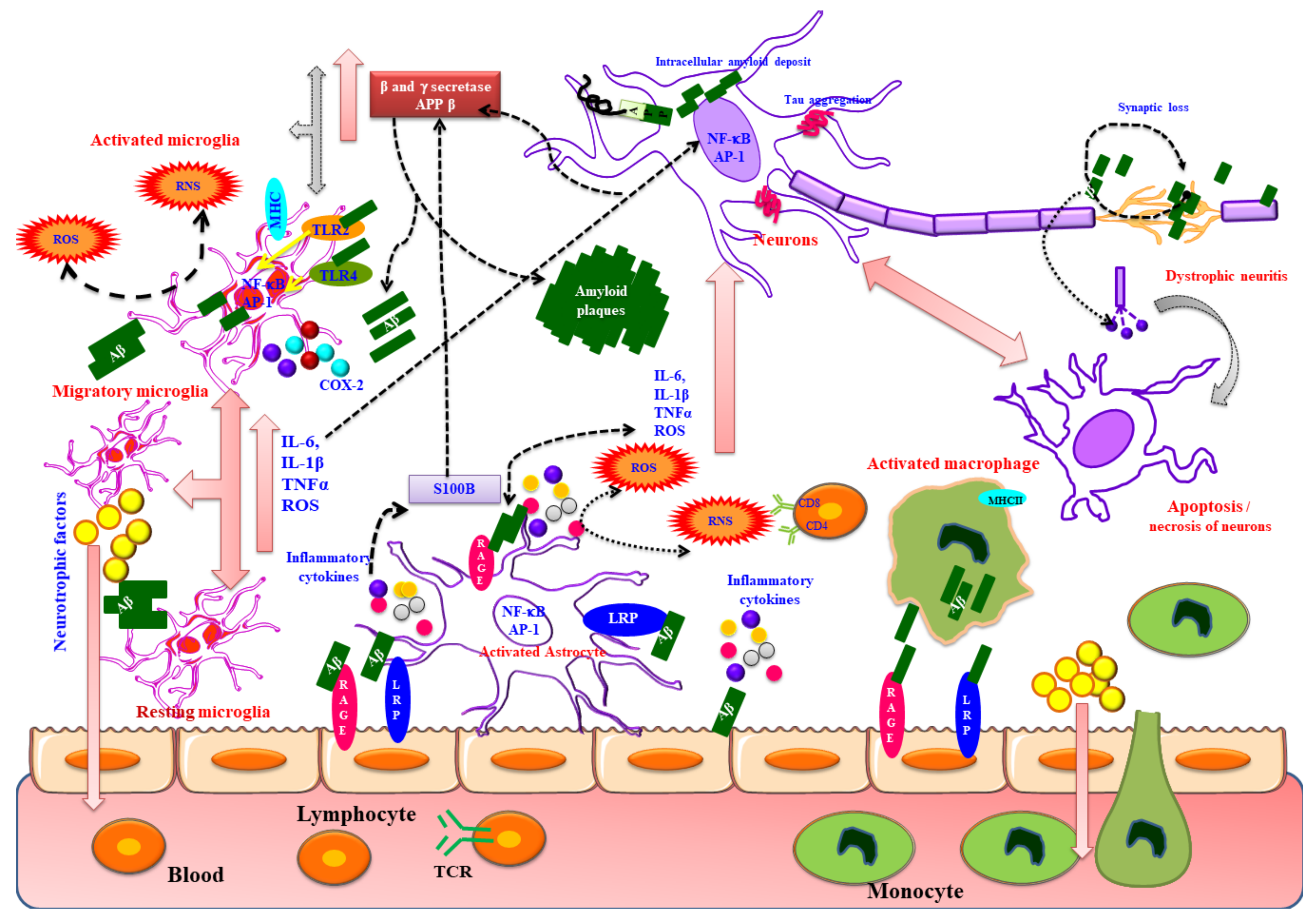
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohamed Asik, R.; Suganthy, N.; Aarifa, M.A.; Kumar, A.; Szigeti, K.; Mathe, D.; Gulyás, B.; Archunan, G.; Padmanabhan, P. Alzheimer’s Disease: A Molecular View of β-Amyloid Induced Morbific Events. Biomedicines 2021, 9, 1126. https://doi.org/10.3390/biomedicines9091126
Mohamed Asik R, Suganthy N, Aarifa MA, Kumar A, Szigeti K, Mathe D, Gulyás B, Archunan G, Padmanabhan P. Alzheimer’s Disease: A Molecular View of β-Amyloid Induced Morbific Events. Biomedicines. 2021; 9(9):1126. https://doi.org/10.3390/biomedicines9091126
Chicago/Turabian StyleMohamed Asik, Rajmohamed, Natarajan Suganthy, Mohamed Asik Aarifa, Arvind Kumar, Krisztián Szigeti, Domokos Mathe, Balázs Gulyás, Govindaraju Archunan, and Parasuraman Padmanabhan. 2021. "Alzheimer’s Disease: A Molecular View of β-Amyloid Induced Morbific Events" Biomedicines 9, no. 9: 1126. https://doi.org/10.3390/biomedicines9091126
APA StyleMohamed Asik, R., Suganthy, N., Aarifa, M. A., Kumar, A., Szigeti, K., Mathe, D., Gulyás, B., Archunan, G., & Padmanabhan, P. (2021). Alzheimer’s Disease: A Molecular View of β-Amyloid Induced Morbific Events. Biomedicines, 9(9), 1126. https://doi.org/10.3390/biomedicines9091126






